Abstract
Feeding a yeast-containing additive (YCA; OmniGen-AF) improves immune responses in ruminant livestock and reduces subsequent production losses. The objective was to identify molecular pathways by which dietary YCA may modify immune responses using a rodent model. Thirty-seven healthy, unchallenged CD rats received a diet containing 0 (control; n = 5, only 28 d), 0.5% (n = 15) or 1% (n = 17) YCA for 7 (n = 4/group), 14 (n = 3 or 4/group), 21 (n = 3 or 4/group) or 28 (n = 5/group) d. At the end of the feeding periods, whole blood was collected and the isolated RNA was analyzed for the expression of 84 genes involved in innate and cell-mediated adaptive immune responses. Three bacterial pattern recognition receptors TLR1 (0.5%: + 2.01; 1%: + 2.38), TLR6 (0.5%: + 2.11; 1%: + 2.34) and NOD2 (0.5%: + 2.32; 1%: + 2.23), two APC surface receptors CD1D1 (0.5%: + 1.75; 1%: + 2.33) and CD80 (0.5%: +2.45; 1%: +3.00), and the cell signaling molecule MAPK8 (0.5%: +1.87; 1%: +2.35) were significantly up-regulated by YCA at both inclusion rates. In conclusion, feeding YCA may potentially increase recognition and responses to bacterial pathogens and T-cell activation and differentiation and thereby maintain health and prevent production losses.
Introduction
Feeding livestock yeast-containing supplements has increased in popularity as a management tool for prevention of diseases and subsequent production losses. Limited information, however, is available on how feed additives affect gene expression profiles associated with the innate and cell-mediated adaptive immune system, and dose–response studies have not been conducted. We documented in several studies that feeding a yeast-containing additive (YCA; OmniGen-AF; Prince Agri Products, Inc., Quincy, IL, USA) at the recommended inclusion rate (0.25–0.50 wt% of diet) promotes immune function and maintains health and production in immune- or stress-challenged ruminant species and rodent models.1–6 For example, Holstein heifers supplemented for 30 d with YCA had significantly increased surface-bound and internalized Escherichia coli in neutrophils. 5 Feeding YCA to the same animals tended to increase binding and uptake of Staphylococcus aureus by neutrophils. Changes in neutrophil function were linked to increased gene expression of the lymphocyte adhesion molecule L-selectin (i.e. CD62L) in blood leukocytes. We had previously confirmed that YCA increased L-selectin in neutrophils on the protein and gene expression level and that those changes were linked to increased circulating concentrations of both neutrophils and lymphocytes. 1 Dairy cows supplemented for 5 wk with YCA had increased milk production, decreased incidence of mastitis and decreased somatic cell counts in milk, 4 and dairy cows supplemented for 60 d with YCA had increased gene expression of the neutrophil-activating chemokine receptor IL-8 R and the lymphocyte adhesion molecule L-selectin in whole blood. 6
After we had observed that feeding YCA increased responses to several bacterial pathogens across various species, we believed that the effect of feeding YCA on the immune system is more general than can be explained solely by the response of L-selectin and IL-8 R to YCA. Therefore, we hypothesized that YCA at 0.5% of diet increased expression of genes involved in several pathways involved in innate and cell-mediated adaptive immune responses in whole blood during a 28-d supplementation period and that higher inclusion rates (1% of diet) than currently recommended are not required in healthy, unchallenged animals. To test our hypothesis, we used healthy, unchallenged male rats as an animal model. Rodents are a cost-effective model for testing the response of the immune system to feed additives. Previously, the efficacy of YCA was evaluated using mice as a bovine mastitis model. 3 We used targeted profiling of 84 innate and cell-mediated adaptive immune response genes in whole blood for identification of molecular immune-response pathways to YCA. Leukocyte-specific microarrays had been previously utilized to evaluate the effect of YCA after 30 d of supplementation in dairy cows. 2
The objective of this study was to identify molecular pathways and immune-response indicators by which dietary YCA may modify immune responses on the gene expression level. This study is novel and important as it helps to explain on a molecular level how dietary YCA may promote the innate and adaptive immune function and thereby maintain health and prevent production losses. We are aware that this study cannot prove causality, which requires linking our observed YCA-induced gene expression changes to functional changes of immune cells in vitro and ex vivo and changes in gene expression in the intestine, where dietary YCA most likely exerts its effect. Our study was focused on discovering potential molecular pathways by which dietary YCA may modify immune responses, which can help to explain our previous findings, and to identify potential dietary immune response indicators, which can be used as an early biomarker to verify the efficacy of immunomodulatory feed supplements (dietary response marker discovery).
Materials and methods
Animals and experimental design
Animals used in this study were maintained and handled according to the guidelines of the National Research Council. 7 Animals had ad libitum access to a standardized diet (8604M; Harlan Teklad, Madison, WI, USA) and water throughout the study. Thirty-seven male, unchallenged, healthy CD rats (180–200 g, 6–7 wk old) were received from Charles River Laboratories (Hollister, CA, USA) and housed in a temperature-modulated room set at 20℃. After a 9-d adaptation period, rats were randomly assigned to three treatment groups: 0 (control; n = 5), 0.5% of YCA (n = 15) or 1% of YCA (n = 17). Experimental diets were fed for 0 (control), 7 (n = 4/YCA group), 14 (n = 3 or 4/YCA group), 21 (n = 3 or 4/YCA group) or 28 (n = 5/YCA group) d such that all rats completed the study at the same time. Feed was prepared by adding 5 or 10 g of YCA to 995 g (0.5% YCA) or 990 g of 8604 M control diet (1% YCA), respectively. The composition of the YCA has been described in detail previously. 2 In short, YCA contains dehydrated Saccharomyces cerevisiae, Trichoderma longibrachiatum fermentation product, B vitamins, choline, vitamin K precursors, diatomaceous earth, aluminosilicate and calcium chloride.
Sample collection and analyses
At the end of the study, a 5–10 ml cardiac whole blood sample was collected from anesthetized rats in tubes containing 1 ml of acid citrate dextrose solution A (cat. no. 15141-215; VWR, Wayne, PA, USA). Blood samples were stored on ice and immediately transported to the laboratory. At the laboratory, 500 μl was transferred into RNAprotect Animal Blood Tubes (cat. no. 76554; Qiagen, Valencia, CA, USA) and stored at –20℃ until RNA isolation.
RNA was isolated using the RNeasy Protect Animal Blood Kit (cat. no. 73224; Qiagen) according to manufacturer’s instructions and stored at –80℃ until cDNA synthesis. All samples had > 500 ng/µl total RNA and 260/280 values above 1.8, as measured by the Multiskan GO microplate spectrophotometer (Thermo Scientific, Pittsburgh, PA, USA). Five hundred ng/µl of RNA was used to synthesize cDNA using reagents and according to the protocol from the RT 2 First Strand Kit (cat. no. 330401; Qiagen). The cDNA samples were either stored at –80℃ or used immediately for RT-PCR analysis. For RT-PCR analysis, the Rat Innate and Adaptive Immune Responses RT 2 Profiler PCR Array (cat. no. PARN-052 Z; Qiagen) was used according to manufacturer’s instructions. The 96-well plates were loaded into a BioRad CFX96 cycler (Hercules, CA, USA) for quantification. The two-step cycling program was as follows: cycle 1 for 10 min at 95℃, followed by 40 cycles for 15 s at 95 ℃ followed by 1 min at 60℃.
Cycle threshold (Ct) values were measured for all 37 array plates at the same location on the lower third of the linear phase of the amplification curve. The array plates included 84 immune response genes and five potential reference genes: β-actin (ACTβ), lactate dehydrogenase A (LDHA), ribosomal protein, large, P1 (RPLP1), hypoxanthine phosphoribosyltransferase 1 (HPRT1); and beta-2 microglobulin (B2M). Of the five reference genes, only HPRT1 and B2M were not significantly affected (P < 0.05) by YCA and thus were used as reference genes. Relative gene expression was calculated by subtracting the average Ct value for the two reference genes from the Ct value of the genes-of-interest (ΔCt-values).
Statistical analyses
The ΔCt-values were analysed using PROC GLM in Statistical Analysis System, version 9.2 (SAS Institute Inc., Cary, NC, USA). The fixed effect in the model was diet (control, 0.5% YCA 7 d, 0.5% YCA 14 d, 0.5% YCA 21 d, 0.5% YCA 28 d, 1% YCA 7 d, 1% YCA 14 d, 1% YCA 21 d and 1% YCA 28 d). Statistical comparisons were done sequentially based on a priori comparisons with increasingly stringent criteria. Comparisons were performed using unpaired t-tests with the ESTIMATE statement. Only genes that showed significance in the previous step were evaluated further. In the first step, the comparison for the effect of YCA supplementation was YCA-fed vs. control rats and for YCA inclusion rate differences was 0.5% vs. 1% YCA-fed rats. In the second step, the comparisons for the effect at both YCA inclusion rates were 0.5% YCA-fed vs. control rats and 1% YCA-fed vs. control rats. In the third step, the comparisons for the effect of supplementation period were YCA 7 d vs. control rats, YCA 14 d vs. control rats, YCA 21 d vs. control rats and YCA 28 d vs. control rats. Results are shown as mean fold changes = 2−ΔΔCT = [(CTgene of interest – CTreference genes)YCA-Diet – (CTgene of interest – CTreference genes)Control Diet] 8 with SEM. All statistical tests were two-sided. To adjust for multiple comparisons, q-values were calculated using the linear step-up procedure. 9 Statistical significance was declared at P ≤ 0.05 and a tendency at 0.05 < P ≤ 0.10.
Results
The gene expression of a total of 84 innate and adaptive immune response genes was analyzed; among these, 73 were above the detection limit in > 90% of samples. The remaining 11 genes, amyloid P component, serum (APCS); chemokine (C-C motif) ligand 12 (CCL12); C-reactive protein (CRP); CSF2; IFNα1 and IFNβ1; IL1A, IL5 and IL13; mannose-binding lectin protein C 2 (MBL2); and myeloperoxidase (MPO) were not further analyzed. In addition, we removed 10 genes that expressed mRNA in low abundance (mean CT-value ≥ 28): chemokine (C-X-C motif) receptor 8 (CCR8), chemokine (C-X-C motif) ligand 10 (CXCL10), IFNγ, IL4, IL6, IL10, recombination activating gene 1 (RAG1), RAR-related orphan receptor C (RORC), and TLR3 and TLR5.
Effect of YCA supplementation on immune gene expression in a rodent model. a
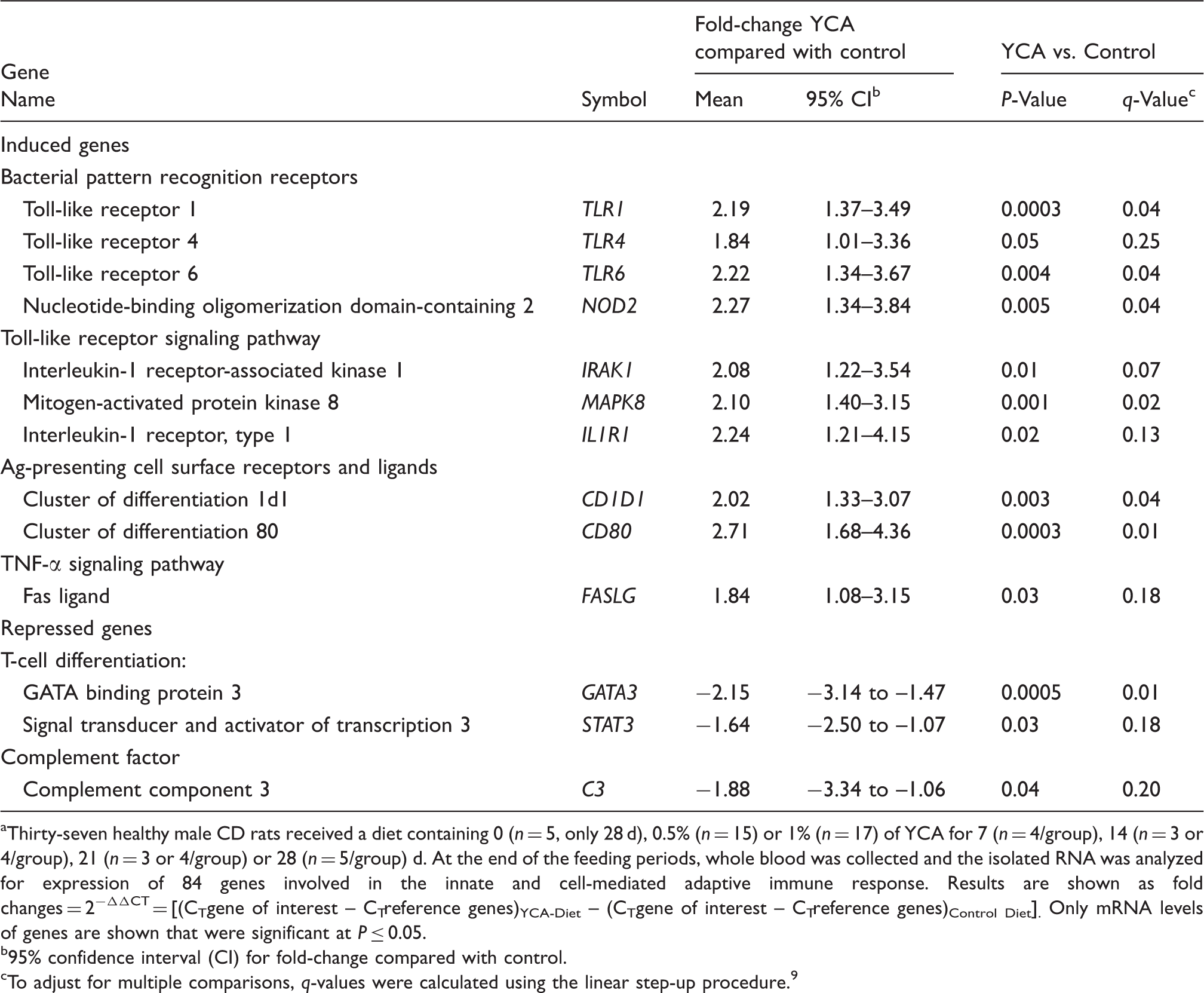
Thirty-seven healthy male CD rats received a diet containing 0 (n = 5, only 28 d), 0.5% (n = 15) or 1% (n = 17) of YCA for 7 (n = 4/group), 14 (n = 3 or 4/group), 21 (n = 3 or 4/group) or 28 (n = 5/group) d. At the end of the feeding periods, whole blood was collected and the isolated RNA was analyzed for expression of 84 genes involved in the innate and cell-mediated adaptive immune response. Results are shown as fold changes = 2−ΔΔCT = [(CTgene of interest – CTreference genes)YCA-Diet – (CTgene of interest – CTreference genes)Control Diet]. Only mRNA levels of genes are shown that were significant at P ≤ 0.05.
95% confidence interval (CI) for fold-change compared with control.
To adjust for multiple comparisons, q-values were calculated using the linear step-up procedure. 9
The 10 induced genes included four bacterial PRRs, the cell surface PRRs TLR1, TLR4 and TLR6, and the cytosolic PRR NOD2, and three genes involved in TLR signal transduction, IL-1 receptor-associated kinase 1 (IRAK1), MAPK8 and IL1R1. Other induced genes were two APC surface receptors cluster of differentiation 1d1 (CD1D1) and 80 (CD80), and the pro-apoptotic protein Fas ligand (FASLG). The repressed genes included the Th2 marker GATA binding protein 3 (GATA3); the Th2 and Th17 marker signal transducer and activator of transcription 3 (STAT3); and the extracellular complement factor protein complement component 3 (C3).
As a second step, we examined whether YCA supplementation altered gene expression at both YCA inclusion rates. At P ≤ 0.05, six genes, TLR1, TLR6, NOD2, MAPK8, CD1D1 and CD80, were induced and one gene, GATA3, was repressed at both inclusion rates (Table 2). As a third step, we examined whether YCA supplementation altered gene expression at each time point. At P ≤ 0.05, four genes, TLR1, NOD2, MAPK8 and CD80, were induced, whereas no gene was repressed at each time point (Figure 1).
Effect of YCA supplementation period on immune gene expression. Thirty-seven healthy male CD rats received a diet containing 0 (n = 5, only 28 d), 0.5% (n = 15) or 1% (n = 17) of YCA for 7 (n = 4/group), 14 (n = 3 or 4/group), 21 (n = 3 or 4/group) or 28 (n = 5/group) d. At the end of the feeding periods, whole blood was collected and the isolated RNA was analyzed for expression of 84 genes involved in the innate and cell-mediated adaptive immune response. Results are shown as average fold changes = 2−ΔΔCT = [(CTgene of interest – CTreference genes)YCA-Diet – (CTgene of interest – CTreference genes)Control Diet] with SEM. Only mRNA levels of genes are shown that were significant at P ≤ 0.05 for all four supplementation periods. Effect of YCA inclusion rate on immune gene expression.
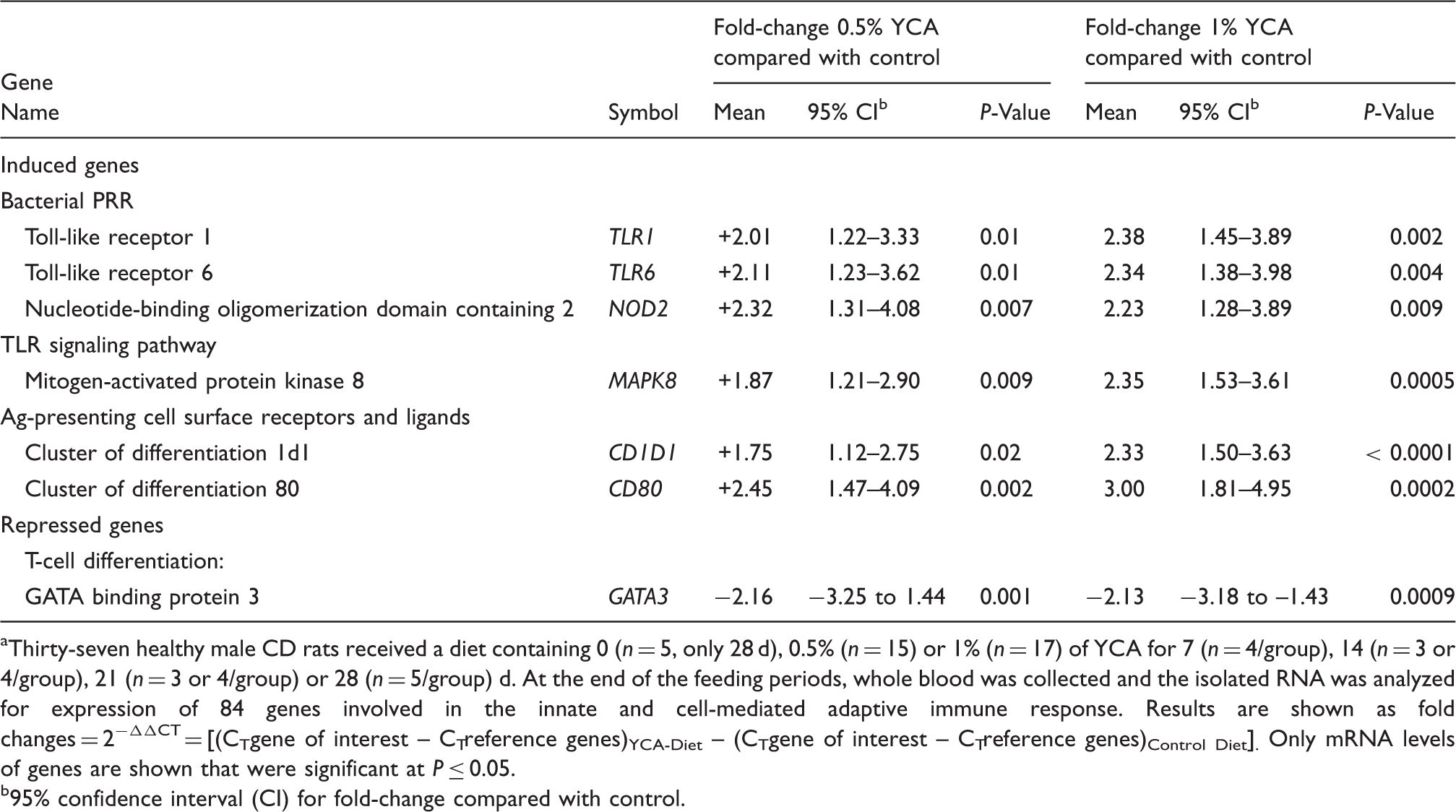
a
Thirty-seven healthy male CD rats received a diet containing 0 (n = 5, only 28 d), 0.5% (n = 15) or 1% (n = 17) of YCA for 7 (n = 4/group), 14 (n = 3 or 4/group), 21 (n = 3 or 4/group) or 28 (n = 5/group) d. At the end of the feeding periods, whole blood was collected and the isolated RNA was analyzed for expression of 84 genes involved in the innate and cell-mediated adaptive immune response. Results are shown as fold changes = 2−ΔΔCT = [(CTgene of interest – CTreference genes)YCA-Diet – (CTgene of interest – CTreference genes)Control Diet]. Only mRNA levels of genes are shown that were significant at P ≤ 0.05. 95% confidence interval (CI) for fold-change compared with control.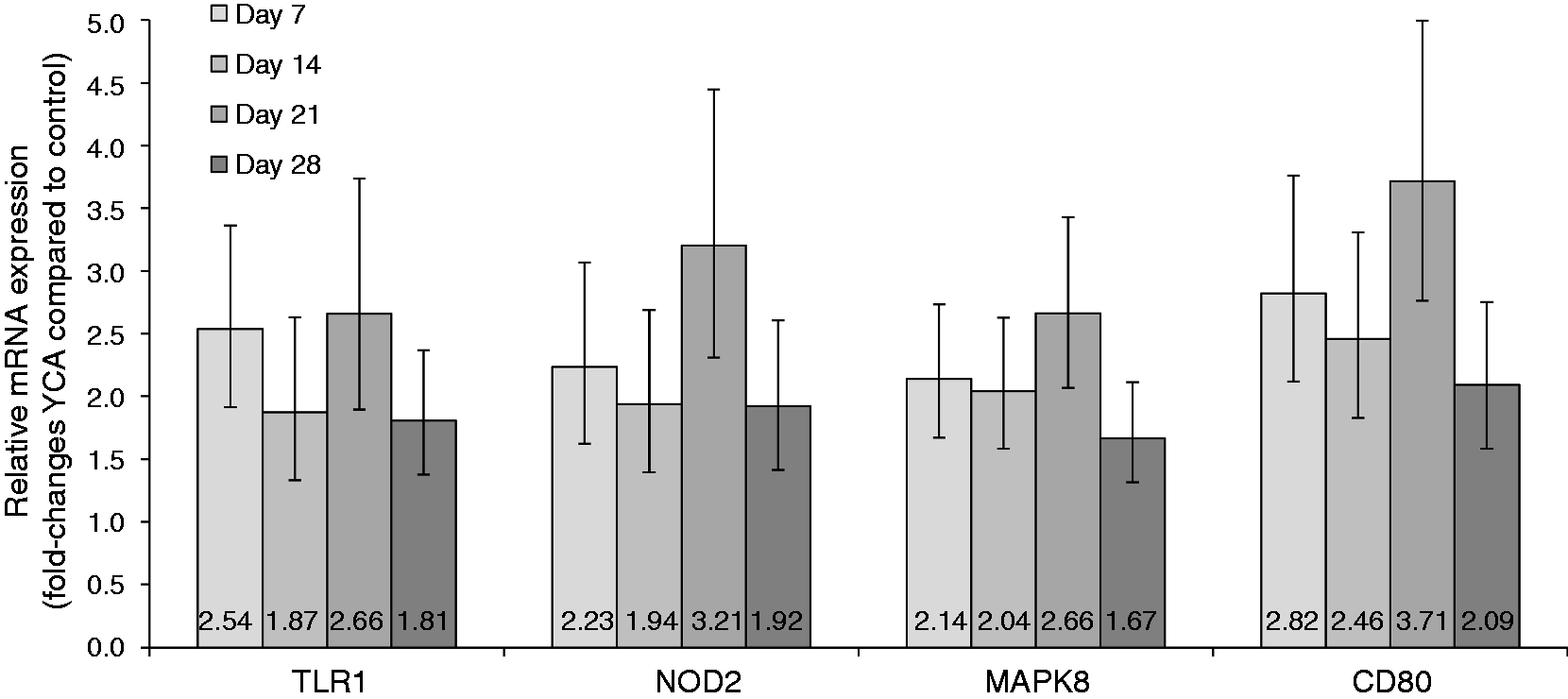
As a fourth step, we examined which pathways were altered by YCA supplementation at the gene expression level. We observed four pathways altered by YCA supplementation. (1) Dietary YCA increased gene expression of bacterial PRRs. Of the 63 examined genes, seven genes coded for bacterial PRR, and four of these, TLR1, TLR4, TLR6 and NOD2, were induced by YCA. (2) Dietary YCA increased gene expression of molecules involved in the TLR signaling cascade. Twenty-two of the 63 examined genes coded for molecules involved in the TLR signaling cascade; eight genes were either at P ≤ 0.05 (IRAK1, MAPK8, IL1R1 and FASLG) or at 0.05 < P ≤ 0.10 induced by YCA supplementation. A statistical tendency was observed for IFNAR1 (+1.69; P = 0.06), IFN regulatory element 3 (IRF3: +1.64; P = 0.08), and caspases 1 and 8 (CASP1: +1.48; CASP8: +1.32; both P = 0.07). 3) Dietary YCA increased gene expression of APC receptors and co-stimulatory signals involved in naïve T-cell activation. Four examined genes coded for APC receptors and co-stimulatory signals; two genes (CD1D1 and CD80) were significantly induced, and CD86 tended to be induced (+1.63; P = 0.10). (4) Dietary YCA increased gene expression of molecules involved in Th-cell differentiation, as transcription factors favoring differentiation toward a Th2 and Th17 profile (GATA3 and STAT3) were repressed, and STAT4, a transcription factor favoring differentiation toward a Th1 profile, tended to be induced (+1.49; P = 0.10).
Discussion
Feeding livestock immunomodulatory supplements has increased in popularity as a management tool for prevention of diseases and subsequent production losses. Limited information, however, is available on how these feed additives act on the immune system. We previously reported that one of the most widely used immunomodulatory feed supplements, YCA, decreased mammary infections in dairy cows and improved milk production, 4 and improved innate and adaptive immune responses function in immune-challenged ruminant livestock.1–6 This study closes an important gap in our knowledge, as it helps to explain the improved innate and adaptive immune responses reported in ruminant livestock fed YCA. At the same time, we want to caution the reader, as gene expression may or may not translate into changes on the protein level and/or functional changes like our previously observed TCA-induced changes in L-selectin gene and protein expression across various ruminant species.1,5
Dietary YCA increased expression of genes involved in bacterial recognition
Immune responses are a complex mechanism involving various cell types. First, immune cells need to detect the presence of a pathogen. This is accomplished through interactions between PAMPs and surface or intracellularly expressed PRRs of immune cells, primarily macrophages and dendritic cells that reside in the tissue or closely adjacent lymphoid tissues. Of the seven bacterial PRRs evaluated in this study, YCA supplementation induced gene expression of three surface-expressed PRRs, TLR1, TLR4 and TLR6, and one cytosolic PRR, NOD2. The aforementioned bacterial PRR recognize a variety of PAMPs, including tri-acylated lipoproteins (most common in Gram-negative bacteria), di-acylated lipoproteins (most common in Gram-positive bacteria), peptidoglycans (found within bacterial cell walls), as well as zymosan, a polysaccharide of yeast cell membranes.10–13 After TLR1 and TLR6 bind to the PAMPs, TLR1 and TLR6 form heterodimers on the immune cell surface with TLR2.14–16 Previous studies in piglets and broiler chickens showed that yeast fermentation product and yeast cell wall components induce TLR2 and TLR4 gene expression in ileal tissue, respectively.17–19 In this study, gene expression of TLR2 was measured in blood and was not induced (+1.17; P = 0.39). Regulation on the gene expression level is one of many factors that have the potential to regulate PRR activity, which also include post-transcriptional regulation of PRR protein levels, surface receptors cycling, internalization and specific location at the cell membrane. In summary, dietary YCA may potentially increase the cell’s sensitivity of bacterial PAMP recognition.
Dietary YCA increased expression of genes involved in TLR signaling
Interactions between PAMPs and bacterial PRR initiate signaling cascades, some of which involve the protein kinases IRAK1 and MAPK8, also known as Jun-kinase. YCA supplementation induced gene expression of both protein kinases. The TLR signaling cascades lead to the activation of transcription factors AP-1 (via MAPK8), NF-κB, and IRFs. 20 YCA supplementation tended to induce gene expression of IRF3. The aforementioned transcription factors can stimulate the production of secreted cytokines and chemokines, including IL-1β and IL-8 (via NF-κB), TNF-α (via AP-1) and IFN (via IRF).13,21 In this study, YCA supplementation did not induce IL-1β gene expression in whole blood (–1.24-fold; P = 0.50). However, we observed that YCA supplementation tended to increase gene expression of the thiol protease CASP1, which activates IL-1β, and increased gene expression of IL1R1, one of the two units of the IL-1 receptor that binds to activated IL-1β.22,23 The other genes affected by YCA supplementation included the pro-apoptotic transmembrane protein FASLG and the protease CASP8, which are molecular targets of TNF-α signaling, and the transmembrane protein IFN-αR1, which is the receptor for IFN-α signaling. In summary, dietary YCA may potentiate TLR signaling.
Dietary YCA increased expression of genes involved in T-cell activation
Naïve T cells are activated when a recognized or cognate antigen is bound to the T-cell antigen receptor TCR. Activation is strictly dependent upon antigen presentation by APC (primarily macrophages and dendritic cells). Binding of APC and the interaction of co-stimulatory signals educates and modulates this activation. This is an important step that links the innate with the cell-mediated adaptive immune response. In this study, YCA supplementation induced gene expression of two APC surface receptors, CD1D1 and CD80, and tended to up-regulate CD86. CD1D1 is responsible for presenting lipid antigens to and activating NK T-cells. 24 CD80 is a membrane ligand on dendritic cells, B-cells and macrophages, 25 which, together with CD86, activates naïve T cells. 26 The fate of activated T cells is complex: after differentiation via co-stimulatory molecules and cytokines, T cells may as one possible route stimulate responsive B cells to proliferate and differentiate into plasma cells. These will secrete Abs to aid in the removal of an infection. 27 Previous studies have shown that yeast fermentation products induce the gene expression of CD80 and CD86 on the cell surface of B lymphocytes.28,29 The effect of these yeast fermentation products were more significant on CD80 than on CD86 expression.28,29 In summary, dietary YCA may promote T-cell activation by increasing gene expression of cell surface receptors of APC.
Dietary YCA increased expression of genes involved in T-cell differentiation
Differentiation of naïve CD4+ T cells is controlled by multiple cytokines and transcription factors. In this study, YCA supplementation repressed gene expression of the transcription factors GATA3 and STAT3, and tended to induce gene expression of Stat4. Differentiation of naïve CD4+ T cells toward a Th1 profile is promoted by STAT4. 30 Conversely, GATA3 and STAT3 promote T-cell differentiation away from a Th1 profile. Stat3 promotes differentiation towards Th2 and Th17 profiles, 31 and GATA3 inhibits differentiation toward a Th1 profile and promotes differentiation toward a Th2 profile.32–34 In summary, dietary YCA may promote T-cell differentiation toward a response that focuses on intracellular pathogens (Th1 response).
TLR1, NOD2, MAPK8 and CD80 are potential response indicators to dietary YCA
The second objective of this study was to identify immune-associated response indicators to dietary YCA supplementation. Four genes, TLR1, NOD2, MAPK8 and CD80, were significantly induced at each time point (d 7, 14, 21 and 28). Therefore, these genes may be used to evaluate the efficacy of dietary YCA for inducing an immune response, similar to early biomarkers in human medicine. Moreover, these four genes play functionally important roles in immune responses and thus could serve as general dietary immune response indicators. Future studies are needed to verify that dietary YCA increases these four genes in ruminant livestock under various environmental conditions, as differences in immune function and gene expression among species are well established, which could potentially impact cross-species translation of our findings.35,36
Other considerations
Given the abundance of intestinal microbial cells (>109 bacterial cells/animal) and the, in comparison, low dosage of dietary YCA (0.5% of diet), one might wonder how can dietary YCA alter immune responses. Besides dehydrated S. cerevisiae, YCA contains fermentation product of the fungus T. longibrachiatum and may induce an immune response either directly or by altering the microbiota. 37 Diatomaceous earth and aluminosilicate are other ingredients of YCA, which may bind toxins and alter the intestinal microbiota or may directly induce an immune response.38–40 Moreover, YCA contains B vitamins, choline and vitamin K precursors that can be utilized as nutrients by the microbiota or may, in sufficient concentrations, alter gene expression directly.
The YCA response was independent of inclusion rate, one of the criteria used to evaluate the efficacy of dietary compounds for disease prevention. Based on our results, YCA promotes pathogen recognition; thus, very low inclusion rates are required. Among the 10 genes that were induced, all but two (IRAK1 and MAPK8) were genes that express extracellular (TLR1, TLR4, TLR6, CD1D1, CD80, IL1R1 and FASLG) or intracellular (NOD2) receptors or ligands.
The small sample size and the use of only male, healthy rats limit the generalizability of our results; however, immune responses in females are known to be stronger than in males. 41 We used whole blood rather than specific immune cell types to get a broad overview on the effect of dietary YCA on immune-associated gene expression. Future studies are needed to verify that dietary YCA increases bacterial recognition and signaling cascades as well as T-cell activation and differentiation in ruminant livestock under various management conditions.
Conclusions
In summary, we identified four molecular pathways whereby YCA supplementation may promote the innate and adaptive immune function: (1) increased bacterial pathogen recognition, (2) increased TLR signaling, (3) increased T-cell activation and (4) increased Th-cell differentiation toward a Th1 response. Four genes, TLR1, NOD2, MAPK8 and CD80, were consistently induced after 7, 14, 21 and 28 d of YCA supplementation and thus could serve as potential dietary immune response indicators to YCA supplementation. In conclusion, YCA supplementation may increase recognition and responses to bacterial pathogens and T-cell activation and differentiation in whole blood. Future studies are warranted to verify these effects in ruminant livestock under various management conditions and evaluate if and how our observed changes on the gene expression level may translate to the protein level and functional in vitro and ex vivo assays examining the suppression and/or clearance of pathogens.
Footnotes
Acknowledgements
We thank Shelby A Armstrong and Tyler H Schell for animal handling and participating in sample collection.
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Several of the authors involved in this study served also as employees of OmniGen Research (Corvallis, OR), which, in part, funded this research. The funders of the study had no role in study design, data collection and analysis, or decision to publish or preparation of the manuscript.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was financially supported by OmniGen Research (Corvallis, OR) and Oregon State University.
