Abstract
Serotonin (5-hydroxytryptamine; 5-HT) transporter (5-HTT) is involved in inflammation and the stress response. In this study, we examined the regulation of 5-HTT expression in macrophage HD11 cells in response to bacterial LPS. Long-term exposure of cells to LPS (6–18 h) produced a decrease in 5-HTT mRNA expression. Accordingly, reduced 5-HTT activity measured by 5-HT uptake was also observed in LPS-treated HD11 cells. Moreover, LPS treatment, as well as co-transfection with an expression vector encoding the chicken CCAAT/enhancer binding protein beta (C/EBPβ), resulted in inhibition of 5-HTT promoter activity. Indeed, sequence analysis revealed several C/EBPβ binding motifs in the upstream region of the 5-HTT gene, which specifically interacted with C/EBPβ both in an in vitro band shift assay and in living HD11 cells. The C/EBPβ binding was activated in cells treated with LPS. The role of C/EBPβ in LPS inhibition of 5-HTT expression was further confirmed by small interfering RNA interference, which demonstrated that knockdown of endogenous C/EBPβ attenuated the inhibition of 5-HTT expression in LPS-treated cells. Taken together, the results suggest that C/EBPβ plays a critical role in regulating the 5-HTT gene in macrophages in response to pro-inflammatory stimuli.
Introduction
Serotonin (5-HT) is widely found in the central and peripheral nervous systems. This neurotransmitter has been shown to be implicated in neurobehavioral processes, and to be involved in intestinal peristalsis and secretion via the enteric nervous system.1,2 Several lines of evidence indicate that 5-HT is present in the immune system. Following injury, 5-HT released from innate immune cells is involved in immune responses including inflammation, chemotaxis and phagocytosis. 3 The membrane-associated 5-HT transporter (5-HTT) is responsible for reuptake of extracellular 5-HT and thus plays an important role in regulating extracellular 5-HT levels at its target cells. 4 Experiments with genetic disruption of the mouse and rat 5-HTT genes have shown that 5-HTT knockout animals have exhibited a variety of phenotypic changes, including increased anxiety, stress-related behaviors, gut dysfunction and so on.5,6 A prospective, longitudinal human study demonstrated that a functional polymorphism in the 5-HTT gene-linked polymorphic region modulates the influence of stressful life events on the development of depression. 7 Furthermore, several studies using knockout mice have also provided increasing evidence indicating that 5-HTT is involved in inflammation and stress responses.8–12 In humans, a recent study using positron emission tomography and radioimmunoassays has revealed that reduced thalamic 5-HTT binding potential was associated with increased cortisol response. 13 In addition, TNF-α, an inflammatory mediator that plays a key role in the initiation of inflammation, has been shown to enhance 5-HTT function. 14
LPS released from the outer membrane of Gram-negative bacteria during bacterial infection or following exposure to antibiotics can activate acute immunological and inflammatory responses by interactions of the lipid A component to stimulate production and secretion of pro-inflammatory mediators such as TNF-α and IL-1β from monocytes and macrophages. 15 During the acute phase of bacterial infection, LPS is known to be responsible for many of the clinical symptoms of sepsis. For instance, LPS has been shown to induce fever and depress locomotor activity in unrestrained mice. 16 Administration of LPS in vivo increases vascular permeability leading to hypotension, hyperemia, intravascular coagulation and so on.17,18 Further, it has been already demonstrated that LPS is responsible for anorexia during bacterial infections. 19
An acute inflammatory response triggered by bacterial infections involves the release of a variety of mediators, including 5-HT, from macrophages, mast cells, basophils and platelets. 20 5-HT has been shown to modulate production and release of cytokines, for example IL-1β, IL-6 and TNF-α, suggesting that 5-HT plays a role in mediating pro-inflammatory responses. 3 In this study, we wished to address the question of whether macrophages could contribute to the regulation of extracellular 5-HT levels in response to LPS. Because extracellular 5-HT levels are dependent on the 5-HT reuptake controlled by 5-HTT, the first goal of this study was to examine whether LPS modulates 5-HTT expression. The 5’-flanking region of the chicken 5-HTT gene contains several putative binding motifs for transcription factors of the chicken CCAAT/enhancer binding protein (C/EBP) family (this study). C/EBPβ has been already shown to activate and repress transcription of a variety of target genes. 21 Therefore, we focused on the role of C/EBPβ in regulating the chicken 5-HTT gene. Our results show that long-term treatment with LPS down-regulates the 5-HTT expression in chicken macrophages and that this negative effect is mediated by interaction of C/EBPβ with the upstream region of the 5-HTT gene.
Materials and methods
Cell culture
The chicken macrophage cell line HD11 was grown in Iscove’s modified Dulbecco’s medium (IMDM) supplemented with 8% FCS, 2% chicken serum, 100 U/ml penicillin and 100 µg/ml streptomycin at 37℃ in humidified 95% air and 5% CO2. 22 For activation, HD11 cells were harvested and seeded at 3 × 106 cells per 9-cm culture dish. On the following day, cells were, when indicated, treated with 10 µg/ml LPS from Salmonella Typhimurium (Sigma-Aldrich, Munich, Germany) for the indicated time periods.
Primary macrophages were generated from chicken blood (Lohmann Brown chickens) as follows. Whole blood from adult hens was collected after decapitation using heparin (400 U/ml) as the anticoagulant. After dilution with one volume of IMDM, the blood was centrifuged in Ficoll–Paque PLUS separating solution with a density of 1.077 g/ml (GE Healthcare, Freiburg, Germany) to separate the mononuclear cell fraction. Collected cells were washed twice with IMDM. After being seeded in 9-cm culture dishes, cells were incubated at 37℃ and fed with fresh IMDM every day. After 5 d, primary macrophages were used for RNA preparation and real-time RT-PCR following activation with 10 µg/ml LPS.
RNA preparation
Total RNA was isolated from HD11 cells as follows. Cells washed twice in PBS were lysed in 1 ml of a denaturing buffer (GTC) containing 4 M guanidinium thiocyanate, 25 mM sodium citrate, 0.5% N-lauroylsarcosine and 0.1 M β-mercaptoethanol. Following scraping off from the dish, cell material was ultrasonicated by three 10-s pulses. After adding 200 µl of 2 M sodium acetate, the sample was extracted once with phenol/chloroform/isoamyl alcohol (25:24:1), followed by cooling for 15 min on ice and centrifugation at 10,000 g for 20 min at 4℃ in a Beckman JS-13.1 rotor (Beckman Coulter, Krefeld, Germany). RNA was then precipitated with isopropyl alcohol, and the RNA pellet was washed once with 80% ethanol. After air-drying, the pellet was dissolved in 440 µl GTC denaturing buffer containing 0.2 M sodium acetate, and the RNA was precipitated with isopropyl alcohol. The RNA pellet was dissolved in 100 µl sterile deionized water following air-drying. To eliminate any remaining genomic DNA, 50 -µg samples of RNA were treated with 15 units of RNase-free DNase (Roche, Mannheim, Germany) in 100 µl of a buffer containing 10 mM Tris-HCl (pH 8.0), 50 mM KCl and 1.5 mM MgCl2 at 37℃ for 30 min, and then extracted twice with phenol/chloroform/isoamyl alcohol. After ethanol precipitation, RNA pellets were dissolved in sterile deionized water at a desired concentration of 1 µg/µl.
Quantitative real-time RT-PCR
Total RNA (1 µg) was reverse-transcribed into cDNA using the Transcriptor First Strand cDNA Synthesis kit (Roche) according to the manufacturer’s instruction. For quantitation of 5-HTT and GAPDH mRNA, quantitative real-time PCR was performed with 6 µl or 0.25 µl of cDNA, respectively, in a 25-µl PCR mixture containing 1 × TaqMan Gene Expression Master Mix (Applied Biosystems, Darmstadt, Germany), gene-specific primers (0.8 µM each) and 0.25 µM TaqMan probe by 45 cycles, each consisting of 20 s at 95℃, 25 s at 55℃ and 35 s at 72℃. To determine the lysozyme expression, real-time PCR was carried out with 5 µl cDNA in a 25-µl PCR mixture containing 1 × Power SYBR Green PCR Master Mix (Applied Biosystems) and primers (0.25 µM each) by 45 cycles of amplification, each consisting of 20 s at 95℃ and 35 s at 60℃. DNA primers and TaqMan probes used for the real-time PCR were as follows:
5-HTT: 5′-CTGGGATTAGACAGCACGTTTG-3′, 5′-CGTGTGGGAATTCAT CCAGTACT-3′, TaqMan probe: 5′-AGGATTAGAGGGAGTGATT-3′; GAPDH: 5′-TGACAAGTCCCTGAAAATTGTCA-3′, 5′-CAAGGGTGCCAGGC AGTT-3′, TaqMan probe: 5′-AATGCATCGTGCACCAC-3′; Lysozyme: 5′-GGGATACAGCCTGGGAAACTG-3′, 5′-TTGTAGCCTGGGTGTTGAAGTTAC 3′.
5-HT uptake assay
HD11 cells were seeded at a density of 1 × 106 cells per well. On the following day, cells were treated with PBS or LPS for the indicated time periods. For uptake assays, cells were washed twice with Hank’s balanced salt solution (HBSS) (Life Technologies, Darmstadt, Germany) and pre-incubated in HBSS at 37℃ for 20 min. Following the addition of 5-HT and [3H]-5-HT (25 nM and 20 Ci/mmol), cells were further incubated for 10 min. The uptake was then stopped by adding 1 ml of ice-cold HBSS and placing the cells on ice. After washing twice with HBSS, the cells were solubilized in 0.1 M NaOH, mixed with a 5-ml scintillation cocktail and counted for radioactivity in a liquid scintillation counter. Nonspecific 5-HT uptake was measured with 0.1 µM fluoxetine (Sigma-Aldrich). Specific 5-HT uptake was calculated by subtracting nonspecific uptake from total 5-HT uptake.
PCR and plasmid construction
The 0.63-kb region upstream of the ATG motif and the 1.3-kb sequence further upstream of the chicken 5-HTT gene were amplified using two specific primer pairs: 5′-TTATGCAGTATGGGGGTCAAG-3′ and 5′-TCTCATTGCTCGTTGCCTTAT-3′; 5′-CCTCAGGATGTGCCAAAAAG-3′ and 5′-GACAAAGCTTGACCCCCATAC-3′ (GenBank Accession NW_001471508)
PCR was performed with PfuUltra high-fidelity DNA polymerase (Agilent Technologies, Waldbronn, Germany) in 50 µl of PfuUltra HF reaction buffer. The PCR products were then cloned into the pJET1.2 plasmid (Fermentas, St. Leon-Rot, Germany). The resulting recombinant plasmid pJET1.2-0.63 was digested with Xho I and partially with Hind III to release the 0.63-kb Hind III-Xho I fragment, and the other (pJET1.2–1.3) was digested with Xba I and Hind III to release the 1.3-kb Xba I-Hind III fragment. The DNA fragments were ligated and placed between Xba I and Xho I sites upstream of the coding sequence for chloramphenicol acetyltransferase (CAT) of pBLCAT6 23 to construct the CAT reporter plasmid containing the 1.93-kb 5′-flanking region. For construction of the CAT reporter plasmid containing the 0.63-kb 5′-flanking region, the Xho I-Xba I fragment isolated from pJET1.2–0.63 was inserted into pBLCAT6.
The expression vector pN3-C/EBPβ encoding the chicken C/EBPβ was constructed as follows. cDNA encoding the full-length C/EBPβ was generated from chicken genomic DNA by PCR using PfuUltra high-fidelity DNA polymerase and the primers 5′-CAGCGCCGTCTTCTCCTC-3′ and 5′-CAATGAAACCCCCAACGAAAC-3′, and cloned into pJET1.2 (Fermentas). The resulting plasmid was digested with Xba I and Xho I, and the Xba I-Xho I fragment containing the C/EBPβ cDNA was introduced into plasmid pN3 containing the CMV promoter.
For construction of C/EBPβ targeting small interfering RNA (siRNA) expression vectors, DNA fragments encoding short hairpin RNA targeting C/EBPβ were placed between the Bam HI and Hind III restriction sites of pRNA-CMV3.1/Neo (GenScript, Piscataway, NJ, USA). pRNA-CMV3.1/Neo-CTL (Genscript) was used as a negative control plasmid. C/EBPβ targeting siRNA sequences designed by siRNA design tools from Genscript were 5′-TCCATGACCGAACTTACCGTA-3′ and 5′-TTTCCGATCTCTTCGCCGAGG-3′.
DNA transfection and CAT assay
Cells were seeded at 3 × 106 cells per 9-cm culture dish in complete IMDM. On the following day, cells were transfected with 20 µg plasmid DNA per 9-cm plate using a calcium phosphate co-precipitation protocol described previously. 24 Twenty-four hours after transfection, cells were harvested for the preparation of protein extracts. CAT assays were performed as described previously. 24
Nuclear extract preparation and electrophoretic mobility shift assay
Nuclear extracts were prepared from exponentially growing cells as described elsewhere.
25
An electrophoretic mobility shift assay (EMSA) was performed as described previously.
26
Briefly, double-stranded oligonucleotide
32
P end-labeled with T4 polynucleotide kinase (10,000 cpm) was incubated with 5 µg of nuclear extracts in a final volume of 20 µl containing 20 mM HEPES (pH 7.9), 30 mM NaCl, 1.75 mM EDTA, 6 mM MgCl2, 2 mM ZnCl2, 2 mM dithiothreitol, 2.5 µg of poly(dI-dC) and 4% (w/v) Ficoll 400 at 4℃ for 45 min. The resulting DNA–protein complexes were resolved on 4% non-denaturing polyacrylamide gels and visualized on X-ray films. EMSA oligonucleotides (altered bases of C/EBPβ binding site are underlined) used were as follows:
M6: 5′-GCTCGGGATGTTGCAATCATA-3′; m1: 5′-GCTCGGGA m2: 5′-GCTCGGGATGTT CE: 5′-AATTCAATTGGG CAATCAGG-3′; κB: 5′-TGGGGACTTTCCAG-3′.
For immunomobility shift assays, radiolabelled oligonucleotide and nuclear extracts were incubated for 15 min, followed by adding Abs, and the reaction continued for another 30 min. The Abs used were anti-C/EBPβ (a gift from Dr. A. Leutz, Max Delbrück Center for Molecular Medicine) and anti-NF-κB p50 (NLS) (Santa Cruz Biotechnology, Santa Cruz, CA, USA).
Chromatin immunoprecipitation assay
Immunoprecipitation of nucleoprotein complexes prepared from HD11 cells was performed as described previously.
26
DNA isolated from immunoprecipitated nucleoprotein complexes was analyzed by quantitative real-time PCR with SYBR Green I using a 7500 Real Time PCR System (Applied Biosystems).
26
The PCR primers used were:
pri1 for amplification of fragment A: 5′-GCTTTGGTTTTTCCAAGAGTTGA-3′, 5′-CGAGCGGAAGAATAAGCGTTA-3′; pri2 for amplification of fragment B: 5′-CCAAACGTGGCAGGGATG-3′, 5′-ACATTCCATGATCACCTTTCTTCA-3′; pri3 for amplification of fragment C: 5′-TAACCCGTATCAAGGCCCATC-3′, 5′-GAAAAATAGCGCTCCTCTGCAC-3′.
Statistical analysis
Statistical analyses were performed using the VassarStats calculator (Vassar College, Poughkeepsie, NY, USA). The significance of statistical differences (P-value) between two indicated groups was verified by an unpaired Student’s t-test. A two-way ANOVA was used to examine the effects of siRNA and drug (LPS vs. PBS). A P-value < 0.05 was considered significant.
Results
Long-term exposure of chicken macrophages to LPS down-regulates 5-HTT gene expression and 5-HT uptake
It has been reported that acute LPS injections in vivo stimulated 5-HTT activity in mouse brain.
27
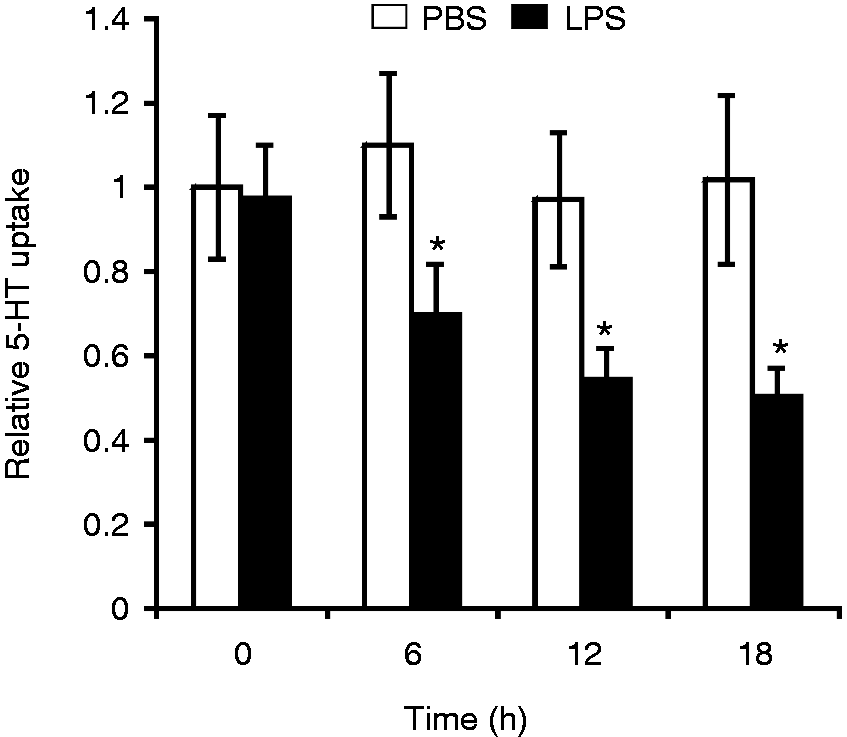
We therefore examined the effect of chronic LPS treatment on the expression of the 5-HTT gene. Chicken macrophage HD11 cells were treated with LPS from S. Typhimurium for 6–18 h, and the 5-HTT mRNA expression was analyzed by quantitative real-time RT-PCR. As a control, lysozyme and GAPDH mRNA levels were measured. As shown in Figure 1A, treatment with LPS significantly reduced the expression of 5-HTT mRNA. Eighteen hours after LPS treatment, the level of 5-HTT mRNA was reduced to 25% in comparison with the mock treatment with PBS. In contrast, the expression of lysozyme mRNA was strongly induced in LPS-treated macrophages (Figure 1B), whereas the GAPDH mRNA level was not significantly affected by LPS (Figure 1C). Exposure to LPS also repressed 5-HTT mRNA expression in primary macrophages isolated from chicken whole blood (Figure 1D). Next, we examined the effect of LPS on 5-HT uptake in HD11 cells using [3H]-5-HT. 5-HT uptake was decreased to 51% in cells treated with LPS for 18 h when compared with the 5-HT uptake in saline-treated cells (Figure 2).
LPS reduces 5-HTT mRNA expression in chicken macrophages. RNA was isolated from (A–C) HD11 cells or (D) primary macrophages isolated from chicken blood treated with PBS for control or with 10 µg/ml LPS for the indicated time periods. The 5-HTT, lysozyme and GAPDH mRNA expressions were analyzed by real-time RT-PCR. The 5-HTT mRNA and lysozyme mRNA levels were normalized to GAPDH. Values are expressed as mean values (n = 3) ± SEM. LPS effect on 5-HT uptake in HD11 cells. Cells were treated with PBS or 10 µg/ml LPS for the indicated time periods. 5-HT uptake was measured following incubation of cells with [3H]-5-HT for 10 min. The values shown are relative to the 5-HT uptake in PBS-treated cells at time zero (4.2 fmol/106 cells). Values are expressed as mean values (n = 3) ± SEM.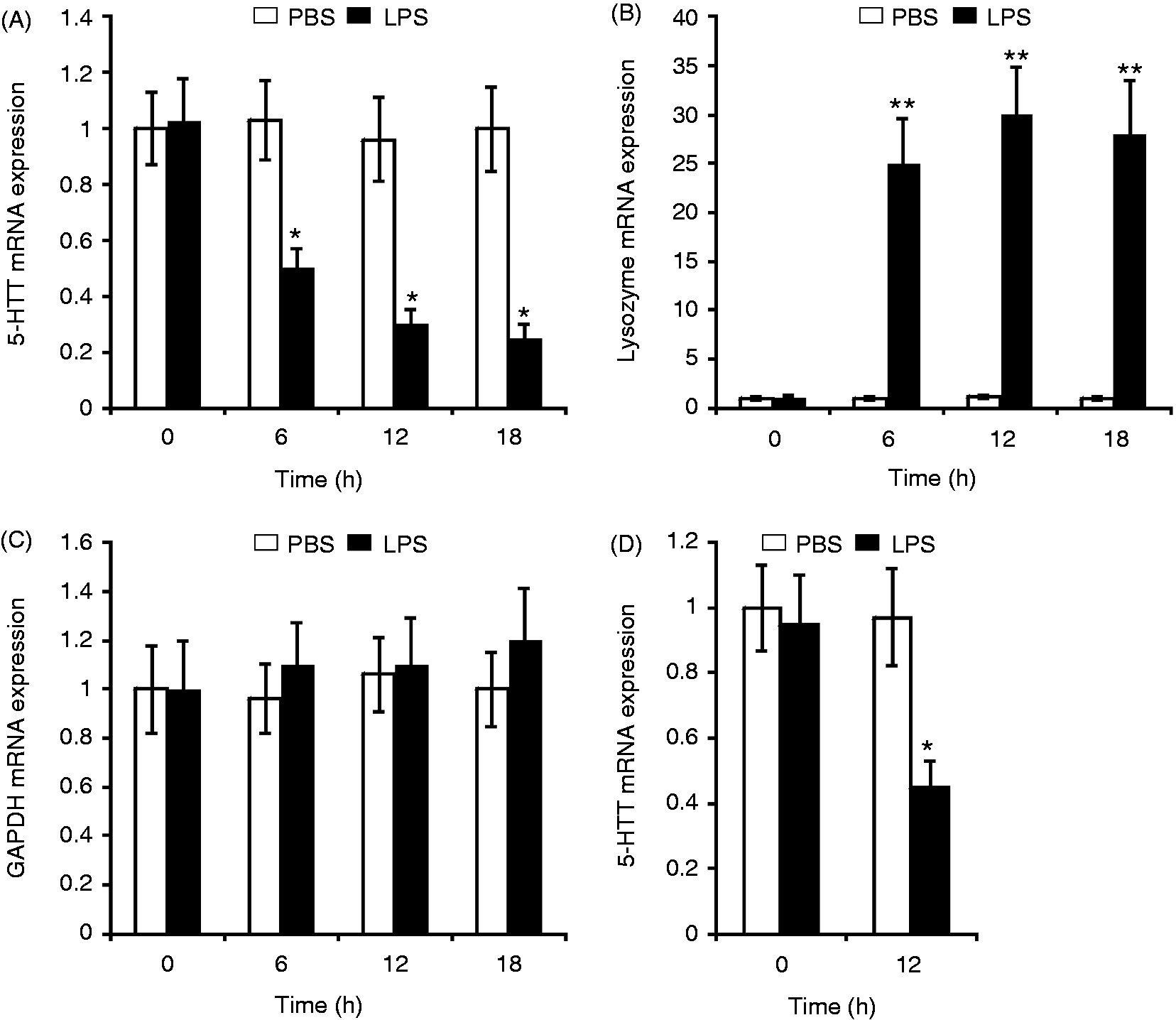
LPS and C/EBPß repress 5-HTT gene promoter activity
To examine whether LPS transcriptionally regulates 5-HTT promoter activity, CAT reporter gene constructs, which were controlled by the upstream region containing the 5-HTT promoter, were transfected into HD11 cells. As a control, we used the CAT reporter gene driven by the chicken lysozyme promoter and the −6.1-kb lysozyme enhancer, which was shown to be activated by LPS.
28
Figure 3A shows that LPS reduced the CAT expression controlled by the 1.93-kb or the 0.63-kb 5-HTT upstream region to 28% or 37%, respectively, in comparison with saline-treated cells. In contrast, the lysozyme promoter under control of the −6.1-kb lysozyme enhancer was strongly activated by LPS (Figure 3B). Thus, the results suggest that the 5-HTT gene is transcriptionally down-regulated by LPS.
LPS represses the 5-HTT promoter activity. HD11 cells were transiently transfected with CAT reporter gene constructs containing (A) the 1.93-kb fragment (p1.93CAT) or the 0.63-kb fragment (p0.63CAT) of the 5-HTT 5’-flanking region or (B) with a CAT construct (pcEPCAT) containing the lysozyme promoter and the far upstream –6.1 kb enhancer. Following transfection, cells were treated with PBS or 10 µg/ml LPS. Twenty-four hours after transfection, cells were harvested for CAT assays. Relative CAT activities are relative to the CAT activity in cells transfected with p1.93CAT or pcEPCAT and treated with PBS. Values are expressed as mean values (n = 3) ± SEM.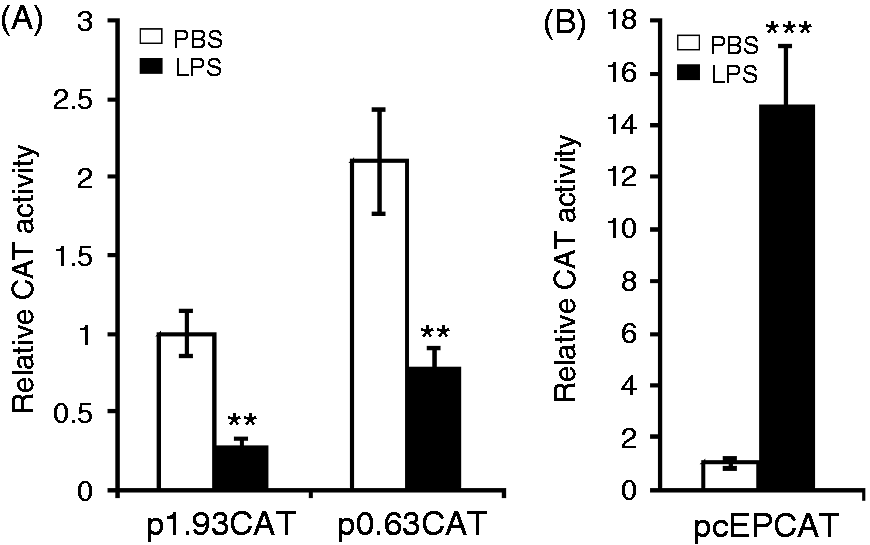
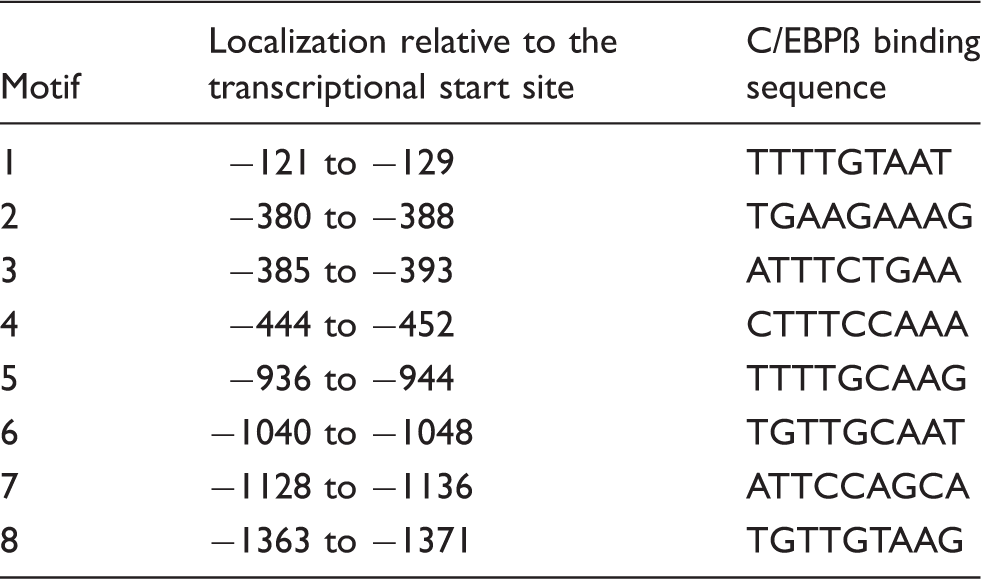
C/EBPβ putative binding sites in the 5-HTT 5’-flanking region. 29
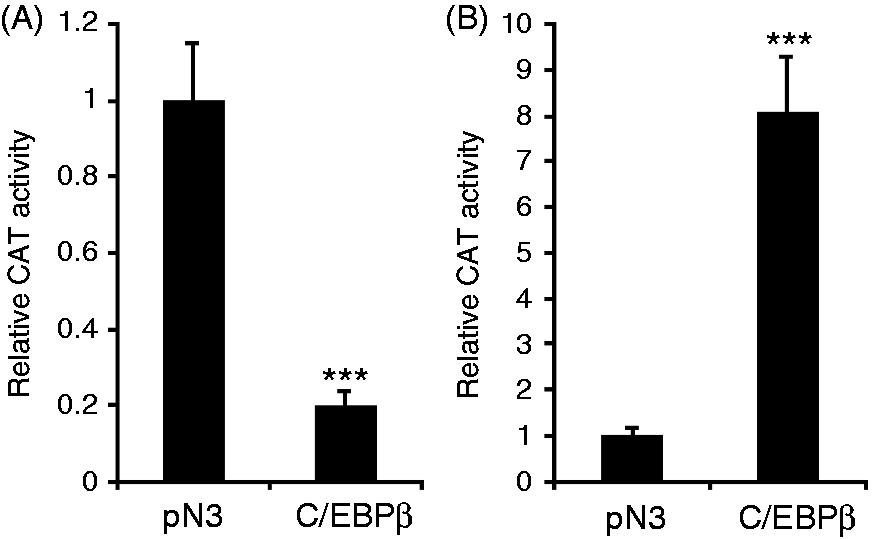
C/EBPβ down-regulates 5-HTT promoter activity. CAT constructs (A) p1.93CAT and (B) pcEPCAT were co-transfected with pN3 for control or with the C/EBPβ expression vector pN3-C/EBPβ into HD11 cells. Twenty-four hours after transfection, cells were harvested for CAT assays. Values are expressed as mean values (n = 3) ± SEM.
LPS induces increased C/EBPß binding to the upstream region of the 5-HTT gene
Analyzing interactions between C/EBPβ and the potential C/EBPβ binding sites in response to LPS, we performed band shift assays with nuclear extracts from HD11 cells treated with LPS and a
32
P-labeled probe containing a putative C/EBPβ motif. As shown in Figure 5A, two prominent binding complexes of M6 containing the putative C/EBPβ binding motif 6 (5′-TGTTGCAAT-3′) (Table 1) were formed with the nuclear extracts. The sequence-specific binding was confirmed by competition experiments with unlabeled oligonucleotides. The formation of the complexes was completely inhibited by the homologous competitor M6 and a double-stranded oligonucleotide containing a well-known C/EBP binding site described by Vinson et al.,
31
but not by double-stranded oligonucleotides containing point mutations within the C/EBPβ consensus sequence or a NF-κB binding motif of HIV-1. The binding complexes were strongly activated with nuclear extracts from cells treated with LPS (Figure 5B). Next, to confirm the presence of C/EBPβ in the binding complexes, we performed immunomobility shift assays using Abs against C/EBPβ. Figure 5C shows that super-shifted bands were caused by C/EBPβ-specific Abs, but not by unspecific IgG or by Abs against NF-кB p50, indicating that C/EBPβ was a component of the binding complexes.
C/EBPβ binds to the 5’-flanking region of the 5-HTT gene. (A) EMSA was performed with
32
P-labeled probe M6 containing the potential C/EBPβ binding motif 6 located −1040 to −1048 relative to the transcriptional start site and nuclear extract from HD11 cells treated with 10 µg/ml LPS for 2 h. Competition assay was carried out with a 100-fold excess of unlabeled oligonucleotides containing motif 6 (M6), two mutated sequences of C/EBPβ motif 6 (m1 and m2), a C/EBPβ binding site (CE) described by Vinson et al.,
31
and a NF-κB binding motif of HIV-1 (κB). Arrows indicate two prominent binding complexes. (B) Nuclear extracts from HD11 cells treated with 10 µg/ml LPS for 0, 2 and 4 h were analyzed by EMSA with the
32
P-labeled probe M6. (C) Immunomobility shift assay.
32
P-Labelled probe M6 was incubated with nuclear extract from HD11 cells treated with 10 µg/ml LPS for 4 h in the presence of unspecific IgG (IgG), Abs against C/EBPβ (α-C/EBPβ) or NF-κB p50 (α-p50).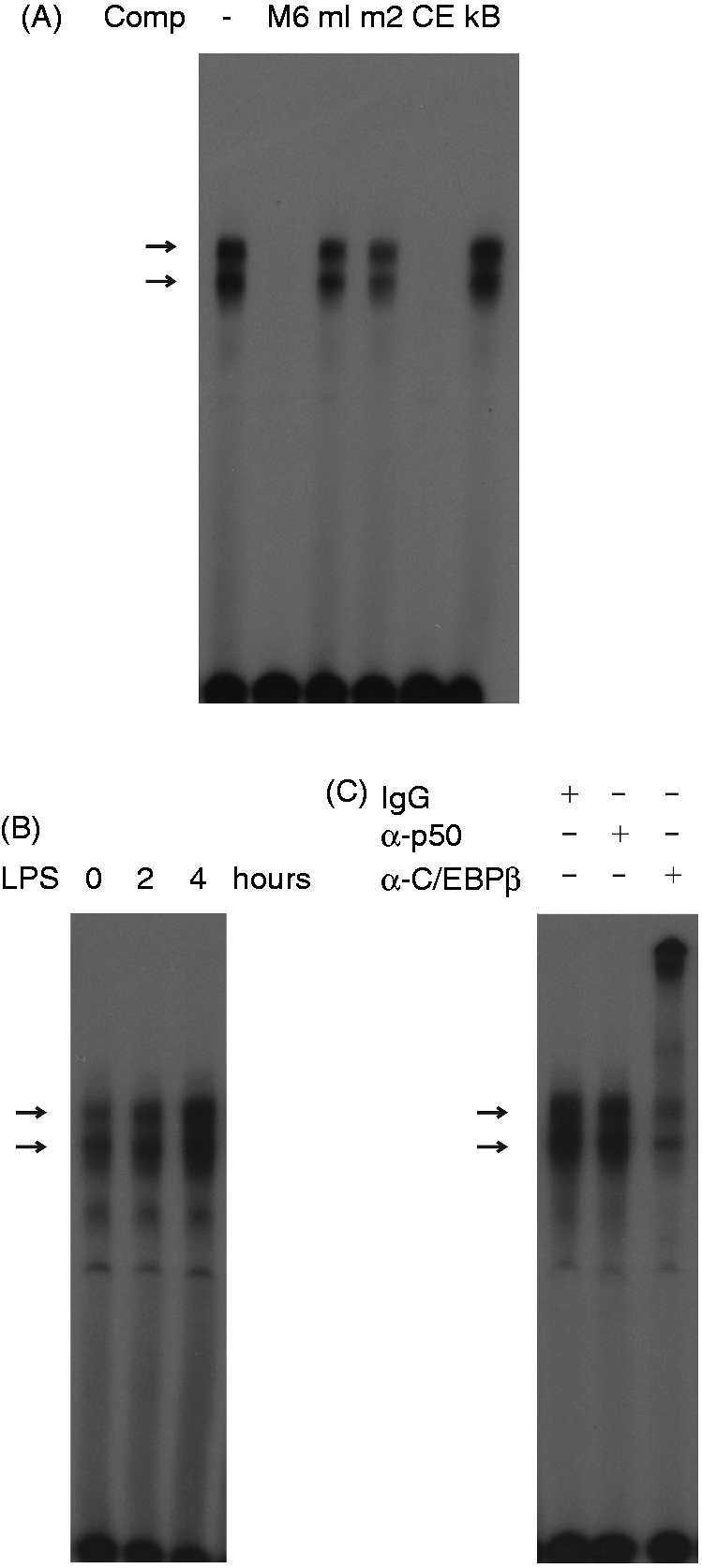
To confirm the interaction of C/EBPβ with the endogeous 5-HTT upstream region in vivo, quantitative chromatin immunoprecipitation (ChIP) assays were performed by immunoprecipitation of fragmented cross-linked chromatin from HD11 cells using Abs against C/EBPβ. DNA was isolated from the immunoprecipitated chromatin and analyzed by quantitative real-time PCR for sequences enriched in chromatin immunoprecipitated with anti-C/EBPβ Abs. For this purpose, primer pairs were used for amplification of DNA fragments A, B and C. Fragments A and B are located between −1103 and −1052 and between −415 and −365 relative to the transcriptional start site, and thus close to the C/EBPβ binding motifs; fragment C is located about 10.2 kb upstream of the transcriptional start site. As shown in Figure 6A, neither fragment A nor fragment B nor fragment C was significantly enriched in chromatin immunoprecipitated by control IgG. In contrast, fragments A and B were several-fold enriched over the control fragment C in C/EBPβ-immunoprecipitated chromatin from untreated cells, as well as in LPS-activated cells (Figure 6B). Figure 6B further shows that treatment of cells with LPS resulted in increased binding of C/EBPβ to the 5-HTT promoter region.
Chromatin immunoprecipitation with C/EBPβ. Cross-linked chromatin fragments from untreated HD11 cells and from cells treated with 10 µg/ml LPS were precipitated with (A) unspecific IgG or (B) Abs against C/EBPβ. DNA purified from the immunoprecipitates was analyzed by quantitative real-time PCR using primers for amplification of the fragments A, B and C. The relative enrichment of the fragments was calculated by comparison to the enrichment of fragment C in IgG-precipitated chromatin from untreated cells. Values are expressed as mean values (n = 3) ± SEM.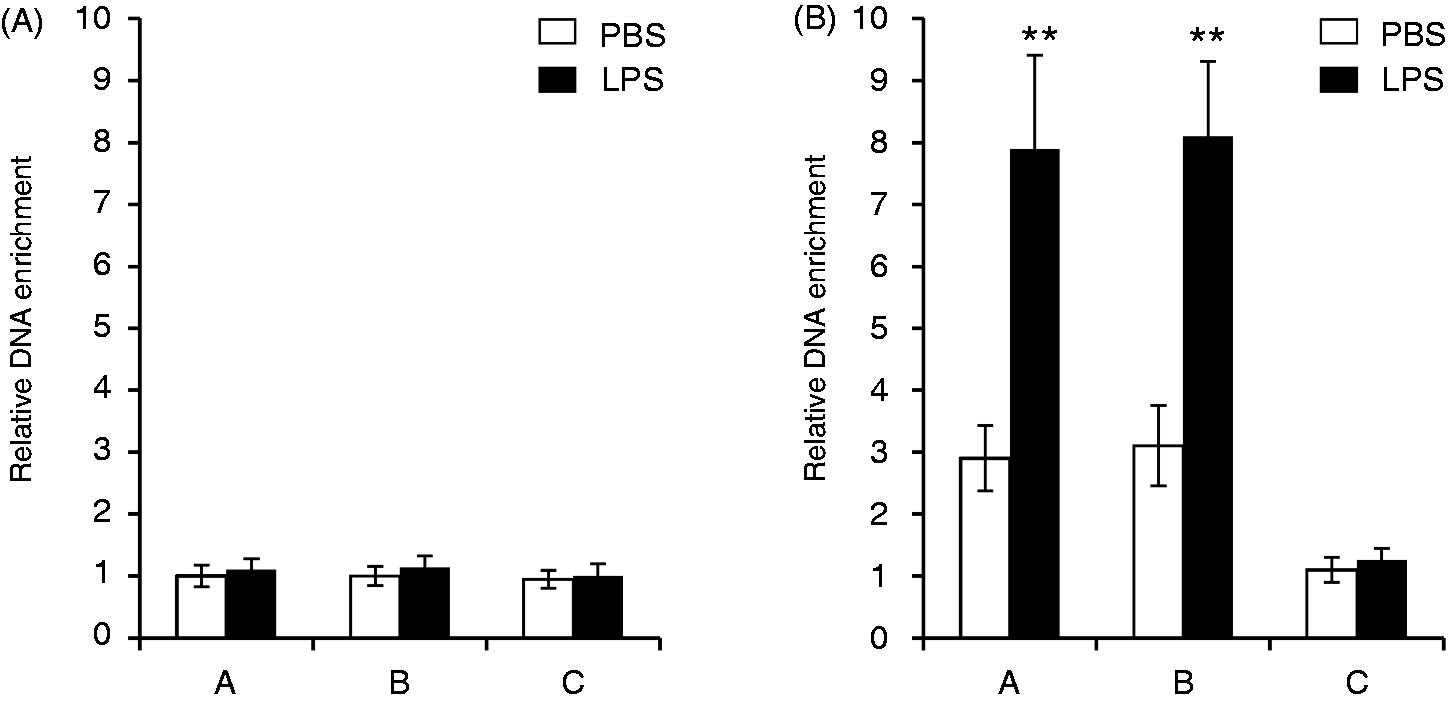
C/EBPß is involved in the LPS down-regulation of 5-HTT gene expression
To examine whether C/EBPβ plays a role in the LPS inhibition of the 5-HTT gene expression, we analyzed the 5-HTT mRNA expression in LPS-treated HD11 cells, in which C/EBPβ was selectively knocked down by transfection with C/EBPβ siRNA expression vectors. EMSA with nuclear extracts from transfected cells showed that C/EBPβ siRNA, when compared with the control siRNA, effectively caused a decrease in C/EBPβ binding to C/EBPβ binding motif 6 (Figure 7A). In contrast, the 5-HTT mRNA expression was increased in cells transfected with C/EBPβ siRNA expression vectors, suggesting that C/EBPβ negatively regulated the basal expression of the 5-HTT gene. As shown above, the treatment of cells with LPS resulted in a strong decrease in 5-HTT expression. However, this negative effect of LPS on 5-HTT expression was attenuated by knockdown of C/EBPβ in cells transfected with C/EBPβ siRNA expression vectors (Figure 7B). Two-way ANOVA also revealed a significant effect of both LPS and C/EBPβ siRNA, but no significant interaction between two variables (P > 0.05). Thus, the data obtained confirm the role of C/EBPβ in LPS inhibiting the 5-HTT expression in macrophage HD11 cells.
Negative regulation of 5-HTT gene expression by C/EBPβ. HD11 cells were transfected with C/EBPβ targeting siRNA expression vectors (siC/EBPβ) or with the negative control plasmid pRNA-CMV3.1/Neo-CTL (siCtrl). Twenty-four hours after transfection, cells were treated with PBS or 10 µg/ml LPS for further 18 h. (A) EMSA was performed with nuclear extracts from transfected cells and
32
P-labeled probe M6 containing the C/EBPß binding motif 6. (B) 5-HTT and GAPDH mRNA expression was analyzed by real-time RT-PCR. The 5-HTT mRNA levels were normalized to GAPDH. Values are expressed as mean values (n = 3) ± SEM.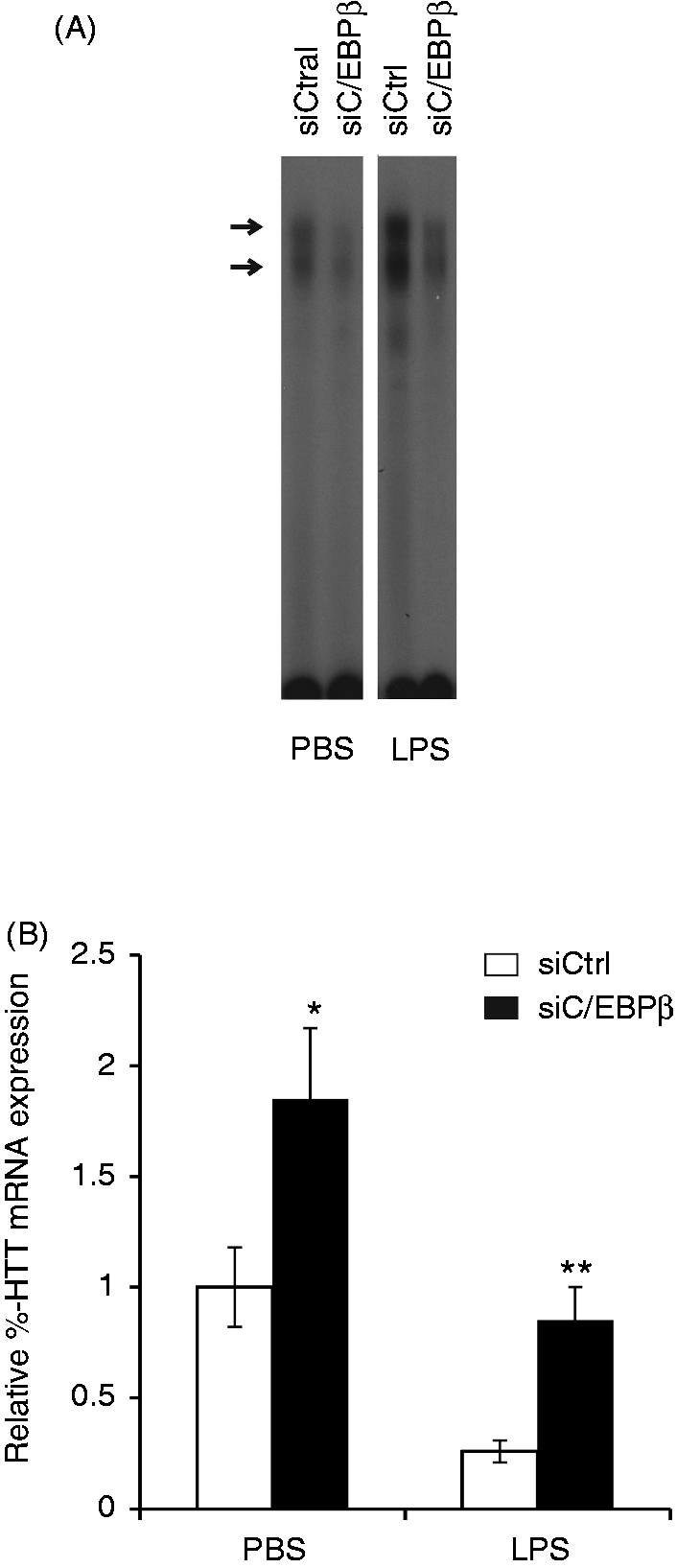
Discussion
Acute peripheral LPS application has been shown to induce rapidly 5-HTT activity in mouse brain, which was mediated by IL-1β and the p38 mitogen activated protein kinase (MAPK) pathway.27,32 On the one hand, our results show that chronic (6–18 h) treatment of chicken primary macrophages and macrophage HD11 cells with LPS produced decreases in 5-HTT mRNA expression, as well as in 5-HTT activity, measured by [3H]-5-HT uptake. On the other hand, short-term exposure of HD11 cells to LPS (1–4 h) did not affect the 5-HTT mRNA levels (data not shown). The decrease in 5-HTT mRNA expression was more pronounced than that in 5-HT uptake in LPS-treated HD11 cells (Figure 1A vs. Figure 2). It is possible that, like murine macrophages, the HD11 cell line also possesses a macrophage-specific active 5-HT uptake system and rapidly metabolizes 5-HT to 5-hydroxyindole acetic acid. 33
The negative effect of LPS on 5-HTT expression was attenuated by knockdown of endogenous C/EBPβ, indicating that this transcription factor plays an active role in down-regulating transcription of the 5-HTT gene in response to LPS. ChIP assays further demonstrated that C/EBPβ directly binds to the 5′-flanking region of the 5-HTT gene and that LPS increases C/EBPβ binding. This increased binding is possibly due to the increase in C/EBPβ gene transcription in LPS-activated HD11 cells. 30 Unusually, the 5-HTT promoter region contains a high number of sequence motifs that match the C/EBPβ consensus binding sequence (Table 1). Using band shift assays, we found that all these eight motifs bound to C/EBPβ (data not shown). Because band shift assays with all the motifs produced similar results, representative results obtained with motif 6 are shown in Figure 5A–C.
Besides being a transcriptional activator, C/EBPβ has been shown to be involved in the transcriptional repression of a variety of genes. For instance, C/EBPβ can functionally interact and repress p53 to promote cell survival downstream of DNA damage, 34 miR-145 35 or let-7i expression by induced binding of a transcriptional C/EBPβ/NFkBp50 silencer complex to the let-7i promoter following microbial infection. 36 At least three isoforms—38 kDa (LAP*), 35 kDa (LAP) and 20 kDa (LIP)—can be produced from C/EBPβ mRNA. 37 In contrast to LAP, LIP lacking the activation domain acts as a transcriptional repressor. For example, LPS has been shown to repress C/EBPα gene expression through increased binding of LIP to the C/EBPα promoter. 38
LPS has been shown to be responsible for several clinical symptoms of sepsis with Gram-negative bacteria. It induces fever, locomotor depression, hyperemia by increased vascular permeability, anorexia and so on.16–19 Interestingly, 5-HT has been suggested, at least in part, to mediate these LPS-induced pathophysiological processes.19,39–41 Thus, our results showing that 5-HTT expression is down-regulated in response to long-term LPS treatment are appropriate in this context. A decrease in 5-HTT activity results in a reduction of 5-HT uptake leading to higher extracellular 5-HT levels, which may, in turn, mediate the LPS-induced pathophysiological effects. Long-term exposure to LPS has also been observed to reduce 5-HTT activity in human enterocyte-like Caco-2 cells. However, the decrease in 5-HTT activity was not attributed to a decrease in 5-HTT mRNA levels, but might be mediated by the protein kinase C pathway. 42 One possible explanation for the discrepancy between our results and those from other studies is the use of different lines (chicken HD11 cells vs. human Caco-2 cells). The activation of C/EBPβ appears to be controlled by different mechanisms, including nuclear translocation following phosphorylation, 43 increased transcription and so on. 30 It is quite possible that different cell types differ in their activation of C/EBPβ in response to inflammatory stimuli. 44 Alternatively, the regulation mechanism of the chicken 5-HTT gene in response to LPS might be different from that of the human 5-HTT gene. Indeed, a careful sequence analysis of the 1.9-kb 5′-flanking region of the chicken 5-HTT gene and the human 5-HTT gene revealed that the sequences considerably differ from each other. For instance, whereas the 1.9-kb chicken sequence contains eight C/EBPβ binding sites (Table 1), its human counterpart has only two sequence motifs that match the consensus sequence for C/EBPβ (data not shown).
The chicken cell line HD11 used in this study expresses 5-HTT, is able to take up 5-HT and responds to LPS. Thus, it may be useful for studies of the innate immune and serotonergic systems. The results we obtained with this cell line will serve to strengthen the hypothesis that 5-HTT may directly contribute to the regulation of extracellular 5-HT levels in the inflammatory response to LPS. These results provide the first evidence indicating that long-term exposure of macrophages to LPS reduces the transcription of the chicken 5-HTT gene and 5-HTT activity by activation of C/EBPβ binding to the 5-HTT 5′-flanking region. This is in comparison to previous studies that demonstrated that short-term administration of LPS rapidly induces 5-HTT activity through IL-1β and the p38 MAPK-dependent signaling pathway.27,32 Thus, depending on the duration of the exposure to LPS, the LPS-induced alteration of 5-HTT activity seems to be regulated by different mechanisms involving transcriptional and post-translational processes.
Footnotes
Funding
This research received no specific grant from any funding agency in the public, commercial or not-for-profit sectors.
Conflict of interest
The authors do not have any potential conflicts of interest to declare.
Acknowledgements
We thank T. Tysper for skillful technical assistance, Dr. L Schrader (Friedrich-Loeffler-Institut, Celle) for critical reading of the manuscript and Dr. A Leutz (Max Delbrück Center for Molecular Medicine, Berlin) for the anti-C/EBPβ antiserum.
