Abstract
Th2 polarization is essential for the pathogenesis of allergic rhinitis (AR). Th2 polarization's mechanism requires further understanding. IL-4 is the primary cytokine involved in Th2 response. Fibroblasts play a role in immune regulation. This study aims to elucidate the role of nasal mucosal fibroblast-derived IL-4 in the induction of Th2 responses. Nasal mucosal tissues were obtained from surgically removed samples from patients with nasal polyps, whether with or without AR. Fibroblasts were isolated from the tissues by flow cytometry cell sorting, and analyzed by RNA sequencing (RNAseq). The data from RNAseq showed that nasal fibroblasts expressed genes of GATA3, CD80, CD83, CD86, STAT6, IL2, IL4, IL5, IL6, IL13 and costimulatory factor. The data were verified by RT-qPCR. The level of gene activity was positively correlated with those of AR-related cytokines present in nasal secretions. Nasal fibroblasts release IL-4 upon activation. Nasal fibroblasts had the ability to transform naive CD4+ T cells into Th2 cells, which can be eliminated by inhibiting IL-4 receptor or CD28 in CD4+ T cells. To sum up, nasal mucosal fibroblasts produce IL-4, which can induce Th2 cell development. The data implicate that nasal fibroblasts are involved in the pathogenesis of nasal allergy.
Introduction
Allergic rhinitis (AR) is a reverse response by the immune system in the nasal mucosa to innocent airborne antigens. The clinical symptoms of AR include paroxysmic nasal itch, sneezing, profound nasal discharge, and nasal congestion [1]. In the early stages, AR attacks are self-limited. In the latter stages, the AR attacks may last longer time [1]. Complications of AR may occur, such as nasal polyposis, chronic rhinosinusitis, and allergic asthma [2,3]. The treatment of AR mainly focuses on the symptom control. Allergen-specific immunotherapy (AIT) is also employed in AR clinics. The effects of AIT need to be improved [4,5]. The pathogenesis of AR is unclear [5]. The prevalence of AR has kept rising in the last decades [4,5,6]. Therefore, it is urgent to further investigate the pathogenesis of AR and develop more effective therapeutic remedies for its treatment.
It is recognized that the T helper (Th)2 cell polarization is a pathological feature of AR [4,5,6]. Th2 cell polarization indicates a condition where more than needed numbers of Th2 cells aggregate in the local tissues. More than needed Th2 cytokines are produced by the Th2 cells to saturate the local tissues [7]. Th2 cytokines cause B cells to differentiate into plasma cells, resulting in the production of IgE, which is the primary mediator of allergic diseases. IgE makes mast cells sensitized. Re-exposure to specific antigens induces sensitized mast cells to release allergic mediators, which trigger allergic attacks [5,8]. Despite the allergy research has been advanced rapidly in the recent years [4,5,6], the initiation of AR is still unclear.
Fibroblasts are one of the major cellular components in the nasal mucosa. Published data indicate that that fibroblasts are involved in the pathogenesis of allergic inflammation [9,10]. For example, nasal fibroblasts can produce eotaxin, one of the eosinophil chemoattractants, in response to exposure to Th2 cytokines [11,12]. Environmental pollutants can induce nasal fibroblasts to produce IL-6 and IL-8 [13]. Thus, we hypothesize that nasal fibroblasts contribute to the development of the Th2 response. In the present study, we found that fibroblasts from the human nasal mucosa expressed IL-4, and could induce the development of Th2 cells.
Materials and methods
Reagents
shRNA kits of IL4RA and CD28, antibodies (Abs) of EpCAM [Cat#: sc-25308, fluorochrome: Alexa Fluor (AF) 488], CD45 (sc-1178, AF546), PDPN (sc-53533, AF594), CD3 (sc-20047, AF648), CD19 (sc-390244, AF680), CD11c (sc-46676, AF700), CD80 (sc-376012, AF594), CD83 (sc-55536, AF648), CD86 (sc-28347, AF700), IL-4 (sc-12723, AF488), CD4 (sc-19641, AF546), IL4RA (sc-28361) and CD28 (sc-70612) were purchased from Santa Cruz Biotech (Santa Cruz, CA). ELISA kits of EPX, tryptase, IL-4, IL-5, IL-13, mite specific IgE, mite specific IgG1 and mite specific IgG2 were purchased from Dakewe BioMart (Shenzhen, China). Reagents and materials for RT-qPCR and Western blotting were purchased from Invitrogen (Carlsbad, CA).
Human subjects
The use of human tissues in the present study was approved by the Human Ethics Committee at our hospital (Approval#: H2022002). A written informed consent was obtained from each person. Patients with nasal polyps with or without perennial AR were enrolled into this study in our hospital from May 2022 to June 2023. The diagnosis of AR was carried out by our doctors following the routine procedures of our hospital, which can be found elsewhere [14]. Briefly, patients had perennial AR history more than two years. Positive results of serum specific IgE and skin prick test (SPT) [14]. Patients with any of the following conditions were excluded from the study, including autoimmune diseases, cancer, severe organ diseases, and those under treatment with immune suppressive agents for any reason. Patients with nasal cancers and healthy subjects were also enrolled into this study used as controls. Control subjects showed negative results of specific IgE and SPT. The demographic data of human subjects are presented in Table 1.
Demographic data of human subjects

Data are presented as mean ± SD or median (IQR).
*, p < 0.01, compared with the AR group.
#, AR patients used corticosteroid spray or/and prednisolone (5–15 mg/day) to control AR attacks (not regularly use).
DME-specific IgE (sIgE) > 35 IU/L was considered as positive.
HC: Healthy control.
Collection of nasal mucosal tissues
Patients with nasal polyps or cancers were undergone surgically procedures to remove the nasal polyps by our doctors following the routine procedures. The surgically removed nasal mucosal tissues were collected in the operating rooms. The nasal mucosal tissues, which were proven by pathologists, were selected for further experiments.
Preparation of single cells of nasal mucosal tissues
The collected tissues were cut into small pieces, incubated with collagen IV (0.5 mg/ml) and DNase I (200 ng/ml) for 30 min at 37°C with mild agitation. Single cells were filtered through a cell strainer (100 μm frist, then 70 μm). The cells were collected by centrifugation at 500 g for 5 min.
Purification of fibroblasts and naïve CD4+ T cells
Single nasal mucosal cells and peripheral blood mononuclear cells (PBMCs) were prepared, labeled with fluorescence labeled Abs. For purification of fibroblasts, cells were incubated with Abs of PDPN, CD45, and EpCAM (0.5 μg/ml each) for 30 min at 4°C. In a flow cytometer (BD Aria), EpCAM + CD45+ cells were gated out. From the remaining cells, PDPN+ cells were gated and sorted to be fibroblasts. For purifying naïve CD4+ T cells, PBMCs were processed using a human naïve CD4+ T cell isolation kit (Myltenyi) following the manufacturer instructions.
Cell culture
RPMI1640 medium was used to culture cells. The medium was supplemented with 10% fetal calf serum, 100 U/ml penicillin, 0.1 mg/ml streptomycin, and 2 mM L-glutamine. Cell viability was assessed using Trypan blue exclusion assay, which exceeded 99%.
RNA sequencing (RNAseq)
RNA samples were extracted from fibroblasts and sent to a biotech company (Shenzhen BGI, China) for processing by the staff members at the company. The data were also analyzed by the staff of the company. The differentially expressed genes (DEGs) were enriched, or which the top 50 DEGs were presented in a heatmap.
Flow cytometry
Cells’ surface markers were stained with fluorescence labeled Abs of interest (Ab types are detailed in figures, diluted to 1 μg/ml) or isotype IgG for 30 min at 4°C. After washing with phosphate-buffered saline (PBS, containing 2% bovine serum albumin, BSA) 3 times, cells were analyzed using a flow cytometer (BD FACSCanto II). To stain the intracellular molecules, brefeldin A (10 μg/ml) was added to the culture for the last 4 h. Cells were fixed with paraformaldehyde (1%, containing 0.05% triton X-100) for 1 h. After washing with PBS, cells were processed using the procedures of surface staining. The data were processed using flowjo software (TreeStar Inc., Ashland, OR) isotype IgG staining as gating references.
Collection of nasal secretions (NS)
A piece of degreasing cotton was gently placed in the middle nasal meatus. The cotton was taken out 5 min later. The NS in the cotton was squeezed out and stored at −80°C for further experiments.
Enzyme-linked immunosorbent assay (ELISA)
The quantity of cytokines and immunoglobulins in samples was determined by ELISA using specific reagent kits, following the protocol provided by manufacturers.
Real-time quantitative Rt-PCR (Rt-qPCR)
RNA samples were extracted from cells harvested from relevant experiments and converted to cDNA using a reverse transcription kit following the manufacturer's instructions. The cDNA samples were amplified using a Bio-Rad CFX96 qPCR device and a SYBR Green Master Mix kit. Primers used in the present study include GATA3 (ctcattaagcccaagcgaag, tttttcggtttctggtctgg), CD80 (agggaacatcaccatccaag, tgccagtagatgcgagtttg), CD83 (cggtctcctgggtcaagtta, ttgcagctggtagtgtttcg), CD86 (gtattttggcaggaccagga, attcctgtggctttttgtg), STAT6 (ctgccaaagacctgtccatt, ggtaggcatctggagctctg), IL2 (acctcaactcctgccacaat, gccttcttgggcatgtaaaa), IL4 (actgcttccccctctgttct, cagatggtgcctgtgtgtct), IL5 (gagaccttggcactgctttc, cagtacccccttgcacagtt), IL6 (tacccccaggagaagattcc, ttttctgccagtgcctcttt), IL13 (gtactgtgcagccctggaat, tttacaaactgggccacctc), Ki67 (cacgagacgcctggttacta, gggcttgcagagcatttatc) and ACTB (ggacttcgagcaagagatgg, agcactgtgttggcgtacag).
Western blotting
SDS-PAGE was used to fractionate protein samples from cells collected from relevant experiments, then transferred them onto a PVDF (polyvinylidene fluoride) membrane. After blocking with 5% skim milk solution for 30 min, the membrane was incubated with primary Abs of interest (diluted to 200 ng/ml) overnight, followed by incubating with HRP (horseradish peroxidase)-labeled second Abs (diluted to 20 ng/ml) for 2 h. Each incubation was followed by washing with TBST (Tris-buffered saline containing 0.05% Tween 20). Immunoblots on the membrane were developed using enhanced chemiluminescence and photographed using an imaging device (UVP, Cambridge, UK).
RNA interference (RNAi)
Commercial RNAi kits and reagents were used to knock down the expression of IL4RA or CD28 in CD4+ T cells according to the manufacturer's instructions. Western blotting was used to assess the results of RNAi in the cells two days after transfection.
Statistics
The difference between two groups was determined by Student's t-test. ANOVA followed by Dunnett's test or Bonferroni test was performed for multiple comparisons. Spearman correlation coefficient test was performed to test correlation between data from multiple groups. p < 0.05 was set as a significant criterion.
Results
Counts of fibroblasts, T cells, B cells, and dendritic cells in nasal mucosal tissues
Surgically removed nasal mucosal tissues were obtained from patients with nasal polyps with or without AR, and nasal cancer (no AR; used as a normal control (NC). Single cells were prepared from the tissues and analyzed by flow cytometry (FCM). Fibroblasts were detected at 16.55 ± 1.22%, 17.49 ± 1.06%, and 18.02 ± 1.12% in the groups of NC, nAR, and AR, respectively. T cells were 6.74 ± 0.77%, 6.97 ± 1.29%, and 8.62 ± 0.73% in NC, nAR, and AR groups. B cells were 4.30 ± 0.86%, 4.51 ± 0.94%, and 6.67 ± 0.77% in NC, nAR, and AR groups. Dendritic cells were 3.61 ± 0.48%, 4.45 ± 0.41%, and 5.21 ± 0.66% in NC, nAR, and AR groups (Fig. 1). The results indicate that besides immune cells, fibroblasts can be detected in the nasal mucosal tissues, which are more abundant than other immune cells.

Assessment of fibroblasts, T cell, B cell, and dendritic cells in nasal mucosal tissues. Single cells were isolated from surgically removed nasal tissues, and analyzed by FCM. A, dead cells were gated out. B, adherent cells were gated out. C, epithelial cells were gated out. D, gated FCM plots show indicated cell types. E, bars show mean ± SD of indicated cell types from 10 subjects per group. Each dot in bars presents one sample. Statistics: ANOVA + Bonferroni test. p values are presented in figures. Abbreviations: NC: Normal control. AR: Allergic rhinitis. nAR: Non-AR. FCM: Flow cytometry.
Fibroblasts from AR nasal mucosa express high levels of IL-4
Surgically removed nasal mucosal tissues were obtained from patients with nasal polyps with or without AR, and non-AR nasal cancer [used as normal control (NC)]. Fibroblasts were isolated from the nasal tissues by flow cytometry (FCM) cell sorting, and analyzed by RNA sequencing (RNAseq). In total of 10,089 genes were analyzed, of which 355 genes were up regulated, and 286 genes were down regulated. In the top 50 DEGs, upregulated DEGs included Gata3, Cd80, CD83, CD86, STAT6, IL2, IL4, IL5, IL6, and IL13 (Fig. 2A). These 10 genes were verified using conventional RT-qPCR (Fig. 2B). The findings show that the expression of IL4 and related molecules in nasal mucosal fibroblasts is elevated in patients with AR.

Nasal fibroblasts express IL4 and related molecules. Surgically removed nasal mucosal tissues were collected, from which fibroblasts were isolated. RNA samples were extracted from the fibroblasts, and analyzed by RNAseq and RT-qPCR. A, a heatmap show the top 50 DEGs. B, bar graphs show mean ± SD of the mRNA amounts of indicated molecules in fibroblasts. Each dot in bars presents one sample. Statistics: ANOVA + Bonferroni test. p values are presented in figures where appropriate. Abbreviations: NC: Normal control. nAR: Non-AR patients. AR: patients with allergic rhinitis. RNAseq: RAN sequencing. DEG: Differentially expressed gene.
IL4 expression in nasal fibroblasts of AR patients is associated with the AR-related factors in nasal secretions
Nasal secretions of each AR, nAR patient and healthy control (HC) subjects were collected, and analyzed by ELISA. We found that the amounts of EPX, tryptase, IL-4, IL-5, IL-13, specific IgE (sIgE), sIgG1 and sIgG2 in nasal secretions were higher in the AR group than that in the nAR group and HC group. The quantity of these parameters didn't significantly differ between the nAR group and the HC group (Fig. 3A). A positive correlation was detected in the data of AR-related parameters in nasal secretions and the mRNA amounts of IL14 and IL4 expression-related molecules (Fig. 3B). The data suggest that the nasal fibroblast-derived IL-4 may play a crucial role in the pathogenesis of AR.
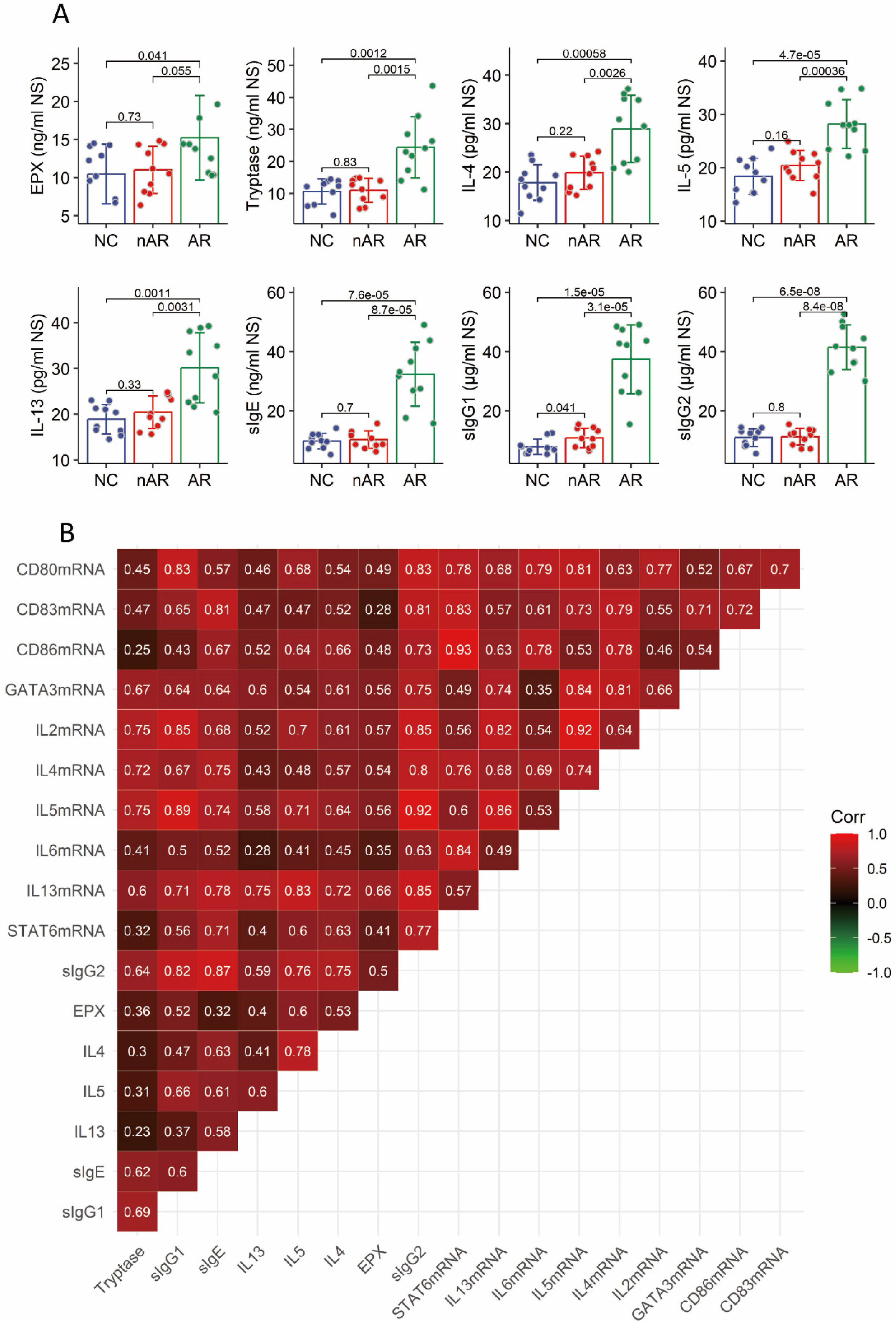
AR-related factors in NS are associated with nasal fibroblast-derived IL4 and related factor expression. A, NS was collected from each patient, and analyzed by ELISA. Bars show mean ± SD or the amounts of AR-related factors. Each dot in bars presents data obtained from one patient. Statistics: ANOVA + Bonferroni test. p values are presented in figures. B, a heatmap show correlation between the AR-related factors in NS and the mRNA amounts of IL4 and IL4-related factors in nasal fibroblasts. The numbers in each square show correlation coefficients between indicated factors. Abbreviations: NC: Normal control. AR: Allergic rhinitis. nAR: Non-AR patients. AR: AR patients.
Activation of nasal fibroblasts express costimulatory factors
Single cells were prepared from nasal mucosal tissues as described above and analyzed by FCM. The results showed that fibroblasts expressed costimulatory factors, including CD80, CD83, and CD86, which were significantly more in the AR group than that in the NC group and nAR group (Fig. 4A-B). To verify the results, fibroblasts were purified from the single cells. Protein samples were extracted from purified fibroblasts, and analyzed by Western blotting. We found that the quantity of CD80, CD83, and CD86 proteins were detected in the samples, which were substantially high in the AR group as compared with that of the NC group and the nAR group (Fig. 4C). The results demonstrate that nasal fibroblasts express the costimulatory factors.
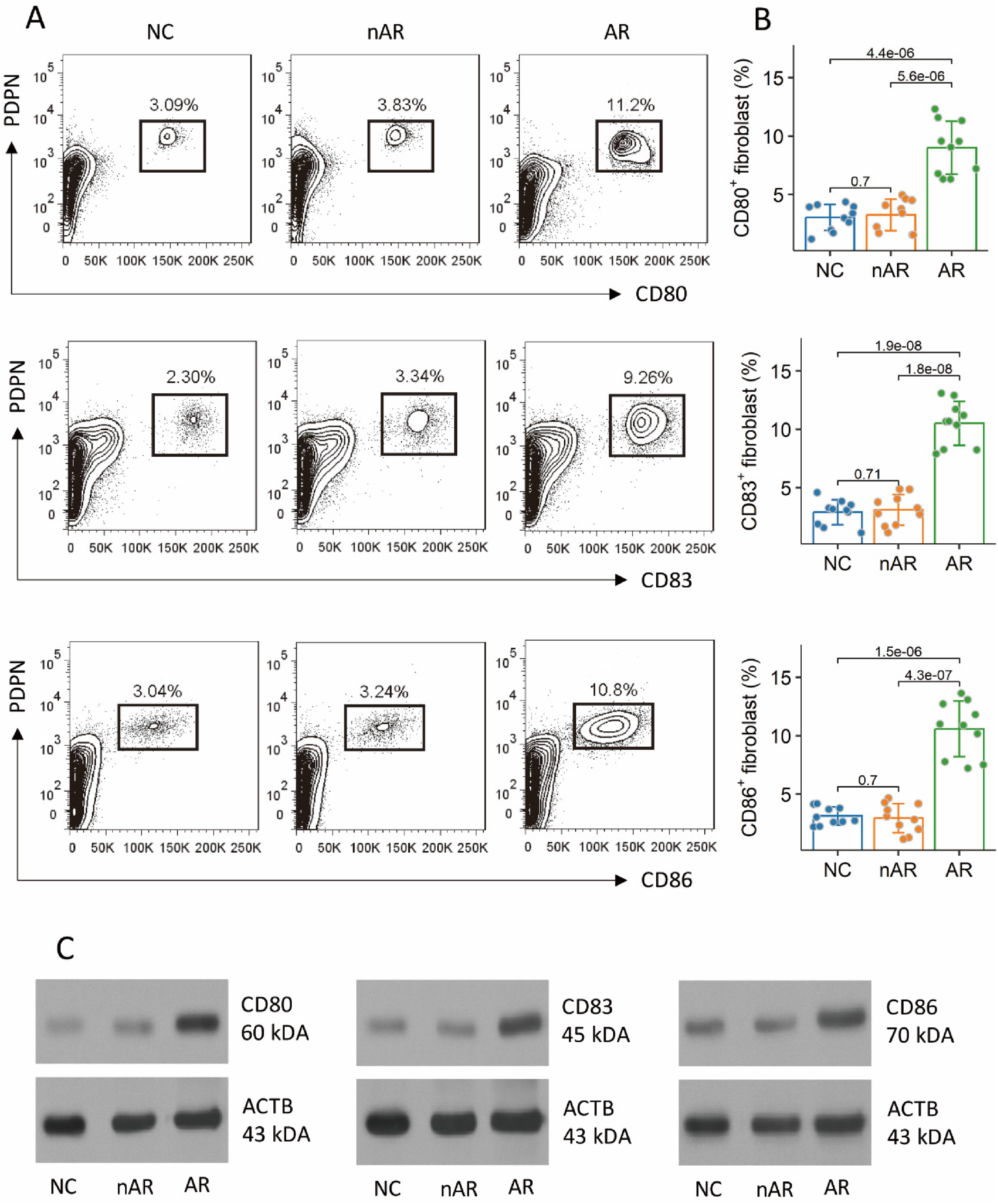
Nasal fibroblasts express costimulatory factors. Surgically removed nasal mucosal tissues were collected, from which fibroblasts were isolated. A, gated FCM plots are CD80+, or CD83+, or CD86+ fibroblasts. B, bars show mean ± SD of CD80+, or CD83+, or CD86+ fibroblasts of 10 samples per group. Each dot in bars presents one sample. Statistics: ANOVA + Dunnett's test. p values are presented in figures. C, protein samples were extracted from purified fibroblasts, and analyzed by Western blotting. Immunoblots show CD80, CD83, and CD86 proteins in fibroblast's extracts. The data of immunoblots are from one experiment that represent three independent experiments with pooled protein from 10 samples per group. Abbreviations: NC: Normal control. AR: Allergic rhinitis. nAR: Non-AR patients. AR: AR patients. FCM: Flow cytometry.
Nasal fibroblasts induce Th2 cell differentiation
Fibroblasts were isolated from surgically removed nasal mucosal tissues of patients with nasal polyps, whether with or without AR. The cells were cultured in the presence of PMA and ionomycin (non-specific cell activators) overnight. Fibroblasts were collected and analyzed by RT-qPCR. We found that exposure to PMA and ionomycin significantly increased the expression of Ki67 in fibroblasts (Fig. 5A). The results indicate that the PMA/ionomycin effectively activated the fibroblasts. Supernatant was collected and analyzed using ELISA. The results showed that the activation markedly increased the amounts of IL-4 in culture supernatant, which were significantly higher in the AR group than that in the nAR and NC group (Fig. 5B). The results demonstrate that nasal mucosal fibroblasts can release IL-4 and costimulatory factors, which are elevated in AR patients.
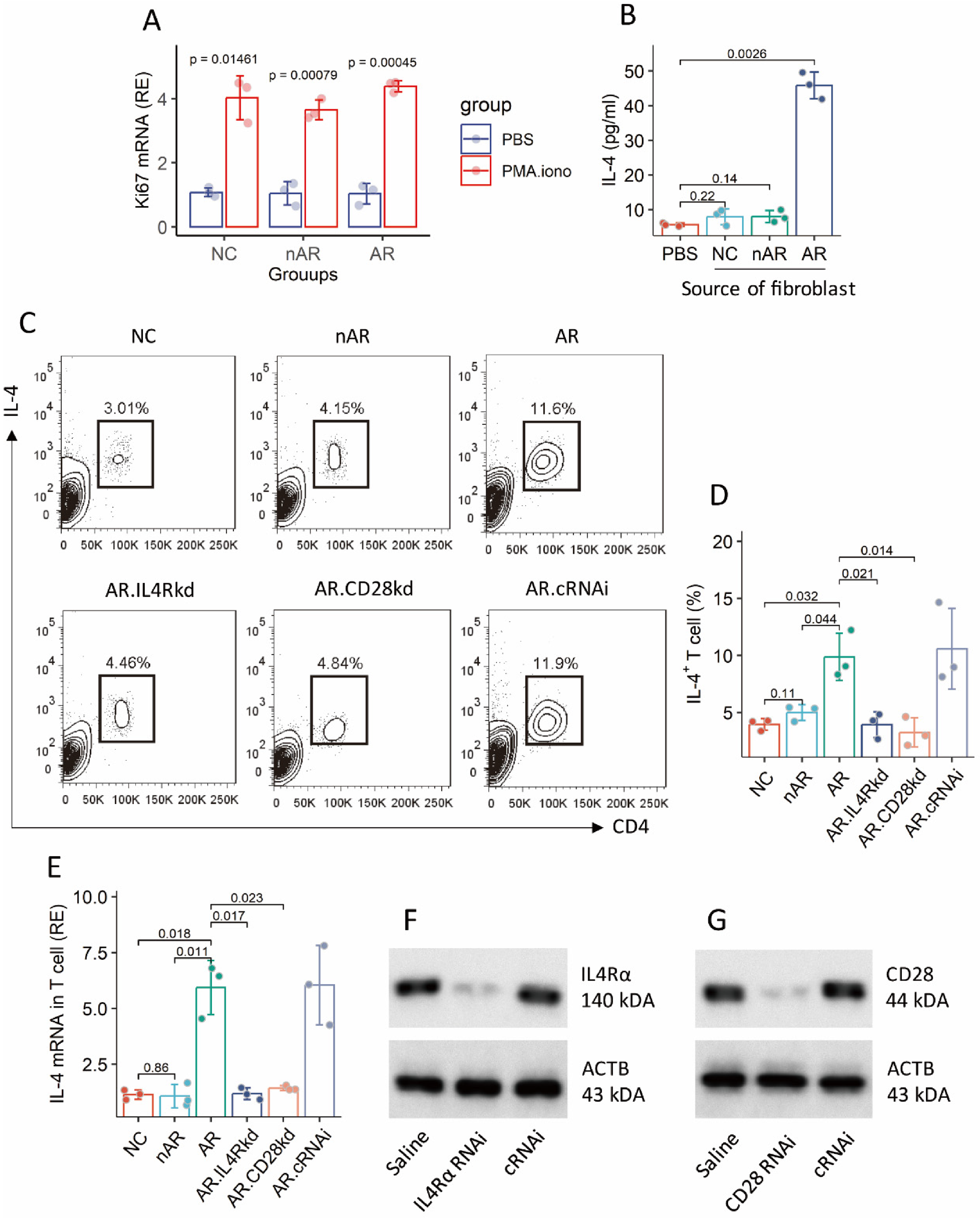
Nasal fibroblast-derived IL-4 induces Th2 cells. A-B, nasal fibroblasts were prepared, and exposed to PMA (50 ng/ml) and ionomycin (100 ng/ml) in culture overnight. A, mean ± SD of Ki67 mRNA quantity. B, mean ± SD of IL-4 quantity in culture supernatant. C, gated FCM plots show IL-4 + CD4+ T cells. D, mean ± SD of IL-4 + CD4+ T cell counts. E, CD4+ T cells were purified from the cell mixture at the end of experiments, and analyzed by RT-qPCR. Bars show mean ± SD of IL4 mRNA quantity in CD4+ T cells. F-G, RNAi results show IL-4Rα (E) and CD28 (F) protein levels in CD4+ T cells. Each dot in bars presents one sample. The data of immunoblots are from one experiment that represent three independent experiments. Abbreviations: NC: Normal control. AR: Allergic rhinitis. nAR: Non-AR patients. AR: AR patients. FCM: Flow cytometry. kd: Knockdown. cRNAi: Control RNA interference.
The data presented suggests that nasal fibroblasts may be responsible for Th2 cell differentiation. To test this, naive CD4+ T cells were isolated from PBMCs, and cultured with nasal fibroblasts in the presence of PMA and ionomycin. Three days later, the cells were analyzed by FCM. We found that NC and nAR fibroblasts generated 3.95 ± 0.49% and 4.99 ± 0.69% Th2 cells, while AR fibroblasts generated 9.87 ± 2.06% Th2 cells (Fig. 5C-D). The IL4 mRNA quantity in CD4+ T cells was markedly up regulated after the coculture with AR fibroblasts (Fig. 5E). The findings indicate that nasal fibroblasts are capable of inducing Th2 cells.
Nasal fibroblast-derived IL-4 and costimulatory factors are required for the induction of Th2 cells
To test the role of fibroblast-derived IL-4 and costimulatory factors in the induction of Th2 cells in this experimental setting, IL4RA or CD28 in CD4+ T cells was knocked down (kd) by RNA interference (RNAi) (Fig. 5F-G). The IL4RAkd or CD28kd CD4+ T cells were cocultured with AR nasal fibroblasts. As expected, the induction of Th2 cells and IL4 expression were abolished (Fig. 5C-E). The results indicate that nasal fibroblast-derived IL-4 and costimulatory factors are required in the fibroblast-inducing Th2 cells.
Discussion
The present study revealed that nasal fibroblasts produced IL-4 and the IL4 expression-related molecules, which were significantly elevated in patients with AR as compared with that of nAR patients and NC subjects. In an ex vivo experiment, we observed that nasal fibroblasts released IL-4 and costimulatory factors upon activation. Coculturing nasal fibroblasts and naive CD4+ T cells induced Th2 cells, in which the fibroblast-derived IL-4 and costimulatory factors played a crucial role.
Fibroblasts are structural cells in the nasal mucosa. The basic role of fibroblasts is to differentiate into fibrocytes, which produce fibrin to maintain the anatomical structure [15]. Current data support this concept by showing that fibroblasts are the major cellular components in the nasal mucosa. Fibroblasts also produce many other molecules. For example, fibroblasts produce CXCL12 (CXC-chemokine ligand 12) and express VCAP1 (vascular cell adhesion protein 1), which bind CXCR4 (CXC-chemokine receptor 4) and integrin α4β1, respectively [16]. Production of IL-1, IL-6, and colony stimulatory factor were also observed in fibroblasts [17]. Fibroblast-derived CCL19, CCL21 and CXCL12, CXCL13 and IL-7 involve in chemoattracting lymphocytes to the local tissues [18]. Our data show that IL4 and the IL4 expression-related molecules can be detected in the nasal mucosa. The findings suggest that nasal fibroblasts may be involved in the pathogenesis of immune inflammation in the nasal mucosa.
The data show elevated expression of IL4 and IL4 expression-related molecules in nasal fibroblasts of AR patients than controls. Costimulatory factors, including CD80, CD83, and CD86, were detected in fibroblasts. The expression of IL4 and costimulatory factors was higher in the AR group than that in the nAR or NC group. IL-4 and costimulatory factors are the key factors in the induction of Th2 cells [19]. Current data implicate that nasal fibroblasts are capable of inducing the development of Th2 cells. Indeed, further experimental data demonstrate that nasal fibroblasts are capable of inducing Th2 cells. AR nasal fibroblasts induced more Th2 cells than nAR or NC nasal fibroblasts. The underlying mechanism is that AR nasal fibroblasts produce significantly amounts of IL-4 and costimulatory molecules. This is in line with the established concept [20]. The data were verified by knocking down the IL4Rα or CD28 in naïve CD4+ T cells, in which the induction of Th2 cells by fibroblasts was abolished.
AR is featured by the Th2 polarization in the nasal mucosa [6]. Current data also show the elevated Th2 cytokine amounts in nasal secretions of AR patients, which were significantly lower in nAR patients and NC controls. Th2 cells are the sources of Th2 cytokines [21]. Bone marrow-derived T cells are the progenitors of Th2 cells. While the factors initiating Th2 polarization in the nasal mucosa have not yet been fully elucidated. Current data reveal that the nasal fibroblasts can be one of the factors inducing Th2 polarization. Supporting evidence of this includes nasal fibroblasts that express IL-4 and costimulatory factors, which are significantly higher in the AR group than in the control groups. AR fibroblasts generate a greater amount of Th2 cells than fibroblasts from control groups.
Taken together, current data indicate that nasal fibroblasts express IL-4 and costimulatory factors, which are significantly higher in the AR group compared to the control group. Th2 cells can be induced by coculturing naïve CD4+ T cells and nasal fibroblasts. The data implicate that nasal fibroblasts are involved in the pathogenesis of nasal allergy.
Limitations of this study: Nasal mucosal tissues of patients with nasal cancer were used as a non-AR and non-nasal polyp control group in the present study. Although these cancer patients did not have nasal allergy, it is for sure these samples are still different from normal nasal mucosal tissues. We need consider this point in the explanation of the results.
Footnotes
Author contributions
Zeng X, Li J, Liu J, Mo L, Liu Y, Zhang A performed the experiments, analyzed data, and reviewed the manuscript. Yang P and Kong H designed the project, organized the experiments, and edited the manuscript. Yang P prepared the manuscript.
Acknowledgements
This study was supported by research grants of the National Natural Science Foundation of China (32090052, 82171120), Shenzhen Key Medical Discipline Construction Fund (No.SZXK062), and Shenzhen science, technology, and innovation committee (KQTD20170331145453160, SGDX20201103095609027).
Data availability
All the data are presented in this paper and the online supplemental materials, or request to the corresponding authors.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
