Abstract
Macrophage autophagy plays a major role in the control and elimination of invading Mycobacterium tuberculosis. However, the function and mechanism of circRNA on macrophage autophagy in tuberculosis remain unclear. Therefore, this study aimed to explore the role of circRNA underlying macrophage autophagy in tuberculosis. Quantitative real-time polymerase chain reaction was used to detect the expression of hsa_circ_0045474, miR-582-5p and TNKS2. Autophagy was detected by LC3B immunofluorescence and transmission electron microscopy. Dual-luciferase reporter assays were used to detect the relationship of miR-582-5p and hsa_circ_0045474 or TNKS2. Western blot was used to detect the expression of LC3-І and LC3-ІІ. The results showed that hsa_circ_0045474 was down-regulated in monocytes from patients with tuberculosis and induced autophagy in macrophages. hsa_circ_0045474 sponged miR-582-5p and negatively regulated miR-582-5p expression. Overexpression of miR-582-5p affected by hsa_circ_0045474 induced autophagy in macrophages. TNKS2 served as a target of miR-582-5p and down-regulation of TNKS2 induced autophagy in macrophages regulated by miR-582-5p. In conclusion, our results demonstrated that hsa_circ_0045474 down-regulation induced macrophage autophagy in tuberculosis via miR-582-5p/ TNKS2 axis, implying a novel strategy to treat the occurrence of active pulmonary tuberculosis caused by immune escape of M. tuberculosis.
Introduction
Tuberculosis (TB), a chronic pulmonary infectious disease caused by Mycobacterium tuberculosis (Mtb), 1 is one of the most common chronic infectious diseases in the world. Due to the complex pathogenesis and untypical clinical symptoms, TB is difficult to early diagnose, which will lead to the difficulty of effective control of the source of TB infection. 2 Therefore, finding an early and accurate diagnostic method is very important for the prevention and treatment of TB.
Macrophages are important immune cells in the body which play a vital role in host anti-infective immunity by phagocytizing and eliminating pathogens and presenting Ags to T cells to initiate protective acquired immune response.3,4 Hence, understanding of the mechanism of Mtb escaping from macrophage killing and the mechanism of growth, dormancy and resuscitation in macrophages is essential for formulating strategies to prevent and cure TB. Increasing studies have demonstrated that macrophage autophagy plays an important role in TB. 5 For instance, Maximiliano et al. has indicated that autophagy is a defense mechanism inhibiting BCG and Mtb survival in infected macrophages. 6 BAG2 ameliorated stress-induced cell apoptosis in Mtb-infected macroendoplasmic reticulumphages through selective autophagy. 7 LncRNA PCED1B-AS1 regulated macrophage apoptosis and autophagy by sponging miR-155 in active TB. 8 However, the regulatory mechanism underlying macrophage autophagy in TB remains unclear.
Circular RNAs (circRNAs) are a new type of endogenous non-coding RNAs that are different from traditional linear RNA. 9 CircRNAs are widely expressed in organisms and have the characteristics of high stability, specificity and conservation. CircRNAs play important regulatory roles in the occurrence and development of inflammation, immune response and tumors.10,11 Studies have found that circRNAs are differentially expressed in the peripheral blood and tissues of TB patients, and is closely related to the occurrence, development and prognosis of TB. 12 Hence, specific circRNA is expected to become a biomarker for TB diagnosis, treatment and prognosis. For instance, Zhang et al. has revealed that hsa_circ_0028883 has a potential value for TB diagnosis by ROC analysis using the gene expression datasets of circRNA GSE117563 and GSE106953. 13 Fu et al. has indicated that some dys-regulated circRNAs are potential biomarkers for the diagnosis of TB and circRNA_101128-let-7a interplay may play a considerable role in PBMCs response to Mtb infection. 14 Recently, another research has showed that up-regulation of circRNA-0003528 promotes Mtb-associated macrophage polarization via down-regulating miR-224-5p, miR-324-5p and miR-488-5p and up-regulating CTLA4. 15
It has been demonstrated that miR-582-5p is up-regulated and can act as a diagnostic biomarker in active TB. 16 Moreover, miR-582-5p inhibited macrophage apoptosis and might play an important role in regulating anti-Mtb directed immune responses. 17 Bioinformatics analysis showed that hsa_circ_0045474 may possess binding sites for bindmiR-582-5p. However, the role of hsa_circ_0045474 underlying macrophage autophagy in TB remains unclear. Therefore, this study aimed to explore the function and mechanism of hsa_circ_0045474 in TB.
Materials and methods
Patients and tissue samples
Human peripheral samples were obtained from 15 patients with TB and 15 healthy donors. All included patients were diagnosed based on typical clinical symptoms, chest radiography, and positive culture and PCR for Mtb. Healthy controls and TB patients with other critical basic diseases (cancer, infection, and immune-compromised conditions) were excluded. All patients received no treatment on TB or only received first-line treatment. This study was approved by the Ethics Review Board of Shanghai Pulmonary Hospital, School of Medicine, Tongji University and performed in accordance with the Declaration of Helsinki. All participants wrote the informed consent agreements.
Cell isolation and culture
The samples were centrifuged using Hypaque-Ficoll (GE Healthcare Bio-sciences AB, Uppsala, Sweden) for isolation of peripheral blood mononuclear cell (PBMC) according to the manufacturer's protocol. The human myelomonocytic leukemia cell line THP-1 was purchased from the Institute of ATCC. Both PBMCs and THP-1 cells were cultured in PRMI-1640 medium (HyClone, Shanghai, China) containing 10% FBS (HyClone). Then, PBMCs and THP-1 were differentiated into monocyte-derived macrophages (MDMs)/THP-1 derived macrophages (TDMs) as previously described.
18
In brief, monocytes were enriched from freshly-isolated PBMCs using MACS Monocyte Isolation Kit II and MACS LS Columns (Miltenyi Biotec), yielding an average 98% purity. To differentiatiate PBMCs into MDMs, 2 × 106 PBMCs were plated in RPMI-1640 medium with 2 mmol/l
Total RNA extraction and qRT-PCR
Total RNA was isolated from cells and tissues by TRIzol Kit (Takara, Japan), and the quantity of total RNA was measured using a NanoDrop (Thermo Fisher Scientific, Waltham, MA, USA). Complementary DNA (cDNA) was synthesized using the cDNA Synthesis kit (Takara, Otsu, Japan). Real-time PCR was performed using SYBR Green PCR Master Mix (Takara) in a Step OnePlus RT-PCR system (Thermo Fisher Scientific). All target genes were normalized to the endogenous reference gene GAPDH. To measure the level of miR-582-5p, a miRNA extraction kit (Vazyme, Nanjing, China) and a stem-loop miRNA Synthesis Kit (Vazyme) were used according to the manufacturer's instructions. U6 snRNA was used as the endogenous control. The expression of the genes was calculated by using an optimized comparative Ct (2−ΔΔCT) method. The primers was listed as follows: hsa_circ_0045474, F: 5′-GCCAACAGTCTCCAAGC-3′, R: 5′-GCGGTAGCAGTCCATACC-3′; miR-582-5p, F: 5′-GCGGTTACAGTTGTTCAACC-3′, R: 5′-CTCAACTGGTGTCGTGGA-3′; TNKS2, F: 5′-CGCGGATCCTGAAGGTATGGTCGATG-3′, R: 5′-CGCGAATTCAATTTAGTACAGACAACCC-3′; U6, F: 5′-GCTTCGGCAGCACATATACTAAAAT-3′, R: 5′-CGCTTCACGAATTTGCGTGTCAT-3′; GAPDH, F: 5′-GAAGGTGAAGGTCGGAGTC-3′, R: 5′-GAAGATGGTGATGGGATTTC-3′.
Western blot assays
RIPA lysis buffer was used to extract the total proteins from cells and tissues, and the protein concentration was quantified by using a BCA protein assay kit (FA016-50G, Amresco, USA,). The 20 μg of protein was separated by 10% SDS-PAGE and the protein bands were then transferred onto polyvinylidene fluoride membranes (IPVH00010, Millipore, USA). The membranes were subsequently blocked with 5% non-fat milk for 2 h and then incubated with primary Abs against TNKS2 (1:1000, Abcam, ab155545), LC3B (1:1000; Abcam, ab192890) or GAPDH (1:2000; Abcam, ab8245) overnight at 4°C. After washing, the membranes were incubated with an HRP-conjugated secondary Ab (1:2000; BOSTER, BA1054) for 50 min. The immunostained proteins were visualized using ECL reagent (Applygen Technologies, China) and X-ray film (SUPER RX-N-C, Japan).
Dual-luciferase reporter assays
The wild type (WT) and mutant (Mut) fragments of hsa_circ_0045474 and 3′untranslated region (3′UTR) of TNKS2 were amplified by PCR and inserted into the pmirGLO vector (Promega). The 293 T cells were seeded into 96-well plates (1 × 104 cells/well) and incubated at 37°C. The cells were co-transfected with miR-582-5p mimics, miR-582-5p NC and WT- or Mut-hsa_circ_0045474 or TNKS2. The nucleotide sequences of all constructs were confirmed by DNA sequencing. Luciferase activity was assessed by using the Dual-Luciferase Assay System (Promega) according to manufacturer's instructions at 48 h after transfection. Firefly luciferase activity was normalized to that of Renilla luciferase activity.
Transmission electron microscopy
Transmission electron microscopy (TEM) was performed according to the instruction carried out previously. 19 The cells were trypsinized and seeded at a density of 1 × 104 cells/well on a Lab-Tek® Chamber Slide (Nalge Nunc International, Naperville, IL, USA) in 1 ml of culture medium with 10% FBS for 48 h. For TEM, the treated cells were washed and fixed with 2.5% glutaraldehyde in 0.1 M of phosphate buffer (pH 7.4) for 2 h and then post-fixed with 1% osmium tetroxide in the same buffer for 2 h. They were dehydrated in a graded series of ethanol and embedded in Epon 812. Ultrathin sections were stained with 4% uranyl acetate and lead citrate. The number of autophagic vacuoles was determined for a minimum of 100 cells, and the autophagic vacuoles were photographed and observed under an electron microscope (JEM-1400, Jeol, Tokyo, Japan; bar = 100 μm) at 80 kV.
Immunofluorescence microscopy
The cells were fixed with 4% paraformaldehyde for 20 min and permeabilized with 0.1% Triton X-100 (Sigma, T9284) for 10 min. Then the cells were blocked by 5% normal goat serum (Jackson laboratories, 005-000-121). Next, the cells were incubated with LC3B primary Ab at room temperature for 2 h, rinsed 4 times with PBS-Tween 20 (0.05%; Sigma, P1379), incubated with secondary Abs produced in mouse or rabbit (diluted 1:500 in 0.5% normal goat serum) for 1 h at room temperature, and washed 4 times with PBS-Tween 20 followed by secondary Ab incubation with Alexa fluor 488 and 568 respectively for 1 h at room temperature. DAPI (Invitrogen, S36938) was used to stain the nuclei. The images were acquired by a Ziess LSM 510 Meta confocal microscope (Jena, Germany; bar = 100 μm). Data analysis was performed using Leica LAS AF Lite software. The number of GFP-LC3 puncta per cell were assessed in 10 non-overlapping fields.
Statistical analysis
Statistical analyses were performed using SPSS 17.0 software (SPSS, Chicago, IL, USA). The data were presented as the mean ± SD. Comparison between two groups or more than two groups was performed using unpaired Student's t-test or one-way analysis of variance, respectively. P < 0.05 acted as the significant difference.
Results
Down-regulation of hsa_circ_0045474 induces autophagy in macrophages
To know the effect of hsa_circ_0045474 on autophagy in macrophages, the expression of hsa_circ_0045474 was investigated in monocytes from patients with TB. Quantitative real-time polymerase chain reaction (qRT-PCR) results demonstrated that the expression of hsa_circ_0045474 in monocytes from patients with TB was significantly down-regulated compared with that in monocytes from healthy controls (Figure 1A). Then, cell autophagy was detected by LC3B immunofluorescence. First, we verified the change of autophagy flux using the autophagy inhibitor, Baf-A1. The results showed that Baf-A1 significantly reduced the number of autophagic puncta (Figure 1B). Moreover, we investigated the function of hsa_circ_0045474 under Baf-A1 treatment. The results indicated that up-regulation of hsa_circ_0045474 dramatically reduced the number of autophagic puncta and inhibited autophagosome formation in macrophages, which showed the similar effect as Baf-A1 did. While down-regulation of hsa_circ_0045474 obviously increased the number of autophagic puncta and induced autophagosome formation in macrophages, which was reversed by Baf-A1 treatment (Figure 1C). The function of hsa_circ_0045474 overexpression or knockdown was further validated by the observation results of TEM (Figure 1D). In addition, LC3-ІІ/LC3-І was dramatically decreased in overexpressing hsa_circ_0045474 group while elevated in hsa_circ_0045474 silencing group than that in control group (Figure 1E). These results demonstrated that down-regulation of hsa_circ_0045474 promoted autophagy in macrophages.
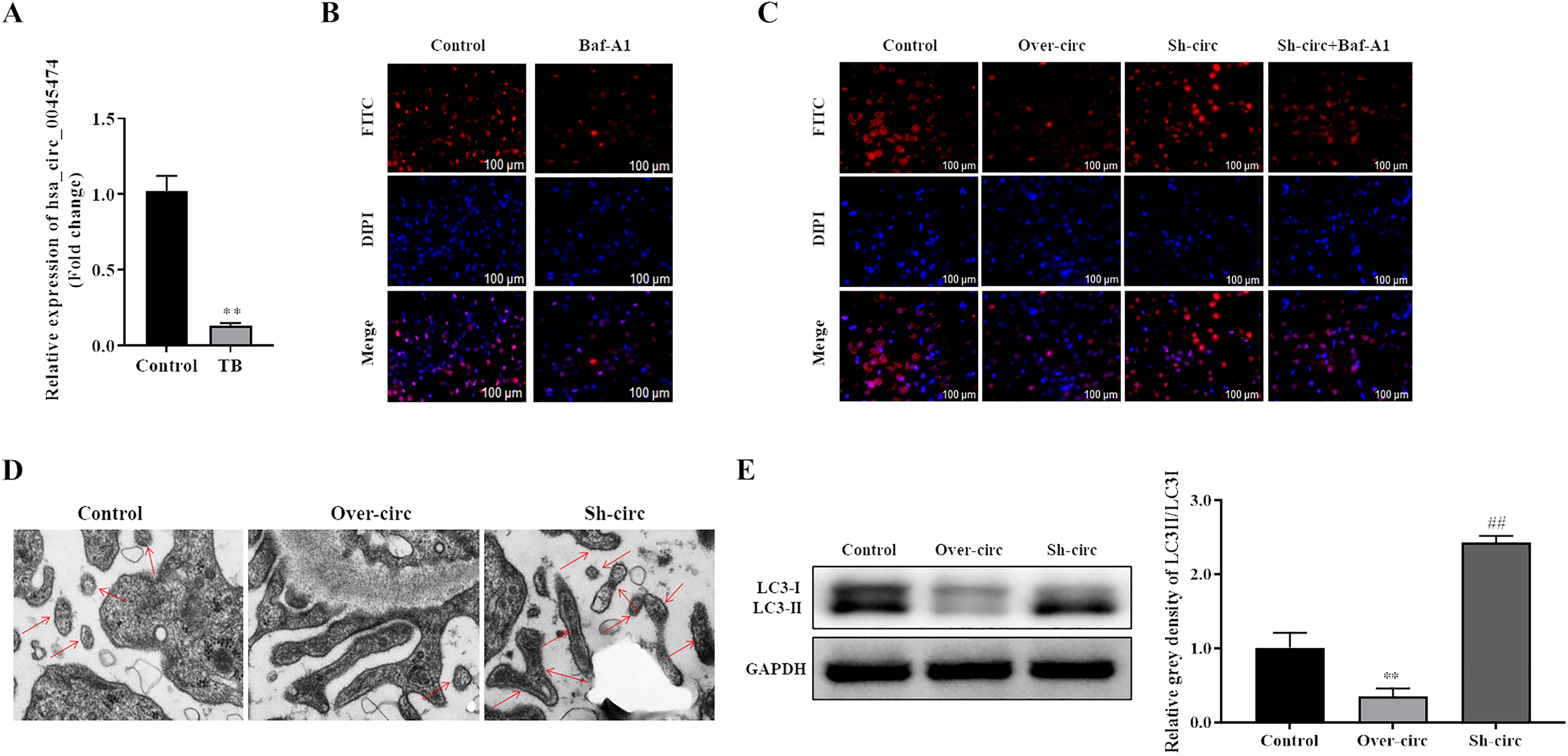
Down-regulation of hsa_circ_0045474 induced autophagy in macrophages. (A) qRT-PCR was used to detect the expression of hsa_circ_0045474 in monocytes from patients with TB, and GAPDH acted as control. The data shown as mean ± SD. **, P < 0.01, TB group versus control group. (B-C) Cell autophagy expression was detected by LC3B immunofluorescence. Bar = 100 μm. (D) Autophagosome was observed by transmission electron microscopy. Bar = 100 μm. (E) Expression of LC3-ІІ and LC3-І was detected by Western blot. The data shown as mean ± SD. **, P < 0.01, Over-circ group versus control group; ##, P < 0.01, Sh-circ group versus control group.
Hsa_circ_0045474 sponges miR-582-5p and negatively regulates miR-582-5p expression
In the previous study, miR-582-5p was up-regulated and could be act as diagnostic biomarker in active TB. 16 In the present study, we also confirmed by qRT-PCR that the expression of miR-582-5p was significantly up-regulated in monocytes from patients with TB compared with that in monocytes from healthy controls (Figure 2(A)). Further bioinformatics analysis indicated that hsa_circ_0045474 could bind with miR-582-5p (Figure 2(B)). Dual-luciferase reporter assays indicated that there was a decreased luciferase intensity under co-transfection with WT- hsa_circ_0045474 and miR-582-5p, while there was no difference between the luciferase intensities of Mut-hsa_circ_0045474 and miR-582-5p (Figure 2(C)). In addition, qRT-PCR assays revealed that up-regulation of hsa_circ_0045474 decreased miR-582-5p expression while knockdown of hsa_circ_0045474 promoted miR-582-5p expression (Figure 2(D)). Our results confirmed that hsa_circ_0045474 could serve as a sponge to directly regulate miR-582-5p expression.
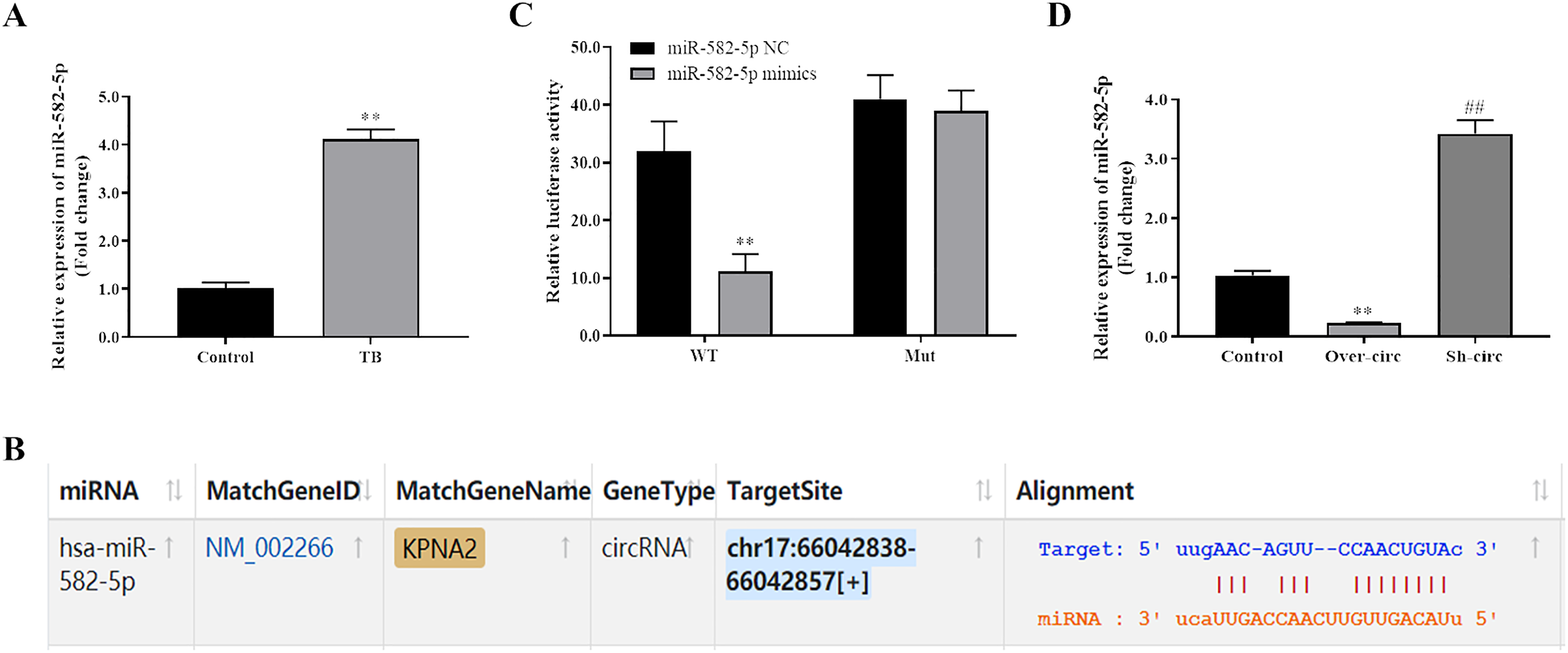
Hsa_circ_0045474 sponged miR-582-5p and negatively regulated miR-582-5p expression. (A) qRT-PCR was used to detect the expression of miR-582-5p in monocytes from patients with TB, and U6 acted as control. The data shown as mean ± SD. **, P < 0.01, TB group versus control group. (B) Starbase was used to predict binding site of miR-582-5p on hsa_circ_0045474. (C) Dual-luciferase reporter assays were used to detect the relationship of miR-582-5p and hsa_circ_0045474. The data shown as mean ± SD. **, P < 0.01, miR-582-5p mimics group versus miR-582-5p NC group. (D) qRT-PCR assays were used to detect the expression of miR-582-5p affected by hsa_circ_0045474. The data shown as mean ± SD. **, P < 0.01, Over-circ group versus control group; ##, P < 0.01, Sh-circ group versus control group.
Overexpression of miR-582-5p affected by hsa_circ_0045474 induces autophagy in macrophages
The influence of miR-582-5p on cell autophagy was detected by LC3B immunofluorescence and TEM. The results indicated that miR-582-5p mimics dramatically increased the number of autophagic puncta and induced autophagosome formation while miR-582-5p inhibitor reduced the number of autophagic puncta and inhibited autophagosome formation in macrophages (Figure 3A and B). In addition, LC3-ІІ/LC3-І was dramatically increased in miR-582-5p mimics group while decreased in miR-582-5p inhibitor group compared to in control group (Figure 3C). In addition, we found that these changes were partially reversed when overexpressing hsa_circ_0045474 in miR-582-5p mimics group or suppressing hsa_circ_0045474 in miR-582-5p inhibitor group (Figure 3A to C). These results demonstrated that overexpression of miR-582-5p affected by hsa_circ_0045474 induced autophagy in macrophages.
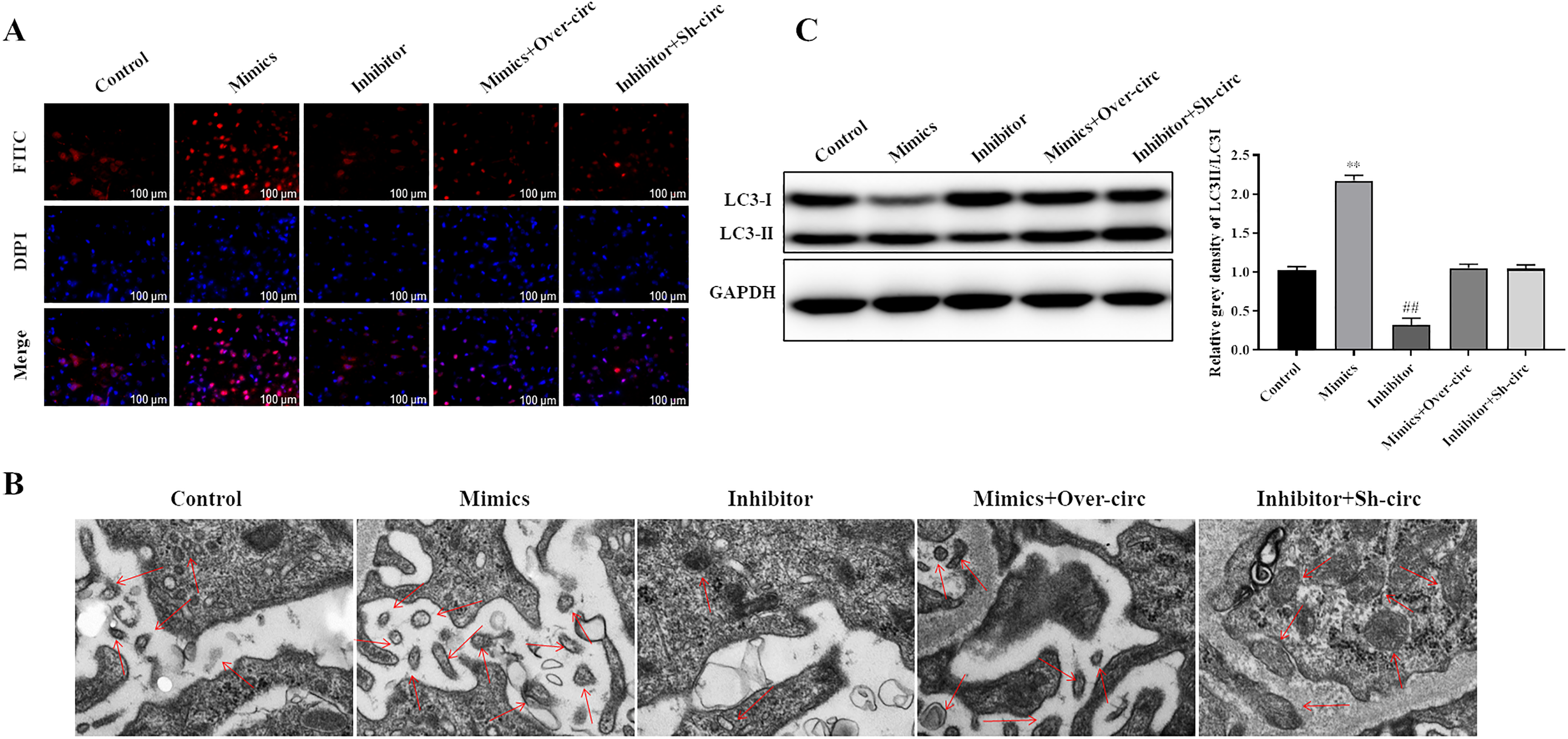
Overexpression of miR-582-5p sponged by hsa_circ_0045474 induced autophagy in macrophages. (A) Cell autophagy expression was detected by LC3B immunofluorescence. Bar = 100 μm. (B) Autophagosome was observed by TEM. Bar = 100 μm. (C) Expression of LC3-ІІ and LC3-І was detected by Western blot. The data shown as mean ± SD. **, P < 0.01, Mimics group versus control group; ##, P < 0.01, Inhibitor group versus control group.
TNKS2 serves as a target of miR-582-5p
To know the downstream molecule of miR-582-5p in TB, TargetScan was used to predict the targets of miR-582-5p. As shown in Figure 4(A), there was a binding site of miR-582-5p in the position 305-312 of TNKS2 3′UTR, suggesting that TNKS2 might have a binding with miR-582-5p. The qRT-PCR and Western blot results showed that the expression of TNKS2 was significantly down-regulated in monocytes from patients with TB compared with that in monocytes from healthy controls (Figure 4B and C). Dual-luciferase reporter assays indicated that there was a decreased luciferase intensity under co-transfection with WT-TNKS2 3′UTR and miR-582-5p, while there was no difference between the luciferase intensities of Mut-TNKS2 and miR-582-5p (Figure 4D). In addition, qRT-PCR and Western blot analysis revealed that miR-582-5p mimics decreased TNKS2 expression while miR-582-5p inhibitor promoted TNKS2 expression (Figure 4E and F). These results indicated that TNKS2 served as a target of miR-582-5p.
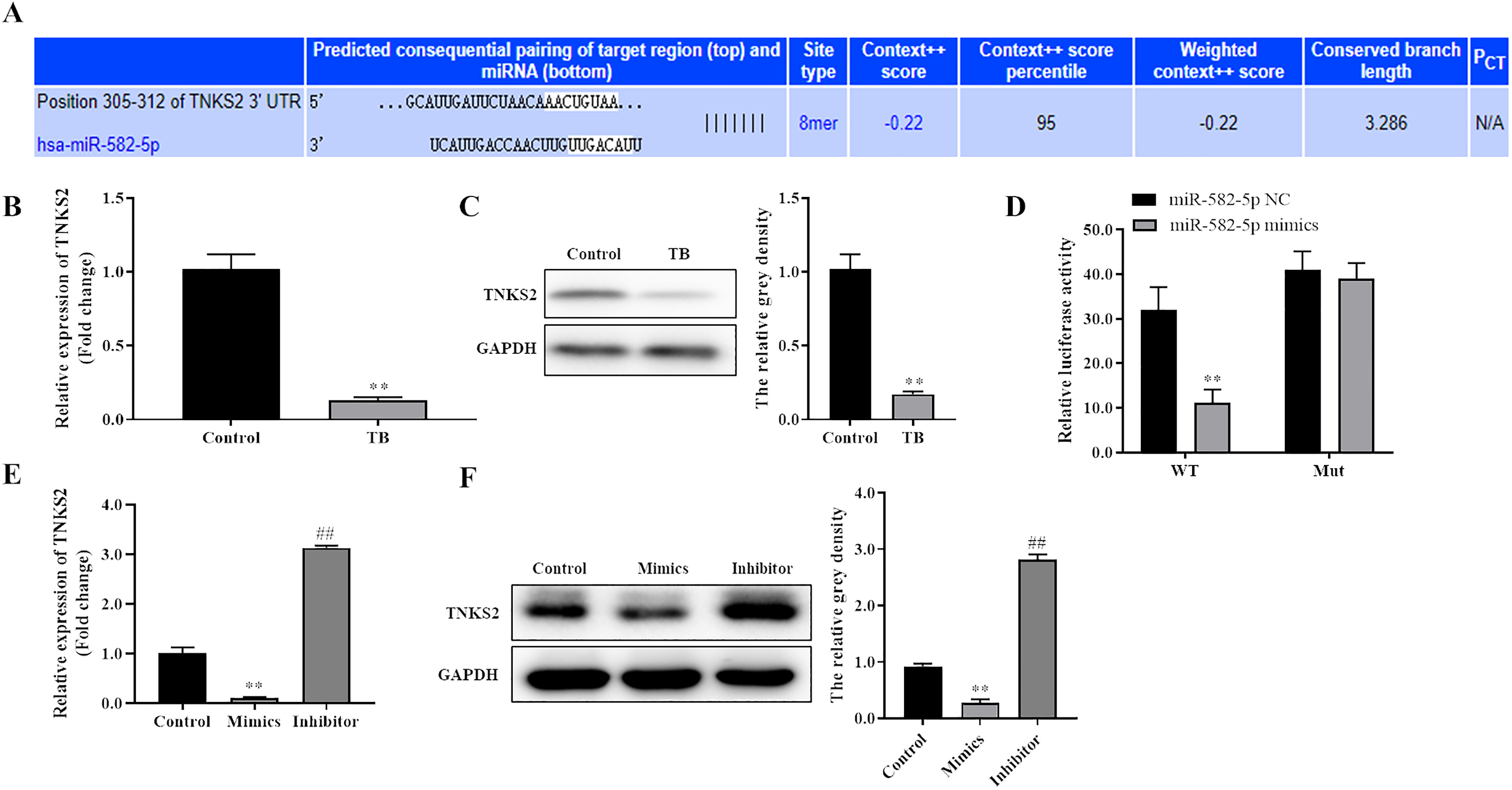
TNKS2 served as a target of miR-582-5p. (A) Targetscan was used to predict the binding site of miR-582-5p on TNKS2 3′UTR. (B-C) qRT-PCR and Western blot were used to detect the mRNA and protein expression of TNKS2 in monocytes from patients with TB, respectively, and GAPDH act as control. The data shown as mean ± SD. **, P < 0.01, TB group versus control group. (D) Dual-luciferase reporter assays were used to detect the relationship of miR-582-5p and TNKS2 3′UTR. The data shown as mean ± SD. **, P < 0.01, miR-582-5p mimics group versus miR-582-5p NC group; (E-F) qRT-PCR and Western blot were used to detect the expression of TNKS2 affected by miR-582-5p. The data shown as mean ± SD. **, P < 0.01, Mimics group versus control group; ##, P < 0.01, Inhibitor group versus control group.
Down-regulation of TNKS2 induces autophagy in macrophages regulated by miR-582-5p
Cell autophagy analysis indicated that up-regulation of TNKS2 dramatically reduced the number of autophagic puncta and inhibited autophagosome formation in macrophages while down-regulation of TNKS2 obviously increased the number of autophagic puncta and induced autophagosome formation in macrophages (Figure 5A and B). In addition, the LC3-ІІ/LC3-І ratio was dramatically decreased in overexpressing TNKS2 group while elevated in suppressing TNKS2 group than that in control group (Figure 5C). In addition, we found that these changes were partially reversed when co-transfection with miR-582-5p mimics in Over-TNKS2 group or co-transfection with miR-582-5p inhibitor in Sh-TNKS2 group (Figure 5A to C). These results demonstrated that down-regulation of TNKS2 induced autophagy in macrophages regulated by miR-582-5p.
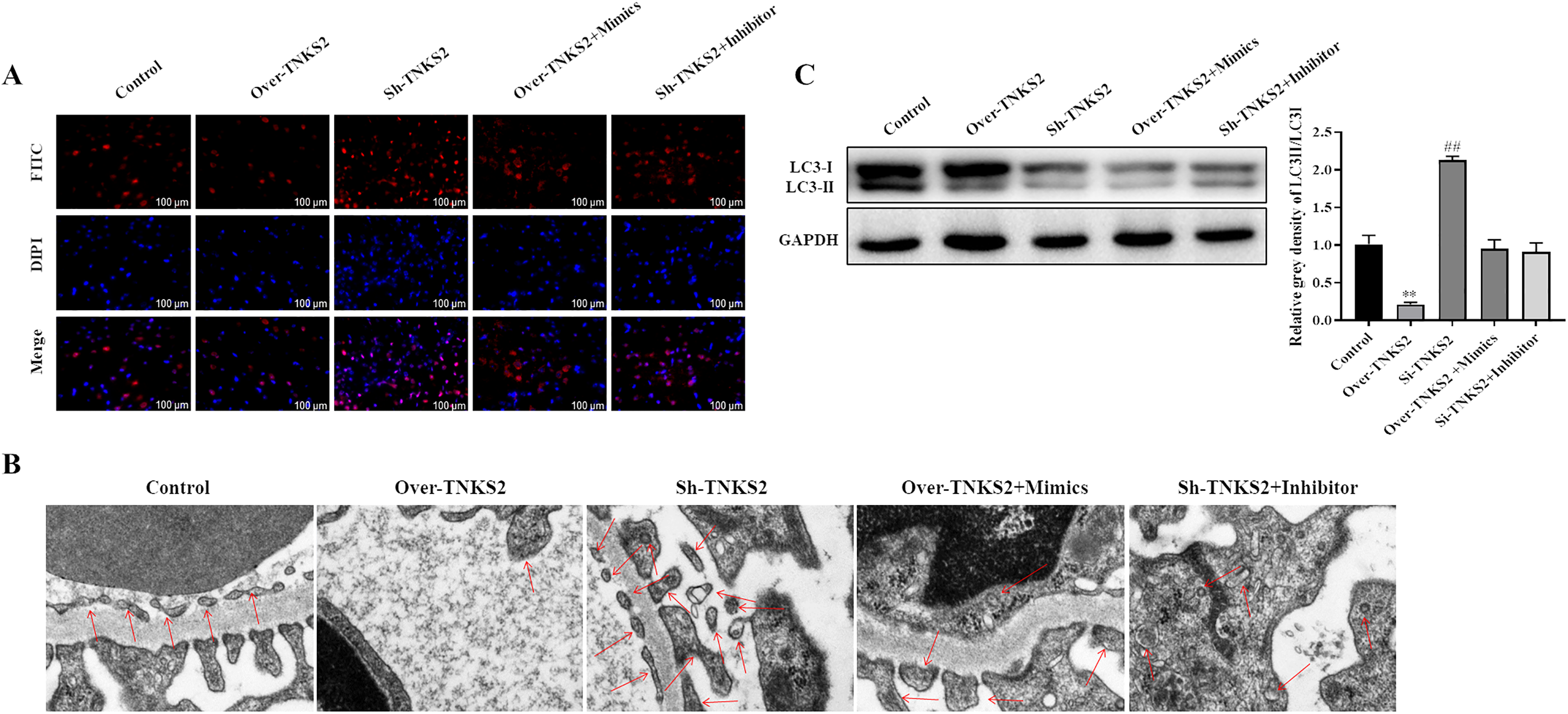
Down-regulation of TNKS2 induced autophagy in macrophages regulated by miR-582-5p. (A) Cell autophagy expression was detected by LC3B immunofluorescence. Bar = 100 μm. (B) Autophagosome was observed by TEM. Bar = 100 μm. (C) Expression of LC3-ІІ and LC3-І was detected by Western blot. The data shown as mean ± SD. **, P < 0.01, Over-TNKS2 group versus control group; ##, P < 0.01, Sh-TNKS2 group versus control group.
Discussion
TB is one of the most common chronic infectious diseases worldwide. It is difficult to make a early diagnosis for TB due to the complex pathogenesis and atypical clinical symptoms. 20 Macrophage autophagy plays a crucial role in the clearance of Mtb in the pathogenesis of pulmonary TB. However, the mechanism of circRNAs underlying macrophages autophagy in TB has not been elucidated yet. In the present study, we demonstrated that down-regulation of hsa_circ_0045474 induced macrophage autophagy in TB via miR-582-5p/ TNKS2 axis.
Mtb is a pathogenic bacterium causing TB. 21 Mtb can escape from the clearance of macrophages through a variety of immune escape mechanisms, and then parasitize in macrophages for a long time. 22 Increasing evidence has revealed that macrophage autophagy plays important roles in TB.23,24 For instance, Franco has indicated that SMURF1 controls Mtb replication in human macrophages and associates with bacteria in lungs of patients with pulmonary TB, suggesting that SMURF1 is required for selective autophagy of Mtb and host defense against TB infection. 25 Hu et al. has indicated that ibrutinib suppresses intracellular Mtb growth by inducing macrophage autophagy. 26
CircRNAs are widely involved in a variety of biological processes and can be used as diagnostic markers and potential therapeutic targets for a variety of diseases. Increasing circRNAs were proved to play crucial roles in TB, 27 such as hsa_circ_0001380, hsa_circ_0001953 and hsa_circ_0009024.28,29 In the present study, we demonstrated that hsa_circ_0045474 was significantly down-regulated in monocytes from patients with TB compared with that in monocytes from healthy controls. In addition, circRNAs are also found to participate in TB-associated macrophage autophagy. For instance, up-regulation of circRNA-0003528 promoted Mtb-associated macrophage polarization via down-regulating miR-224-5p, miR-324-5p and miR-488-5p and up-regulating CTLA4. 15 CircAGFG1 modulated autophagy and apoptosis of macrophages infected by Mtb via the Notch signaling pathway. 30 In the present study, we indicated that down-regulation of hsa_circ_0045474 induced autophagy in macrophages, suggesting that autophagy in macrophages was regulated by hsa_circ_0045474 in TB.
Several studies have demonstrated that circRNAs can act as ceRNAs of miRNAs which play crucial roles in different diseases. 31 It was also proved that up-regulation of circRNA-0003528 promoted Mtb-associated macrophage polarization via down-regulating miR-224-5p, miR-324-5p and miR-488-5p and up-regulating CTLA4. 15 It has been demonstrated that miR-582-5p is up-regulated in TB. 17 In the present study, we demonstrated that hsa_circ_0045474 sponged miR-582-5p and TNKS2 served as a target of miR-582-5p. However, the function of miR-582-5p or TNKS2 on autophagy in macrophages remains unclear. Further analysis indicated that overexpression of miR-582-5p and down-regulation of TNKS2 induced autophagy in macrophages. However, hsa_circ_0045474 or miR-582-5p partially reversed the function of miR-582-5p or TNKS2 on autophagy in macrophages, suggesting that down-regulation of hsa_circ_0045474 induced macrophage autophagy in TB via miR-582-5p/ TNKS2 axis.
It was revealed that macrophages from TB patients have higher autophagic activity. However, whether these cells can improve capacity to control mycobacteria remains unclear. In our future study, we will explore whether these cells can control intracellular growth of Mtb.
In conclusion, we demonstrated that down-regulation of hsa_circ_0045474 induced macrophage autophagy in TB via miR-582-5p/TNKS2 axis, indicating a potential biomarker and novel strategy for diagnosis, treatment and prognosis of TB caused by immune escape of Mtb.
Footnotes
Acknowledgments
We thank technical support from Shanghai Pulmonary Hospital.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received the following financial support for the research, authorship and/or publication of this article: This work was supported by Natural Science Foundation of Huai'an (NMUB2019342).
