Abstract
Background:
Changes in gastric mucosa caused by Helicobacter pylori (H. pylori) infection affect the observation of early gastric cancer under endoscopy. Although previous researches reported that computer-aided diagnosis (CAD) systems have great potential in the diagnosis of H. pylori infection, their explainability remains a challenge.
Objective:
We aim to develop an explainable artificial intelligence system for diagnosing H. pylori infection (EADHI) and giving diagnostic basis under endoscopy.
Design:
A case–control study.
Methods:
We retrospectively obtained 47,239 images from 1826 patients between 1 June 2020 and 31 July 2021 at Renmin Hospital of Wuhan University for the development of EADHI. EADHI was developed based on feature extraction combining ResNet-50 and long short-term memory networks. Nine endoscopic features were used for H. pylori infection. EADHI’s performance was evaluated and compared to that of endoscopists. An external test was conducted in Wenzhou Central Hospital to evaluate its robustness. A gradient-boosting decision tree model was used to examine the contributions of different mucosal features for diagnosing H. pylori infection.
Results:
The system extracted mucosal features for diagnosing H. pylori infection with an overall accuracy of 78.3% [95% confidence interval (CI): 76.2–80.3]. The accuracy of EADHI for diagnosing H. pylori infection (91.1%, 95% CI: 85.7–94.6) was significantly higher than that of endoscopists (by 15.5%, 95% CI: 9.7–21.3) in internal test. And it showed a good accuracy of 91.9% (95% CI: 85.6–95.7) in external test. Mucosal edema was the most important diagnostic feature for H. pylori positive, while regular arrangement of collecting venules was the most important H. pylori negative feature.
Conclusion:
The EADHI discerns H. pylori gastritis with high accuracy and good explainability, which may improve the trust and acceptability of endoscopists on CADs.
Plain language summary
Helicobacter pylori (H. pylori) is the main risk factor for gastric cancer (GC), and changes in gastric mucosa caused by H. pylori infection affect the observation of early GC under endoscopy. Therefore, it is necessary to identify H. pylori infection under endoscopy. Although previous research showed that computer-aided diagnosis (CAD) systems have great potential in H. pylori infection diagnosis, their generalization and explainability are still a challenge. Herein, we constructed an explainable artificial intelligence system for diagnosing H. pylori infection (EADHI) using images by case. In this study, we integrated ResNet-50 and long short-term memory (LSTM) networks into the system. Among them, ResNet50 is used for feature extraction, LSTM is used to classify H. pylori infection status based on these features. Furthermore, we added the information of mucosal features in each case when training the system so that EADHI could identify and output which mucosal features are contained in a case. In our study, EADHI achieved good diagnostic performance with an accuracy of 91.1% [95% confidence interval (CI): 85.7–94.6], which was significantly higher than that of endoscopists (by 15.5%, 95% CI: 9.7–21.3%) in internal test. In addition, it showed a good diagnostic accuracy of 91.9% (95% CI: 85.6–95.7) in external tests. The EADHI discerns H. pylori gastritis with high accuracy and good explainability, which may improve the trust and acceptability of endoscopists on CADs. However, we only used data from a single center to develop EADHI, and it was not effective in identifying past H. pylori infection. Future, multicenter, prospective studies are needed to demonstrate the clinical applicability of CADs.
Introduction
Gastric cancer (GC) is one of the most common malignancies, accounting for over 1,000,000 new cases and an estimated 783,000 deaths in 2018. 1 Helicobacter pylori (H. pylori) is the leading risk factor for GC, which induces atrophic gastritis and intestinal metaplasia, ultimately leading to the development of GC.2–6 Furthermore, H. pylori eradication improves gastric mucosal atrophy and inhibits the development of intestinal metaplasia.3,7 Therefore, early detection and eradication of H. pylori infection are essential to avoid the development of GC.
The most important tool for early GC (EGC) screening is white light endoscopy (WLE). 8 However, the risk stratification of EGC is related to endoscopic findings of H. pylori infection status. Atrophy, intestinal metaplasia, nodularity, etc., in H. pylori-positive patients are related to the risk of EGC, whereas atrophy plays a more important role in H. pylori-negative patients. 9 In addition, depressed macroscopic EGC lesions are more common in the infected cases than in the uninfected cases, 10 whereas flat elevated lesions are more common in uninfected cases than in the infected cases. 11 The mucosal hyperemia, edema, and redness caused by H. pylori infection make the surface and edges of EGC more difficult to observe. 12 Thus, recognizing H. pylori infection under endoscopy is critical for the diagnosis of EGC. H. pylori infection does not produce detectable specific lesions, making it difficult to diagnose using endoscopy. 13 In addition, the accuracy of endoscopists in diagnosing H. pylori gastritis based on previous experience was approximately 70% under WLE.14,15 Fortunately, an increasing number of H. pylori infection-related mucosal features has been identified, allowing the diagnosis of H. pylori gastritis using WLE.16–19 However, this approach requires advanced skills and knowledge.20,21
Recent studies have shown that artificial intelligence (AI) uses deep learning to achieve feature expression. 22 Furthermore, it plays a vital role in identifying upper gastrointestinal diseases, including esophageal cancer and EGC.23,24 Researchers have made great efforts with the help of AI to diagnose H. pylori infection using WLE. Shichijo et al. collected 32,208 images from 1750 patients for convolutional neural network (CNN) model development and achieved an accuracy of 88.9%, which was higher than that of endoscopists. 25 Zheng et al. 26 developed the CNN model using 11,729 images with an accuracy of 81.4% for a single image per patient, and 93.8% for multiple images (8.3 ± 3.3) per patient, suggesting that CNN using multiple gastric images achieved higher diagnostic accuracy for the evaluation of H. pylori infection. However, previous diagnosis systems return a final decision result without explanation, making it difficult for endoscopists to learn from the models. 27 The poor explainability of these black-box models undermines the physicians’ trust and puts patients at risk, severely limiting AI systems’ clinical applications.28,29 Therefore, improving the explainability of AI systems is necessary for their applications.
In this study, we developed an explainable AI system for diagnosing H. pylori infection (EADHI) based on feature extraction using ResNet-50 and long short-term memory network (LSTM). The performance of EADHI under WLE was evaluated in internal and external test sets and further compared with endoscopists of different levels. To the best of our knowledge, this is the first study to concretize abstract diagnostic theories through feature extraction, providing diagnostic results and a diagnostic basis for endoscopists.
Methods
Patients and esophagogastroduodenoscopy protocol
We retrospectively reviewed patients undergoing esophagogastroduodenoscopy (EGD) with gastric biopsies or H. pylori breath test at Renmin Hospital of Wuhan University (RHWU) between June 2020 and July 2021. We included 1826 patients (881 H. pylori positive and 945 H. pylori negative) for the development of EADHI. Table 1 shows the patient characteristics. Exclusion criteria include (1) patients with a history of GC, peptic ulcer, gastric surgery, or submucosal tumor and (2) patients who received H. pylori eradication or administered antibiotics within a month or proton pump inhibitor within 2 weeks of H. pylori breath test.
Baseline characteristics according to dataset.
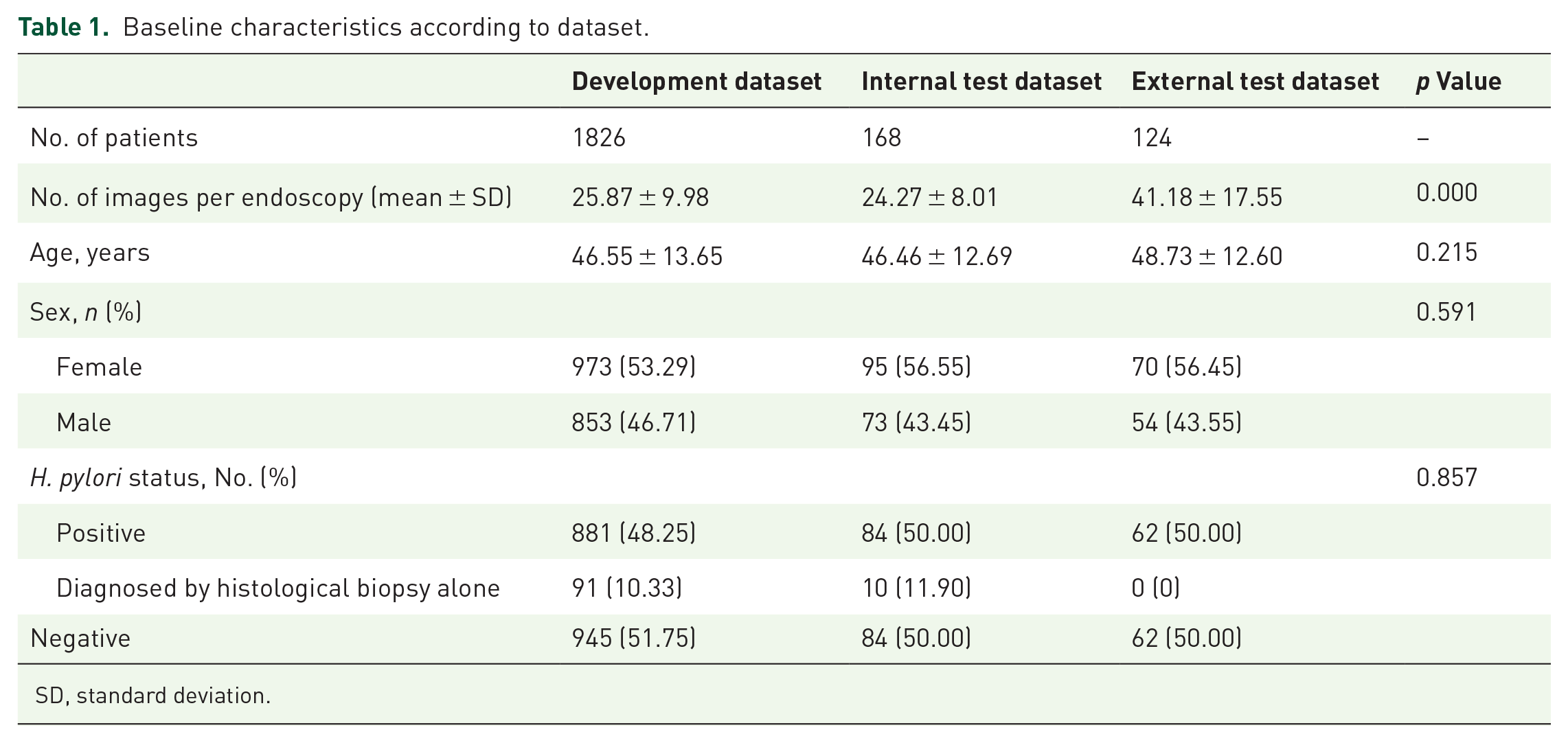
SD, standard deviation.
EGD was performed using a standard endoscope (GIF-HQ290, GIF-H260; Olympus, Tokyo, Japan; EG-L590ZW; Fujifilm, Tokyo, Japan) and the images were captured during high-definition, white-light examination of the antrum, angularis (retroflex), body (forward and retroflex), and fundus (retroflex). Gastric biopsies were performed in the antrum and body at the endoscopist’s discretion.
Establishment of diagnostic feature for H. pylori infection with prior knowledge
Atrophy, intestinal metaplasia, nodularity, diffuse redness, spotty redness, mucosal swelling, enlarged folds, and sticky mucus are positive endoscopic findings for H. pylori. At the same time, regular arrangement of collecting venules (RAC), fundic gland polyp (FGP), red streak, and hematin are predictive features for H. pylori negative.16–19 These endoscopic features in the patients’ images were evaluated by two experts, who had performed more than 5000 examinations. Before evaluating the images, the two experts were educated on the Kyoto classification of gastritis 16 using PowerPoint presentation. This pre-study training was conducted to avoid inter-observer variance, making the endoscopist labels more objective and accurate. The two experts were blinded to the results of patients’ H. pylori infection and were asked to evaluate the same images independently. We included those in which they both agreed and had distinct mucosal features. Mucosal swelling, enlarged folds, and sticky mucus frequently appear in the same images, and these features were collectively referred to as mucosal edema. Because spot redness and diffuse redness are similar and the background of spotty redness is mostly diffuse redness, 16 the two features were classified as mucosal redness. Supplemental Table S1 shows the results of the endoscopic findings in patients. A logistic regression model 30 was employed to evaluate the predictive capabilities of each endoscopic feature (Supplemental Table S2). According to the similarity between endoscopic features and their performance, nine features were finally included. Figure 1 shows the typical images.
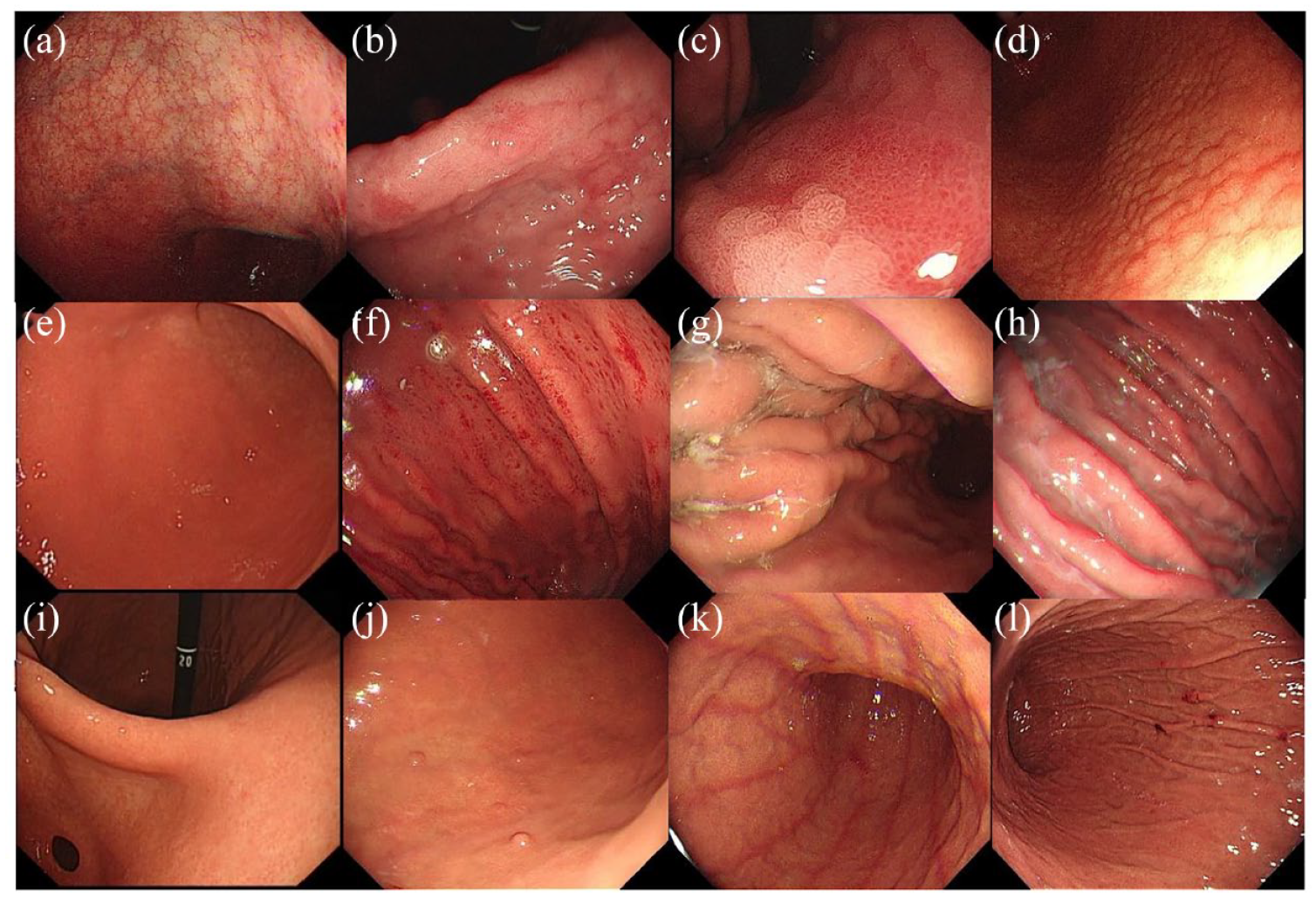
Evaluated endoscopic features: (a) atrophy, (b–c) intestinal metaplasia, (d) nodularity, (e–f) mucosal redness, (g–h) mucosal edema, (i) RAC, (j) FGP, (k) red streak, and (l) hematin.
Datasets and data preprocessing
Figure 2 shows the study’s workflow. EGD images retrospectively obtained from RHWU were used for the development and internal test of EADHI. Meanwhile, to verify the robustness of the system, EGD images were collected from Wenzhou Central Hospital for the external test.
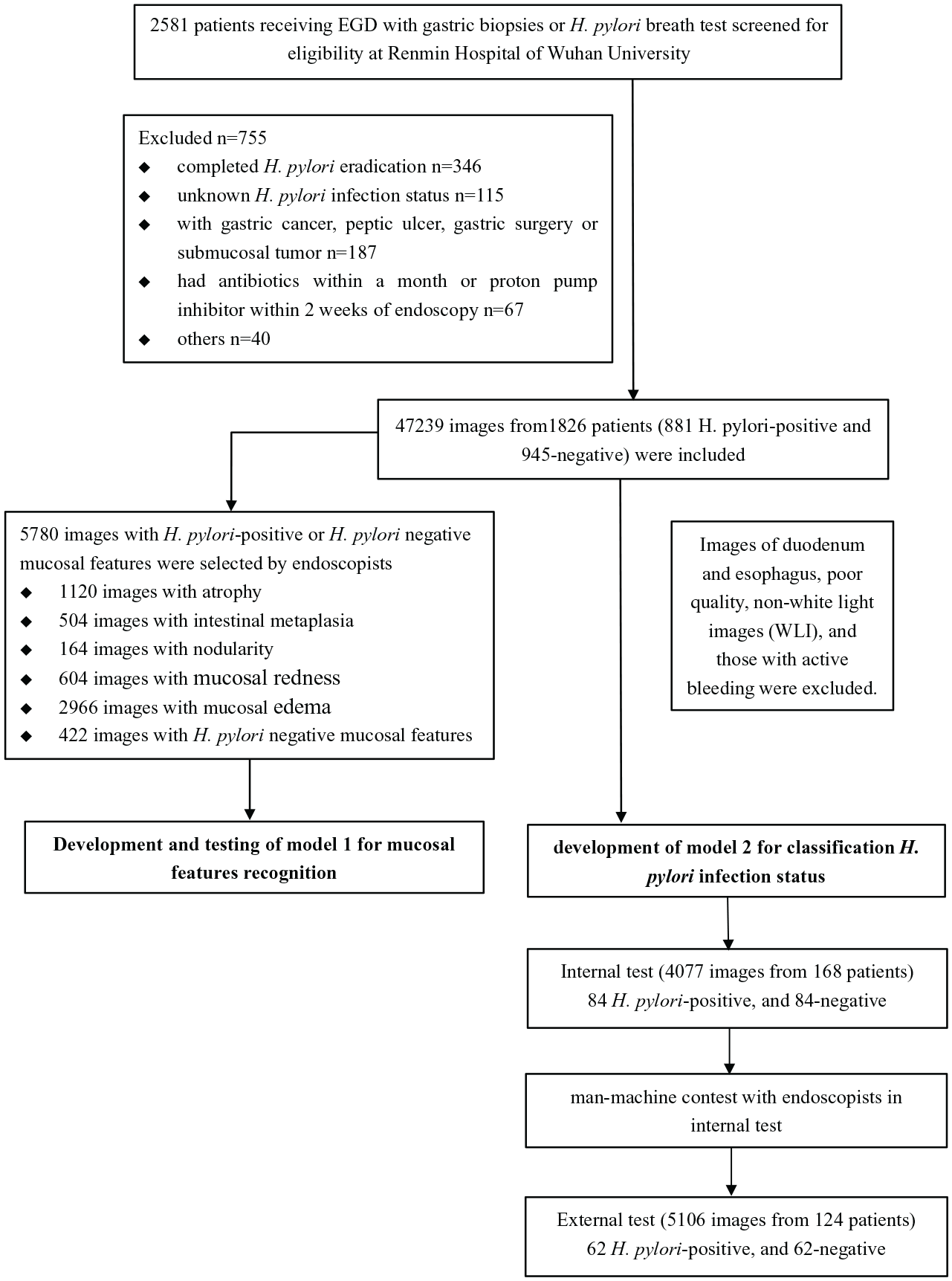
Workflow chart for the development and evaluation of EADHI.
Two gastroenterology doctoral students who had mastered the basic operation and diagnosis knowledge of EGD removed images of duodenum and esophagus, poor quality, non-white light images, and those with active bleeding. Finally, 47,239 images from 1826 patients were used for the development of EADHI. Two experts independently screened 5780 images with distinct mucosal features for developing a model to enhance its ability to identify mucosal features. The images were randomly allocated to the training validation and testing dataset at a ratio of 8:1:1. Random projective transformations such as scaling, shearing, zooming, and horizontal flipping were applied to selected images. Furthermore, in AI model, training projective transformation is a commonly used data augmentation technique that can increase the robustness, stability, and generalization of the model.24,31
The prepared endoscopic images (about 26 images per case) of 881 H. pylori-positive patients and 945-negative patients (Table 1) were used for the development of model 2 to detect H. pylori infection based on identified mucosal features.
Internal testing dataset
A separate testing dataset was developed to evaluate the diagnostic accuracy of EADHI, and compare it with endoscopists. In August 2021, the EGD images (about 24 images per case) of 168 patients (84 H. pylori positive and 84 H. pylori negative) were included as the testing dataset and 71 patients were excluded using the exclusion criteria (Figure 3). Table 1 shows the patient demographics, and there was no overlap between the testing and the development datasets. In all, 10 endoscopists, trained on the Kyoto classification of gastritis 16 before assessing cases using PowerPoint presentation, of varying experience blinded to the patients’ H. pylori infection status were independently asked whether a patient was H. pylori positive or H. pylori negative. Four of the 10 endoscopists were classified as follows: ‘expert group’, with EGDs > 5000. The other endoscopists were further classified as the ‘relatively experienced group’, EGDs > 1000 (n = 3); and the ‘beginner group’, EGDs < 1000 (n = 3). All 10 endoscopists were not involved in the selection of the data.

Flow chart for the enrolled patients.
External testing dataset
Images from the patients diagnosed with H. pylori positive or H. pylori negative (62 H. pylori positive and 62 H. pylori negative) were obtained from Wenzhou Central Hospital as an external testing dataset to evaluate the robustness of the EADHI (Table 1).
Gold standard of H. pylori infection
Patients who tested positive for H. pylori using histological examination of biopsy specimen or breath test were classified as H. pylori positive. However, only 9.8% (101/1027) of H. pylori-positive cases were diagnosed using histological examination of biopsy specimen alone (Table 1). Moreover, patients who tested negative on H. pylori breath test in the absence of eradiation treatment were classified as H. pylori negative.
Construction of the explainable AI system
The EADHI developed in the study contained two models, model 1 developed with ResNet-50 to extract mucosal features, and model 2 developed by combining model 1 with an LSTM network to detect H. pylori infection.
Model 1 was developed with ResNet-50 using six types of EGD images. Figure 4 shows a typical ResNet-50 architecture with these layers, including convolution layers, max-pooling Layers, 16 residual blocks, average pooling, and fully connected layers. High-level features obtained from the average pooling layer of trained ResNet-50 are fed into fully connected layers for classification. We further attempted to understand how model 1 recognized the input images by applying a Gradient-weighted Class Activation Map (Grad-CAM) 32 to determine which area of the images was most essential to the classification result. We developed heatmap images from the location map data. Supplemental Table S3 and Supplemental Figure S1 show the performance of model 1.

A typical architecture of model 1.
Model 2 was developed by combining ResNet-50 and LSTM networks, which were developed by the Keras deep learning framework (TensorFlow backend). 33 In model 2, ResNet-50, which incorporated all layers of model 1 except fully connected layers, was used for feature extraction, while LSTM was used to classify H. pylori infection status based on identified features. And the proposed model 2 developed in the study contains two phases, in phase one, ResNet-50 extracts all the mucosal features in a case from input case-based EGD images and generates feature vectors, whereas in phase two, LSTM receives the feature vectors to extract time information, which is then passed to the dense layer for classification (Figure 5).

A typical architecture of model 2.
To make the diagnosis of the system more specific, we added the information about mucosal features when training model 2, and trained each feature separately. Supplemental Table S4 shows their performance in the validation dataset. As shown in Figure 6, EADHI could identify and output which mucosal features were present in a case, and then output the final result based on the identified mucosal features and their weights calculated using a gradient-boosting decision tree. 34 The sensitivity was plotted against the false-positive rate (i.e. 1-specificity) for all thresholds in the range [0, 1]. In addition, the receiver operating characteristic (ROC) curves were obtained.
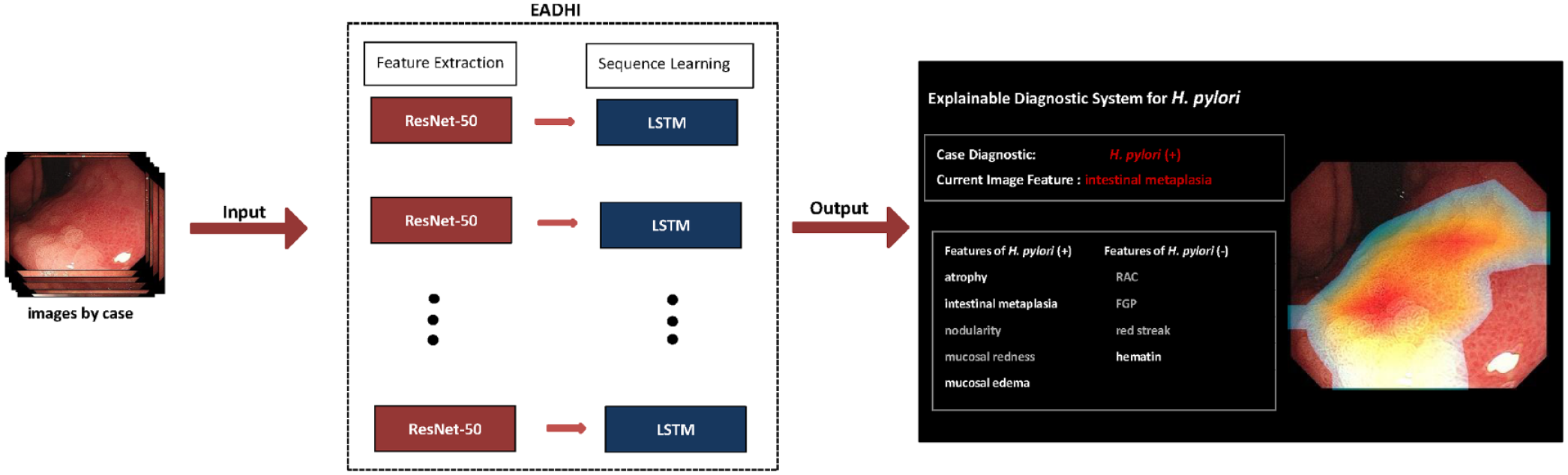
A schematic illustration of how EADHI diagnoses H. pylori infection in images by case. The ResNet-50 could identify and output mucosal features in a case, and the LSTM could diagnose H. pylori infection status based on identified mucosal features.
Sample size
Sample size calculations were performed according to Exact Clopper–Pearson, with assumptions that EADHI has a sensitivity of 90% and a specificity of 90% for detecting H. pylori infection. A prevalence of 44% was estimated, based on recent demographic meta-analysis for China. 35 A sample size of 79 patients was calculated using a two-sided 95% confidence interval (CI) with a width of 0.1.
Ethics
We de-identified all patient information before data analysis to keep patients anonymous. Patient details were not accessible to any of the endoscopists involved in the study. The reporting of this study adheres to the Strengthening the Reporting of Observational Studies in Epidemiology statement. 36
Statistical analysis
Demographic data were expressed as mean with standard deviation (SD). The performance of EADHI was evaluated using the following metrics: accuracy, specificity, sensitivity, positive predictive value, negative predictive value, and area under the curve (AUC) using ROC. Optimal cutoff values to obtain the highest AUC were calculated using the Youden index. The performance of the EADHI and endoscopists were compared using a two-tailed unpaired heteroscedastic Student’s t test. Two-sided p < 0.05 was considered statistically significant.
Results
Performance of EADHI for mucosal features
Table 2 shows that EADHI identified various mucosal features with an overall accuracy of 78.3% (95% CI: 76.2–80.3%), and the accuracy for atrophy, intestinal metaplasia, nodularity, mucosal redness, mucosal edema, RAC, FGP, red streak, and hematin were 78.6% (95% CI: 71.7–84.1%), 81.6% (95% CI: 75.0–86.7%), 91.7% (95% CI: 86.4–95.1%), 83.3% (95% CI: 76.9–88.3%), 85.7% (95% CI: 79.6–90.3%), 82.7% (95% CI: 76.3–87.8%), 69.6% (95% CI: 62.3–76.1%), 72.6% (95% CI: 65.4–78.8%), and 58.9% (95% CI: 51.4–66.1%). The AUC of EADHI for each mucosal feature ranged from 0.66 to 0.95 (Figure 7(a)). And the corresponding weights of each mucosal feature were 0.109, 0.019, 0.024, 0.163, 0.386, 0.275, 0.012, 0.006, and 0.007, respectively (Figure 7(b)).
The performance of EADHI for mucosal features.

CI, confidence interval; EADHI, explainable artificial intelligence system for diagnosing H. pylori infection; FGP, fundic gland polyp; NPV, negative predictive value; PPV, positive predictive value; RAC, regular arrangement of collecting venules.

The corresponding weights of each mucosal feature, the ROC curves of EADHI, and the performance of endoscopists. (a) ROC curves of various mucosal features. (b) The corresponding weights of each feature. The ROC curve for H. pylori infection in internal testing dataset. (c) The ROC curve for H. pylori infection in internal and external testing dataset, and the performance of endoscopists.
Performance of EADHI for H. pylori infection on the internal test set in patients
Figure 7(c) shows the AUC of EADHI for H. pylori infection was 0.96, and at the optimal threshold of 0.26, the accuracy, sensitivity, and specificity were 91.1% (95% CI: 85.7–94.6%), 92.9% (95% CI: 85.0–97.0%), and 89.3% (95% CI: 80.7–94.5%) in the internal test set, respectively (Table 3).
Diagnostic accuracy in internal test data: EADHI versus endoscopists.

Comparison between other groups and experts, *indicates p < 0.05, **indicates p < 0.01.
EADHI, explainable artificial intelligence system for diagnosing H. pylori infection; NPV, negative predictive value; PPV, positive predictive value; SD, standard deviation.
In the internal test set, 94.0% (158/168) cases were diagnosed by H. pylori breath test. EADHI achieved an accuracy of 92.4% (95% CI: 87.1–95.7%), a sensitivity of 93.2% (95% CI: 84.8–97.4%), and a specificity of 91.7% (95% CI: 83.5–96.2%) in these cases (Table 3).
Comparison between EADHI and endoscopists
Table 3 shows the results of H. pylori infection evaluation of the internal test data by the 10 endoscopists. The overall accuracy, sensitivity, and specificity for the diagnosis of H. pylori infection were 75.6% (SD 8.1%), 78.7% (12.7%), and 72.4% (15.5%). The expert group was found to have significantly higher accuracy (83.6% versus 73.2%, p < 0.01) than the relatively experienced group. Similarly, a significant difference in the accuracy was observed between the expert group and the beginner group (83.6% versus 67.3%, p < 0.01). However, there was no statistical difference in the accuracy between the relatively experienced group and the beginner group (73.2% versus 67.3%, p = 0.618).
In addition, the EADHI was found to have a significantly higher accuracy (by 15.5%; 95% CI: 9.7–21.3%), sensitivity (by 14.2%; 95% CI: 5.1–23.3%), and specificity (by 16.9%; 95% CI: 5.8–28.0%) than the endoscopists. When compared with the expert group, the EADHI had higher accuracy (by 7.5%; 95% CI: 2.5–12.4%), although their sensitivity and specificity were comparable. The comparison results are summarized in Figure 7(c) and Table 3.
Performance of EADHI for H. pylori infection on the external test set in patients
To verify the robustness of the EADHI, we examined its performance on an external test set. It achieved an accuracy of 91.9% (95% CI: 85.6–95.7%), a sensitivity of 90.3% (95% CI: 80.1–95.8%), and a specificity of 93.6% (95% CI: 84.1–97.9%), respectively. And the AUC value of 0.96 was achieved at the optimal threshold (Figure 7(c)).
Discussion
We developed an explainable AI system, EADHI, for auto-identification of H. pylori infection by identifying multiple mucosal features. We compared EADHI’s diagnostic ability with endoscopists by examining its internal and external test data performance, and EADHI performed better than that endoscopists and showed good robustness.
In recent years, AI has remarkable improved in diagnosing H. pylori infection based on CNN.25,26 However, because CNN models were developed with a few endoscopic images per patient, mucosal features associated with H. pylori infection status may have been missed. In addition, there have been concerns about the black-box nature and lack of explainability of AI models, which greatly limits their clinical application. 27 In this study, two models were integrated into the system to make a diagnosis of EADHI more specific and comprehensive. Model 1 was first constructed with ResNet-50 to enhance its ability to identify mucosal features. In addition, we confirmed which parts of the images model 1 focuses on using heatmaps. Model 2 was developed by combining ResNet-50 and LSTM networks using case-based images (about 26 images per case). Furthermore, ResNet-50 in model 2 incorporated all layers of model 1 except the fully connected layer to maximize the ability of model 2 to recognize mucosal features. In addition, the information on mucosal features in the cases was added during the training to strengthen further the ability of the system to identify mucosal features.
Previous studies show that an estimated 40–50% of the global population is infected with H. pylori, which is the main risk factor for GC.2,37,38 In addition, H. pylori infection affects the morphology of EGC and makes it difficult to diagnose.10–12 Therefore, early detection of H. pylori infection using endoscopy is crucial. As more endoscopic findings associated with H. pylori infection status are identified, WLE may be used to diagnose H. pylori gastritis.16–19 However, this approach requires advanced skills and knowledge, and the diagnostic process is highly subjective.20,21 In contrast, an AI-aided diagnosis system could provide an objective second opinion and help endoscopists avoid over-reliance on prior experience in the diagnosis process. 39 Herein, the EADHI developed by us could effectively identify mucosal features and comprehensively detect H. pylori infection based on identified features. Our results indicate its potential to assist endoscopists in screening H. pylori infection in real clinical work.
Nodularity is considered to be the strongest evidence to support current H. pylori infection, with extremely high specificity (95.8–98.8%).40–42 A recent study reported that the diagnostic odds ratios (DOR) of diffuse redness, mucosal swelling, sticky mucus, and enlarged serpentine for judging H. pylori positive were 26.8, 13.3, 10.2, and 8.6, respectively. 42 Moreover, most patients with atrophic gastritis have evidence of H. pylori infection.43,44 In addition, Zhao et al. reported that when the positive signs were combined, the ROC/AUC of two or more features had the highest value (0.723). 40 Among these negative mucosal features, the DOR of RAC, FGP, and red streak were 32.2, 7.7, and 4.7, respectively. 42 Furthermore, when these negative features were combined, one or more had the highest ROC/AUC (0.701). 40 These mucosal features make the diagnosis of H. pylori infection status easier than before using WLE, but endoscopists’ skill levels vary significantly.22,40 In our study, the expert group performed significantly better than the other groups. Therefore, an effective method for identifying various mucosal features is required. Model 1 was developed in EADHI to improve its ability to identify mucosal features, and it achieved good diagnostic performance.
The EADHI for detecting H. pylori infection combines ResNet-50 and LSTM networks, with ResNet-50 being used for feature extraction and LSTM to classify H. pylori infection based on these features. The feature extraction network (ResNet-50) is a residual deep learning network (with 50 layers) which attempts to address the problem of vanishing gradients that occur during back-propagation of CNN to effectively extract and recognize the local and global features of images. 45 Furthermore, a recent study reported that ResNet-50 using image generation techniques based on an 80% training set resulted in nearly perfect multi-class prediction accuracy (98.99%) in a 20% validation set. 46 The LSTM network is capable of learning from imperative experiences with long-term states by introducing gate functions into the cell structure. In the case of LSTM, nodes are connected from a directed graph along a time series that is treated as an input with a specific order. 47 Hence, ResNet-50 and LSTM layout feature combination effectively extracted and integrated mucosal features from about 26 endoscopy images per patient, significantly improving the classification.
There are some limitations to this study. First, we invited two doctoral students and two experts to preprocess the data to obtain high-quality labeled data at a low cost. However, it would be better if more endoscopists participated in data preprocessing. Second, the EADHI was developed using data collected retrospectively from a single center and was not prospectively validated. However, we examined EADHI performance in the external testing dataset and found it accurate. Third, we excluded patients with a history of H. pylori eradication from the study. Further research should be conducted to identify previous infections. Finally, H. pylori infection status was confirmed in most patients using one test. However, H. pylori breath test, which has high sensitivity (>95%) and specificity (95%) for the diagnosis of H. pylori infection, was used to diagnose >90% of the positive cases and all the negative cases in this study. 48 Furthermore, we excluded patients who received H. pylori eradication or had antibiotics within a month or proton pump inhibitor within 2 weeks of H. pylori breath test. Thus, the possibility of a false positive or negative H. pylori diagnosis was considered insignificant.
In conclusion, the explainable AI system, EADHI, outperformed the endoscopists in diagnostic accuracy. Furthermore, its diagnostic logic is similar to that of endoscopists, which may increase endoscopists’ trust and acceptability. It has a high potential for assisting endoscopists in screening for H. pylori infection in clinical settings. Further research should be conducted to validate and globally apply the AI-based diagnostic system.
Supplemental Material
sj-docx-1-tag-10.1177_17562848231155023 – Supplemental material for An explainable artificial intelligence system for diagnosing Helicobacter Pylori infection under endoscopy: a case–control study
Supplemental material, sj-docx-1-tag-10.1177_17562848231155023 for An explainable artificial intelligence system for diagnosing Helicobacter Pylori infection under endoscopy: a case–control study by Mengjiao Zhang, Jie Pan, Jiejun Lin, Ming Xu, Lihui Zhang, Renduo Shang, Liwen Yao, Yanxia Li, Wei Zhou, Yunchao Deng, Zehua Dong, Yijie Zhu, Xiao Tao, Lianlian Wu and Honggang Yu in Therapeutic Advances in Gastroenterology
Supplemental Material
sj-tif-2-tag-10.1177_17562848231155023 – Supplemental material for An explainable artificial intelligence system for diagnosing Helicobacter Pylori infection under endoscopy: a case–control study
Supplemental material, sj-tif-2-tag-10.1177_17562848231155023 for An explainable artificial intelligence system for diagnosing Helicobacter Pylori infection under endoscopy: a case–control study by Mengjiao Zhang, Jie Pan, Jiejun Lin, Ming Xu, Lihui Zhang, Renduo Shang, Liwen Yao, Yanxia Li, Wei Zhou, Yunchao Deng, Zehua Dong, Yijie Zhu, Xiao Tao, Lianlian Wu and Honggang Yu in Therapeutic Advances in Gastroenterology
Footnotes
Acknowledgements
We thank the fund support from Innovation Team Project of Health Commission of Hubei Province (Grant no. WJ202C003); The Fundamental Research Funds for the Central Universities (Grant no. 2042021kf0084). We thank the endoscopists and machine-learning engineers for their hard work.
Declarations
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
