Abstract
Background:
Achieving endoscopic and histological remission is a critical treatment objective in ulcerative colitis (UC). Nevertheless, interobserver variability can significantly impact overall assessment performance.
Objectives:
We aimed to develop a deep learning algorithm for the real-time and objective evaluation of endoscopic disease activity and prediction of histological remission in UC.
Design:
This is a retrospective diagnostic study.
Methods:
Two convolutional neural network (CNN) models were constructed and trained using 12,257 endoscopic images and biopsy results sourced from 1124 UC patients who underwent colonoscopy at a single center from January 2018 to December 2022. Mayo Endoscopy Subscore (MES) and UC Endoscopic Index of Severity Score (UCEIS) assessments were conducted by two experienced and independent reviewers. Model performance was evaluated in terms of accuracy, sensitivity, and positive predictive value. The output of the CNN models was also compared with the corresponding histological results to assess histological remission prediction performance.
Results:
The MES-CNN model achieved 97.04% accuracy in diagnosing endoscopic remission of UC, while the MES-CNN and UCEIS-CNN models achieved 90.15% and 85.29% accuracy, respectively, in evaluating endoscopic severity of UC. For predicting histological remission, the CNN models achieved accuracy and kappa values of 91.28% and 0.826, respectively, attaining higher accuracy than human endoscopists (87.69%).
Conclusion:
The proposed artificial intelligence model, based on MES and UCEIS evaluations from expert gastroenterologists, offered precise assessment of inflammation in UC endoscopic images and reliably predicted histological remission.
Plain language summary
Why was this study done? This study aimed to develop a real-time and objective diagnostic tool to reduce subjectivity when evaluating ulcerative colitis (UC) endoscopic disease activity and to predict histological remission without mucosal biopsy.
What did the researchers do? We developed and validated a deep learning algorithm that uses UC endoscopic images to predict the Mayo Endoscopic Score (MES), US Endoscopic Index of Severity Score (UCEIS), and histological remission.
What did the researchers find? The constructed MES- and UCEIS-based models both achieved high accuracy and performance in predicting histological remission, outperforming human endoscopists.
What do the findings mean? The efficiency and performance of the deep learning algorithm rivaled that of expert assessments, which may assist endoscopists in making more objective evaluations of UC severity and in predicting histological remission.
Graphical abstract

Introduction
Ulcerative colitis (UC) is an idiopathic chronic inflammatory disorder featuring recurrent inflammation of the colonic and rectal mucosa, manifesting as diffuse and continuous superficial inflammation and corresponding histological changes. 1 Given its increasing incidence worldwide,2–4 it has become even more important to achieve early diagnosis and induce rapid remission. Treatment choice depends on disease severity, with mild to moderately active UC usually treated with oral/topical 5-aminosalicylic acid or oral glucocorticoids and severe UC usually requiring intravenous glucocorticoid therapy, even immunosuppressants, and expensive biological reagents.5,6 Assessment of UC patients and their response to therapy primarily involves colonoscopy and histological analysis. Several metrics have been proposed to evaluate the endoscopic activity of UC, 7 including the Mayo Endoscopic Subscore (MES) 8 and Ulcerative Colitis Endoscopic Index of Severity (UCEIS). 9 The MES system, which is based on four grades, remains the most widely employed in clinical practice due to its simplicity. However, it lacks the ability to distinguish superficial from deep ulcers and has yet to be formally validated. By contrast, the UCEIS system outperforms the MES in assessing disease activity in UC by providing finer details to distinguish endoscopic severity. Nevertheless, its application is mostly restricted to clinical trials due to the relative complexity of the scoring system. 10 Furthermore, the reliance on subjective interpretation by individual endoscopists for endoscopic scoring raises concerns regarding interobserver variability and subsequent treatment planning for UC. In addition, the evaluation of histological sections is critical for predicting long-term remission and cancer prevention. 11 However, obtaining the necessary mucosal specimens imposes financial strains and psychological burdens on patients, extends waiting times for pathological diagnoses, and poses potential risks during colonoscopy procedures. Furthermore, different histological interpretations can also be a challenge. Therefore, the implementation of objective assessment techniques for evaluating disease conditions in UC patients could enhance treatment options and efficacy and provide a more accurate prognosis.
Recently, the application of artificial intelligence (AI) in colonoscopy has attracted attention as an endoscopist-independent tool for predicting UC disease activity.12–14 Studies have shown that deep learning models trained on specific medical images can achieve expert-level evaluations. For instance, Ozawa et al. 15 assessed the performance of a convolutional neural network (CNN) in differentiating between active inflammation (defined as MES 2 or 3) and remission (defined as MES 0 or 1) using a large number of endoscopic images of UC patients, yielding encouraging results. Similarly, Stidham and Takenaka 16 reported on the ability of a CNN model to distinguish between MES 0 or 1 disease and MES 2 or 3 disease, showing excellent performance and good agreement with human reviewers. However, these studies did not discriminate against each category. More recently, Bhambhvani and Zamora 17 developed a CNN for automated classification of individual MES grades, while Byrne et al. 18 advanced a deep learning model to enhance and accelerate the evaluation process, demonstrating strong agreement with the MES and UCEIS systems. Remarkably, deep learning approaches have also shown potential in predicting histological remission using endoscopic images only, without necessitating a mucosal biopsy specimen. For example, Maeda et al. 19 established a real-time AI system that automatically predicted histologically active inflammation, achieving an accuracy of 81.5%. Furthermore, Takenaka et al. 20 developed a deep neural network system that predicted histological remission with an accuracy of 92.9% and a kappa coefficient of 0.859. Nevertheless, despite the notable contributions of existing research and applied AI solutions, various challenges remain to be addressed for successful integration into daily clinical practice, particularly in the context of UC. As such, we developed a computer-aided diagnosis (CAD) system containing two CNN modules based on the MES (MES-CNN) and UCEIS systems (UCEIS-CNN) to evaluate endoscopic remission and activity, differentiate individual MES and UCEIS scores, and predict histological remission based on endoscopic images of UC patients.
Materials and methods
Data collection
Clinical data from patients who underwent endoscopic procedures from January 2018 to December 2022 at the Department of Gastroenterology, First Affiliated Hospital of Kunming Medical University, Kunming, Yunnan, China, were reviewed. All imaging procedures utilized standard colonoscopy and endoscopy systems (Olympus, Tokyo, Japan). Using the Lennard-Jones criteria, a total of 1 124 UC patients were diagnosed based on the typical clinical course of the disease, endoscopic examination, and histological confirmation. 21 Exclusion criteria included the following: (1) patients with prior colon surgery, unclassified inflammatory bowel disease (IBD), or Crohn’s disease; (2) patients diagnosed with neoplasm, concomitant infectious colitis, or who were pregnant or lactating; and (3) patients for whom colonoscopy was contraindicated. Disease activity and severity were categorized using Truelove and Witts’ classification of UC. After excluding unclear images due to the presence of stool, blurriness, or halos, a total of 12,257 endoscopic images were collected. 22
The MES evaluation criteria range from 0 to 3: MES = 0 (MES 0) indicates the absence of obvious active lesions; MES = 1 (MES 1) indicates mild lesions, with endoscopic features of erythema and reduced blood vessel texture; MES = 2 (MES 2) indicates moderate lesions, with endoscopic features of obvious erythema, blood vessels, texture loss, and erosion; and MES = 3 (MES 3) indicates severe lesions, with endoscopic features of spontaneous bleeding and ulcer formation. 8 The UCEIS scoring system ranges from 0 to 8 and consists of three descriptors (calculated as a simple sum): vascular pattern (scored as 0–2), bleeding (scored as 0–3), and erosions and ulcers (scored as 0–3), which are further stratified into four grades: that is, remission (0), mild (1–3), moderate (4–6), and severe (7–8). 9 For this study, images were evaluated based on MES and UCEIS endoscopic severity by two expert gastroenterologists, QN and YM, both with over 10 years of experience. If scores between the two differed, a third independent reviewer, JM, with over 15 years of experience, rendered the final determination. Both MES = 0 and UCEIS = 0 were defined as mucosal healing.
Histological findings from the same patient cohort were also analyzed. The Geboes score 23 was used to evaluate the histological severity of inflammation, defining remission as Geboes ⩽ 3.0 and active inflammation as Geboes > 3.0. 24 Histological grade scoring was not performed given the difficulty in determining the grade solely from endoscopic images. All histological images were examined and interpreted by two pathologists, each with over a decade of experience. In instances where assessments differed for a particular biopsy, consensus was reached through discussion. Both pathologists and gastroenterologists conducted assessments blind to any clinical information.
We initially reviewed 1124 patients diagnosed with UC from January 2018 to December 2022. A total of 9807 images from 872 patients met the selection criteria and were used as a training set. These images were annotated using the MES and UCEIS scoring systems, respectively. To verify the effectiveness of the network, 2450 endoscopic images from 252 patients with UC, obtained from July 2021 to December 2022, were used as a verification set. Prior to training, the images underwent data augmentation, including horizontal flip, vertical flip, random zoom, and random rotation. The study was conducted in compliance with the Ethics Committee of the First Affiliated Hospital of Kunming Medical University (No. 2022-L-126). The reporting of this study conforms to the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) statement. 25 Written informed consent was obtained from all participants or their guardians in the case of patients under the age of 18. All accompanying patient information was annotated before data analysis. No patient information, including text, images, and tables, is presented in the paper, and all research protocols were conducted following relevant guidelines and regulations. Based on the endoscopic images taken by different machines (named ‘dataset’), the endoscopic features and inflammation grade were categorized based on the MES and UCEIS scoring systems. Detailed information on the dataset is shown in Figure 1 and Table 1.
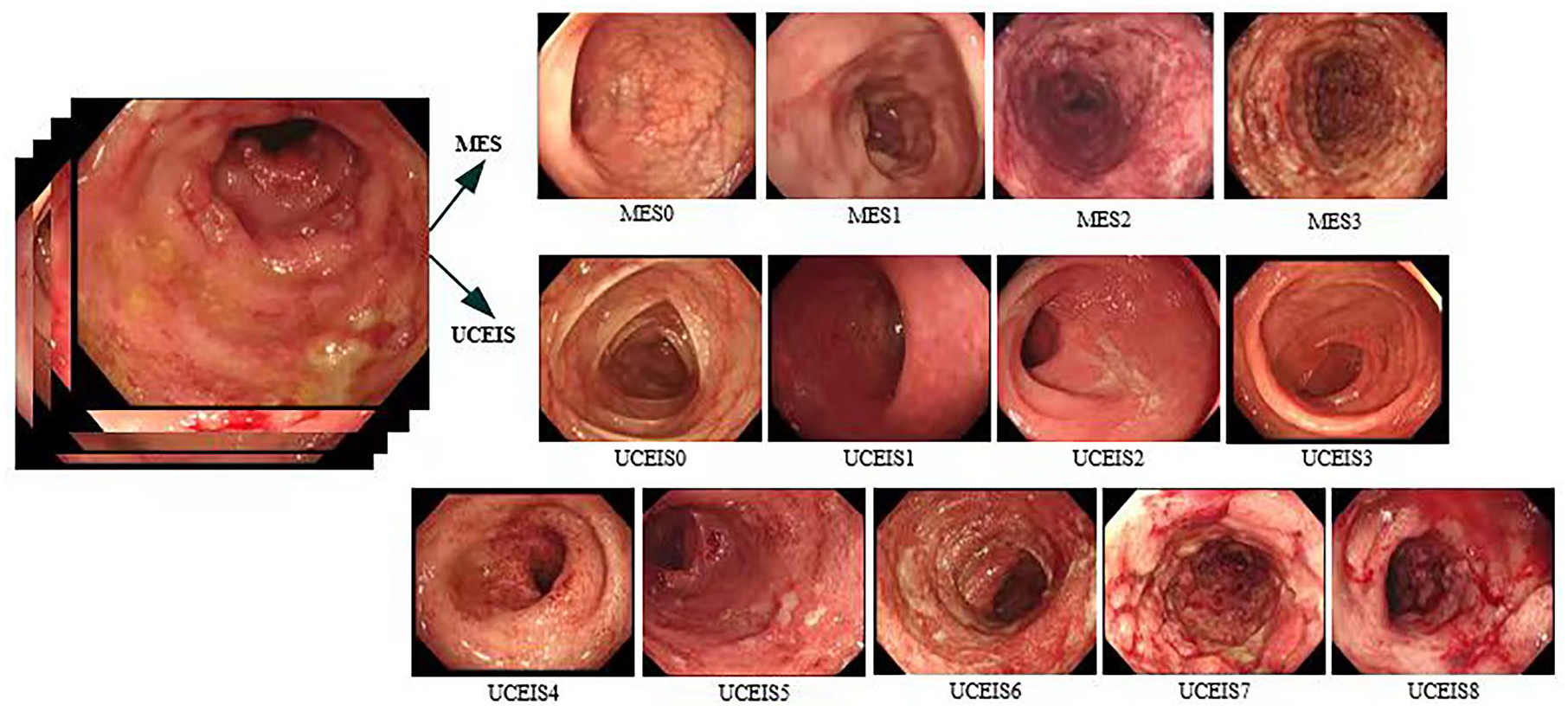
Endoscopic features of MES and UCEIS systems.
Statistics of MES and UCEIS datasets.

MES, Mayo Endoscopy Subscore; UCEIS, UC Endoscopic Index of Severity.
Construction of CNN models
With the rapid advancements in high-performance computing, CNNs have become increasingly important in the fields of computer vision and medical image processing. In the current study, we used Inception-ResNet-v2 26 architecture as a skeleton network, composed of stem, Inception-ResNet, reduction, and SoftMax layers. By integrating the ‘residual’ structure proposed in ResNet 27 into the Inception module, we accelerated training and improved performance. The inception module allowed us to capture both sparse and non-sparse features of the same layer and utilize 1 × 1 convolution to reduce parameter number, improve recognition speed, and facilitate faster network convergence. We also used dropout 28 to reduce weight and improve network robustness. Finally, the corresponding probability was calculated using the SoftMax classifier. The MES and UCEIS scores of each endoscopic image were judged and the relationship with histological images was constructed to predict histological remission. The overall network structure is shown in Figure 2.

Overall network structure.
Software, hardware, and evaluation indices
The experiment was conducted using Windows 10, Spyder editor, and SPSS (v26.0) software. 29 The computational setup included the following: CPU model AMD Ryzen 7 and GPU model NVIDIA GeForce RTX 2080Ti. All programs were implemented using the open-source framework Keras, 30 with TensorFlow backend and Python port.
Accuracy was used to measure the proportion of samples for which the diagnostic predictions aligned with actual outcomes. Sensitivity was applied to represent the percentage of patients correctly identified as positive among the total number of patients. Positive predictive values (PPVs) were used to indicate the proportions of true-positive and true-negative results, in statistical and diagnostic testing. The formulas for calculating the outcomes mentioned above are provided in Table 2, where TP refers to true positive (correctly recognizing a positive sample), TN represents true negative (correctly recognizing a negative sample), FP indicates false positive (incorrectly identifying a sample as positive when it is negative), and FN stands for false negative (incorrectly identifying a sample as negative when it is positive).
Formulas of evaluation metrics.

PPV, positive predictive value.
Results
Clinical, endoscopic, and histological features in validation sets
After excluding patients with unclassified IBD, colorectal neoplasia, infectious disease, or contraindicated for colonoscopy, a total of 2450 images from 252 patients were collected from July 2021 to December 2022 as a verification set to validate the effectiveness of the network. Clinical features of the UC patients in the verification set are shown in Table 3.
Clinical, endoscopic, and histological features of UC patients in validation set.
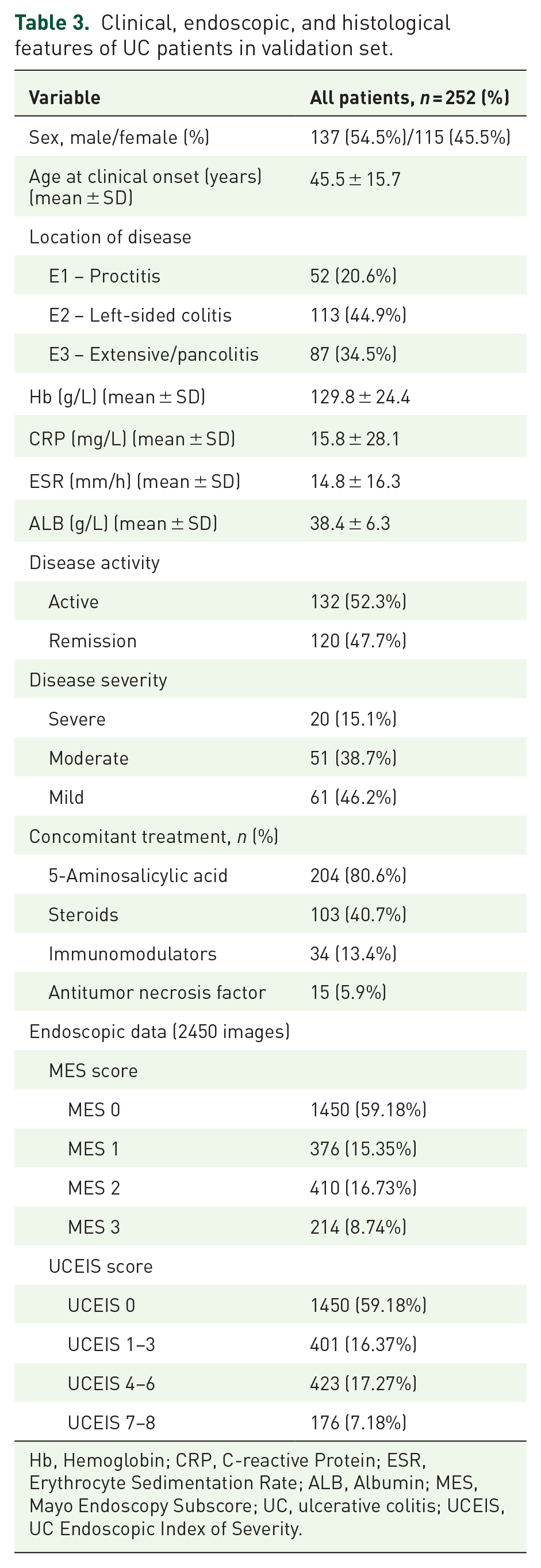
Hb, Hemoglobin; CRP, C-reactive Protein; ESR, Erythrocyte Sedimentation Rate; ALB, Albumin; MES, Mayo Endoscopy Subscore; UC, ulcerative colitis; UCEIS, UC Endoscopic Index of Severity.
Diagnostic capabilities of CNN models
In this experiment, the training phase spanned 30 epochs, after which changes in accuracy and loss were observed. Loss refers to the ‘disagreement’ between the obtained and ideal outputs and loss function refers to the mathematical functions that measure this deviation. 31 Thus, 30 passes of the entire training dataset of the deep learning algorithm were completed before weights were updated in the network. The models with the highest accuracy on the validation set were saved. Data augmentation was used to prevent overfitting and improve the generalization capabilities of the models. Each image was adjusted to a 299 × 299 input network and Adam optimization was performed. 32 The initial learning rate of the optimizer, which controls the rate or speed with which the model parameters are altered during training, was set to 1e−4. When evaluating endoscopic remission, the trained MES-CNN model achieved 97.04% accuracy, 98.43% sensitivity, and 96.36% PPV. When assessing endoscopic disease activity, the trained MES-CNN model achieved 90.15% accuracy, 83.66% sensitivity, and 85.60% PPV. Similarly, when evaluating endoscopic disease activity, the trained UCEIS-CNN model yielded 85.29% accuracy, 77.75% sensitivity, and 67.07% PPV. The corresponding results and 95% confidence intervals (CIs) are presented in Table 4. Figures 3 and 4 display the box plots for the diagnosis and classification of endoscopic images, as well as the confusion matrix of the dataset.
Diagnostic capabilities of MES-CNN and UCEIS-CNN models.
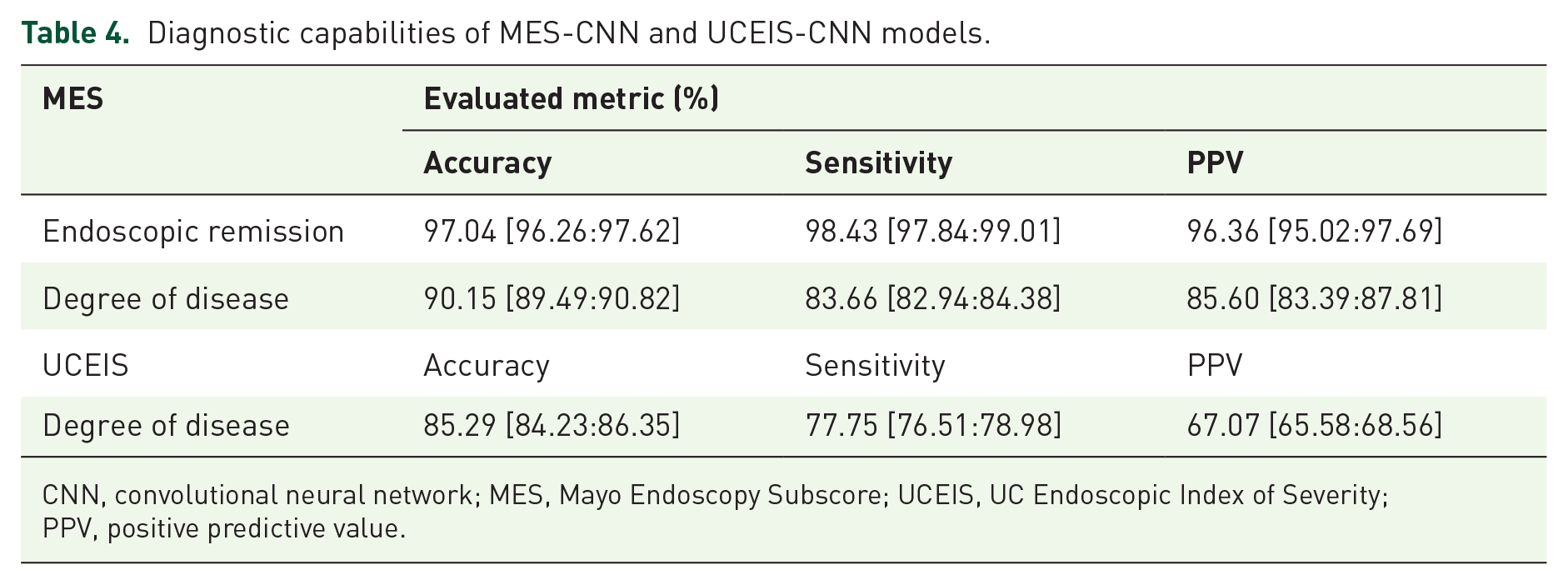
CNN, convolutional neural network; MES, Mayo Endoscopy Subscore; UCEIS, UC Endoscopic Index of Severity; PPV, positive predictive value.

Box plot of endoscopic image diagnosis and classification (accuracy). Panel (a) represents two categories under MES, Panel (b) represents four categories under MES, and Panel (c) represents nine categories under UCEIS.

Confusion matrix of the dataset. Panel (a) represents two categories under MES, (b) represents four categories under MES, and (c) represents nine categories under UCEIS.
We evaluated the accuracy of the CNN models for each MES and UCEIS category. When assessing UC severity using the dataset, the diagnostic accuracies for MES = 1, 2, and 3 were 71.80%, 85.85%, and 77.57%, respectively. Notably, the diagnostic accuracies for UCEIS were lower than those for MES-CNN, as shown in Figure 4 and Table 5.
Accuracy of different MES and UCEIS scores.

MES, Mayo Endoscopy Subscore; UCEIS, UC Endoscopic Index of Severity.
The trained model output probability scores (between 0 and 1) for each category for each image. The category with the highest probability was used as the final classification of the model and Gradient-Weighted Class Activation Mapping (Grad-CAM) 33 was applied to visualize the endoscopic images, as shown in Figure 5.
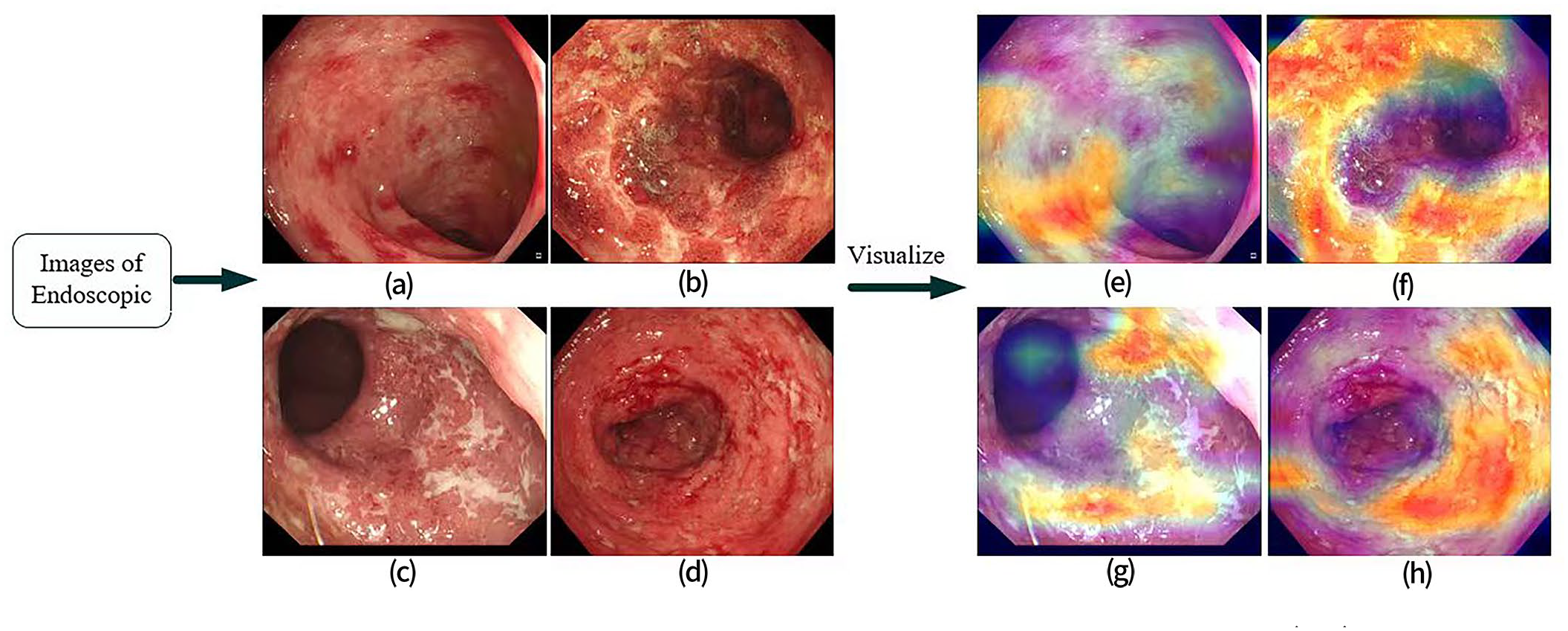
Grad-CAM visualizations of endoscopic images. Grad-CAM was performed on endoscopic images (a-d) for the visualized heatmaps(e-h), which provided visual cues about the areas in the images that the model focuses on for making its classification decision. Panel (a) corresponds to (e), (b) corresponds to (f), (c) corresponds to (g), and (d) corresponds to (h).
Prediction of histological remission
Clinical data from 218 out of the 252 UC patients in the verification set, who underwent biopsies, were utilized to assess the performance in predicting histological remission. In total, 2127 endoscopic images and 1763 biopsy images from 218 patients were used as the verification set. Images diagnosed with MES = 0 and UCEIS = 0 were indicative of endoscopic remission. A one-to-one correspondence was established to compare histological results with endoscopic images. Among the 1763 biopsy images, 55.64% (981 cases) showed histological remission (Geboes⩽3.0), while 44.36% (782 cases) exhibited active disease. Of these, 1546 images were consistent with the endoscopic data, while 217 images were inconsistent. The coincidence rate between endoscopic and histological measures was 87.69% (Table 6). The CNN systems performed well in predicting histological remission, with both the MES-CNN and UCEIS-CNN models yielding consistent histological results (accuracy rate of 91.28%, Table 7).
Results of endoscopic images and biopsy specimens.

Diagnostic performance of CNN for histological remission.

CNN, convolutional neural network.
Discussion
Diagnosis of UC is based on clinical presentation, endoscopic evaluation, and histological parameters. Recent studies have highlighted the importance of mucosal healing in UC, suggesting it should be considered alongside clinical findings for effective long-term treatment strategies.34,35 At present, endoscopy and mucosal biopsy serve as the primary methods to assess mucosal lesions and therapeutic efficacy. Thus, objective evaluation of UC remains essential for diagnosis, treatment, and monitoring of disease. Nonetheless, both endoscopic and histological assessments are prone to interobserver variability, and achieving proficiency in discerning MES and UCEIS scores requires rigorous training, especially for individuals new to endoscopy. The American Society for Gastrointestinal Endoscopy currently recommends the collection of nearly 10 biopsy specimens for this purpose, which can impose considerable time and cost burdens as well as increase the potential for adverse complications during colonoscopy.
Recent advancements in AI offer a promising avenue to improve the quality of endoscopy. In this context, we constructed a CAD-based system containing two CNN models. The models accurately diagnosed UC severity by analyzing features from endoscopic images, including mucosal membrane conditions and blood vessel states, to assess endoscopic activity and inflammation and to predict histological remission. Inception-ResNet-v2 was used as a skeleton network to diagnose and classify the degree of activity in the endoscopic images. Given the limited dataset, various data augmentation techniques were also applied to extract additional image features. The trained MES-CNN model showed a robust diagnosis of endoscopic remission, with high accuracy (97.04%), sensitivity (98.43%), and PPV (96.36%). The trained CNN model also performed well in diagnosing endoscopic severity, with high accuracy (90.15%), sensitivity (83.66%), and PPV (85.60%). Under the UCEIS score system, the trained UCEIS-CNN model also performed well in severity diagnosis, with accuracy, sensitivity, and PPV of 85.34%, 77.75%, and 67.07%, respectively (Table 4). In their previous study, Stidham et al. 36 applied a CNN to distinguish endoscopic remission from moderate-to-severe disease, achieving a PPV of 87% (95% CI: 0.85–0.88), sensitivity of 83.0% (95% CI: 80.8–85.4%), and specificity of 96.0% (95% CI: 95.1–97.1%). Furthermore, Sutton et al. 37 achieved moderate to good performance in mild versus moderate-to-severe UC on a public dataset of endoscopic images. Our results also showed high consistency between the two MES- and UCEIS-based CNN models for endoscopic images from clinical datasets, although the MES-CNN model performed slightly better than UCEIS-CNN due to the relatively high similarity between images of adjacent categories.
Most previous research has focused on two-level classification studies, that is, remission (MES 0 or 1) and moderate to severe disease (MES 2 or 3).15,36,37 However, in clinical settings, determining the exact MES or UCEIS scores is critical, given their direct relevance to evaluation, treatment, and prognosis. In this context, the accuracy of the CNN models was assessed for individual MES and UCEIS categories, as illustrated in Figure 4 and Table 5. In evaluating UC severity, the diagnostic accuracies for MES scores 1, 2, and 3 were 71.80%, 85.85%, and 77.57%, respectively. The diagnostic accuracy for MES 1 was notably reduced, primarily due to the tendency to incorrectly classify endoscopic images as either MES 0 or 2. This misclassification was also evident with MES 3 images, often labeled as MES 2, resulting in decreased accuracies for MES scores of both 1 and 3. A comparable pattern was evident within the UCEIS scoring system. Real-world data often exhibit long-tailed distributions. As our data were obtained from a single center, there was an overrepresentation of MES or UCEIS 0 scored images and an underrepresentation of other scored images. Neural networks, when trained on these imbalanced databases, tend to perform well on head classes but worse on tail classes. Therefore, a larger sample containing MES 1–3 and UCEIS 1–8 images from additional endoscopy centers is needed to improve the performance of the CNN models. The diagnostic accuracies for UCEIS were lower than those of MES-CNN, attributed to the relatively high similarity between images of adjacent grades in the UCEIS scoring system. The ability to discern subtlety may be challenged by the smaller differences among adjacent UCEIS scores (ranging from 0 to 8) compared to MES scores (ranging from 0 to 3). Thus, there is potential for further improvement in the CNN model behavior.
In recent years, histological assessment has played a significant role in evaluating inflammatory activity and monitoring treatment responses in UC. Histological remission is related to decreases in relapse rate, hospitalization rate, steroid use, surgery rate, and risk of UC-associated colorectal cancer.38,39 Previous studies have highlighted a relationship between endoscopic mucosal healing and histological remission, with several AI systems utilized to predict such remission.19,20,40 Therefore, we tested the capability of the CNN models in predicting histological disease activity using endoscopic images, intended to reduce the disadvantages associated with biopsy collection and assessment. Our results showed that the CNN models achieved high accuracy in predicting histological remission (91.28%), showing better performance than human endoscopists (87.46%), as well as a high kappa value (0.826). White-light imaging can assess the surface structure and vessel pattern of the mucosa, but it does not sufficiently evaluate the inflammatory infiltrate in the lamina propria. This suggests that endoscopy may underestimate the degree of inflammation in UC,41,42 which can, in turn, limit the congruence between histological and endoscopic inflammatory activity. Presently, MES and UCEIS are the primary endoscopic scoring systems for mucosal inflammation in clinical practice, yet neither aligns perfectly with histological inflammation.5,43 Score systems that show higher concordance with histological severity are in development. The capability of AI to identify and analyze details that may be missed by clinicians highlights its potential role in improving the correlation between endoscopic scores and histological inflammation and in predicting histological remission.
The AI models developed in this study, based on MES and UCEIS and evaluated by expert gastroenterologists, exhibited high accuracy and consistency. Nevertheless, the research has several limitations. First, accuracy was significantly lower for MES 1–3 and UCEIS 1–8 in contrast to MES 0 or UCEIS 0. To address the lack of training examples for these tail classes, future work will implement advanced distribution calibration strategies such as label-aware distribution calibration. 44 Furthermore, other solutions, such as the generative adversarial network, may be employed to decrease misclassification probabilities in subsequent studies. Second, while using video materials may better replicate real-world clinical scenarios and enhance CAD utility in clinical settings, current automated video analysis systems yield suboptimal accuracy, as evidenced in recent studies.45,46 In general, there is potential for enhancing CAD systems using both still images and videos. Thus, further studies are needed, incorporating more real-world video content to corroborate and expand on our observations. Lastly, our research was conducted using data from a single center and a retrospective design, which may constrain the broader applicability of the results. To mitigate this limitation, a prospective, multicenter, large-scale trial will be initiated to evaluate the efficacy and enhance the accuracy of CNN-based AI models in real clinical settings.
Conclusion
In conclusion, we successfully trained and compared two CNN models (MES-CNN and UCEIS-CNN). The CAD system demonstrated expert-level judgment in evaluating mucosal inflammation and predicting histological remission in UC patients. These findings have practical implications in medical settings and may assist inexperienced endoscopists in improving diagnostic accuracy. Furthermore, the proposed CAD system and models may serve as auxiliary tools for clinical teaching and research purposes.
Supplemental Material
sj-pdf-1-tag-10.1177_17562848231215579 – Supplemental material for Application of deep learning in the diagnosis and evaluation of ulcerative colitis disease severity
Supplemental material, sj-pdf-1-tag-10.1177_17562848231215579 for Application of deep learning in the diagnosis and evaluation of ulcerative colitis disease severity by Xinyi Jiang, Xudong Luo, Qiong Nan, Yan Ye, Yinglei Miao and Jiarong Miao in Therapeutic Advances in Gastroenterology
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
