Abstract
The genetically tractable organism Caenorhabditis elegans is a powerful model animal for the study of host innate immunity. Although the intestine and the epidermis of C. elegans that is in contact with pathogens are likely to function as sites for the immune function, recent studies indicate that the nervous system could control innate immunity in C. elegans. In this report, we demonstrated that protein kinase A (PKA)/KIN-1 in the neurons contributes to resistance against Salmonella enterica infection in C. elegans. Microarray analysis revealed that PKA/KIN-1 regulates the expression of a set of antimicrobial effectors in the non-neuron tissues, which are required for innate immune responses to S. enterica. Furthermore, PKA/KIN-1 regulated the expression of lysosomal genes during S. enterica infection. Our results suggest that the lysosomal signaling molecules are involved in autophagy by controlling autophagic flux, rather than formation of autophagosomes. As autophagy is crucial for host defense against S. enterica infection in a metazoan, the lysosomal pathway also acts as a downstream effector of the PKA/KIN-1 signaling for innate immunity. Our data indicate that the PKA pathway contributes to innate immunity in C. elegans by signaling from the nervous system to periphery tissues to protect the host against pathogens.
Introduction
The innate immune system is the first line of defense against microbial infection and it is evolutionarily conserved in both vertebrate and invertebrate animals. 1 Activation of the innate immune system upon pathogen infection results in a species-specific antimicrobial response to invading microbes. Because several powerful molecular genetics tools are available in Caenorhabditis elegans, the worm is being developed as a tractable model for investigating the molecular basis of interactions between hosts and microorganisms.1–4 When worms encounter pathogens, the cuticle serves as a natural mechanical barrier to microbial infection, 5 and the nervous system has been shown to contribute to the recognition and behavioral avoidance of pathogenic bacteria.6–11 When pathogens cannot be avoided, worms synthesize different antimicrobial peptides, such as lysozymes, lectins, caenopores (amoebapore-like enzymes), antibacterial factor and neuropeptide-like peptides, to resist pathogens.12–15 Using the useful model, several signaling components such as PMK-1/p38 MAPK, 16 DAF-16, 17 HSF-1, 18 HIF-1, 19 the extracellular signal-regulated kinase MAPK MPK-1, 20 the protein kinase D DKF-2, 21 the G protein-coupled receptor FSHR-1 22 and the G protein Gqα EGL-30, 23 and SKN-1 24 have been identified to be involved in innate immunity.
The cAMP–protein kinase A (PKA) pathway is a second messenger-mediated signaling cascade that is evolutionarily conserved from highly evolved vertebrates to the simplest metazoans. In C. elegans, the PKA catalytic and regulatory subunits are encoded by kin-1 and kin-2, respectively. 25 The binding of cAMP to KIN-2 leads to its dissociation from the inactive holoenzyme and the release of active KIN-1. PKA/KIN-1 is involved in a variety of biological processes, including locomotion, exocytosis, rhythmic behavior, lipid distribution and reproduction in C. elegans.26–31 Previously, Alper et al. 32 screened more than 1000 genes by RNAi, and identified that 11 genes positively regulate the innate immune response both in murine macrophages and C. elegans. One of these genes encodes for adenyl cyclase (acy-1 in nematode and Adcy-9 in mouse), which is involved in production of cAMP, suggesting that the downstream molecule, PKA/KIN-1, is involved in the innate immune response to pathogens in C. elegans.
In this study, we investigated the role of the PKA/KIN-1 signaling in innate immunity in C. elegans. We found that the PKA/KIN-1 signaling protects worms against pathogenic bacterium Salmonella enterica. KIN-1 in the nervous system promotes the expression of a set of antimicrobial and lysosomal genes, which are required for innate immune responses to S. enterica.
Materials and methods
Nematode strains and cultivation
Standard conditions were used for C. elegans growth at 20℃.33,34 Mutant nematodes used in this study included strains kin-1(ok338)/mIs13I, BC14629(kin-1p::gfp), BC14666(dpy-5(e907)I;sEx14666)(lip1-1p::gfp), = and TU3401(sid-1(pk3321)V;uIs69V) for neuronal-specific RNAi and were kindly provided by the Caenorhabditis Genetics Center, which is funded by the NIH Office of Research Infrastructure Programs (P40 OD010440). The nematode strain for intestinal-specific RNAi (sid-1(qt9); Is[vha-6pr::sid-1];Is[sur-5pr:: GFPNLS]) were kindly provided by Dr. Gary Ruvkun (Massachusetts General Hospital, Harvard Medical School). Mutants were backcrossed three times into the wild type (WT) strain (N2) used in the laboratory. All strains were maintained on nematode growth medium (NGM) and fed with Escherichia coli OP50.
Construction of transgenic nematode strains
ilys-3p::gfp was generated by integrating 2 kb of the ilys-3 promoter fused to pPD95.75 that includes the unc-54 3'-UTR. The amplified primers were 5'-ACA TGC ATG CCG GGA AGC ATA AAA TCA TAA -3' and 5'-GCG TCG ACA ATA ACT GCA GAT GAA TTG ACA A-3 contained SphI and SalI restriction site. This construct was transformed into E. coli DH5α. The vector expressing ric-19p::kin-1 was constructed as follows. A 700 bp of ric-19 promoter fragment was obtained by PCR on C. elegans genomic DNA using primers 5'-ACA TGC ATG CCA TCG TTT TCG TGT GTT TTA TTC T-3' and 5'-GCG TCG ACA TCC CGC TGA AAT TTG GT-3' followed by SphI and SalI digestion. The fragment was inserted into pPD95.75 vector, resulting in the plasmid pPric. This construct was transformed into E. coli DH5α. kin-1 cDNA was amplified by PCR using primers 5'-GCG TCG ACA TGC TCA AGT TTC TGA AAC CAA-3' and 5'- GGG GTA CCT TAA AAC TCG GCA AAC TCT T-3'followed by SalI and KpnI digestion. The fragment was inserted into pPric vector, resulting in the plasmid expressing ric-19p::kin-1. This construct was transferred into E.coli DH5α. kin-1 overexpression was generated by integrating 2 kb of the kin-1 promoter and the kin-1 cDNA fused to pPD95.75 that includes the unc-54 3'-UTR. The primers for amplified kin-1 promoter were 5'-CCC AAG CTT CTA TTT CAG TAT TTC TTA CT-3' and 5'-CAT GCA TGC TGT TTT GCT GGT AGA CGA AG-3' and contained HindIII and SphI restriction sites. The primers for amplified kin-1 cDNA were 5'-CAT GCA TGC ATG CTC AAG TTT CTG AAA CC-3' and 5'-CGC GGA TCC AAA CTC GGC AAA CTC TTT GG-3' and contained SphI and BamHI restriction sites. These constructs were co-injected with the marker plasmid pRF4 containing rol-6(su1006) into the gonads of WT worms and kin-1(ok338) mutant worms by standard techniques. 35
RNA interference
The strains of E. coli used for RNAi were obtained from the Ahringer library.
36
RNAi feeding experiments were performed on synchronized L1 to L2 larvae at 20℃. Briefly, E. coli strain HT115(DE3) expressing dsRNA was grown overnight (12–16 h) in LB broth containing 100 μg/ml ampicillin at 37℃ and were then spread to NGM plates containing 100 μg/ml ampicillin and 5 mM isopropyl 1-thio-β-
Infection with pathogenic bacteria
E. coli OP50, S. enterica SL1344, and Pseudomonas aeruginosa PA14 were grown overnight in LB broth at 37℃, and Staphylococcus aureus NCTC8325 (a gift from Dr. Wen-Hui Lee, Kunming Institute of Zoology, Chinese Academy of Sciences) was grown overnight in tryptic soy broth (BD, Sparks, MD, USA) at 37℃ and then spread to NGM plates. Synchronized populations of worms were cultivated on E. coli OP50 at 20℃ until the young adult stage (i.e. within 12 h beyond the L4 stage). Fifty to 60 worms were transferred to NGM agar plates containing S. enterica SL1344, S. aureus NCTC8325 (gifts from Dr. WH Lee, Kunming Institute of Zoology, Chinese Academy of Sciences) and P. aeruginosa PA14 at 25℃. The number of living worms was counted at 12-h intervals. Immobile adult worms unresponsive to touch were scored as dead. Three plates were tested per assay and all experiments were performed three times independently.
Fluorescence microscopy analysis of GFP-labeled worms
After 24 h of S. enterica exposure, the transgenic worms carrying lipl-1p::gfp, and ilys-3p::gfp were immediately mounted in M9 onto microscope slides. The slides were viewed using a Zeiss Axioskop 2 plus fluorescence microscope (Carl Zeiss, Jena, Germany) with a digital camera. Fluorescence intensity was quantified by using the ImageJ software (NIH, Bethesda, MD, USA). Three plates of about 30 bacteria per plate were tested per assay and all experiments were performed three times independently.
Quantitative real-time PCR
Total RNA was extracted from worms with TRIzol Reagent (Invitrogen, Carlsbad, CA, USA) as previously described. 37 Random-primed cDNAs were generated by reverse transcription of the total RNA samples with SuperScript II (Invitrogen) and qPCR analysis was conducted using SYBR Premix-Ex TagTM (Takara, Dalian, China) on an Applied Biosystems Prism 7000 Sequence Detection System (Applied Biosystems, Foster City, CA, USA). The internal control pmp-3 was used as previously described. 37 The primers used for PCR are listed in Table S6.
Microarray analysis
Total RNA from each sample was quantified using the NanoDrop ND-1000 and RNA integrity was assessed using standard denaturing agarose gel electrophoresis. For microarray analysis, an Agilent Array platform was employed. The sample preparation and microarray hybridization were performed based on the manufacturer's standard protocols. Agilent Feature Extraction software (version 11.0.1.1) was used to analyze acquired array images. Quantile normalization and subsequent data processing were performed using the GeneSpring GX v12.1 software (Agilent Technologies). Differentially expressed genes were identified through Fold Change filtering. Gene Ontology (GO) analysis was applied to determine the roles of these differentially expressed genes played in these biological pathways or GO terms. The GEO accession number is GSE86342.
Quantification of intestinal bacterial loads
Synchronized populations of worms were cultivated on E. coli OP50 at 20℃ until the young adult stage. S. enterica GFP were grown in LB liquid medium containing ampicillin (100 μg/ml) at 37℃ overnight and plated onto NGM plates. Worms then were transferred to NGM agar plates containing S. enterica GFP (a gift from Dr. XP Qi, Kunming Institute of Zoology, Chinese Academy of Sciences) for 48 h at 25℃. 10 To eliminate the S. enterica GFP around the surface of worms, worms were transferred to NGM agar plate seeded with E. coli OP50 for 20 min for three times. 10 Ten worms were transferred into 50 μl PBS plus 0.1% Triton and ground. 10 The lysates were serially diluted by 10-fold in sterilized water and spread onto LB agar plates/ampicillin at 37℃. After 1 d of incubation at 37℃, colonies of S. enterica/GFP were counted. Five plates were tested per assay and all experiments were performed three times independently.
Western blotting
After worms were homogenized in liquid nitrogen, the homogenate was lysed on ice for 60 min in lysis buffer (BioTeKe, Beijing, China). The lysates of total protein were loaded (40 μg per well) and separated on a 10% SDS-PAGE. Proteins were then transferred to immobilon-PSQ transfer PVDF membrane (Millipore, Bedford, MA, USA). Primary Abs were monoclonal GFP Ab for detection of P62::GFP (#ab183734, 1:10000 dilution; Abcam, Shanghai, China), and anti-α tubulin Abs (#ab52866, 1:1000 dilution; Abcam). The secondary Ab was a peroxidase-coupled anti-rabbit IgG (1:20000 dilution; Abmart, Shanghai, China). Blots were developed using SuperSignal chemiluminescence substrate (Thermo Fisher Scientific, Waltham, MA, USA). An imaging system (Amersham Imager 600) was used for documentation of the western results. Band intensities were measured using ImageJ software (NIH).
Statistics
Differences in survival rates were analyzed using the log-rank test. Differences in gene expression, fluorescence intensity and the numbers of GFP::LGG-1 puncta were assessed by performing a one-way ANOVA followed by a Student–Newman–Keuls test. Data were analyzed using SPSS 16.0 (IBM, Armonk, New York, USA).
Results
PKA/KIN-1 is crucial for host defense against pathogenic bacteria
We found that kin-1(ok338) mutants were more sensitive to killing by S. enterica than WT worms (Figure 1a). H89 (10 μM), a specific inhibitor of PKA,
38
significantly reduced the survival of WT worms after S. enterica infection (Figure 1a). Furthermore, overexpression of kin-1 in the whole body enhanced the immune responses to S. enteric (Figure 1a). Taken together, our results demonstrated that KIN-1 is required for innate immunity of C. elegans to S. enterica infection. Furthermore, the PKA/KIN-1 pathway is also required by the innate immunity in C. elegans to resist other bacterial pathogens, such as P. aeruginosa PA14 and S. aureus NCTC8325 (Figure S1A, B).
PKA/KIN-1 functions in neurons to promote innate immune response to S. enterica. (a) kin-1(ok338) mutants were hypersensitive to killing by S. enterica. H89 (10 μM) reduced the survival rate of WT worms after S. enterica infection. Overexpression of kin-1 enhanced the immune response against S. enterica P < 0.001, relative to WT. (b) Intestinal-specific knockdown of kin-1 by RNAi did not affect sensitivity to S. enterica infection. (c) Neuronal-specific knockdown of kin-1 by RNAi reduced the survival rate of WT worms after S. enterica infection. P < 0.001 relative to WT. (d) Expression of kin-1 under the neuronal-specific ric-19 promoter restored resistance against S. enterica infection in kin-1(ok338) mutants. Neuronal kin-1overexpression increased the survival of S. enterica infection. P < 0.001, relative to WT kin-1(ok338).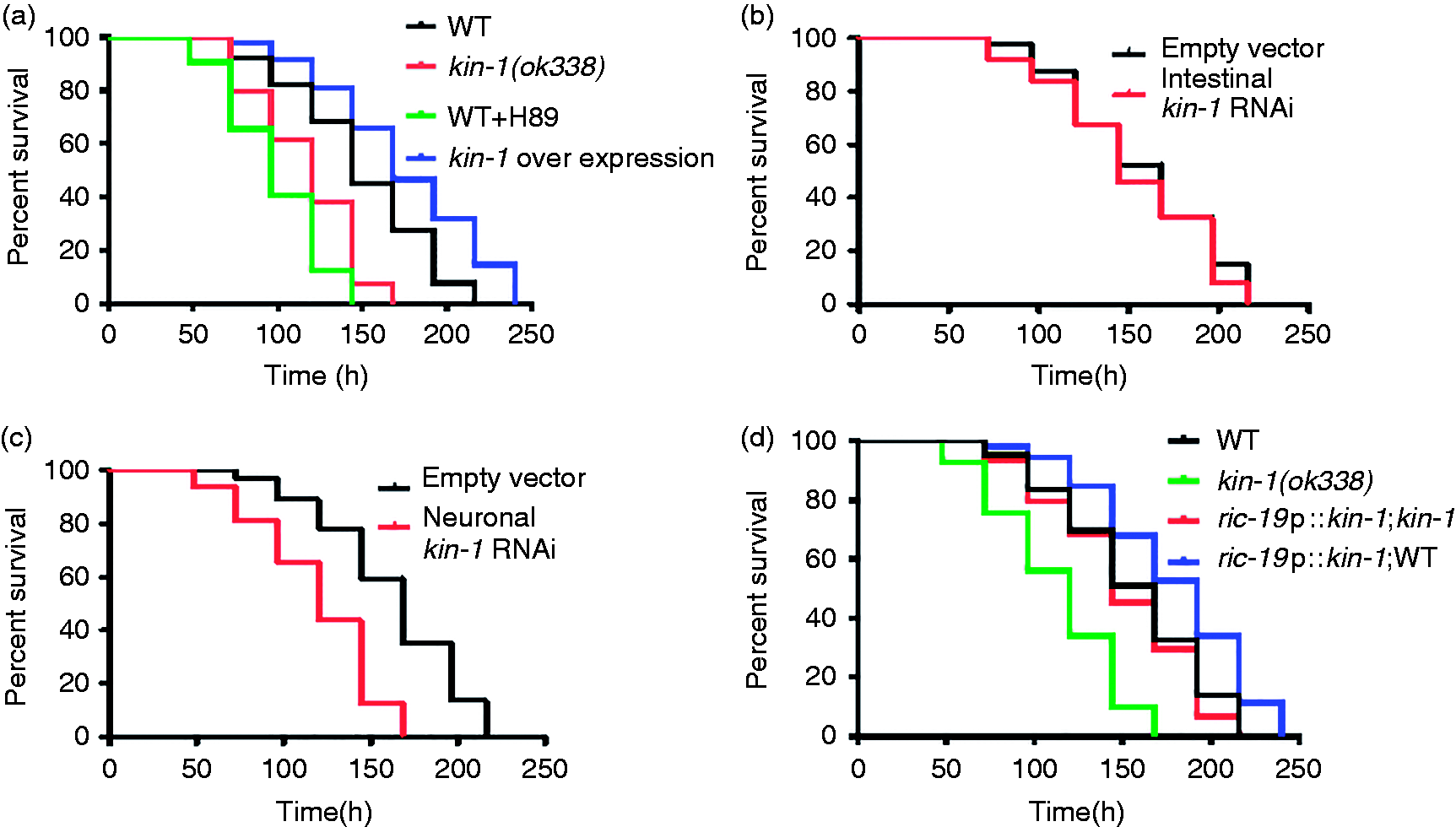
KIN-1 is expressed in the intestine, excretory cells and all neurons of nematodes.39,40 Using transgenic worms expressing kin-1p::gfp, we found that S. enterica infection did not influence the transcription activity of kin-1 (Figure S2). We found that intestinal-specific knockdown of kin-1 by RNAi did not affect sensitivity to S. enterica infection (Figure 1b). In contrast, neuronal RNAi of kin-1 reduced survival under S. enterica infection (Figure 1c). To further determine whether kin-1 acts in the neurons to regulate immunity, we expressed kin-1 under the control of the neuronal ric-19 22 in kin-1(ok338) mutants. We found that expression kin-1 in the neurons increased the survival of WT worms and fully rescued the survival of kin-1(ok338) mutants exposure to S. enterica (Figure 1d). In addition, we tested the lifespan of these strains. We found that the kin-1(ok338) mutants exhibited shortened lifespan, whereas overexpression of kin-1 under its promoter did not influence the lifespan in WT worms (Figure S3). Meanwhile, expression of kin-1 under the neuronal-specific ric-19 promoter did not influence the lifespan in WT worms (Figure S3). In contrast, neuronal-specific expression of kin-1 rescued the shorten lifespan in kin-1(ok338) mutant worms. These results suggest that KIN-1 function in the neurons is required for innate immunity and for longevity determination.
PKA/KIN-1 controls the expression of a set of antimicrobial genes involved in host defense
To gain further insight into the mechanism underlying PKA/KIN-1-mediated innate immunity, we used microarray analysis to study the transcriptional profile of WT relative to kin-1(ok338) animals after 24 h of S. enterica infection. The microarray analysis revealed that 4092 genes were up-regulated and 3232 genes downregulated in WT animals, defining the transcriptional host response to S. enterica infection (Tables S1–S4). Of these up-regulated genes, 488 (11.9%) were kin-1-dependent (Figure 2a; Tables S3 and S5). Of these down-regulated genes, 538 (16.6%) were kin-1-dependent (Figure 2a; Tables S4 and S5).
KIN-1 is involved in expression of host-response components. (a) Venn diagrams of genes that are up-regulated (left) or down-regulated (right) by S. enterica infection and positively regulated by kin-1. (b) Over-represented functional categories of KIN-1-dependent S. enterica-up-regulated genes. FDR: false discovery rate.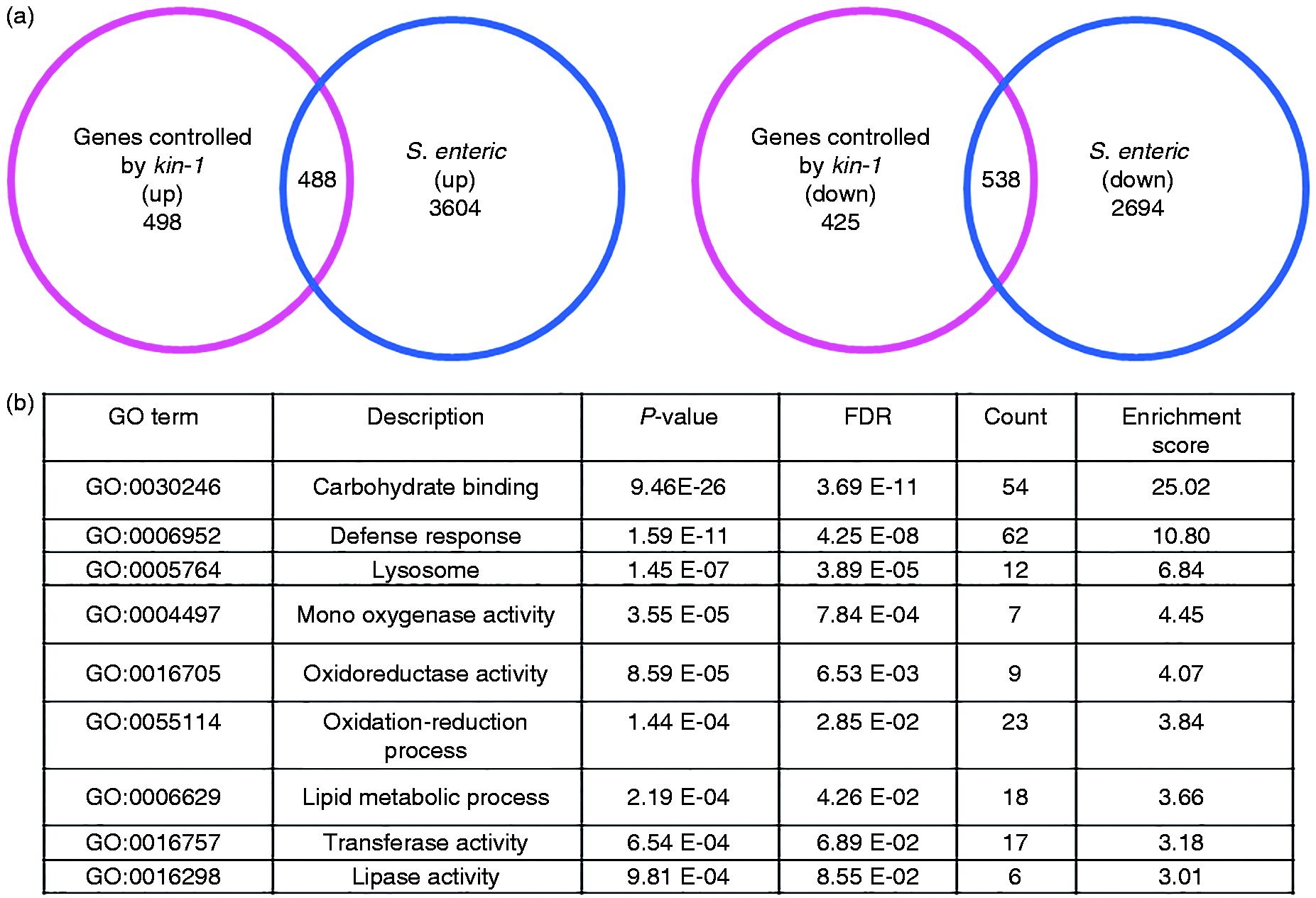
Gene Ontology (GO) cluster analysis of kin-1 up-regulated genes revealed that carbohydrate binding, defense responses, lysosome, mono-oxygenase activity, oxidoreductase activity, lipid metabolic process, transferase activity and lipase activity were more enrichment (Figure 2b; Table S3). KEGG pathway analysis indicated that the most over-represented KEGG pathway is the one categorized as ‘lysosome’ (Figure S4). Of these genes, kin-1 regulated a ‘defense response’ component, including genes that encode proteins with antimicrobial activity, such as lysozymes, caenopores, pqn/abu genes, C-type lectins and caenacins (Figure 3a; Tables S3).10,12 Using quantitative real-time PCR (qRT-PCR), we detected four of these antimicrobial genes (ilys-3, spp-1, spp-6, spp-20) in this group and demonstrated that the mRNA levels of these antimicrobial genes were increased after S. enterica infection (Figure 3b). Mutation kin-1 suppressed the up-regulation of these antimicrobial genes induced by S. enterica infection. Furthermore, rescue of kin-1 in the neurons markedly restored their expression (Figure 3b). Neuronal-specific knockdown of kin-1 also inhibited the expression of these antimicrobial genes by infected S. enterica (Figure S5). Finally, using transgenic animals expressing ilys-3p::gfp, we found that exposure to S. enterica led to the induction of ilys-3p::gfp in the intestine (Figure 3c). In contrast, the expression of ilys-3p::gfp was inhibited by knockdown of kin-1.
KIN-1 regulates a set of antimicrobial genes after S. enterica infection. (a) Hierarchical clustering of kin-1-dependent antimicrobial genes in worms after S. enterica infection. (b) The expression of four antimicrobial genes was determined by qPCR in worms in the presence and absence of S. enterica (SM). Expression of kin-1 under the neuronal-specific ric-19 promoter restored expression of these genes in kin-1(ok338) mutants. These results are means ± SD of four experiments. aP < 0.05 vs. WT (WT)+OP50; bP < 0.05 vs. kin-1+SM. (c) Expression of ilys-3::gfp was reduced by kin-1 RNAi after S. enterica infection. The right panel shows quantification of GFP levels. These results are means ± SD of three experiments. aP < 0.05 vs. EV (empty vector); bP < 0.05 vs. SM.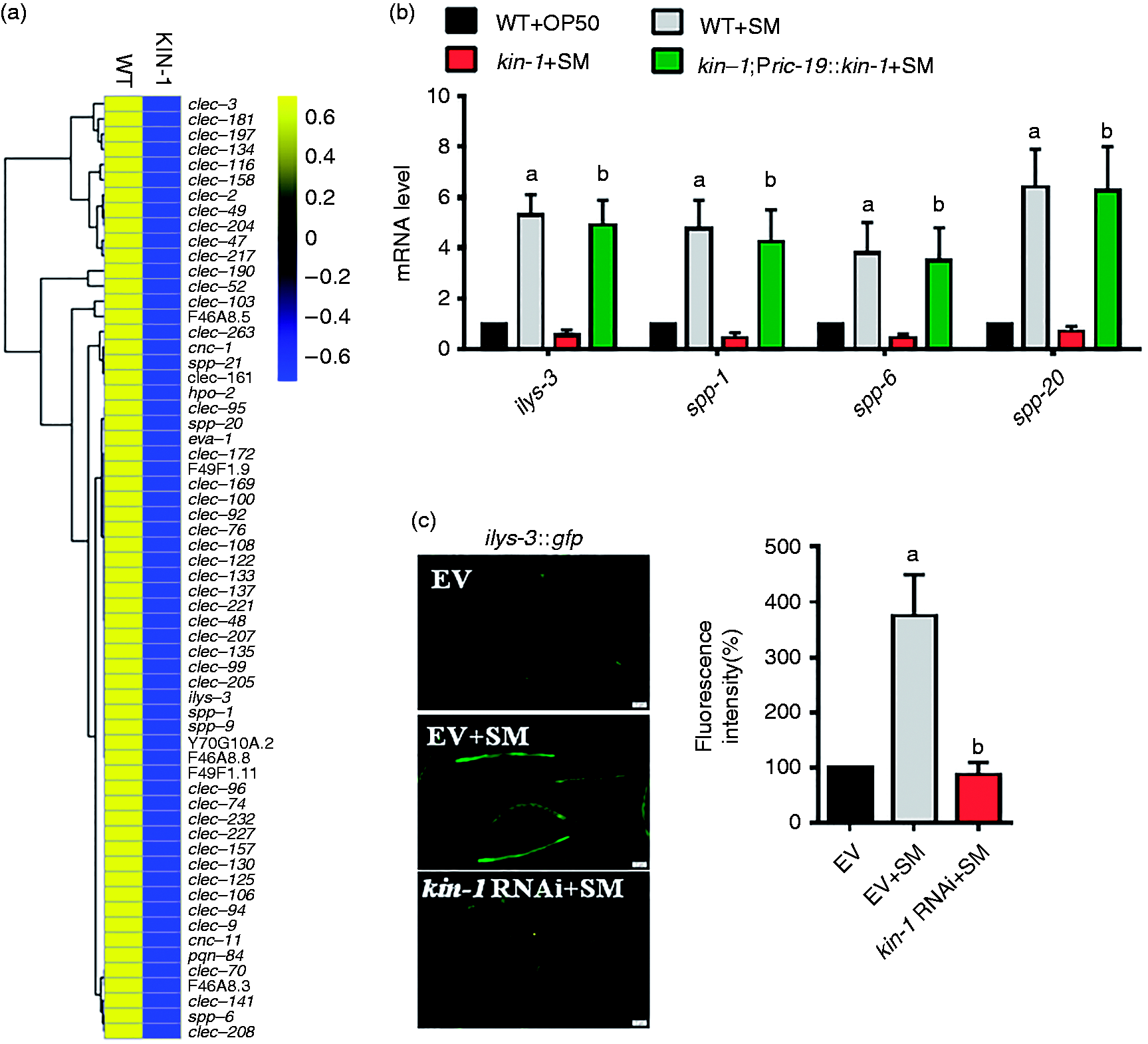
Next, we found that knockdown of ilys-3, spp-1, spp-6 and spp-20 by RNAi in WT increased the susceptibility to S. enterica, but not further shorten the survival in kin-1(ok338) mutant worms (Figure 4a–d). Taken together, these results suggest that antimicrobial genes controlled by kin-1 are essential to host defense response.
Antimicrobial genes are required for innate immunity. (a–d) Knock-down of (a) ilys-3, (b) spp-1, (c) spp-6 and (d) spp-20 resulted in significantly enhanced sensitivity to S. enterica infection, but not further shorten the survival in kin-1(ok338) mutant worms. P < 0.05 relative to WT + empty vector (EV).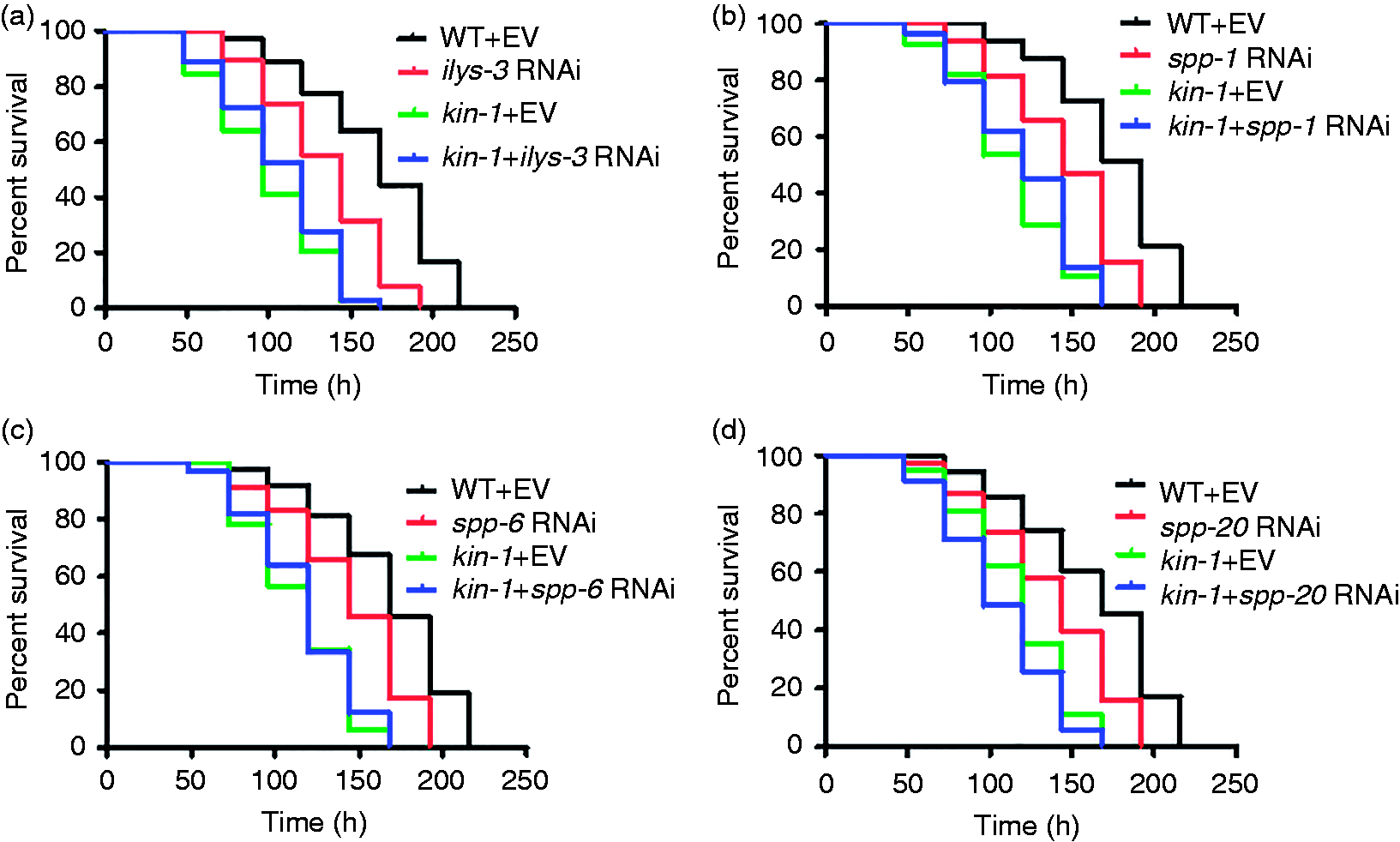
Lysosomal signaling pathway is required for host defense
In addition to antimicrobial genes, kin-1 was involved in the induction of lysosomal genes after S. enterica infection (Figure 5a; Table S3). These genes included lipl-1 and lipl-3 (ortholog of human lipase LIPF), asm-2 and asm-3 (ortholog of human sphingomyeline phosphodiesterase SMPD1), cpr-2, cpr-5, cpr-8/W07B8.1 and F32H5.1 (ortholog of human lysosomal cathepsin B CTSB), gln-3 (ortholog of human glutamate-ammonia ligase GLUL), hex-1 (β-N-acetylhexosaminidase), tre-5 (ortholog of human trehalase TREH) and hrg-1 (ortholog of vertebrate heme transporters). Next, we used qRT-PCR to verify the induction of lysosomal genes, such as lipl-1, lipl-3, cpr-5, asm-2, asm-3, tre-5, hrg-1, gln-3, hex-1 and F32H5.1. The mRNA levels of these lysosomal genes in kin-1(ok338) mutants were significantly lower than those in WT worms after S. enterica infection (Figure 5b). Rescue of kin-1 in the neurons significantly restored their expression in kin-1(ok338) mutants (Figure 5b). Neuronal-specific knockdown of kin-1 also inhibited the expression of these lysosomal genes induced by S. enterica (Figure S6). Furthermore, the expression of lipl-1p::gfp in the intestine was markedly induced by S. enterica (Figure 5c). In contrast, the expression of lipl-1p::gfp was inhibited by knockdown of kin-1.
KIN-1 regulates a set of lysosomal genes after S. enterica infection. (a) Hierarchical clustering of kin-1-dependent lysosomal genes in worms after S. enterica (SM) infection. (b) The expression of 10 antimicrobial genes was determined by qPCR in worms in the presence and absence of S. enterica. Expression of kin-1 under the neuronal-specific ric-19 promoter restored expression of these genes in kin-1(ok338) mutants. The results are means ± SD of four experiments. aP < 0.05 vs. WT (WT)+OP50; bP < 0.05 vs. kin-1+SM. (c) Expression of lipl-1p::gfp was reduced by kin-1 RNAi relative to WT animals after S. enterica infection. The right panel shows quantification of GFP levels. These results are means ± SD of three experiments. aP < 0.05 vs. EV (empty vector); bP < 0.05 vs. SM.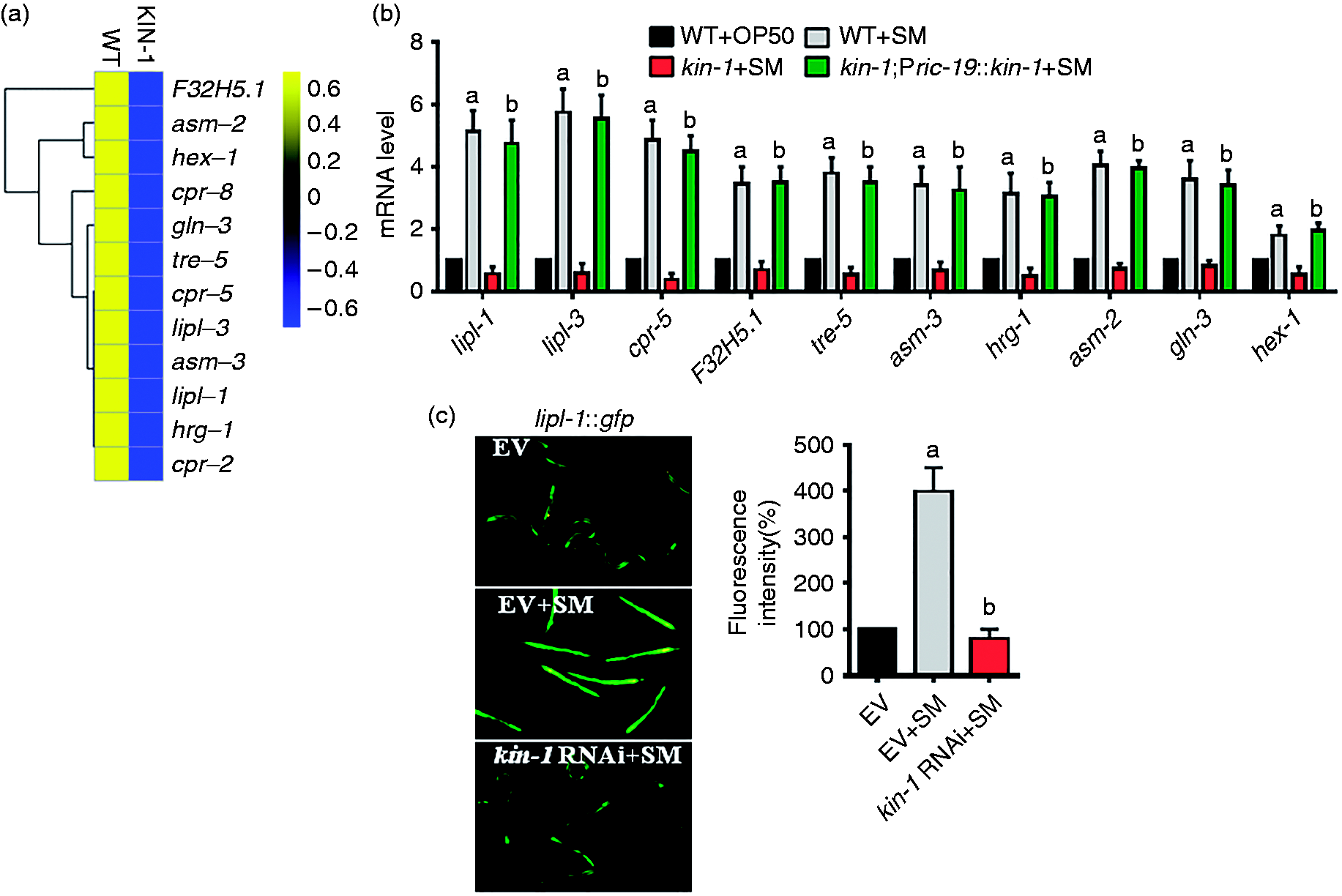
To test the role of the lysosomal signaling in C. elegans defense against S. enterica, we silenced some of these genes by RNAi. We found that knockdown of lipl-1, lipl-3, cpr-5, F32H5.1, asm-3, tre-5 and hrg-1 reduced the survival of WT worms but did not further shorten the survival in kin-1(ok338) mutants (Figure 6a–g). In contrast, knockdown of hex-1 did not alter the innate immunity in C. elegans (Figure 6h). Taken together, these results indicate that the lysosomal signaling is required for innate immunity in C. elegans.
Lysosomal genes are required for innate immunity. (a–g) Knockdown of (a) lipl-1, (b) lipl-3, (c) cpr-5, (d) tre-5, (e) asm-3, (f) hrg-1 and (g) F32H5.1 reduced the survival of WT worms but did not further shorten the survival in kin-1(ok338) mutants after S. enterica infection. P < 0.05 relative to WT + empty vector (EV). (h) Knockdown of hex-1 did not alter the innate immunity in C. elegans.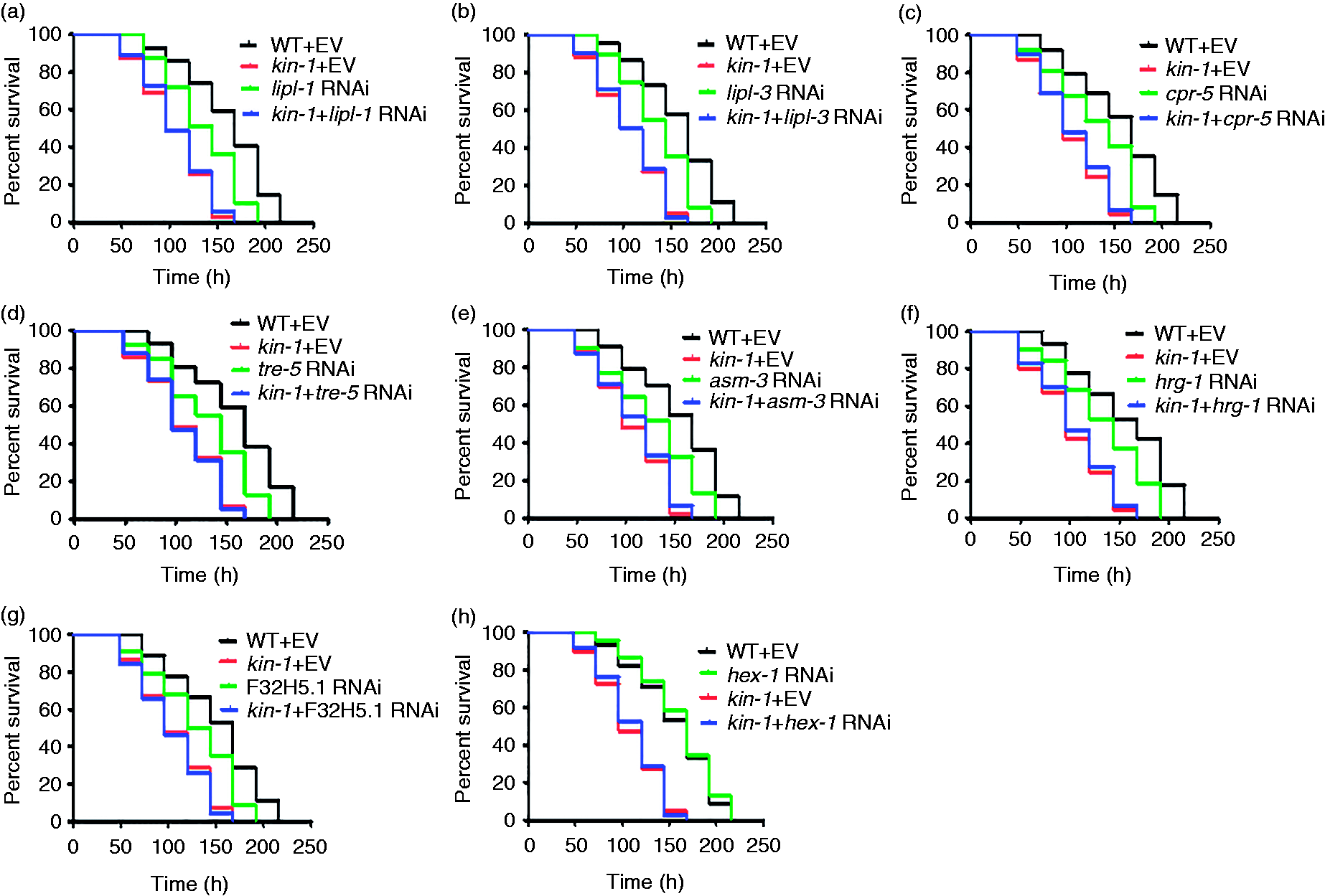
PKA/KIN-1 is involved in innate immunity by regulating autophagic flux
Autophagy plays a crucial role in host defense against S. typhimurium infection in C. elegans.
41
As lysosome plays an essential role in autophagy, we monitored autophagy using transgenic worms carrying GFP::LGG-1, a reliable indicator of autophagy.
41
As expected, exposure of WT worms to S. enterica increased the abundance of GFP::LGG-1-positive punctae in the seam cells and intestine compared with WT worms fed E. coli OP50 (Figure 7a, b). However, knockdown of kin-1 did not influence the number of GFP::LGG-1 punctae in worms in the presence or absence of S. enterica. These results suggest that kin-1 is not required for formation of autophagosomes.
KIN-1 is involved in autophagy by regulating autophagic flux. (a, b) Knockdown of kin-1 did not affect formation of autophagosomes. The numbers of GFP::LGG-1-positive puncta were counted in the (a) intestinal cells and the (b) seam cells of worms exposed to S. enterica (SM). *P < 0.01 vs. empty vector (EV). (c) Knockdown of kin-1 suppressed p62 degradation. Expression of P62::gfp was enhanced by kin-1 RNAi relative to WT animals after S. enterica infection. The blot is typical of three independent experiments. The lower panel shows quantification of p62 levels. P < 0.05 kin-1 RNAi vs. EV; P < 0.05 kin-1 RNAi+SM vs. EV+SM. These results are mean ± SD of three independent experiments. (d) The CFUs of S. enterica in kin-1(ok338) worms were markedly increased compared with those in WT worms at 48 h post-infection. Overexpression of kin-1 under its promoter or the neuronal-specific ric-19 promoter did not influence CFU in WT worms. In contrast, the neuronal-specific expression of kin-1 reduced CFU in kin-1(ok338) mutant worms. These results are mean ± SD of three independent experiments. aP < 0.05 vs. WT; bP < 0.05 vs. kin-1(ok338).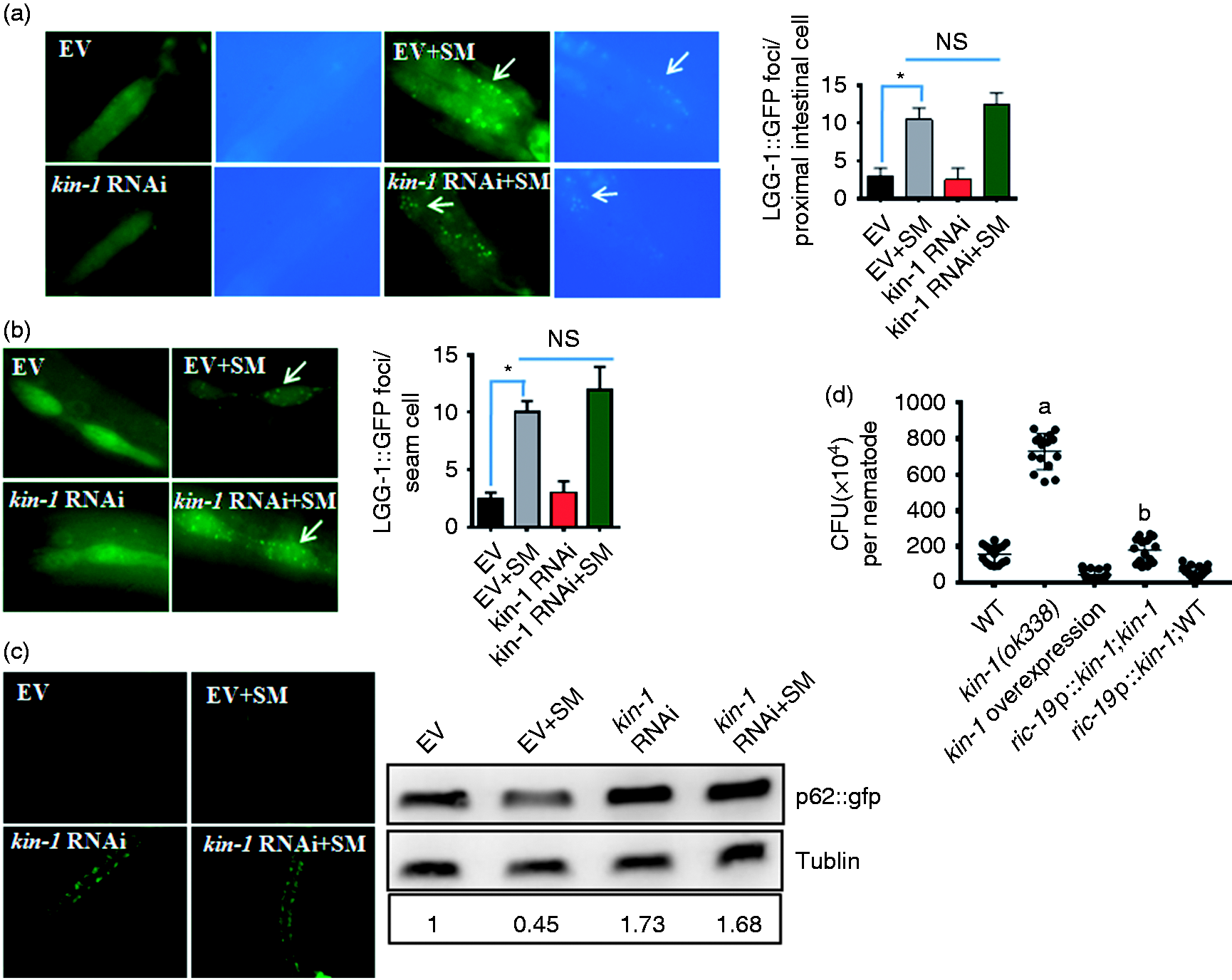
As kin-1 regulates the expression of lysosomal genes, we tested the effect of kin-1 on autophagic flux by detecting p62/SQSTM1 in transgenic worms carrying p62::GFP. The amount of p62/SQSTM1 increases when autophagy process is impaired and decreases when autophagy is induced. 42 We found that exposure of WT worms to S. enterica reduced the expression of p62::gfp vs. WT worms fed E. coli OP50 (Figure 7c). Interestingly, kin-1 RNAi significantly increased the expression of p62::gfp, both in worms with and without S. enterica infection, similar results confirmed by Western blotting, implicating a defective autophagic flux.
As autophagy is crucial for S. typhimurium clearance in worms, we tested whether impaired autophagic flux by kin-1 mutation influences pathogen load. The colony forming units (CFU) of bacteria cells in kin-1(ok338) worms were markedly higher than those in WT worms (Figure 7d). Overexpression of kin-1 under its promoter or the neuronal-specific ric-19 promoter did not influence CFU in WT worms after S. enteric infection. In contrast, expression of kin-1 under the neuronal-specific ric-19 promoter reduced CFU in kin-1(ok338) mutant worms (Figure 7d). Thus, KIN-1 in the neurons inhibits pathogen accumulation to enhance innate immunity in C. elegans.
Discussion
We identified PKA/KIN-1 as a critical kinase for host defense against pathogenic bacteria. PKA/KIN-1 mediates the upregulation of a set of antimicrobial genes upon exposure to S. enterica, indicating that the kinase is responsive to the presence of pathogen. Meanwhile, PKA/KIN-1 controls the expression of a set of lysosomal genes. These genes are involved in worm resistance to S. enterica, probably by controlling autophagic machinery.
In C. elegans, antimicrobial genes are regulated by several innate immune pathways after bacterial infection, such the p38 MAPK/PMK-1 pathway, 43 the DKF-2/PKD pathway, 21 and the HLH-30 signaling. 44 In this study, we identified that PKA/KIN-1 up-regulation a set of antimicrobial genes such as lysozymes, caenopores, pqn/abu families, C-type lectins and caenacins. The corresponding proteins are involved in many facets of innate immunity. Caenopores, which belong to the SAPLIP (saposin-like protein) superfamily, exhibit antimicrobial activities, probably through forming pores in bacterial membranes. 45 ILYS-3 can lyse both Gram-positive and Gram-negative bacteria by hydrolyzing β-(1→4)-glycosidic bonds in the glycan moiety of peptidoglycan. 46 C-Lectins, which can bind to bacterial oligosaccharides, are probably involved in the recognition of pathogens and the initiation of a protective immune response in C. elegans.47,48 Our results demonstrate that silence of lysozyme or caenopore genes is more susceptibility to S. enterica than WT animals, indicating that antimicrobial genes are required for enhancing endurance to pathogen infection. Thus, antimicrobial genes act in downstream effects of PKA/KIN-1 signaling in response to pathogen infection.
Autophagic machinery is capable of sequestering invading bacteria, and targeting these pathogens for lysosomal degradation, thus providing a mechanism for the clearance of intracellular bacteria. For example, cellular infection by a pathogenic bacterium Listeria monocytogenes promotes an autophagic response, thereby inhibiting the bacterial growth in fibroblasts. 49 Autophagy in the intestinal epithelium, which is activated by S. Typhimurium infection, is critical for limiting extra-intestinal spread of the bacterium in mice. 50 In C. elegans, autophagy is involved in innate immunity by limiting intracellular infection with S. Typhimurium. 41 Our data demonstrate that kin-1 is not required for formation of autophagosomes; instead, it controls the expression of a set of lysosomal genes. Lysosomes are ubiquitous organelles, which play a crucial role in sustaining cellular homeostasis. As a stage of autophagic process, the autolysosome is generated by fusion of autophagosomes with lysosomes. 42 Consistent with the observation, a mutation in kin-1(ok338) led to a marked increase in accumulation of S. enterica. Meanwhile, knockdown of the lysosomal genes reduces the survival of worms exposed S. enterica. These results support the idea that the mechanism underlying autophagy-mediated innate immunity in C. elegans is the clearance of the intracellular bacteria. 41 Collectively, the lysosomal pathway functions as the downstream effectors of the PKA signaling for defense response.
C. elegans does not have specialized immune cells, and the tissues that are in contact with pathogens are likely to act as sites for immune function.7,10 It is believed that the pathogenic bacteria infect the intestine of nematode. Although PKA/KIN-1 is expressed in multiple tissues, including intestine, excretory cell and neurons,39,40 the kinase performs its immune function in the neurons but not in the intestine. Of the antimicrobial and lysosomal genes up-regulated by PKA/KIN-1, most of them are expressed in the intestine, epidermis, and muscle. As C. elegans expresses at least 12 different isoforms of the catalytic subunit of PKA,51,52 the specific isoform(s) in the neurons probably is involved in innate immunity. Accumulating evidence indicates nervous systems that respond to environmental cues have the potential to control innate immune responses to pathogens.7,10 For example, OCTR-1, a putative octopamine G protein-coupled catecholamine receptor, functions in sensory neurons to suppress innate immune responses by down-regulating the expression of pqn/abu genes in non-neuronal tissues. 10 How does PKA/KIN-1 act in the nervous system to promote pathogen resistance? PKA/KIN-1 in the neurons probably releases an unknown signal to control the expression of these antimicrobial and lysosomal genes, which are mainly expressed in non-neuronal body tissues. Thus, our results provide an insight into the mechanism that a non-autonomous cell signal from the nervous system may control the downstream signaling to boost innate immunity in the tissues that serve the primary role in host defense against pathogens.
Footnotes
Acknowledgements
We thank the Caenorhabditis Genetics Center, and Dr. G Ruvkun for nematode strain, and Drs. W.H. Lee and X.P. Qi for bacterial strains.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported, in part, by grants (2013CB127500) from National Basic Research Program of China (to KQZ), grants from the National Natural Science Foundation of China (81471916) and Ministry of Education of the People's Republic of China (213035A) (to CGZ).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
