Abstract
Species differences in the structure and function of the immune system of laboratory animals are known to exist and have been reviewed extensively. However, the number and diversity of wild and exotic species, along with their associated viruses, that come into contact with humans has increased worldwide sometimes with lethal consequences. Far less is known about the immunobiology of these exotic and wild species. Data suggest that species differences of the mechanisms of inflammation, innate immunity and adaptive immunity are all involved in the establishment and maintenance of viral infections across reservoir hosts. The current review attempts to collect relevant data concerning the basics of innate and adaptive immune functions of exotic and wild species followed by identification of those differences that may play a role in the maintenance of viral infections in reservoir hosts.
Introduction
Species differences in the structure and function of the immune system of laboratory animals have been previously reviewed.1,2 The purpose of that review was to identify and characterize species differences of the immune system with the goal of aiding in the extrapolation of animal data to humans. Correlating a specific functional difference with a specific structural difference was recognized to be beyond the scope of that review. It was concluded that species differences do exist, but determining which differences are biologically and functionally meaningful requires additional data and effort. Nevertheless, an important purpose of the earlier and the present review is to identify, describe and categorize structural and functional differences in order to assist researchers in knowing what to expect in a normal individual of a given species, and place such differences in the right perspective when attempting to assign them relevance.
Species differences have not occurred serendipitously, but rather have resulted from multiple evolutionary pressures across species within highly variable and changing environments. Such evolutionary pressures have culminated in different solutions for the same problems across species, such as the generation and maintenance of immunological protection from bacteria, viruses and parasites.
More specifically, this manuscript attempts to collectively review and begin to build a compendium of the literature concerning the immune structure and function of those wild and/or exotic species that are currently having a significant worldwide impact as reservoir hosts for a range of dangerous viruses, such as rabies, MERS-CoV, H1N1 influenza, SARS-CoV1, SARS-COV2, EBOLA, Hendra, Nipah, and Marburg viruses. As an example, in 1999, Nipah virus spilled-over from fruit bats (Pteropus sp.) in Malaysia, leading to 105 human deaths and the necessary slaughter of one million pigs. 3 Of greater pertinence has been the postulated spillover of SARS-CoV2 from Chinese horseshoe bats, and possibly other zoonotic hosts, into humans in China with subsequent worldwide spread and numerous human deaths beginning in 2019. 4 The exact source of SARS-COV2 is still hotly debated.
The number and diversity of wild and exotic species, along with their associated viruses, that now come into contact with humans has increased worldwide. A key factor for successfully confronting these devastating viral outbreaks is the recognition that the innate and adaptive immune systems of reservoir species sometimes function differently, as compared to other species, and thereby may help the reservoir host to tolerate otherwise lethal microbes without pathologic outcomes.5–10 However, it is unclear which immune mechanisms different species use to tolerate viral infections without overt disease, and if they are the same across reservoir species. Data suggest that species differences of the mechanisms of inflammation, innate immunity and adaptive immunity are all involved in the establishment and maintenance of viral infections across reservoir hosts.
To aid in this review the following terms will be used. A
To understand such pathogen and host relationships, a number of questions need to be considered. Do reservoir hosts have unique, potent and efficient immunologic mechanisms that suppress but not eliminate viruses, thus maintaining active viral infections without overt disease? Why do viruses that have been “controlled” by the reservoir host‘s immune system, suddenly spill-over and spread to new and vulnerable species? Are the mechanisms for achieving and maintaining a reservoir state the same for each of the different reservoir species? And lastly, can an understanding of such immunological differences and mechanisms shed light on how to therapeutically manage potentially lethal virus infections in humans? Unfortunately, given the state-of-art regarding the immunology of these species, some of these questions cannot be answered.
The animal species reported to play a role as primary, reservoir or intermediate hosts include bats, pangolins, palm civets, raccoon dogs, swine, and camelids.1–13 Because most, but not all, of these species are wild, a complete characterization of their immune system is not possible due to the lack of availability of published data. This is especially true for palm civets and raccoon dogs, therefore, the immunology and anatomy for these two species will not be addressed, beyond the recognition of their close association with domestic animals, for which data are abundant.
The dromedary camel (Camelus dromidarius) has been considered a natural reservoir and/or intermediate host for MERS-CoV, based on the seroprevalence rate for MERS-CoV in dromedaries (70% to nearly 100%) and that approximately 55% of primary MERS-CoV human cases are a result of direct contact with dromedary camels or camel products. 7 However, based on the high sequence similarity of MERS-COV to coronavirus sequences in bats, it is possible that MERS-CoV likely originated in bats. 14 For this reason, camelids will not be discussed in this review.
Luckily, not all reservoir hosts are wild. For one fully domesticated species, swine, much data are available, and have been reviewed in detail elsewhere.1,15,16 Despite being a domesticated species, swine are relevant to this discussion because of their pivotal role in the origination and transmission of influenza viruses, and because they are the only livestock species presently know to be naturally infected with ebolaviruses, although they are not likely the original source.13,17,18 Information from humans and mice will be used as the benchmark for comparisons when appropriate.
Importantly, there are significant differences in the quality of data available from wild and exotic species, and such data are more variable compared to that generated under laboratory conditions. For example, bat behavioral and environmental needs make it extremely difficult to maintain controlled bat colonies, although a few have been successful in doing so, while others have used pre-existing bat populations in large colonies in the wild or worked with individually caught wild bats.19–24 As a result, significant bat-to-bat variations make data interpretation of even the simplest of observations highly challenging. Also, research tools such as species-specific cell lines, and reagents (i.e. antibodies) are very limited. Some monoclonal antibodies have been found to be satisfactorily cross reactive in bats. 25
Part 1. The Innate Immune System
The innate, natural or nonspecific immune system acts as a first line of defense against viral infections. Components of the innate immune response include physical (skin, mucus, gut flora, etc), molecular (cytokines, chemokines) and nonspecific cellular defenses (macrophages, neutrophils). An excellent review by Majzoub et al., 2020 is available. 26
Innate immunity does not require prior exposure and/or immunological memory to attack the invading organism. Upon microbe penetration of the host, nonspecific cascades of chemokines, cytokines and messenger molecules are triggered to neutralize and eliminate the microbe from the host. Of the molecular components involved, type I interferons (IFN), highly conserved regulators of cellular functions, are the most central to the initiation of the innate immune response. A wide variety of cells, including epithelia, endothelia, stroma, and immune system cells glycosolate IFNs in response to viral infections, bacteria, parasites, inflammation, and cancer. 27 This subsequent expression of IFN induces a diverse range of biological functions and responses, including cell proliferation and differentiation, inflammation, chemotaxis, immune cell (natural killer cells and macrophages) activation, and apoptosis.
In the case of virus infection, viral components, such as viral RNA, DNA, or intermediate products, otherwise designated as pathogen-associated molecular patterns (PAMP), interact with pattern recognition receptors (PRRs) on host cells which then induce type I IFNs followed by other pro-inflammatory cytokines in the infected and immune cells. Type I IFNs bind to the interferon alpha cell-surface receptor and initiate signal transduction leading to the activation and transcription of the JAK–STAT pathway within cells. Subsequently, hundreds of IFN-stimulated genes (ISGs) are induced and drive the cell-autonomous antiviral defense that inhibits virus replication.26–30 Mammalian IFNs are classically grouped into three major types (I, II, and III). These types are comprised of several IFN families allowing for numerous regulatory actions, that include antiviral, anti-inflammatory, and immunomodulatory activity.30–32
While IFNs play a critical role in innate immune defenses, they also appear to factor in the maintenance of persistent viral infections in some species. Species differences of IFN have been identified including the number genes and type of IFNs.
Human type I IFNs, including 13 IFN-α subtypes, are encoded on chromosome 9, as are single members each of IFN-β, IFN-ω, IFN-κ, and IFN-ε.30,33 Mice have about 14 functional IFN genes, located on chromosome 4, except for IFN-ω which is absent; see Table 1. 27 Some bats and pangolins, on the other hand, have only three IFN-α genes, the fewest number of IFN genes of any mammalian species sequenced so far to date.8,34,35 However other bat species have >20 IFN-ω genes that factor significantly in their innate immune responses. In comparison, there are 60 porcine type I IFN functional genes, the highest number in one species (paralleled only by domestic cows), divided into eight subtypes, plus about 20 pseudogenes.36–41 Pseudogenization is a process in which a gene acquires a spontaneous mutation that prevents transcription or meaningful translation. This may be due to a frame shift or introduction of a premature stop gene codons. DNA sequences that are pseudogenes are not translated, but may still be transcribed.
Number of IFN genes identified in humans, mice and four species of bats .
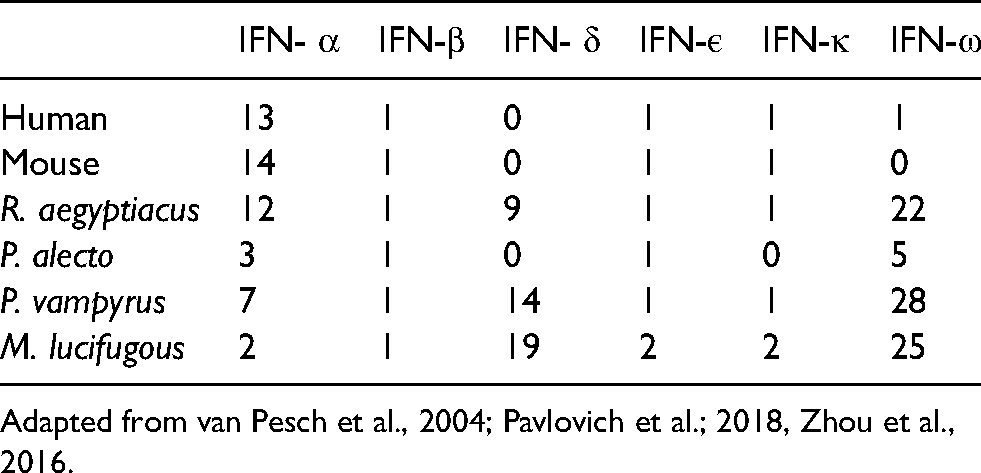

Adapted from van Pesch et al., 2004; Pavlovich et al.; 2018, Zhou et al., 2016.
Bats
Given that there are >1300 species of bats spread across two suborders, studies of multiple bat species will be required to accurately delineate chiropteran innate and adaptive immune responses. For this review, data will be sampled from reports on 22 species, nine that are considered megabats, and 13 that are considered microbats; see Table 2.
List of common and scientific name of bat species used in this review.

The very high number of bat species across two orders precludes making generalizations across all bat species. Despite the study of immunology of bats still being at a very early stage, efforts in recent years to better understand bat immune responses, the genetic profile of bats, and bat behavior have markedly improved. Genome and transcriptome analyses including full genome sequencing, RNA-Seq and real time PCR have begun to fill in the knowledge gaps. Examples of ongoing multiple worldwide efforts concerning bats include, but are not limited to, the Bat1 K Project, the Wild Comparative Immunology Consortium, the efforts of the Wuhan Institute of Virology, and many other individual laboratories.8–10,20,42,43
Bats are the predominant reservoir host of many of the most serious viral infections of man (rabies, H1N1 influenza, SARS-CoV1, SARS-CoV2, EBOLA, Hendra, Nipah, and Marburg viruses).8–10 These viruses usually do not cause discernible disease or pathological consequence in bats but often cause lethal disease in man.9,10,19,43 Eighty-four percent of some species of horseshoe bats have antibody to bat SARS-CoV but lack discernible pathology. 43 This dichotomy makes the bat an excellent animal model to investigate the role of antiviral innate and adaptive immunity in a reservoir species, and to possibly provide insight into potential antiviral therapeutic approaches.
Bat-specific qualitative and quantitative differences in their innate immune system have resulted in identification of several functional characteristics in P. alecto not found in other mammals.44,45 Zhang et al., 2017 identified novel and bat-specific
Gene regions associated with type I IFN, MHC-1 and MHC-II in bats are condensed compared with the corresponding region from other mammals, with a few exceptions. As an example, while the type I IFN locus in P. alecto is contracted, it is expanded in P. vampyrus, M. lucifugus, and R. aegypticus.8,9 In human cells, IFN signaling functions through the IFN α/β receptors, IFNAR1 and IFNAR2, but IFN signaling in P. alecto cells does so via IFNAR2 only. 45
One aspect of the innate immune responses of bats that has received considerable attention is the constitutive expression of IFN and ISGs. IFN-α genes, along with associated ISGs, such as IFN regulatory factor 7 (IRF7), have been identified as being constitutively expressed in unstimulated tissues and cells from P. alecto, P. vampyrus, M. lucifugus, C. brachyotis and A. jamaicensis.8,32,46
In theory, the constitutive expression of IFN limits inflammation-induced damage to tissues by suppression, but not elimination of viral infection.8,47,48 Another interpretation for constitutive expression of IFN is that a continual high level expression of IFN in bats allows for a more rapid and robust activation of IFN by keeping the system “revved-up” or “always on”.8,48–50 Constitutive expression of IFN would thus allow bats to more rapidly respond to microbial infection and provide a continuous front-line defense against infection.8,46
However, constitutive expression of IFN genes is not unique to bats. Constitutive expression of IFNs has not been reported for most of the other reservoir species discussed herein, with one exception: swine.32,36,51 In addition, constitutive expression of IFN, albeit a low level, has been identified in humans and mice.52–57 It has been suggested that even the weak constitutive expression of IFN-α/β in mice connotes a priming of cells that then contributes to initiation of strong responses to IFN-γ and IL6, and the amplification of type 1 IFN induction followed by an efficient and more robust production of IFN-α/β in response to virus infection.30,50,58–60
Not all bats have constitutive expression of IFN-α as it is found lacking in R. aegyptiacus, P. vampyrus, and M. lucifugus.8,9,61
This lack of constitutive expression of IFN-α in primary cells from these bat species shows that there are also species differences within the order of Chiroptera and that high baseline levels of IFN-α may not be a feature of all bats. Moreover, because all of these bat species, regardless of constitutive expression of IFN, are reservoirs for different viruses of high virulence to humans, such intra-order species differences suggests that there may be additional mechanisms capable of maintaining or suppressing virus-induced pathology within a reservoir host, or that constitutive expression of IFN plays only a limited role in the persistence of viral infections.
Other molecular mechanisms that might contribute to enhanced tolerance to virus infection in bats, most of which reduce viral-induced inflammatory damage to the host‘s tissues include the following:
Zhou et al., 2014 showed that bat IRF7 is inducible by double-strand RNA and has a unique and broad tissue expression pattern across both immune and non-immune related tissues that may allow more rapid activation of the IFN response in a wider range of tissues of bats compared to other mammals.
46
Constitutively expressed ISG transcripts in type I IFN-α treated P. alecto cells increase rapidly to higher levels compared to humans followed by a subsequent rapid decline.34,45,62 These kinetics may correspond to a quicker and more aggressive control of virus replication and reduced secondary cellular damage. Elevated ISG transcript levels in IFN-treated human cells however, while of lower magnitude, remain elevated for longer times.62,63 This lesser inflammatory profile of ISGs may be key to the ability of bats to tolerate higher IFN expression without adverse consequences. Bat ISGs associated with U-ISGF3 are enriched and lead to extended resistance to virus infection and prevent DNA damage. ISGs that are well-characterized intracellular antiviral factors include BST-2 (tetherin), which has been reported to restrict replication of HIV, ebolaviruses, and Marburg viruses, and Mx1. Therefore, ISGs may provide an additional defense mechanism that is “switched-on” to blunt virus replication in bats.
8
Mammals possess ISGs that are common to all species, as well as others that are species or lineage specific.
32
Interestingly, the basal transcription level of type I IFNs and antiviral ISGs are higher, and expansion of ISGs is greater, in both microbats (M. lucifugus) and macrobats (P. vampyrus) compared to other mammals.
32
This observation is in-line with the observation of constitutive activity of IFNs and ISGs in bats that may confer a selective advantage to the host. Bat ISG profiles are characterized by a subset not associated with the acute inflammatory responses in other species.8,62 These include a bat-specific novel subset of IFN inducible ISGs, ribonuclease L (RNaseL) that is different from the RNaseL found in humans.
63
Members of PYHIN family, which are DNA sensors that recognize foreign and damaged self-DNA generated by RNA viral infection, are absent in bats and may be another mechanism by which bats limit activation of the innate immune response and lessen tissue damage.
64
Subudhi et al., 2019 identified an inhibitor molecule (cRel) binding site in the promoter region of tumor necrosis factor alpha (TNFα) in E. fuscus, P. alecto and M. davidii. Activation of cRel binding site actively suppresses TNFα expression.
49
PolyI:C-stimulated cell lines are associated with induction of high levels of pro-inflammatory cytokines, TNFα and IL8 in humans, but not in E. fuscus, suggesting that some bats may regulate this aspect of their immune response differently compared to other species.
65
Differences in interferon regulatory factor expression have also been noted for bats based on the finding that bat IRF3 sequences are evolutionarily distinct from their mammalian counterparts.23,66 Stimulator of interferon genes (STING) sense damaged DNA or dsDNA from viruses and then induce an interferon response. Several bat species have a mutation at a highly conserved serine residue (replacement of serine at position 358 by other amino acids) in STING that reduces its functionality and contributes to the lessening of inflammation.
11
Primary immune cells of some bats have a significantly dampened activation of the NOD-LRR-and pyrin domain-containing protein 3 (NLRP3) inflammasome, compared to human or mouse cells.
10
NLRP3 senses and recognizes cellular stress (such as extracellular ATP, mitochondrial damage and oxidized DNA) and viral and bacterial infections. The reduction of NLRP3 would contribute to reducing tissue inflammation. The unique organization, structure, and increased signaling complexity of the NK cell receptors in multiple bat species point to adaptations that are consistent with an infection tolerance hypothesis. NK cell receptors in bats are uniquely regulated.
9
KLRC (NKG2) and KLRD (CD94) gene families of natural killer cell receptors are expanded and diversified in R. aegypticus resulting in high baseline expression of inhibitory KLRC genes that thereby contribute to inhibition of both NK and T cells. Bats have reduced apoptosis-associated speck-like protein containing CARD (ASC) and interleukin-1B (IL-1B). The induction of ASC normally recruits and activates caspase-1 which promotes inflammatory cell death via apoptosis and cleavage/secretion of IL-1B.
10
The reduction of ASC would reduce apoptosis and secretion of IL-1B. IFN-α3, is the most abundant IFN-α found in P. alecto tissues and it induces robust responses of genes associated with anti-virus and antigen presentation (such as Mx1/2, OAS1-3, HLA-DRB1, CIITA).
45
the IFN-ω subfamily, which has significant antiviral activity, is the greatest contributor to the marked expansion and diversification type I IFN locus in R. aegyptiacus. High expression levels of the IFN-ω subfamily are not observed in other mammals, with the exception of swine.47,51 IFN-ω is absent in mice.
The combination of the differences of the innate immune response in bats noted above likely contributes to the ability of bats to tolerate constitutive IFN-α expression without pathology. Thus, RNA virus infections of bats induce a strong IFN response while controlling an exaggerated pro-inflammatory response, thus limiting virus-induced immunopathology. In addition, bats appear to rapidly control viral replication, resulting in short periods of virus replication and shedding, and low antibody titers. Type I IFN responses to viral infections in bats are otherwise similar to that of other mammals.
25
Pangolins
After the COVID-19 outbreak in Wuhan, China, many efforts to identify and characterize potential reservoir hosts were initiated and are still ongoing. Among these efforts is the creation of the Pangolin Gene Database, as well as the work of Zang et al., 2020 and others.6,67 Pangolins, like bats, have been identified as a possible natural reservoir and source of SARS-CoV2.6,68 It has been determined that pangolin-CoV is ∼90% identical to SARS-CoV-2 receptor binding domain and is its second closest relative. Moreover, pangolin ACE2 has been shown to have better affinity to SARS-CoV2 than human ACE2. 68
While the role of pangolins in the emergence of SARS-CoV2 remains inconclusive, Malaysian pangolins have been shown to host two sub-lineages of SARS-CoV2-related coronaviruses: pangolin-CoV-GDC which is 90% genetically similar to SARS-CoV2, and pangolin-CoV-GXC which is 85% similar to SARS-CoV2. Based on these observations, the WHO concluded that the pangolin should be considered as a potential natural or intermediate host for novel coronaviruses. 4
Intact functional gene copies of IFN families IFN-α, IFN- δ, IFN-L, IFN-κ, IFN1H1 and IFN-β, are found in Malayan (Manis javanica) and Chinese (Manis pentadactyla) pangolins. 69 As previously mentioned, while humans have 13 functional IFN-α genes, pangolins have only 10, of which only three were found in Malayan pangolin and only two in Chinese pangolin. 69 This suggests that some species of pangolins, like bats, have a marked contraction of the IFN families.
Interferon epsilon (IFN-ε), which is important for skin and mucosal immunity in most mammals, was pseudogenized (presumably loss-of-function) in epithelial cells in all African and Asian pangolin species examined. 69 This reduction of IFN-mediated innate immunity contributes to the contraction of the interferon gene family in pangolins and suggests that innate immunity may be compromised in the skin and mucosa-protected organs. 69
IFIH1, otherwise called melanoma differentiation-associated protein 5 (MDA5), binds to cytoplasmic double-stranded RNA of coronaviruses and other viruses, and signals through mitochondrial antiviral-signaling protein (MAVS). This signaling subsequently activates expression of interferons and induces inflammation. 70 IFIH1/MDA5 has been inactivated by numerous disruptive mutations in pangolins (GenBank gene ID: 108398082), and IFIH1 is inactivated in the Malayan pangolin by more than 10 frameshift and in-frame stop mutations. 70
Like IFN-ε, Z-DNA-binding protein (ZBP1), which senses both Z-DNA and Z-RNA, is a pseudogene. ZBP1 is inactivated in pangolins by multiple in-frame stop codons in all ZBP1 segments. Taken together, these data indicate that pangolins have evolutionarily lost functional IFN-ε, IFIH1 and ZBP1 resulting in the down-regulation of innate defense against viruses
It is clear that there are many gaps in the understanding of pangolin immunology and further delineation and understanding of it represents another opportunity to investigate the relationship of SARS-CoV2 and host innate and adaptive immunity. In addition, determining how the pangolin fulfills the role of a reservoir host will be highly valuable to unraveling the ecology of SARs-CoV2.
Swine
Swine, while being an important animal reservoir for influenza A viruses (influenza viruses are principally hosted by waterfowl), also act as ‘mixing vessels’ for a myriad of other viruses, including ebolaviruses and coronaviruses
As noted earlier, swine have the largest type I IFN family (paralleled only by domestic cows) consisting of nearly 60 IFN-coding genes and eight subtypes that include multigene subtypes of IFN-α, -δ, and –ω. There are fourteen functional IFN-α genes and two IFN-α pseudogenes. Swine IFN genes are coded on chromosomes 1 and 10, and include 17 IFN-α, 11 IFN-δ, 7 IFN-ω genes and single genes for IFN-αω, IFN-β, IFN-ε, and IFN-κ.36,40,41 Swine IFNs include unconventional subtypes that are functionally novel and act at least partly through non-canonical IFN signaling in immune regulation. 51 After cloning 16 of 17 IFN-α genes, Zanotti et al., 2015 reported antiviral activity of 4 IFN-α subtypes (IFN-α2, -α5, -α9, and -α10) in a porcine pseudorabies virus yield reduction assay, while Sang and others reported the antiviral activity of 12 IFN-α subtypes against porcine reproductive and respiratory syndrome virus and vesicular stomatitis virus.31,36,39,40 IFN-α3, -α7, -α13, -α4, and -α15 had little-to-no antiviral activity using the pseudorabies virus assay.
All swine IFN-α subtypes, except IFN-α2, -α7, -α9, and -α11 have been shown to have significant anti-inflammatory activities and down-regulate the expression of IL-8, TNF-α, IL-6, Toll-like receptor 4 (TLR4), bD1, and nuclear factor-kB (NF-kB) genes. 31 Moreover, healthy pigs have constitutive expression of IFN-α, and IFN-β in some tissues without any evidence of microbial infection or inflammation. 36 Epithelial and constitutive expression of unconventional IFNs, IFN-ε, -κ, -δ, and -ω, are present in normal intestine, lymph nodes, and lung. It is tempting to speculate that this constitutive expression of IFNs in pigs leads to suppression, but not elimination of viral infection, thereby limiting pathogen and inflammation-induced damage to tissues, as it is proposed to do in bats and pangolins.36,69
In addition, all swine IFN-α subtypes, except IFN-α11, -α15, and -α16, significantly upregulate class I and/or class II MHC, while IFN-α11, -α15, and -α16, significantly downregulate class I MHC.
As noted above, swine have an unconventional and constitutively expressed IFN-ω. IFN-ω has several unusual features that include (1) it is a signature multi-gene subtype identified in bats, cattle, horses and pigs, (2) its isoforms have superior antiviral potency compared to typical IFN-α, and (3) it shows highly cross-species antiviral (but little anti-proliferative) activity exerted in cells of humans and other mammalian species. 51 Additional potential novel molecular and functional properties are being investigated further. 71
Part 2. The Adaptive Immune Response
Given the considerable amount of literature available on mammalian adaptive immune responses in general, this review will forego repeating it and instead focus on observations relevant to species differences.
The mammalian immune system has evolved unique genetic mechanisms that enable it to generate an almost unlimited number of different light and heavy chains. During B cell development, the V and J gene segments (for the light chain) and the V, D, and J gene segments (for the heavy chain) are recombined before they are transcribed to form a functional VL- or VH-region coding sequence by a process of site-specific recombination, called
Alternatively, combinatorial diversity is generated by the random formation of many different VJL and VDJH combinations. Combinatorial diversity is increased by the ability of any VH region to pair with any VL region to bind antigen. Random pairing of 320 different VL with almost 11,000 different VH results in about 3.5 × 106 different possible antibody specificities.
The large number of inherited V, J, and D gene segments available for encoding antibody chains makes a substantial contribution on its own to antibody diversity, but the combinatorial joining of these segments greatly increases this contribution.
Two other mechanisms for achieving antibody diversification include junctional diversification during gene segment joining, and somatic hypermutation. Junctional diversification is the random loss and gain of nucleotides at joining sites that markedly increases the diversity of V-region coding sequences created by recombination. Somatic hypermutation, which occurs in the recombined VDJH and VJL genes, involves mature B cells that have migrated to the secondary lymphoid organs where they proliferate following antigen contact. B cells with mutations then produce membrane Ig with mutated antigen-binding sites. If these altered antibodies fail to bind antigen, the B cell dies. If the altered binding sites bind antigen better than the original, the B cells receive stronger signals to proliferate and mature into plasma cells. As the immune response progresses, the average affinity of the responding B cells and the antibody produced increases, a process called affinity maturation. Repeated cycles of somatic hypermutation, followed by antigen-driven proliferation of selected clones of memory B cells results in antibodies of increasingly higher affinity becoming abundant during an immune response, thus providing progressively better protection against the pathogen. 73
Bats
Bats can generate IgM, IgD, IgA, IgE as well as subclasses of IgG.74–76 Highly specific heavy chain variable (VH) segments are encoded in the genomes of bats, with P. alecto and M. lucifugus possibly having the greatest number of VH gene segments among mammals75,76 Somatic hypermutation, while being the primary tool for generating antibody diversity in most mammals, appears lacking in some bats based on cDNA sequences in M. lucifugus suggesting combinatorial diversity may play a bigger role in bats for the generation of antibody diversity.77,78
The adaptive immune response of bats has functional differences compared to other species, such as delayed isotype switching from IgM to IgG in E. fuscus, and delayed generation of antibodies to red blood cells in E. fuscus, P. alecto and P. giganteus.78–81 The delay in generation of a primary humoral responses and the failure to sustain them may be relevant to the development and maintenance of chronic low-grade and/or persistent viral loads in bats. The mechanism of these delayed responses may be of interest to scientists working to achieve secondary antibody responses in human given vaccines for COVID-19.
Nevertheless, in most cases, viruses trigger humoral immune responses in bats that follow the same progression as seen in other mammals beginning with the early appearance of IgM followed by IgG antibodies.81–83 As an example, naïve R. aegyptiacus inoculated with Marburg virus seroconvert with IgG antibodies peaking between 14–28 days post infection (dpi) but then rapidly decline within 1 month and become undetectable after 2 months. 84 Long-term protective immunity, as indicated by virus-specific secondary antibody, has been achieved in some but not all bats rechallenged with Marburg virus 17–24 months after the primary experimental infection. 84 It has been repeatedly shown that while wild-caught bats develop neutralizing antibodies to a variety of viruses, these antibodies do not always provide protection following re-exposure.34,84–86
Notably, secretory IgA is seen at surprisingly low levels in P. alecto, raising questions about mucosal immunity of some bats. 83 A lack of gut associated lymphoid tissue (GALT), to be discussed later, has been noted in at least two species of bats, P. pipistrellus and R. hilebrandtii, suggesting a correlate between the lack of GALT and the absence of IgA.87,88 However, the combination of both low IgA and an absence of GALT have not been made in a single species of bat.
Cellular mediated immune (CMI) responses have been measured in T. brasiliensis using skin sensitivity tests to phygtohemagglutin (PPD) and in D. rotundus and C. perspicillata using BNSA.89,90 T. brasiliensis exhibit peak leukocyte traffic consisting of lymphocytes and neutrophils, as well as eosinophils, basophils, and macrophages from 6 to 24 h after injection which is faster than that for most mammals that display a peak multicellular response from 24 to 48 h. 1 CMI responses are also seen in P. giganteus to the contact allergen 2–4 dinitrofluorobenzene in skin sensitivity tests with a peak response observed 48-h after the second application, as observed in other mammals. However, the peak CMI response as measured by mixed lymphocyte culture (MLC) assay was delayed in P. giganteus (7 days) compared to mice (4–5 days). 91
A significantly higher percentage of CD8 + T cells in secondary lymphoid organs (i.e. spleen) of some bat species supports the premise that cell mediated immune responses of bats are geared toward controlling intracellular pathogens such as viruses. 25
Swine
Adaptive immune responses of swine have been described in detail previously and only a short synopsis will be provided here.1,15,16 Research into porcine immunobiology is extensive and detailed.15,16 In most ways the structure and function of the immune system in pigs is very similar to that of other mammalian species, including man. Porcine genes share 87% sequence similarity with their human counterparts and as a result swine Igs have the highest similarity to those of human, among all nonprimate sequences available. Swine have 5 IgG subclasses that can be grouped into two clusters: one contains IgG1 and IgG3; the other contains IgG2a, IgG2b (which differ by only 3 amino acids), and IgG4. However, swine show a limited usage of variable heavy chain genes to generate antibody diversity. Five cDNA clones from swine kappa variable region immunoglobulin transcripts have been identified and sequence analysis shows this species use two families of VL genes (SwVK1 and SwVK2) homologous to human VK1 and VK2. The JK sequences in swine are homologous to human JK2 and JK4 gene families.
Part 3. Anatomy of the Immune System
The limited data available on the histomorphology of the immune system for reservoir species underscores the need to further our understanding of the “structure” of the immune system along with the “functional capability” of lymphoid organs. In humans, progressive lymphopenia has been described in SARS-CoV2 infection and atrophy of lymph nodes, spleen and Peyer‘s patches, and loss of germinal centers has been seen in deceased SARS patients.92–97 Splenic white pulp atrophy also occurs in humans with Ebola and Marburg disease and with H5N1 influenza.98–101 The recognition of the loss of germinal centers in peripheral lymphoid organs (lymph nodes and spleen) secondary to SARS and COVID19 infection in humans provides clues to understanding how viruses might curb lasting immunity. Similar observations linking morphological effects on lymphoid organs with virus infections have not been consistently identified in reservoir or intermediate hosts but doing so would add significantly to our understanding of the host-parasite relationship.
Lymph nodes - general
A major challenge with using lymph nodes as indicators of immune status across species is the recognition of the considerable differences in number, location, size and macroscopic and microscopic anatomy of lymph nodes across and within species.102,103 The accurate assessment of lymph node biology and pathophysiology is compounded by comparison of the simplistic lymph node diagrams, as they occur in medical texts and biomedical references, with the high variability of lymph nodes in nature. Moreover, the considerable naturally occurring variations encountered across species and within individuals is magnified following the use of non-standardized, inconsistent protocols for collection and sectioning. Without careful application of standardized protocols, the ability to accurately identify and make comparisons of increases (hyperplasia or hypertrophy) or decreases (atrophy, lymphocytosis, depletion) of lymphocytes, infiltration of abnormal cells, perturbations of blood flow, etc. becomes severely hampered. This is clearly a case of ‘one-size (shape) does not fit all‘. While a few lymph nodes may fit the description of a ‘kidney-bean shaped‘ or ‘spherical‘ node, more often one encounters irregular lobulation that breaks the node into uneven, lumpy structures. In addition, the tissue that is being drained by a regional lymph node significantly affects the nodes macroscopic and microscopic morphology.
Strict attention to collection and processing details of lymph nodes are needed to accurately characterize lymph node anatomy. Histomorpological variability can be exaggerated by collection and sectioning techniques. Because lymph nodes do not have a simple, consistent shape, (both across species or within an individual) serial sections of a node can reveal areas of markedly different proportions of cortex, paracortex and medulla (Figure 1).104–105 The challenges imposed by normal nodal variations combined with poor processing techniques have been addressed eloquently by two groups and it is highly recommended that all the work by Belisle and Sainte-Marie 1981, and Hoshi et al., 1987 be thoroughly reviewed in order to correctly identify, characterize and understand both normal and altered lymph node anatomy of any species.104–108
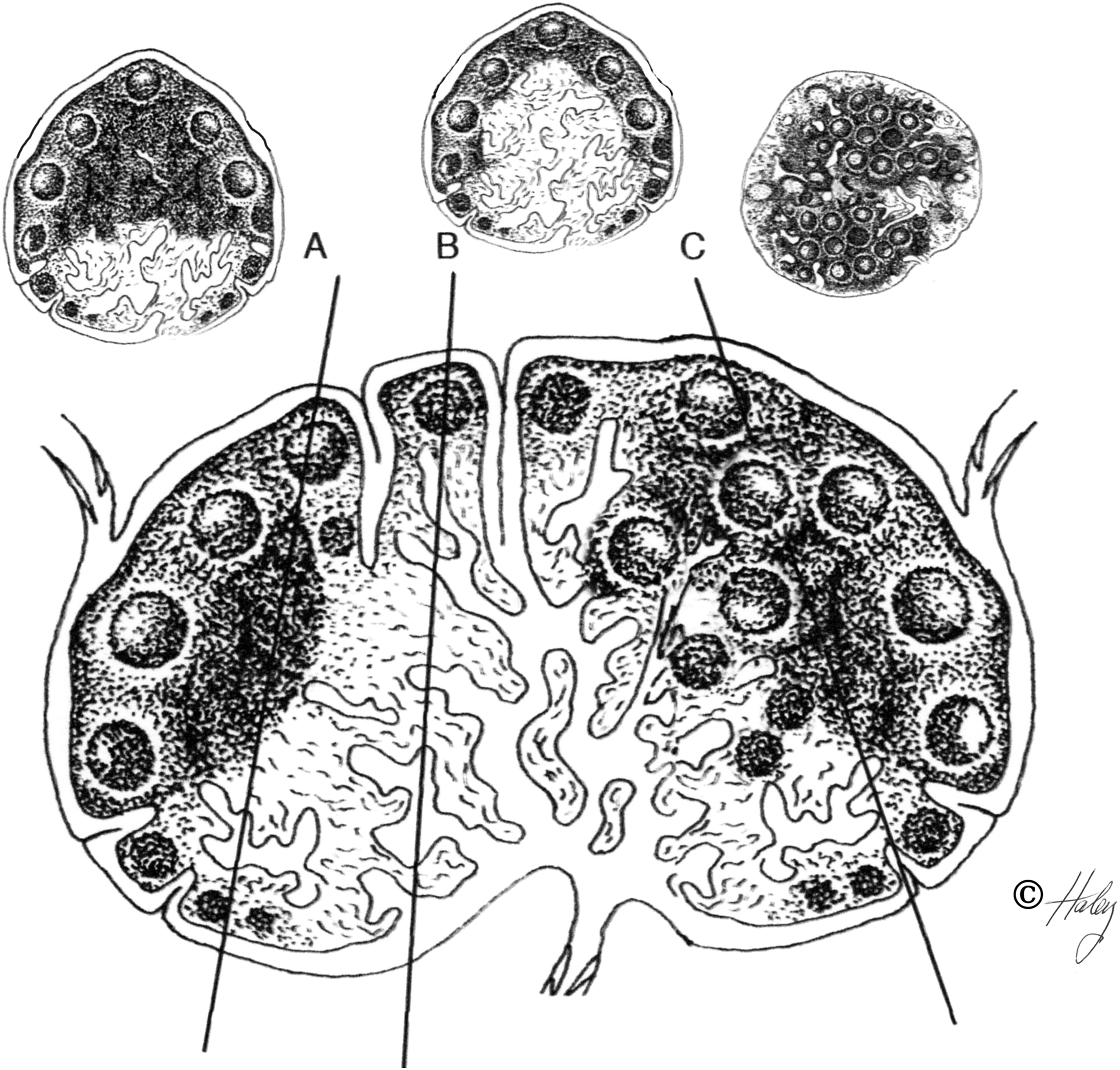
Diagram of a standardized lymph node demonstrating how histologic sections taken from different angles and locations from the node can give remarkably variable presentations of nodal components and hence lead to different interpretations.
It is the experience of this author that the histology of lymph nodes from a single animal can vary significantly depending of their location and the tissues drained, especially peripheral nodes, compared to deep nodes such as mediastinal nodes. 1 Peripheral lymph nodes draining the feet or oral cavity tend to have numerous lymphoid follicles with large germinal centers that may obscure the anatomy of the cortex and paracortex, while mesenteric nodes tend to have wider and thicker medullary cords filled with macrophages and fewer lymphocytes. Eosinophils may be abundant in peripheral lymph nodes that drain the skin because of ectoparasites, i.e. mange mites, whereas mediastinal nodes of the same animal will be devoid of eosinophils.
Bats
The anatomical distribution of lymph nodes in three different common bat species: R. aegyptiacus, T. perforatus and the R. microphyllum have been reported. 109 The lymph node number, shape and size were remarkably variable across all individuals of the three species of bats examined, especially of R. aegyptiacus and T. perforatus. These species demonstrated significant variation in the number and location of the facial, posterior cervical, brachial, inguinal and popliteal lymph groups (Table 1).
Anatomical location and number of lymph nodes in three species of bats.
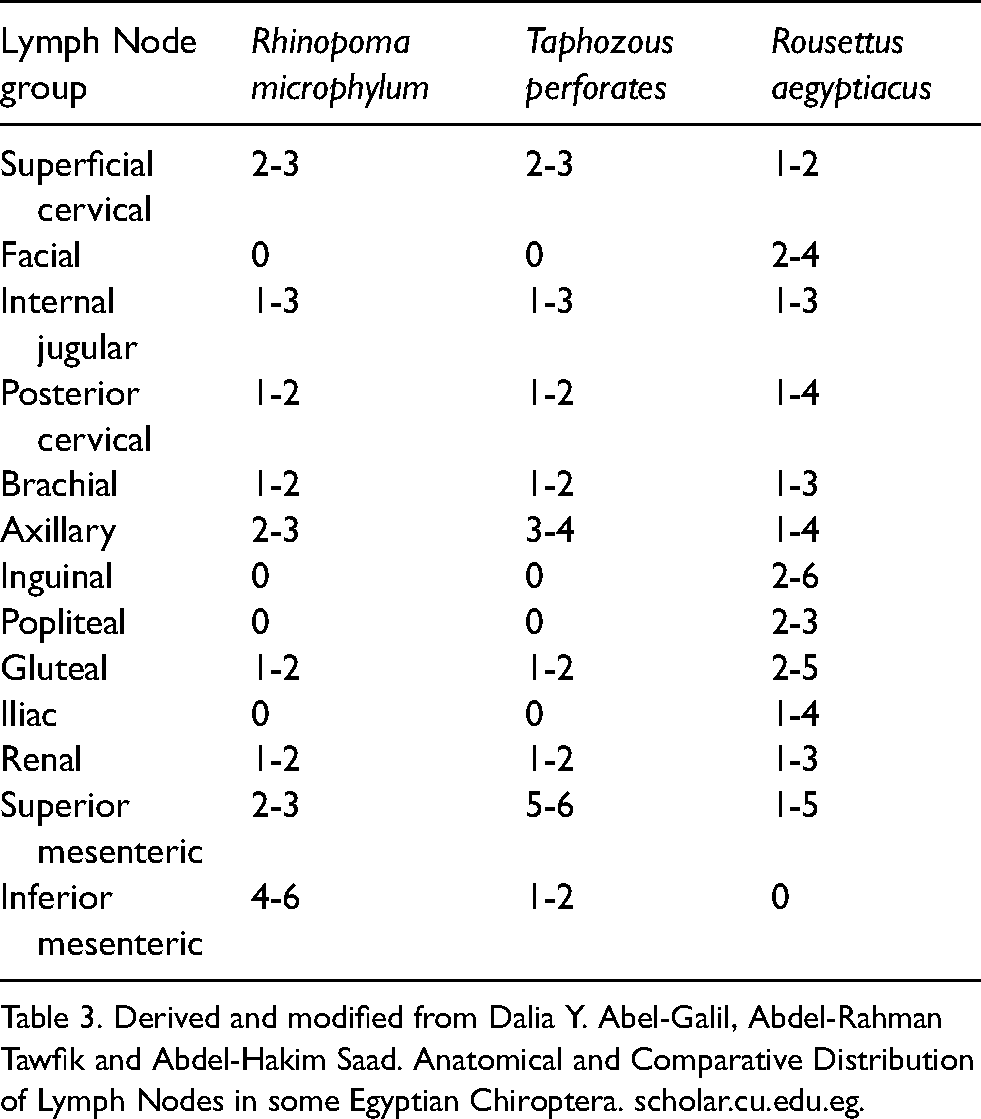
Table 3. Derived and modified from Dalia Y. Abel-Galil, Abdel-Rahman Tawfik and Abdel-Hakim Saad. Anatomical and Comparative Distribution of Lymph Nodes in some Egyptian Chiroptera. scholar.cu.edu.eg.
The presence or absence of some lymph nodes also varies across bat species. 109 The facial, popliteal, iliac and inguinal lymph nodes are absent in T. perforatus and R. microphylum, but are present in R. aegyptiacus. However, the inferior mesenteric nodes are not observed in R. aegyptiacus, but are present in T. perforatus and R. microphyllum. In some cases the lack of a group of lymph nodes is compensated for by the presence of a different node that drains the same area. For example, the lack of iliac lymph nodes in T. perforatus and R. microphyllum appears to be compensated for by the presence of a gluteal node. The number and size of lymph nodes in bats mimics the observation in mice and supports the premise that smaller species have fewer and smaller lymph nodes as compared to larger mammals. 1
A “non-symmetry” of bat lymph nodes in two species, R. aegyptiacus and T. perforatus, has been noted in which the left side nodes were larger compared to those on the right. 109 Additional variations were noted for male, female, and pregnant females with males and non-pregnant females having lymph nodes that are similar in size and numbers, and pregnant females having larger and more numerous lymph nodes (specifically inguinal, renal and superior mesenteric).
The specific histologic morphology of normal lymphoid tissues from multiple bat species have not been described in the literature. A rigorous examination and documentation of the histology of lymphoid tissue, using standardized protocols, special stains and electron microscopy, of multiple bat species would be of significant value.
The majority of other primary and secondary lymphoid organs observed in other mammals appear to be present in bats, including the thymus, bone marrow, and spleen, but descriptions of the histology of these organ is not available.109,110
Swine
The lymph nodes of swine have been described in detail previously.1,15,16,111 Swine have an anatomical reversal of the classical cortical and medullary components. The central area of the node contains cortical lymphoid follicles with B-cell dependent areas, and the paracortical T cell-dependent area. The medullary sinusoids and cords are located peripherally and penetrate irregularly into the center or occupy the majority of one pole of the node (Figure 2). Lymph usually flows into the node centrally and leaves the node through efferent vessels located on the capsular surface.
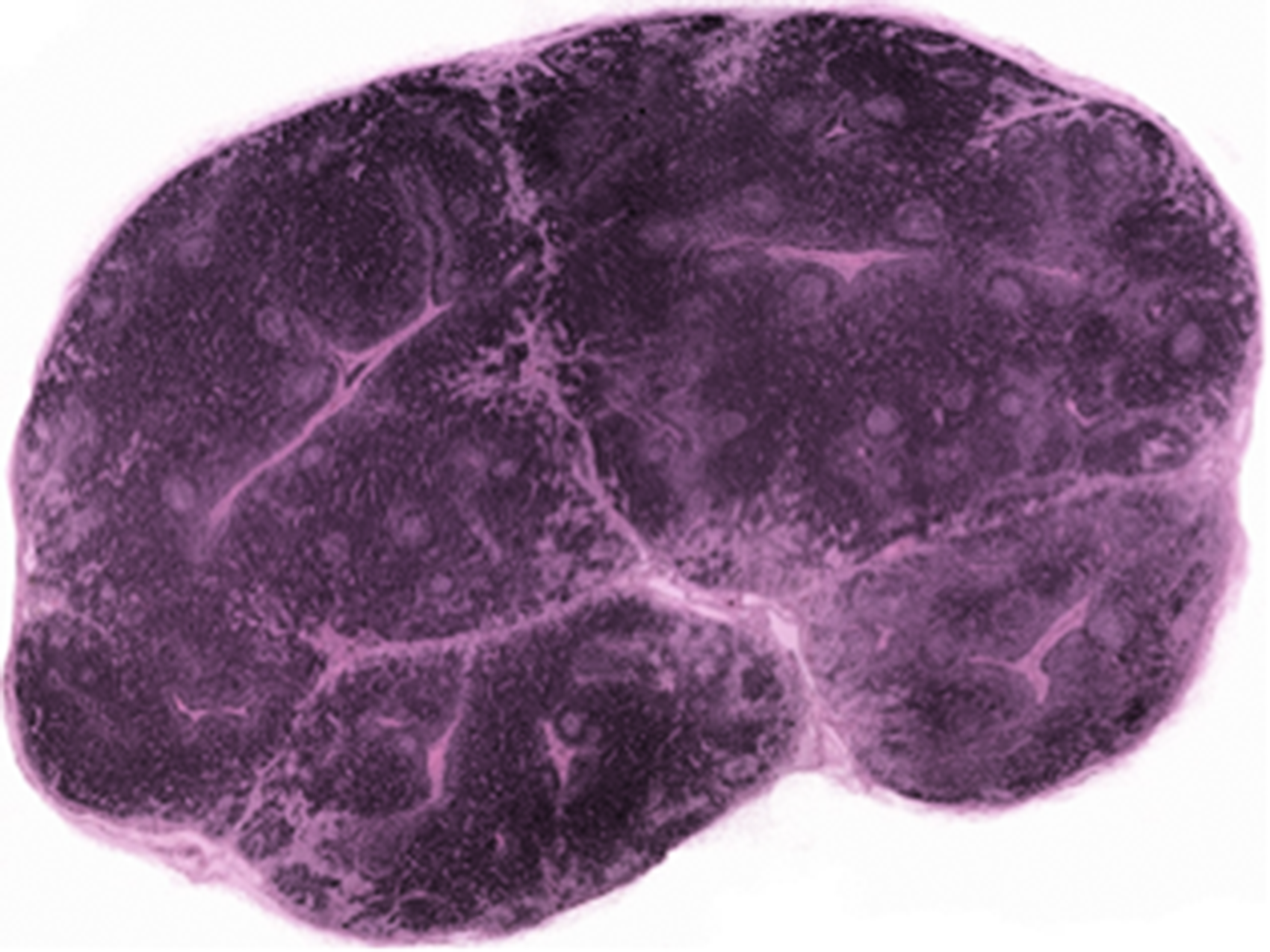
Photomicrographs of mandibular (A) and mesenteric (B) lymph nodes from a pig. There is a reversal of cortical and medullary regions, as well as the remarkable difference in the amount of cortical tissue present in each node. In (A) the cortical tissue is confluent throughout the node with the medullary tissue compressed against the outer capsule. In the mesenteric node (B) the peripherally located medullary tissue is abundant and surrounds the centrally located cortical tissue.
Spleen - general
The three variations of splenic architecture are encountered in the species described herein and include storage, defensive, and intermediate types.1,15,112,113 The storage spleen has a thick capsule with many trabeculae of well-developed smooth muscle that provides for contractility. It also has relatively sparse lymphoid tissue, and includes trapping and removing effete erythrocytes as part of its function. The defensive spleen provides cells for immunologic defense, has less smooth muscle in the capsule and trabeculae, and more lymphoid tissue than does the storage type. Lymphoid follicles and marginal zone are well delineated. The intermediate type has trabecular and lymphoid development intermediate to the other two types.
Swine
The spleen of swine is long and narrow but highly variable in size depending on strain of pig. A thick capsule of smooth muscle and elastic fibers gives rise to heavy muscular trabecular that penetrate the parenchyma. Swine have well developed periarteriolar lymphoid sheaths (PALS) but small less distinct lymphoid follicles. Branches of pulp arteries called sheathed capillaries are surrounded by concentric layers of macrophages [pericapillary macrophage sheaths (PAMS)] that are easily identified as ellipsoids within the marginal zone. The vascular sinuses of swine are lined by flat, elongated endothelial cells and are categorized as nonsinusal; the human and dog spleen is sinusal. 114 Therefore, similar to ruminants, swine have a spleen type characterized by trabecular and lymphoid development intermediate between defensive (humans, mice, rats and rabbits) and storage (cats and dogs) types. Venous sinusoids are present.
MALT - general
Mucosal-associated lymphoid tissue (MALT) refers to diffuse lymphocytes or nodular lymphocyte aggregates found in membranes lining mucosal surfaces. It is characterized as dispersed aggregates of nonencapsulated lymphoid tissue within the mucosa that play a pivotal role in the generation of local immune responses at mucosal surfaces.
There are multiple named accumulations of these lymphocytes that include, but are not limited to GALT (gut-associated lymphoid tissue), GMALT (gastric mucosa-associated lymphoid tissue) BALT (bronchus-associated lymphoid tissue), and NALT (nasal-associated lymphoid tissue). Of considerable interest for this discussion is, whether or not MALT, particularly GALT, exists in bats.
BALT consists of organized bronchial lymphoid aggregates, or follicles, morphologically similar to other components of MALT and are located within the bronchial submucosa. The amount of BALT present is highly variable across species with increased BALT reflecting the occurrence of pulmonary contact with antigens.
GALT represents the largest accumulation of MALT and plays a very significant role in generating and maintaining humoral immune responses in mammals. GALT is characterized by lymphocytes organized into discrete follicles within the mucosa along the free wall of the small intestine (Peyer‘s patches: PP), as well as lymphocytes scattered throughout the lamina propria (lamina propria lymphocytes - LPL), and intraepithelial lymphocytes (IEL). GALT has been found in all laboratory and domestic animal species previously described. 1 The mass of LPL in some mammals is thought to be greater than the mass of lymphocytes within the spleen.
Bats
An intriguing observation in bats is that, in at least two species of microbat, P. pipistrellus and R. hilebrandtii, GALT has been reported to be lacking.88,115 R. hildebrandti was reported to have only a few lymphoid aggregates in the rectal mucosa. 115 Coincidently, as noted before, it has been shown that some bats have very low levels of secretory IgA and this may be reflective of the absence of the lymphoid tissue primarily responsible for secreting IgA. 76 No other bat species have been investigated for the presence of MALT. Detailed characterization of MALT, especially GALT and all of its components (LPL, IEL PP), in bats should provide valuable insights into the role of IgA in mucosal immunity as well as its possible role in the establishment of virus reservoirs.
Swine
As noted above, MALT in swine has been extensively reviewed.1,15,16 Many forms of MALT have been identified in swine including BALT, GMALT, and GALT (PP, LPL and IEL). Similar to other forms of MALT, BALT in swine increases with antigen exposure and age. Germ-free pigs are devoid of BALT.
Summary and conclusions
As stated, information on the species considered here-in is highly variable with regard to which immune structures and functions are annotated in detail. For example, the details of innate immunity in bats has been thoroughly investigated and from it has emerged new insights into the relationships of reservoir hosts and to the viruses that infect them. But there is limited-to-no published data on the structure of the immune system of bats, or characterization of that species’ adaptive immunity that might provide additional insight and clarity to the biology of the immune responses of bats. With the exception of swine, there are large knowledge gaps concerning the structure and function of bats, civets, raccoon dogs, pangolins and camelids that of the immune system warrant investigation. It is clear that there are many research opportunities that would extend and refine our understanding of the immune structure and system of these species, and perhaps provide insight into novel therapeutic strategies.
Patterns of relative lymphoid tissue paucity in some species suggest that a reservoir species might maintain chronic infections based on the reduced percentage of cells attacking the infections thereby ‘anatomically blunting‘ the immune response and contributing to the ability of reservoir hosts to spread infection. Thus, a naturally occurring absence of GALT and its associated germinal centers in bats (if verified) may be the root cause of a limited production of IgA secretory antibody. Likewise, limited observations of bat lymph nodes suggest that there may be a normal reduction in lymphocytes in peripheral lymph nodes as compared to other mammals. A decreased capability to generate a robust humoral immune response might contribute to the ability of bats to function so effectively as reservoir hosts.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
