Abstract
Although liver ischaemia–reperfusion (I/R) injury remains the primary underlying reason for liver transplant failure or post-transplantation liver dysfunction, the underlying mechanism is still largely elusive. MicroRNAs (miRNA) are involved in multiple physiological and pathological processes, including inflammation. Here, we identified that the miR-128-3p/Rho family GTPase 3 (Rnd3)/NF‐κB axis might play a critical role in liver I/R injury. Our results demonstrated that the level of miR-128-3p was negatively correlated with the Rnd3 level during liver I/R. Dual luciferase reporter assay results proved that Rnd3 mRNA was a direct target of miR-128-3p. Additionally, Western blotting and quantitative RT-PCR analyses revealed that knock-down of miR-128-3p could up-regulate Rnd3 mRNA and protein levels, thereby suppressing the NF-κB pathway through down-regulating NF‐κB p65. Consequently, the serum levels of NF-κB–associated inflammatory factors and aspartate aminotransferase/alanine aminotransferase were decreased. Moreover, overexpression of Rnd3 could reverse the activation of NF-κB caused by miR-128-3p agomir during liver I/R injury. Overall, our study results suggest that repression of miR-128-3p can alleviate liver I/R injury through the miR-128-3p/Rnd3/NF‐κB axis and may facilitate the development of novel protective approaches against liver I/R injury.
Introduction
As a major cause of organ damage occurring during surgical procedures such as liver transplantation, liver ischaemia–reperfusion (I/R) injury remains a key reason for transplant failure and liver dysfunction after transplantation.1–3 After the first stage of ischaemic damage and the followed lesion of restoration of oxygen delivery, a wave of inflammation, oxidative stress and calcium overload caused by I/R injury eventually leads to irreversible cell damage.4–6 However, the specific underlying cellular and molecular mechanisms of liver I/R injury are still unclear in both basic research and clinical practice.
NF-κB is an aggregative name for inducible dimeric transcription factors, which are considered the main factors in inflammatory responses following multiple diseases.7,8 RelA/p65 is one of the main subunits of NF-κB. 9 When the NF-κB signal is activated, the expression level of p65 and its move into the nucleus acts as a crucial role in the progress of inflammation during liver I/R injury. 10 MicroRNAs (miRNAs), a set of non-coding RNA molecules, are endogenous, highly conserved and 20–24 nucleotides in length. 11 Decades of research have shown that they could participate in multiple biological and pathological processes, such as inflammation. 12 miR-128-3p is a newly found miRNA first reported in 2015. Recently, Xia et al. showed that decreased miR-128-3p could markedly restrain the inflammation response of rheumatoid arthritis by inhibiting NF-κB pathway ignition. 13 Wu et al. claimed that miR-128-3p might participate in the damnification of bone marrow mesenchymal stem cells (BMSCs) during TNF-α-induced inflammation. 14 Nevertheless, whether miR-128-3p participates in I/R injury and how miR-128-3p functions remain a mystery.
In this study, we considered Rho family GTPase 3 (Rnd3), an important member of the Ras superfamily and recently reported as a NF-κB pathway associated negative regulator, 15 to be a latent target of miR-128-3p through an online bioinformatics platform. We found that Rnd3 could be targeted by miR-128-3p directly, and silencing of miR-128-3p in mice increased the Rnd3 protein level and inhibited the protein expression level of p65, resulting in alleviating liver injury during I/R and reducing TNF-α and IL-6. In contrast, overexpression of miR-128-3p could decrease the Rnd3 protein level, thus leading to activation of the NF-κB pathway. However, this activation derived from miR-128-3p augmentation could be reversed by Rnd3 overexpression. In conclusion, the miR-128-3p/Rnd3/NF‐κB axis may provide new insight into the mechanisms of liver I/R injury.
Methods
Animals
Male C57BL/6 mice (mass 25–28 g; age 6–8 wk) were acquired from and fed in a specific-pathogen-free facility at Chongqing Medical University (Chongqing, PR China). All animal experiments were performed according to AVMA Guidelines for the Euthanasia of Animals (2013 Edition). According to the Guide for the Care and Use of Laboratory Animals, animal health and behaviour were monitored twice a d before operation and once an h after operation. The animal protocols were approved by the Animal Care and Use Committee of Chongqing Medical University.
Cell culture
Human embryonic kidney 293T (HEK293T) cells were bought from Cell Bank of the Type Culture Collection of the Chinese Academy of Sciences and cultured in DMEM (Gibco, Grand Island, NY) supplemented with 10% FBS and 1% penicillin and streptomycin (Gibco). The cultivation atmosphere of the cell incubator (Thermo Fisher Scientific, Waltham, MA) was set at 5% CO2 and a temperature of 37°C.
Target prediction and luciferase activity
Potential target genes and binding sites of miR-128-3p were predicted using the online tool TargetScan (www.targetscan.org). To verify whether miR-128-3p directly binds to Rnd3 mRNA, a dual luciferase reporter assay was performed according to the manufacturer’s instructions. Briefly, miR-128-3p agomir or miR-128-3p negative control (NC; Genepharma, Shanghai, PR China) was first transfected into 293T cells with the plasmid containing wild type (WT) 3′-UTR sequences of Rnd3 (p-MIR reporter plasmid; Ambion, Inc., Austin, TX) simultaneously for 6, 12, 24 and 48 h before estimating luciferase activity. Then, miR-128-3p agomir or miR-128-3p NC (Genepharma) was transfected into 293T cells with the plasmid containing WT or mutant 3′-UTR sequences, respectively, of Rnd3 (p-MIR reporter plasmid; Ambion, Inc.) simultaneously for 48 h before estimating luciferase activity.
Establishment of mouse liver I/R injury model
A 70% mouse liver I/R injury model was established according to a previous study. 16 Briefly, mice were anaesthetised with 0.1% pentobarbital sodium, and the arterial and portal vein of the middle and left lobe of the liver were interrupted with a vascular clamp for 1 h before reperfusion. Body temperature was maintained at 37°C using a warming pad. Sham groups were treated in the same way but without liver vascular clamping. Each mouse was 6–8 wk old and was treated with 1, 3, 6 and 9 h of reperfusion after 1 h of ischaemia. All animal-associated experiments were confirmed by the Animal Laboratorial Ethics Committee, Chongqing Medical University.
Transfection
Agomir, antagomir and NC of miR-128-3p (Genepharma) were transfected into mice via the tail vein according to the mouse experimental grouping 12 h before I/R treatment. Overexpression of Rnd3 using the adenovirus Rnd3 (ADV-Rnd3) was built according to NCBI Reference Sequence NM_028810.2 and was synthesised commercially (Genepharma). Transfection of Rnd3 was performed 1 wk before I/R treatment via mouse tail-vein injection. Blood and liver tissues were gathered after the reperfusion.
Quantitative RT-PCR
Liver tissues of I/R-treated mice were used for total RNA extraction using Trizol reagent (Takara, Dalian, PR China) following the manufacturer’s protocols. Complementary DNA (cDNA) was synthesised from total RNA using All-in-One cDNA Synthesis SuperMix (Bimake, Shanghai, PR China) or miRNA First Strand cDNA Synthesis (Sangon Biotech, Shanghai, PR China). Quantitative RT-PCR analysis of miR-128-3p and Rnd3 mRNA expression levels were performed following the manufacturer’s protocols. β-Actin was used as an internal normalisation control for mRNA levels, and U6 was used as an internal control for normalisation of miRNA levels. Primers for quantitative RT-PCR were designed and synthesised as follows: U6 forward, 5′-AGAGAAGATTAGCATGGCCCCTG-3′, reverse, 5′-ATCCAGTGCAGGGTCCGAGG-3′; miR-128-3p forward, 5′-CGGGCTCACAGTGAACCGG-3′, reverse, 5′-CAGCCACAAAAGAGCACAAT-3′; Rnd3 forward, 5′-CTATGACCAGGGGGCAAATA-3′, reverse, 5′-TCTCTTTGTCCTTTCGT-3′; β-actin forward, 5′-GGCTGTATTCCCCATCG-3′, reverse, 5′-CCAGTTGGTAACAATGCCATGT-3′. All data were analysed using the 2–ΔΔCT method.
Western blotting assay
Western blotting assay was performed following the manufacturer’s instructions. Briefly, total protein was extracted from supernatants of mouse liver tissues homogenised in RIPA Lysis Buffer (Beyotime, Shanghai, PR China). Then, after protein samples were loaded evenly and separated by 10% SDS-PAGE, the protein was transferred to polyvinylidene fluoride (PVDF) membranes. Subsequently, the PVDF membranes were blocked with skim milk (5%) dissolved in TBST for 2 h and washed with TSBT three times before overnight incubation a 4°C with primary Abs as follows: Rnd3 (1:1000; Abcam, Cambridge, UK), anti-NF-κB p65 (1:1000; Abcam) and β-actin (1:2500; Abcam). Finally, the membranes were washed and incubated with secondary Abs for 1 h as follows: goat anti-rabbit (1:4000; Beyotime) and goat anti-mouse (1:4000; Beyotime). The protein bands were visualised with an enhanced chemiluminescence system (Beyotime) in the dark and quantitated using Fusion (Bio-Rad, Hercules, CA).
ELISA for the detection of IL-6 and TNF-α levels
Serum levels of mouse IL-6 and TNF-α were detected using ELISA kits (Neobioscience, Beijing, PR China) following the manufacturer’s instructions.
Detection of aspartate aminotransferase and alanine aminotransferase levels
Aspartate aminotransferase (AST) and alanine aminotransferase (ALT) levels in mouse serum were detected according to the manufacturer’s instructions (JianCheng Bioengineering Institute, Nanjing, PR China).
Hematoxylin and eosin staining
Mouse liver tissues were fixed by formalin and then embedded with paraffin. Each sample was cut into 5 μm sections and stained with hematoxylin and eosin (H&E). The results were observed using a light microscope (Olympus, Tokyo, Japan) under 200× magnification. Suzuki’s criteria were used to evaluate the grade of the hepatic injury. 17
Statistical analysis
All data were analysed using SPSS Statistics for Windows v22.0 (IBM Corp., Armonk, NY) and are expressed as means ± SEM. Student’s t-test was used to compared samples between two groups, while ANOVA was used to compare differences among multiple groups. The correlation between two groups was measured using Pearson’s correlation analysis. For all experiments, each group involved five samples, and experiments were repeated five times. Significant differences were defined as P < 0.05.
Results
Rnd3 is negatively associated with miR-128-3p in a mouse liver I/R-treated model
The relationship between the Rnd3 and miR-128-3p mRNA expression level was investigated by quantitative RT-PCR using mouse liver tissues after I/R treatment. The results showed that 1 h after ischaemia, miR-128-3p mouse liver mRNA expression decreased as the reperfusion time went by, and eventually reached the bottom at 6 h (Figure 1a). The level of Rnd3 mRNA increased as the reperfusion time extended, and finally peaked after 6 h of reperfusion (Figure 1b). Then, Pearson’s correction test revealed that the Rnd3 mRNA expression level was negatively correlated with miR-128-3p (Figure 1c and d). Furthermore, the mouse serum IL-6, TNF-α, AST and ALT levels which underwent I/R treatment were demonstrated, and the results showed that 1 h ischaemia within 6 h of reperfusion was the optimal time for the I/R model for use in subsequent experiments (Figure 1e–h).
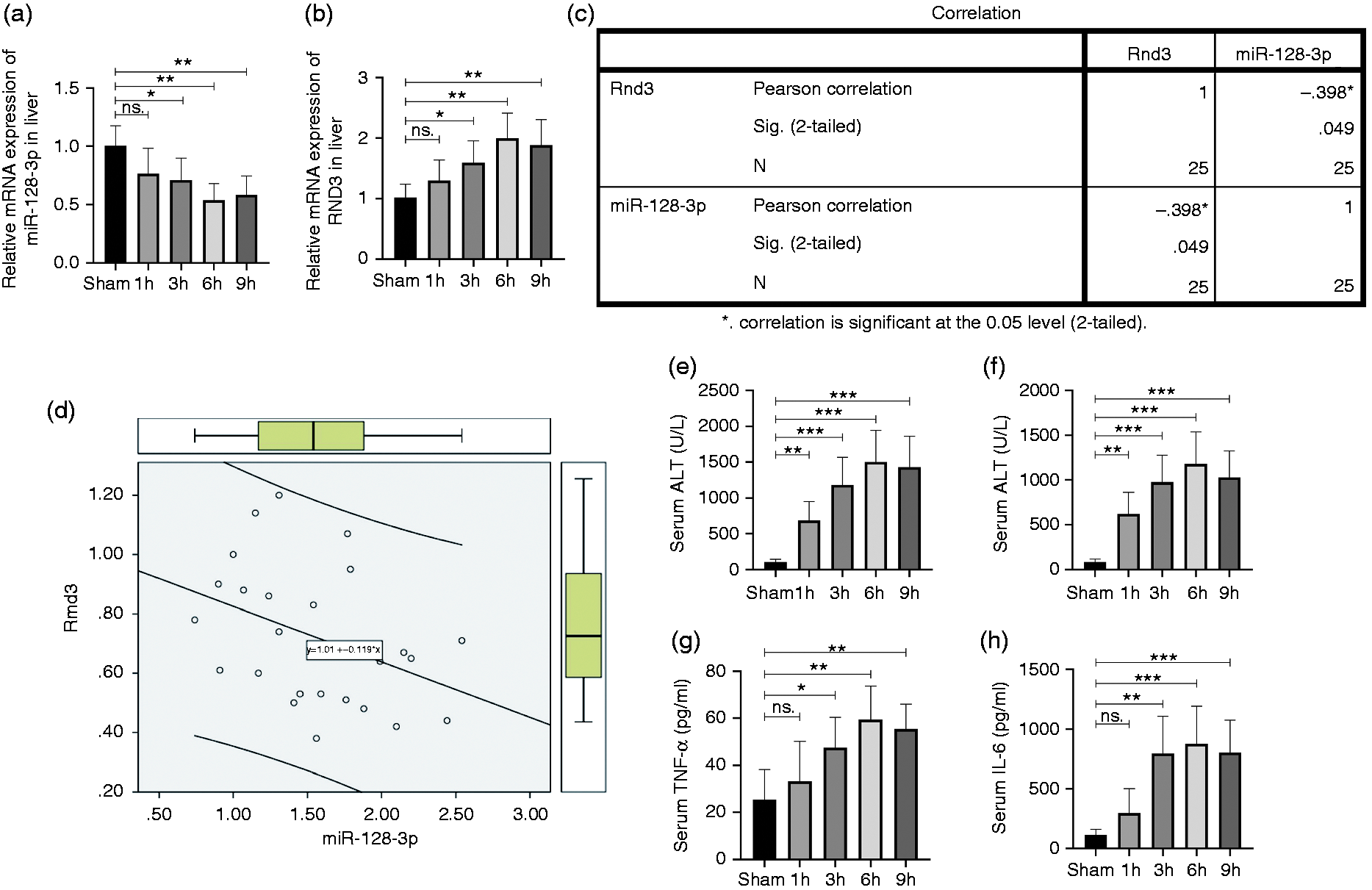
Rho family GTPase 3 (Rnd3) is negatively correlated with miR-128-3p in a mouse liver ischaemia–reperfusion (I/R)-treated model. (a) and (b) Relative miR-128-3p and Rnd3 microRNA (mRNA) levels in liver tissues. (c) and (d) Correlation analysis of mRNA expression levels of Rnd3 and miR-128-3p. (e) and (f) Serum level of aspartate aminotransferase (AST) and alanine aminotransferase (ALT) in a mouse liver I/R-treated model. (g) and (h) Serum level of TNF-α and IL-6. n = 5. *P < 0.05; **P < 0.01; ***P < 0.001; ns P>0.05.
Rnd3 is a direct target of miR-128-3p
Next, in order to examine whether miR-128-3p directly targets Rnd3, the online tool TargetScan v7.2 (www.targetscan.org) was used (Figure 2a). After mutating the supposed miR-128-3p binding site of Rnd3 from ACUGU to UGACA (Figure 2b), a luciferase reporter assay showed that in the WT group, the inhibitory ability of miR-128-3p agomir increased with time (Figure 2c), while this kind of suppression was completely erased in the mutant group (Figure 2d).
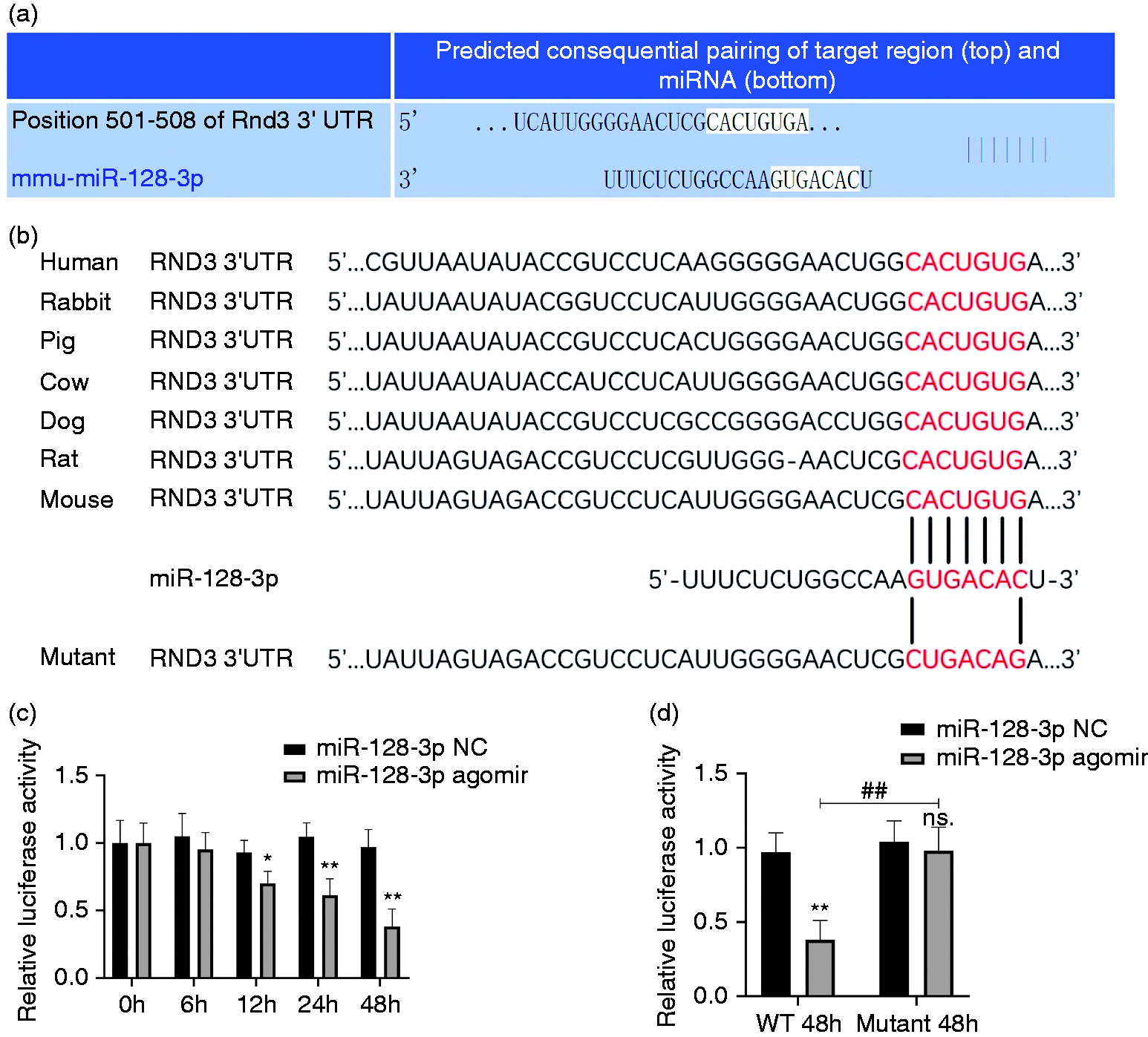
Rnd3 is a direct target of miR-128-3p. (a) Rnd3 was forecast as a potential target of miR-128-3p through the online tool TargetScan v7.2. (b) Wild type (WT) and mutant Rnd3 mRNA as a potential binding site of miR-128-3p. (c) Luciferase activity of the WT group was restrained in a time-dependent manner using miR-128-3p agomir. (d) A luciferase reporter assay demonstrated that miR-128-3p could bind straight to Rnd3 mRNA. *P < 0.05; **P < 0.01 vs. miR-128-3p negative control (NC) in WT group; ns P < 0.05 vs. miR-128-3p NC in mutant group; ##P < 0.01.
Inhibition of miR-128-3p could alleviate liver I/R injury in mice
Liver I/R-treated mice were separated in four groups – sham, I/R, miR-128-3p NC and miR-128-3p antagomir – in order to explore the effects of miR-128-3p in the model. Data from quantitative RT-PCR revealed that knockdown of miR-128-3p could significantly increase the mRNA level of Rnd3 (Figure 3a and b). Next, we found that the high mouse serum levels of TNF-α, IL-6, AST and ALT could be reversed by the inhibition of miR-128-3p (Figure 3c–f). Results of Western blotting showed that inhibition of miR-128-3p led to an increase in Rnd3 and a decrease in NF‐κB (p65) protein levels (Figure 3g–i). H&E staining was performed to assay liver histopathological characteristics, and the results proved that miR-128-3p inhibition decreased the histopathological changes (Figure 3j and k).
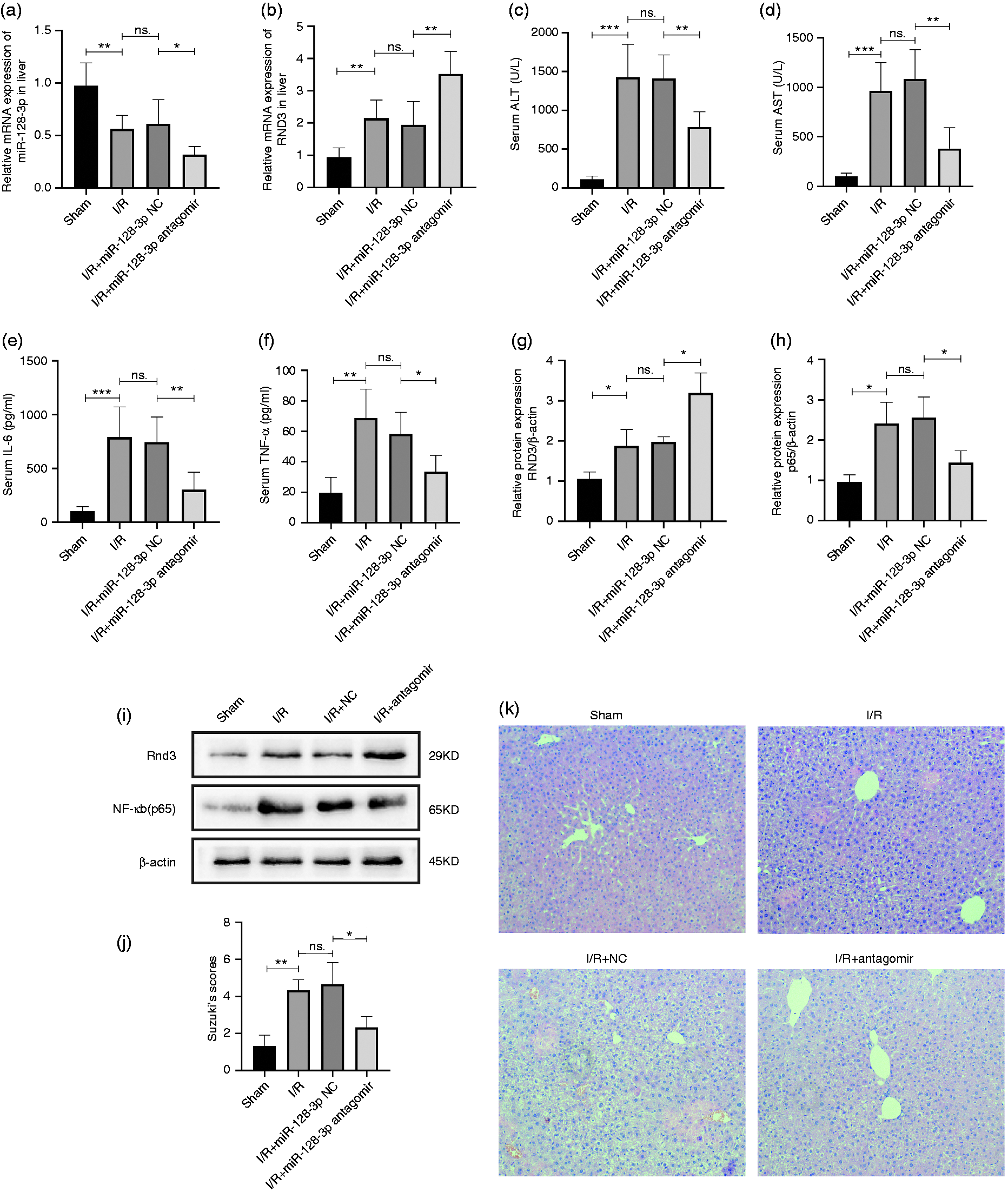
Inhibition of miR-128-3p could alleviate mouse liver I/R injury. (a) and (b) Relative miR-128-3p and Rnd3 mRNA levels in a mouse liver I/R model served with miR-128-3p antagomir. (c) and (d) Serum ALT and AST levels after being treated with miR-128-3p antagomir. (e) and (f) Serum TNF-α and IL-6 levels after being treated with miR-128-3p antagomir. (g)–(i) Relative Rnd3 and NF-κB p65 protein expression after being treated with miR-128-3p antagomir. (j) and (k) Histological changes of the liver in the sham, I/R, I/R + miR-128-3p NC and I/R + 3miR-128-3p antagomir groups. Original magnification ×200. n = 5. *P < 0.05; **P < 0.01; ***P < 0.001; ns P > 0.05.
miR-128-3p activates the NF-κB pathway by inhibiting Rnd3 expression in a mouse liver I/R model
Furthermore, miR-128-3p agomir, miR-128-3p NC, ADV-Rnd3 and ADV control were injected through the mouse caudal vein to verify their effectiveness (Figure 4a and b). After that, liver I/R-treated mice were separated in four groups: sham, I/R, I/R + miR-128-3p agomir and I/R + miR-128-3p agomir + ADV-Rnd3. Quantitative RT-PCR data revealed that ADV-Rnd3 could eliminate the inhibiting effect of miR-128-3p agomir on Rnd3 expression (Figure 4c). Further, Western blotting showed that Rnd3 overexpression could reverse the low expression of NF‐κB (p65) caused by miR-128-3p agomir (Figure 4h–j). The detection of mouse serum IL-6, TNF-α, AST and ALT also underlined the protective role of Rnd3 against miR-128-3p (Figure 4d–g). Finally, H&E staining showed that Rnd3 could decrease the histopathological alterations caused by miR-128-3p overexpression (Figure 4k and l).
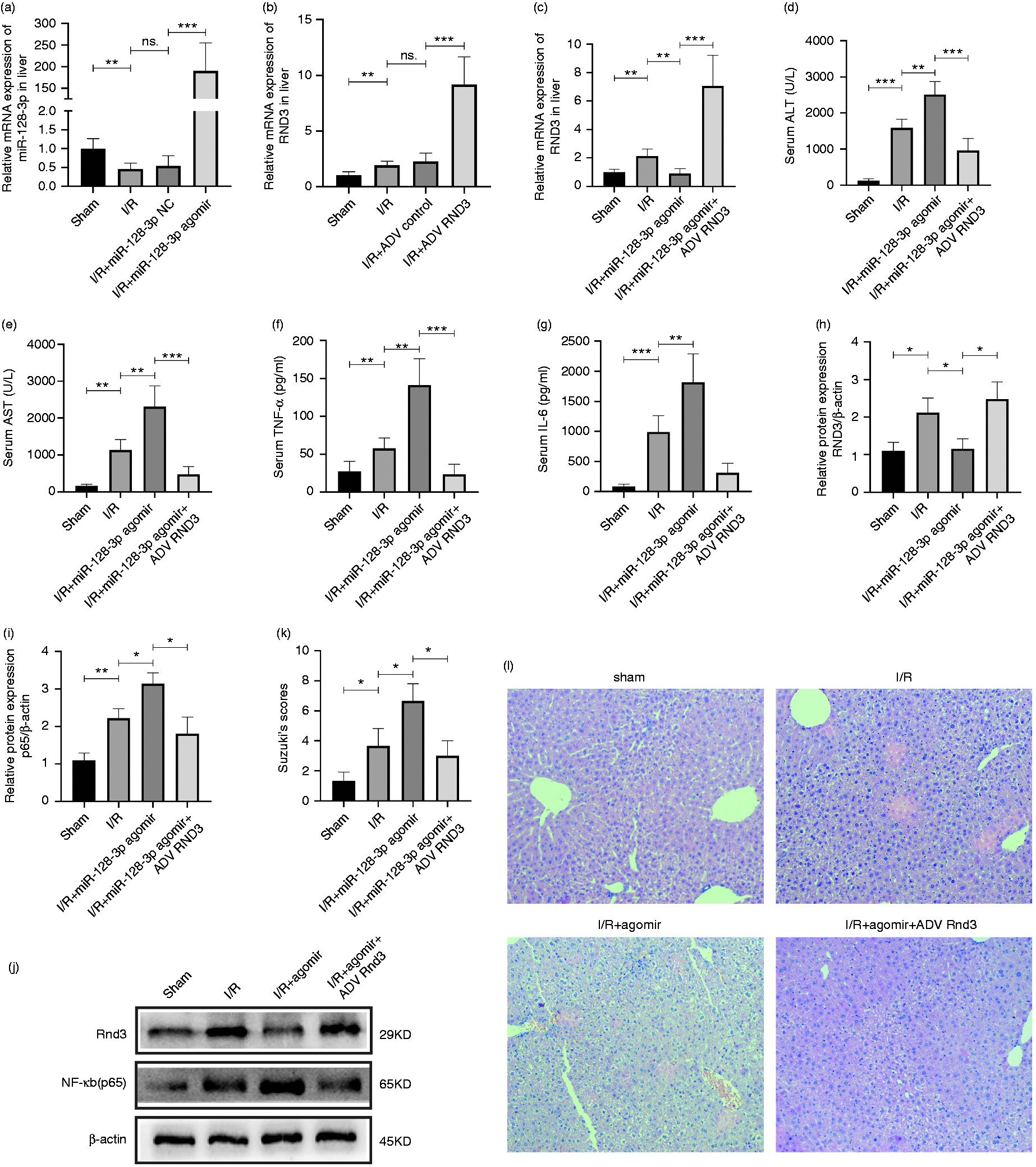
MiR-128-3p activates the NF-κB pathway by inhibiting Rnd3 expression in a mouse liver I/R-treated model. (a) Relative miR-128-3p mRNA level after being treated with miR-128-3p agomir and miR-128-3p NC. (b) Relative Rnd3 mRNA level after being treated with ADV-Rnd3 and ADV control. (c) Relative Rnd3 mRNA level in a mouse liver I/R model treated with miR-128-3p agomir and ADV-Rnd3. (d) and (e) Serum ALT and AST levels after being treated with miR-128-3p agomir and ADV-Rnd3. (f) and (g) Serum TNF-α and IL-6 levels after being treated with miR-128-3p agomir and ADV-Rnd3. (h)–(j) Relative Rnd3 and NF-κB (p65) protein expression in a mouse liver I/R model after being treated with miR-128-3p agomir and ADV-Rnd3. (k) and (l) Histological changes of the liver in the sham, I/R, I/R+miR-128-3p agomir and I/R+miR-128-3p agomir+ADV-Rnd3 groups. Original magnification ×200. n = 5. *P < 0.05; **P < 0.01; ***P < 0.001; ns. P > 0.05.
Discussion
Multiple researchers have shown that the mechanisms of I/R injury might provide a possible therapeutic target against fatal clinical problems such as liver dysfunction after transplantation.18–20 Herein, we offer some evidence that miR-128-3p could participate in mouse liver I/R injury progress by targeting Rnd3 while activating NF‐κB signalling. Our data showed that the knockdown of miR-128-3p led to the elevation of Rnd3 and silencing of NF‐κB signalling, which provides a possible underlying mechanism for liver I/R injury.
Recently, multiply miRNAs have been shown to be involved in liver I/R injury progress. 21 Tang and Tan reported that the lack of miR-155 could relieve liver I/R injury through targeting SOCS1, and may be associated with activation of M2 macrophage.22,23 The circulating level of miR-122 was recently reported as a novel biomarker of liver I/R injury24,25 and could inhibit apoptosis through the IGF-1R/AKT pathway in LO2 cells. 26 MiR-128-3p is a novel miRNA associated with multiple physiological processes. Cai et al. claimed that during non-small-cell lung cancer, TGF-β and the β-catenin pathway could be activated by miR-128-3p simultaneously to facilitate metastasis. 27 However, there has been increasing evidence to show that miR-128-3p also plays a part in the inflammatory response. Chen et al. declared that the inhibition of miR-128-3p could regulate the phosphorylation of p70s6k1 to protect human cardiomyocytes from I/R injury. 28 Zhao et al. reported that miRNA-128-3p could aggravate liver injury by promoting oxidative stress via targeting sirtuin-1. 29 Moreover, Wu et al. also found that miR-128-3p was contained in the inflammatory responses of BMSCs, 14 while Surmiak et al. uncovered that miR-128-3p was significantly reduced in granulomatosis with polyangiitis during the active phase. 30 Our study is the first to report the connection between miR-128-3p and liver I/R injury, and we went a step further to seek the underlying mechanism involved.
Rnd3, also known as RhoE, is a novel target gene reported in several mechanisms that could regulate physiological and pathological processes. Dankel et al.’s 31 research showed that obesity and insulin resistance might be partly associated with the diverse expression of adipose Rnd3. Zhu et al. 32 found that Rnd3 was an important regulator of the Notch1 pathway. Meanwhile, in the inflammatory area, Rnd3 was reported as a new fine-tuning factor regulating myocardial infarction-induced inflammation and promoted cardiac cell recovery. 15 Neubrand et al. 33 showed that Rnd3 might be an underlying neuroprotective phenotype in microglia, while Sun et al. 34 further showed that Rnd3 could inhibit NF‐κB signalling to induce glioblastoma multiforme cell apoptosis by binding p65 and promoting its ubiquitination, resulting in decreased p65.
NF-κB plays a well-known role in the survival and death of liver cells, the development and progression of cancer and the regulation of immune responses and inflammation.35,36 Recently, it was found that increased NF-κB expression led to a Forkhead box M1 (FOXM1) overexpression, which could enhance hepatic tumourigenesis. When FOXM1 is inhibited, it could restrain liver cancer cell growth in vitro and in vivo by regulating the expression of NF-κB element p65. 37 Another factor, hydrogen peroxide-inducible clone-5, was found to aggravate inflammation during hepatic I/R injury through the NF-κB signalling pathway. 38 In our study, we proved that by overexpression of miR-128-3p, the protein levels of Rnd3 were significantly reduced, while NF-κB element p65, NF‐κB signalling associated inflammatory factor TNF-α and IL-6, AST and ALT were significantly increased. Furthermore, these effects by miR-128-3p up-regulation could be reversed by Rnd3 overexpression. Our experimental data suggest that miR-128-3p might be capable of managing liver I/R injury through NF‐κB signalling by targeting Rnd3 and also hinted at the potential for liver cancer therapy by targeting NF‐κB signalling. However, the current study presents a negative-regulated relationship between Rnd3 and p65. Whether Rnd3 really reduces p65 expression in a p65-binding-dependent manner during liver I/R injury and whether there are other underlying mechanisms such as phosphorylation of NF-κB inhibitor alpha in the miR-128-3p/Rnd3/NF‐κB axis are subjects for future study.
Footnotes
Acknowledgements
We thank the Laboratory Research Centre in the First Affiliated Hospital of Chongqing Medical University for technical support and guidance.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The present study was funded by the National Natural Science Foundation of China (No. 81873592), the National Natural Science Foundation of China (No. 81703063) and the graduate tutor team construction project of Chongqing Municipal Education Commission Foundation, China (No. dstd201801).
