Abstract
The IL-1 family consists of 11 members that play an important role as key mediators in inflammation and immunity. Here, we report the identification of a new member of the IL-1 family (IL-1Fm2) that is present in species belonging to the most evolutionarily advanced group of teleost fish (Series Percomorpha), including Perciformes, Beloniformes, Gasterosteiformes, Cyprinodontiformes and Pleuronectiformes. However, IL-1Fm2 seems to be absent in Tetraodontiformes, which also belong to the Percomorpha. The expression pattern of gilthead seabream IL-1Fm2 revealed that although it was hardly induced by PAMPs, the combination of PAMPs and recombinant IL-1Fm2 synergistically induced its expression in macrophages and granulocytes. In addition, recombinant IL-1Fm2 was able to activate the respiratory burst of seabream phagocytes and to synergistically induce the expression of IL-1β, TNF-α, IL-8 and IL-10 when combined with PAMPs. Finally, although gilthead seabream IL-1Fm2 did not show a conserved caspase-1 processing site, macrophages processed IL-1Fm2 before being released. However, both pan-caspase and caspase-1 inhibitors failed to inhibit the processing and release of IL-1Fm2. These results demonstrate an important role of IL-1Fm2 in the regulation of fish immune responses, shed light on the evolution of the IL-1 family in vertebrates and point to the complexity of this cytokine family.
Introduction
The IL-1 family consists of 11 members, five of which are relatively well characterized, namely IL-1α, IL-1β, IL-1ra, IL-18 and IL-33. 1 They play an important role as key mediators in inflammation, microbial invasion, immunological reactions and tissue injury. 1 IL-1β was the earliest cytokine to be discovered and cloned in fish2,3 and has now been found in many bony fish, including salmonids, cyprinids, gadoids, perciforms and anguilliforms, and cartilaginous fish.4,5
Although fish IL-1β has some similarities to its mammalian counterparts, there are also differences, the most intriguing being the absence of a conserved caspase-1 processing site in the non-mammalian vertebrate IL-1β sequences. 5 In mammals, caspase-1 is activated in a cytosolic molecular platform called the inflammasome.6,7 Interestingly, while it has recently been found that zebrafish (Danio rerio) inflammatory caspase A and caspase B 8 and European sea bass (Dicentrarchus labrax) caspase-1 9 can process IL-1β, caspase-1 is not involved in the processing of gilthead seabream (Sparus aurata) IL-1β. 10 In addition, seabream macrophages fail to respond to classical NLRP3 activators, such as ATP, nigericin, and alum and monosodium urate crystals.10–12 Even most puzzling is the observation that invasive Salmonella typhimurium, which activates NLRC4 inflammasome in mammals,13,14 and cell swelling, which activates NLRP3 inflammasome in mammals and fish, 11 both are able to activate caspase-1 in gilthead seabream, but also fail to trigger the processing and secretion of IL-1β.10,11
In some primitive fish species, such as salmonids and cyprinids, more than one il1b gene has been found,4,5 but this is not common to all species of these phylogenetic groups, as the zebrafish, which is also a cyprinid, has a single il1b gene. However, the finding of several il1b genes in some of the species is not totally unexpected, as they have a tetraploid ancestry and are likely to have multiples of certain genes. In addition, the presence of multiple copies of il1 family genes is not restricted to fish because a number of novel homologue members of the IL-1 family have also been found in humans. 1
Here, we report the identification of a new member of the IL-1 family (IL-1Fm2) that is present in species belonging to the most evolutionarily advanced group of teleost fish (Series Percomorpha), including the gilthead seabream (S. aurata), European sea bass (D. labrax) and tilapia (Oreochromis niloticus) (Perciformes); medaka (Oryzias latipes, Beloniformes); stickleback (Gasterosteus aculeatus, Gasterosteiformes); platyfish (Xiphophorus maculatus, Cyprinodontiformes); and Japanese flounder (Paralichthys olivaceus,; Pleuronectiformes). Interestingly, on investigation of the genomes, IL-1Fm2 seems to be absent in pufferfishes (Takifugu rubripes and Tetraodon nigroviridis, Tetraodontiformes), which also belong to Percomorpha. The expression pattern of gilthead seabream IL-1Fm2 and its biological activity demonstrate its importance in the regulation of the immune response of fish.
Materials and methods
Identification of teleost IL-1Fm2 and sequence analysis
A partial sequence of gilthead seabream IL-1Fm2 was identified by BLAST searching in the Expressed Sequence Tag (EST) database of the European Nucleotide Archive (ENA, accession number AM957809). A 3’-Rapid Amplification of cDNA Ends (RACE) strategy using the primers indicated in Supplementary Table S1 was used to obtain the full coding sequence, 15 which was deposited in ENA with accession number HF936678. The deduced IL-1Fm2 protein was analyzed for similarity with other known sequences using the BLAST program within the National Center for Biotechnology Information server (http://blast.ncbi.nlm.nih.gov/Blast.cgi). A direct comparison between two sequences was performed using the ALING program, while multiple sequence alignment was carried out with the CLUSTALW2 program from the European Bioinformatics Institute (http://www.ebi.ac.uk/).. A phylogenetic tree was constructed based on the amino acid sequence alignments with the CLUSTALW2 program using the neighbor-joining inference and was then displayed with the TreeView X program. 16 The domains of the protein deduced from the nucleotide sequence were determined by comparison with the PFAM database. 17 Finally, synteny analysis was performed by searching the available genomes and using GENSCAN (http://genes.mit.edu/GENSCAN.html) 18 to predict gene sequences within them. The accession numbers of all genes analyzed are indicated in Supplementary Table S2.
Animals
Healthy specimens (300 g mean mass) of the hermaphroditic protrandrous marine fish gilthead seabream were kept at the Spanish Oceanographic Institute (Mazarrón, Murcia) in 14 m3 running seawater aquaria (dissolved oxygen 6 ppm, flow rate 20% aquarium volume/h) with natural temperature and photoperiod, and fed twice a day with a commercial pellet diet (Skretting, Burgos, Spain). Fish were fasted for 24 h before sampling.
Head kidney, spleen, thymus, liver, peritoneal exudate cells, gills, blood, brain, gut and skin were collected from healthy individuals. For experimental infections, fish were injected i.p. with 1 ml of PBS alone or containing a sublethal dose (108) of exponentially growing Vibrio anguillarum R82 cells (serogroup 01). 19 Head kidney, spleen, thymus, liver, peritoneal exudate cells, gills and blood were obtained 4 h after bacterial challenge and processed for subsequent reverse transcription-quantitative PCR (RT-qPCR) (see below).
The experiments performed comply with the Guidelines of the European Union Council (86/609/EU) and the Bioethical Committee of the University of Murcia (approval no. #537/2011) for the use of laboratory animals.
Production of recombinant seabream IL-1Fm2
Recombinant seabream IL-1Fm2 was produced in Escherichia coli. Briefly, the open reading frame (ORF) of IL-1Fm2 encoding the putative mature protein (residues 118–270) was synthesized, cloned in vector E3, produced as an N-terminal 6 × His fusion protein in E. coli, obtained from inclusion bodies with 6 M guanidine hydrochloride and purified by immobilized-metal affinity chromatography using Ni-HiTrap columns (GenScript, Piscataway, NJ, USA).
Cell culture and treatments
Seabream head–kidney leukocytes (bone marrow equivalent of fish) obtained and cultured as described elsewhere 20 were stimulated for 4 h at 23℃ with 50 µg/ml phenol-extracted genomic DNA from V. anguillarum ATCC19264 cells (VaDNA). 21 Some experiments were conducted using purified acidophilic granulocytes obtained by magnetic-activated cell sorting using a mAb specific to these cells 21 and macrophage monolayers. 22 Macrophages and acidophilic granulocytes were stimulated for 0.5 to 24 h at 23℃ with 50 µg/ml VaDNA, 1 ng/ml flagellin (Invivogen, Madrid, Spain), 25 µg/ml poly I:C (Invivogen, San Diego, CA, USA), 10 µg/ml LPS from E. coli (EcLPS; Sigma-Aldrich, Madrid, Spain), 10 µg/ml Porphyromonas gingivalis LPS (PgLPS; Invivogen) or 10 µg/ml muramyldipeptide (MDP; Sigma-Aldrich).21,23
To examine the effect of recombinant seabream IL-1Fm2 on the macrophages and acidophilic granulocytes, they were stimulated for 3 (macrophages) or 16 h (acidophilic granulocytes) at 23℃ with 50 µg/ml VaDNA and 10 ng/ml recombinant seabream IL-1Fm2. cDNA was produced from the cells, and expression of IL-1β, IL-1Fm2, TNF-α, IL-8 and IL-10 was then analyzed using RT-qPCR. Protein levels of IL-1Fm2 were also measured within macrophage extracts and supernatants using the mAb against IL-1Fm2 (see below). In some experiments, macrophages were pretreated for 1 h with the pan-caspase inhibitor Q-VD-OPh (50 µM; Sigma-Aldrich) or the specific caspase-1 inhibitor Ac-YVAD-CMK (100 µM; Peptanova, Sandhausen, Germany)10,24 before being stimulated.
Respiratory burst assays
Respiratory burst activity was measured as the luminol-dependent chemiluminescence produced by acidophilic granulocytes after different stimulation times. 21 This was brought about by adding 100 µM luminol and 1 µg/ml PMA (both from Sigma-Aldrich), while the chemiluminescence was recorded every 117 s for 1 h in a FLUOstart luminometer (BGM, LabTechnologies, Ortenberg, Germany). The values reported are the average of quadruple readings, expressed as the slope of the reaction curve from 117 to 1170 s, from which the apparatus background was subtracted.
Cell viability
Aliquots of cell suspensions were diluted in 200 µl PBS containing 40 µg/ml propidium iodide. The number of red fluorescent cells (dead cells) from triplicate samples was analyzed using flow cytometry (BD Biosciences, Madrid, Spain).
RT-qPCR
Total RNA was extracted from tissues or cell pellets with TRIzol reagent (Life Technologies, Madrid, Spain) following the manufacturer's instructions and treated with DNase I, amplification grade (1 unit/µg RNA; Life Technologies). The SuperScript III RNase H- ReverseTranscriptase (Life Technologies) was used to synthesize first-strand cDNA with oligo-dT18 primer from 1 µg of total RNA at 50℃ for 50 min. qPCR was performed with an ABI PRISM 7500 instrument (Life Technologies) using SYBR Green PCR Core Reagents (Life Technologies). Reaction mixtures were incubated for 10 min at 95℃, followed by 40 cycles of 15 s at 95℃, 1 min at 60℃, and, finally, 15 s at 95℃, 1 min at 60℃ and 15 s at 95℃. For each mRNA, the gene expression was normalized to the ribosomal proteins S18 content in each sample using the Pfaffl method. 25 The primers used are shown in Supplementary Table S1. In all cases, each PCR was performed with triplicate samples and repeated with at least two independent samples.
Western blot
Cells were lysed at 4℃ in lysis buffer (10 mM Tris-HCl pH 7.4, 150 mM NaCl, 1% Triton X-100, 0.5% NP-40, and a 1:20 dilution of the protease inhibitor cocktail P8340; Sigma-Aldrich). The protein concentrations of cell lysates were estimated by the BCA protein assay reagent (Thermo Fisher Scientific, Rockford, IL, USA) using bovine serum albumin (BSA) as a standard. Cell extracts (40 µg protein) and supernatants (clarified with a 0.45-µm filter and concentrated by precipitation with 20% trichloroacetic acid; Sigma-Aldrich) were analyzed on 15% SDS-PAGE and transferred for 50 min at 200 mA to nitrocellulose membranes (BioRad, Alcobendas, Spain). The blots were developed with a rabbit monospecific polyclonal Ab to seabream IL-1β (1:5000)10,26 or a purified mouse mAb generated against recombinant sbIL-1Fm2 (6D1E4, 1:5000; GenScript), 27 and enhanced chemiluminescence reagents (GE Healthcare, Barcelona, Spain) according to the manufacturer's protocol. Membranes were then re-probed with a 1:5000 dilution of an affinity purified rabbit polyclonal to histone H3 (#ab1791; Abcam, Cambridge, UK).
Statistical analysis
Data were analyzed by one- or two-way ANOVA followed by a Tukey multiple range test to determine differences between groups using GraphPad Prism 5.01. Statistical significance was defined as P < 0.05.
Results
Molecular characteristics of seabream IL-1FM2
A partial sequence of gilthead seabream IL-1Fm2 was identified by BLAST searching in the EST database of the ENA (accession number AM957809). A 3’-RACE strategy was used to obtain the full coding sequence, which was deposited in ENA with accession number HF936678. The IL-1Fm2 cDNA consisted of 1383 nucleotides, including a 813 bp single ORF, with a 77 bp 5' untranslated region (5'UTR) and a 493 bp 3'UTR containing a poly(A) tail and a putative polyadenylation signal AATAAA located 11 nucleotides upstream of the poly(A) tail (Figure 1). The translation of the ORF predicted a 270-amino acid polypeptide (calculated molecular mass: 30.6 ku) that lacked a signal peptide and a conserved caspase-1 cut site (Figures 1 and 2). A potential glycosylation site (NXT) was predicted at residues 106–108 (Figure 1).
Molecular characteristics of gilthead seabream IL-1Fm2. Full-length cDNA sequence and deduced amino acid sequence of gilthead seabream IL-1Fm2. The start and stop codons, the polyadenylation signal (AATAAA) in the 3' UTR, and a potential glycosylation site (NFT) are highlighted in gray. The IL-1 family signature [FC]-x-S-[ASLV]-x(2)-P-x(2)-[FYLIV]-[LI]-[SCA]-T-x(7)-[LIVM] (PDOC00226), is underlined. ENA accession number is HF936678. Molecular characteristics of teleost IL-1Fm2 proteins. Multiple alignment of the teleost IL-1Fm2 proteins with known vertebrate IL-1β proteins. Identical (*) and similar (. or :) residues identified in all the proteins are indicated. The IL-1 family signature [FC]-x-S-[ASLV]-x(2)-P-x(2)-[FYLIV]-[LI]-[SCA]-T-x(7)-[LIVM] (PDOC00226) is underlined. The arrow shows the caspase-1 cut site of human IL-1β sequence. The residues that form the secondary structure of 12 β-sheets in human IL-1β are highlighted in gray. The accession numbers are indicated in Supplementary Table S2.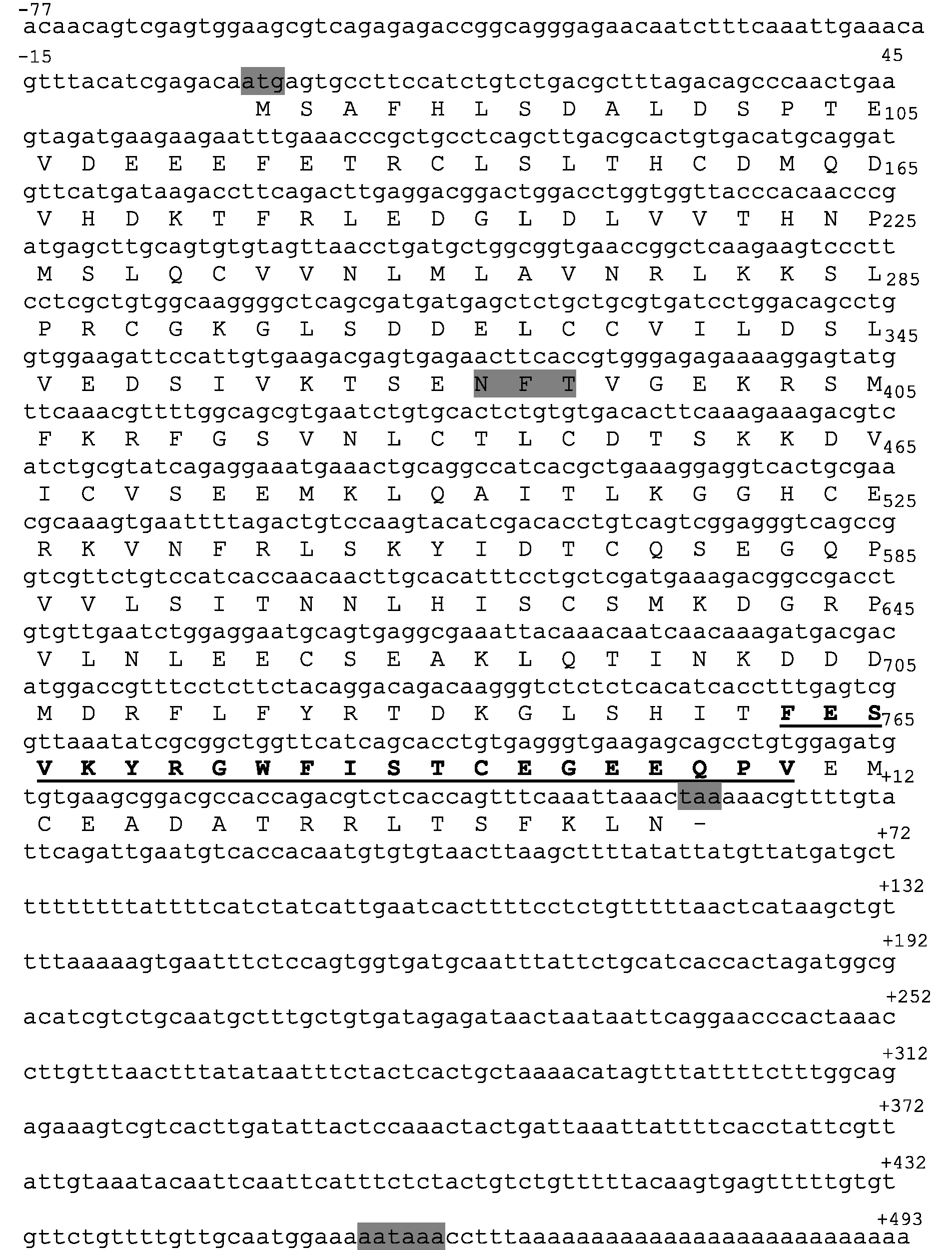
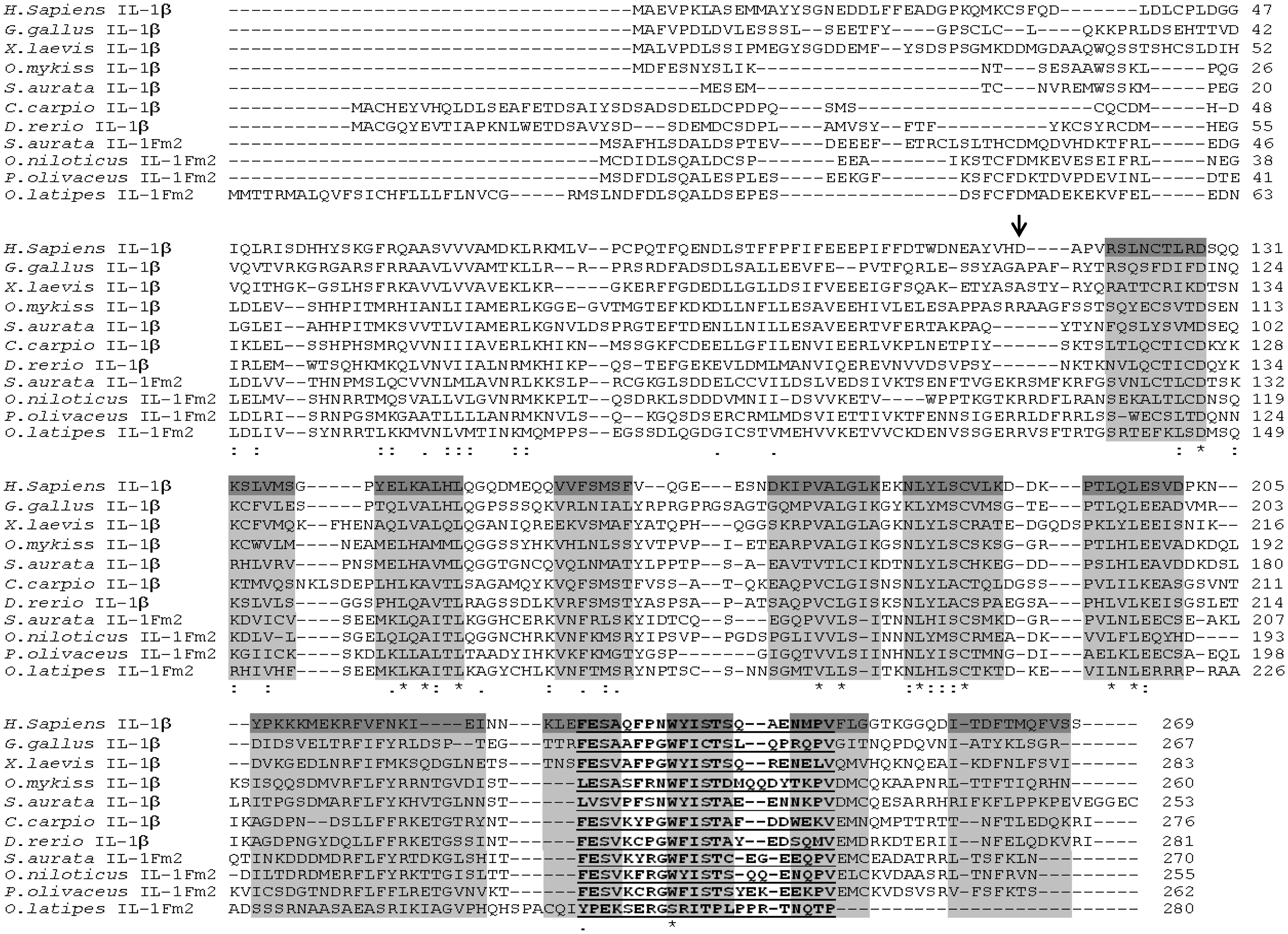
A BLASTP search versus non-redundant protein sequence databases retrieved as most significant hits several teleost genes annotated as il1b-like (E-value < 10−64, 45–48% amino acid identity) followed by teleost il1b genes (E-value < 10−43, 32–39% amino acid identity) and mammalian IL1B genes (E-value < 10−15, 27–31% amino acid identity). Notably, seabream IL-1Fm2 showed 26.4% and 47.1% amino acid identity and similarity, respectively, with seabream IL-1β. In addition, analysis of the domains with PFAM showed that the seabream IL-1Fm2 had an IL-1 propeptide domain (PF02394) from residues 6 to 100 and an IL-1 domain (PF00340) between positions 149 and 269. A multiple alignment generated by CLUSTALW showed the highest amino acid conservation in the predicted β-strands regions (Figure 2). In addition, the IL-1 family signature, [FC]-x-S-[ASLV]-x(2)-P-x(2)-[FYLIV]-[LI]-[SCA]-T-x(7)-[LIVM] (PDOC00226), was also well conserved (Figures 1 and 2). These results, taken together, confirm that gilthead seabream IL-1Fm2 belongs to the IL-1 family.
IL-1Fm2 is present in most evolutionary advanced teleost fish
Using the gilthead seabream IL-1Fm2 sequence as query in TBLASTN search of different fish genomes, we identified a IL-1Fm2 orthologue only in several teleost species belonging to the Percomorpha Series, including the Orders Perciformes [D. labrax (HF936682) and O. niloticus (XM_003452995)], Beloniformes (O. latipes, XM_004074606), Gasterosteiformes (G. aculeatus, HF936679), Cyprinodontiformes (X. maculatus, HF952169), and Pleuronectiformes (P. olivaceus, AB720985). However, IL-1Fm2 seemed to be absent in Tetraodontiformes (T. rubripes and T. nigroviridis), which also belong to the Percomorpha. All IL-1Fm2 sequences formed a separate clade from teleost IL-1β and nIL-1Fm sequences (Figure 3). Notably, the IL-1Fm2 from P. olivaceus has very recently been identified and named novel IL-1β-like1 (nIL-1β-like1).
28
Collectively, these results further support that IL-1Fm2 is a new member of the IL-1 family.
Phylogenetic relationships of teleost IL-1Fm2. Phylogenetic tree of fish IL-1 family members. The tree was generated by the cluster algorithm using amino acid sequences. Numbers shown are percentages of 100 bootstrap replicates in which the same internal branch was observed. The horizontal lines are drawn proportional to the inferred phylogenetic distances. Vertical lines have no significance. The accession numbers are indicated in Supplementary Table S2.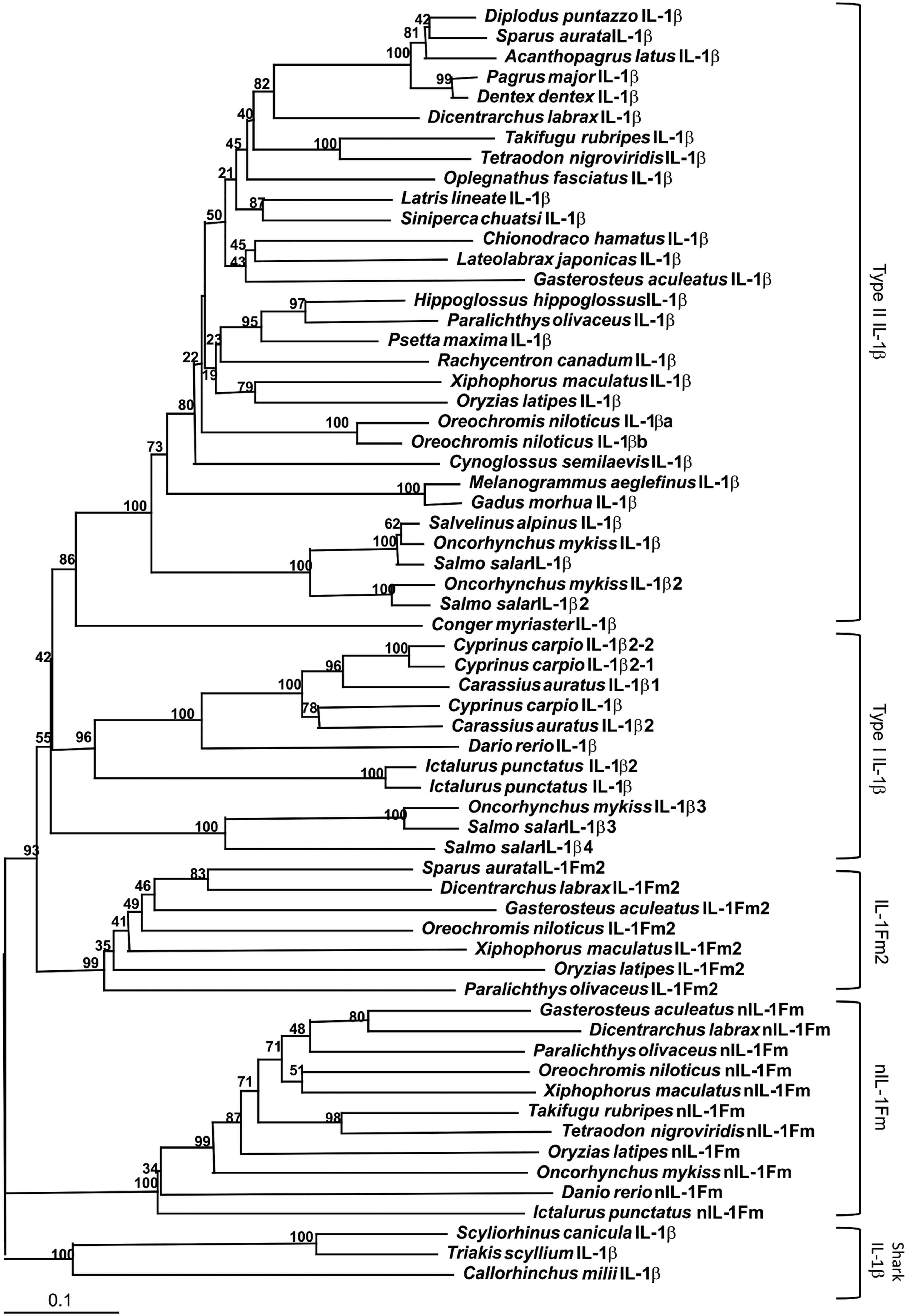
Analysis of the O. latipes, X. maculates, O. niloticus and G. aculeatus genomes revealed the IL-1Fm2 locus in chromosome 12, and scaffolds JH557184, GL831259 and 293, respectively. We also performed synteny analysis and found that the IL-1Fm2 locus of the four species was common in most of the genes, including cnmm3, neu3, cnnm4, sec14l2, ube2l3, ckmt2, rasgrf2, purb, hesx1 and polm (Figure 4). Importantly, the genes cnmm4 and purb were syntenically conserved in the teleost IL-1β and IL-1Fm2 loci. These results suggest that IL-1β and IL-1Fm2 probably arose from the fish-specific genome duplication (FSGD).
29
Synteny analysis of teleost type II IL-1β and IL-1Fm2 loci. Schematic diagrams showing the syntenically conserved genes in the type II IL-1β and IL-1Fm2 loci in stickleback (G. aculeatus), tilapia (O. niloticus), platyfish (X. maculatus) and medaka (O. latipes). The human IL-1β locus is also shown for comparison. Conserved genes are indicated in different colors and the direction of gene transcription is indicated with arrows.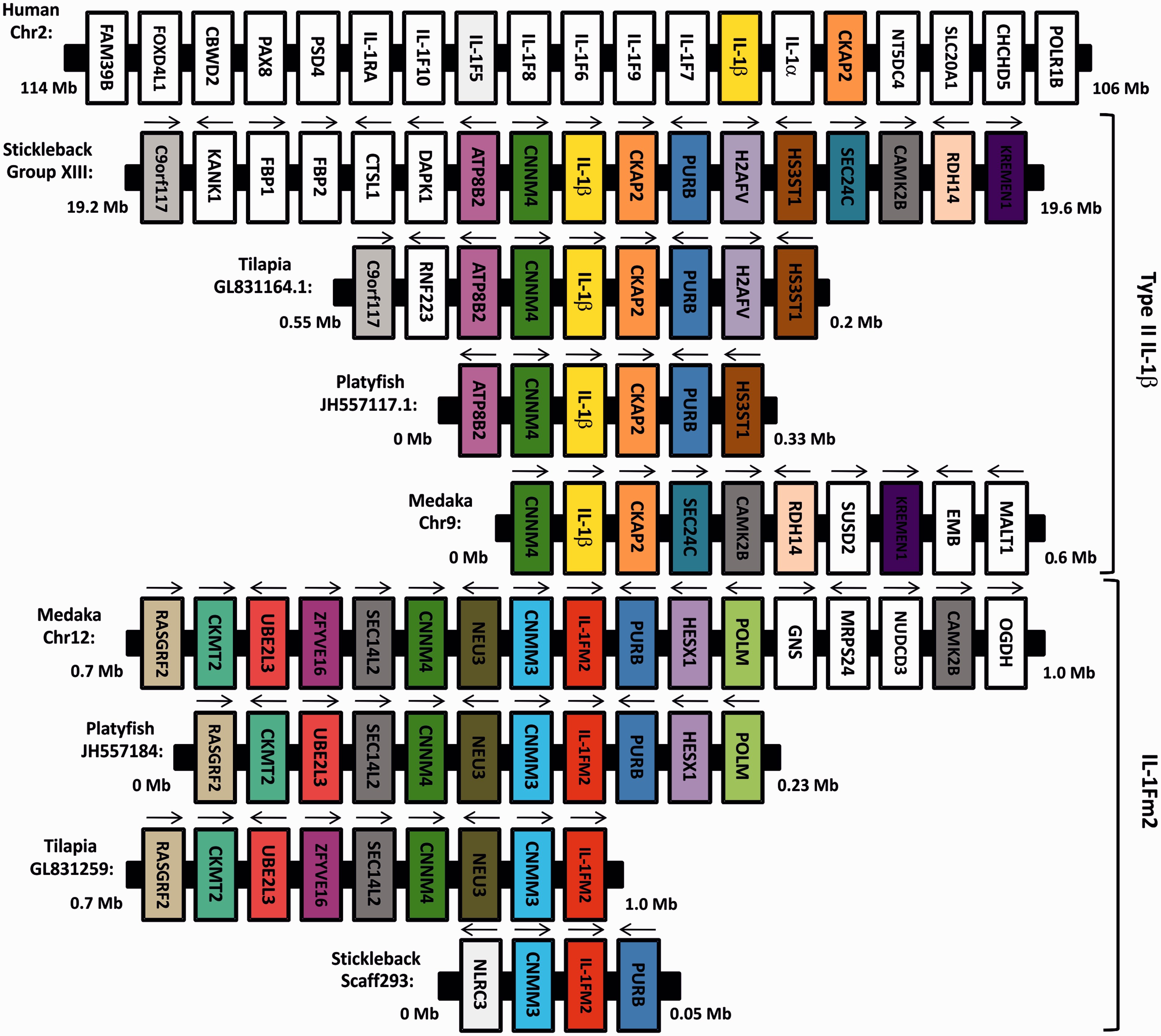
IL-1Fm2 is ubiquitously expressed in S. aurata and weakly induced after infection
Although seabream IL-1Fm2 was constitutively expressed in all tissues examined, the lower expression was found in several organs involved in the immune response, namely the spleen, head–kidney and blood, as well as the skin and peritoneal exudate (Figure 5A). Conversely, the highest expression was found in intestine, liver, gill, brain and thymus. We next examined the in vivo expression of IL-1Fm2 during a bacterial infection and found that the mRNA levels of IL-1Fm2 increased slightly in the tissues showing the lowest basal expression, namely head–kidney, spleen, peritoneal exudate and blood, but decreased within the liver, gills and thymus (Figure 5B).
IL-1Fm2 is ubiquitously expressed in gilthead seabream and weakly induced after infection. The mRNA levels of IL-1Fm2 were determined by RT-qPCR in the indicated immune tissues of control adult specimens (A) and at 4 h after challenge with 108 cells of V. anguillarum (B). Gene expression is normalized against ribosomal protein S18 (A, B) and is shown as relative to the mean of uninfected fish (B). Each bar represents the mean ± SEM of triplicate biological samples. Different letters denote statistically significant differences among the groups according to a Tukey test. The groups marked with ‘a’ did not show statistically significant differences from non-infected fish.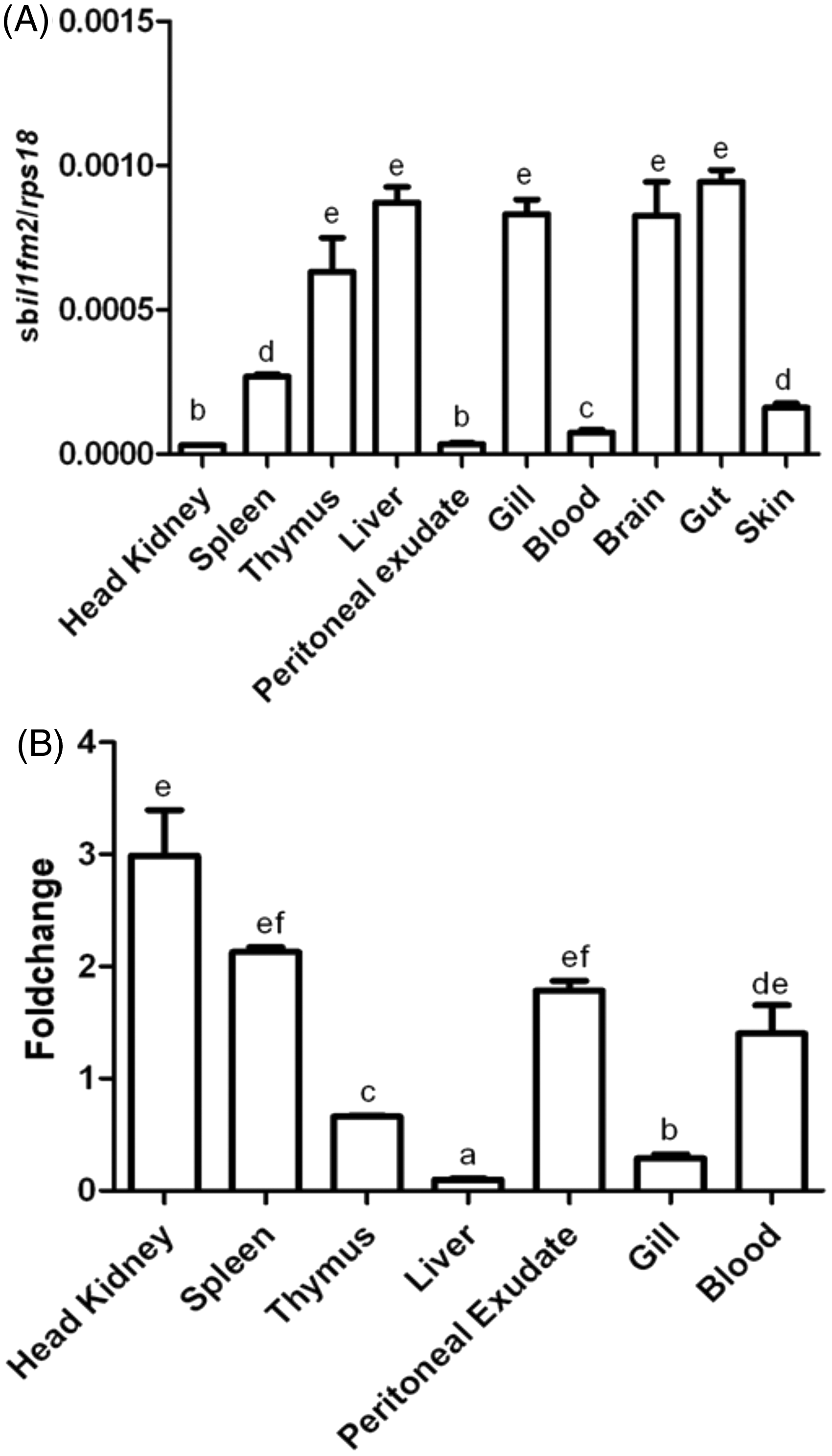
IL-1Fm2 is weakly induced by S. aurata phagocytes after pathogen-associated molecular pattern activation
The two professional phagocytic cell types of the gilthead seabream were purified and stimulated with optimal concentrations of different PAMPs.
21
It was found that only bacterial DNA and E. coli LPS were able to increase in a time-dependent manner (P < 0.0001 for the interaction according to a two-way ANOVA) the mRNA levels of IL-1Fm2 in acidophilic granulocytes (Figure 6A) and macrophages (Figure 6B). These results contrast the drastic induction (> 1000-fold) of IL-1β in the same samples,
21
suggesting that the expression of il1b and IL-1Fm2 genes is differentially regulated.
IL-1Fm2 is weakly induced by gilthead seabream phagocytes after PAMP activation. The mRNA levels of IL-1Fm2 were determined by RT-qPCR in macrophages (A) and acidophilic granulocytes (B) incubated for the indicated times in medium alone or containing 50 µg/ml VaDNA, 1 µg/ml flagellin, 25 µg/ml poly I:C, 10 µg/ml EcLPS, 10 µg/ml MDP or 10 µg/ml PgLPS. Gene expression is normalized against ribosomal protein S18 and is shown as relative to the mean of non-stimulated cells. Each bar represents the mean ± SEM of triplicate biological samples. Different letters denote statistically significant differences among the groups according to a Tukey test. The groups marked with ‘a’ did not show statistically significant differences from control cells. ND: not determined.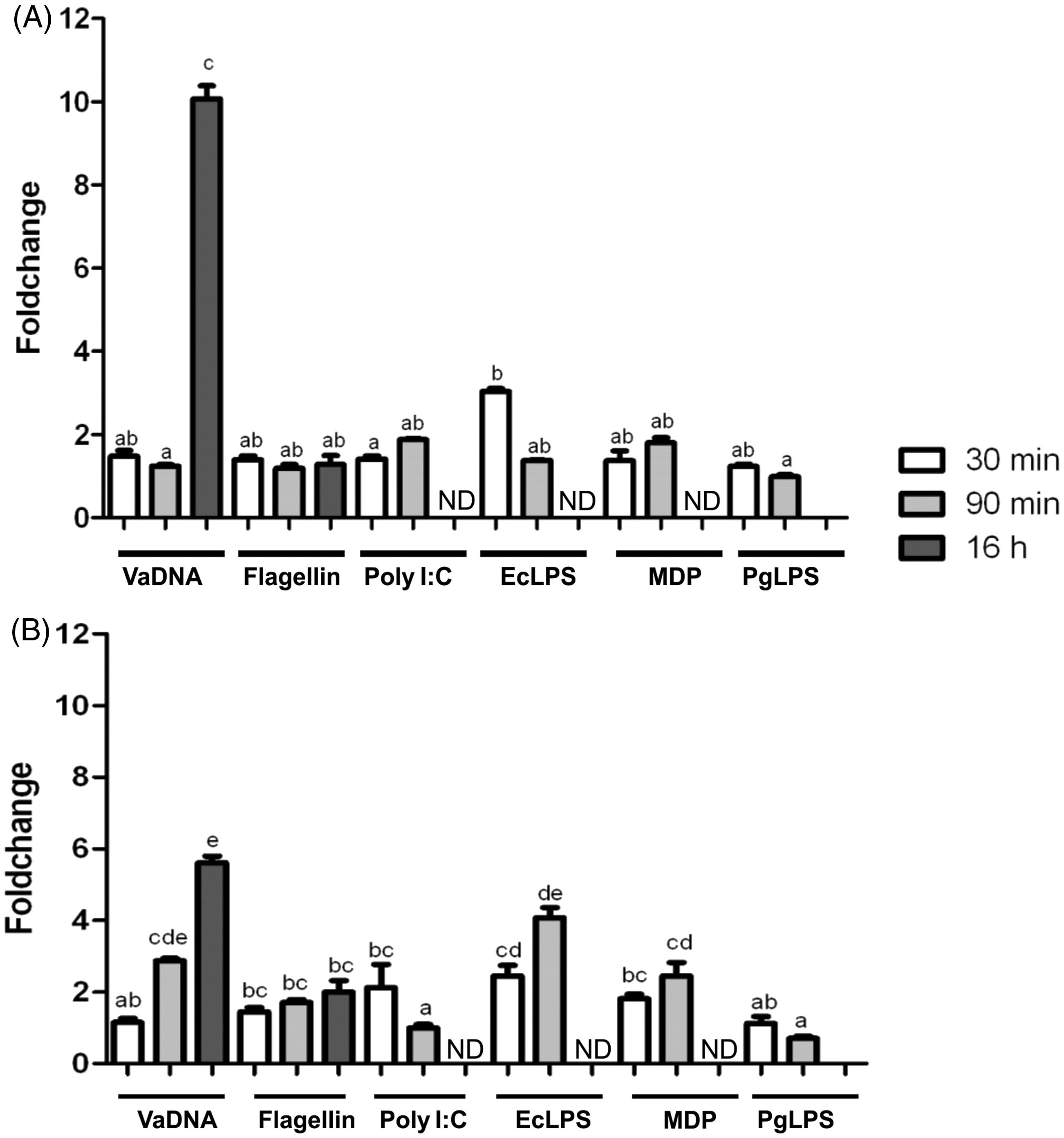
Recombinant S. aurata IL-1Fm2 activates phagocytes
The putative mature seabream IL-1FM2 was produced in E. coli, and four different mAbs were generated (Figure 7A, B). The recombinant IL-1Fm2 was able to prime, in a dose-dependent manner, the respiratory burst of head–kidney leukocytes triggered by PMA (Figure 7C), while it had no effects on cell viability (data not shown). However, no additive or synergistic effects were observed with the combination of IL-1Fm2 and bacterial DNA (Figure 7C). This priming effect was specific, as the purified 6D1E4 mAb neutralized IL-1Fm2 activity, while the mAb 10E2G6, which was raised against zebrafish IL-1β, failed to do so (Figure 7D). In addition, the seabream IL-1Fm2 activity was completely abolished by heat treatment (Figure 7D).
Recombinant gilthead seabream IL-1Fm2 activates the respiratory burst of phagocytes. (A) SDS-PAGE analysis of IL-1Fm2 recombinant protein (rsbIL-1Fm2). Lane 1: BSA (2 µg); lane 2: rsbIL-1Fm2 (2 µg). (B) Western blot analysis of rsbIL-1Fm2 (200 ng/lane) using four different mAbs raised against rsbIL-1Fm2. The monomer and putative dimers are indicated. (C, D) Respiratory burst triggered by PMA of gilthead seabream head–kidney leukocytes primed for 16 h with 50 µg/ml VaDNA and/or 1–10 ng/ml rsbIL-1Fm2 (untreated or preheated at 95℃ for 10 min) in the presence of 100–1000 ng/ml purified 6D1E4 mAb against rsbIL-1FM2 or purified 10E2G6 mAb against zebrafish IL-1β (zfIL-1β). Data are presented as mean ± SEM and are representative of three independent experiments. Different letters denote statistically significant differences among the groups according to a Tukey test. The groups marked with ‘a’ did not show statistically significant differences from control cells.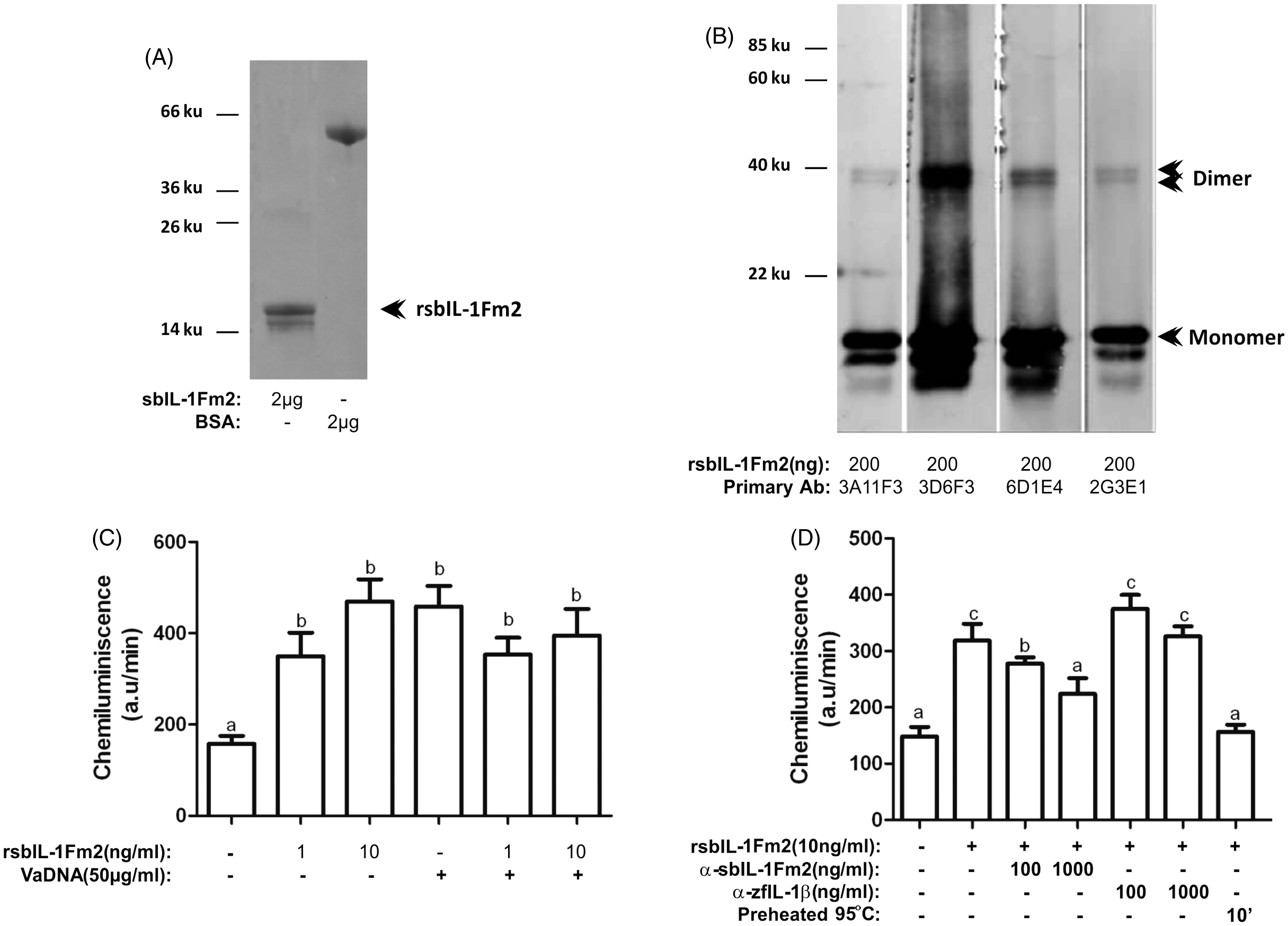
We next examined the effect of recombinant seabream IL-1Fm2 in purified macrophage monolayers (Figure 8A) and sorted acidophilic granulocytes (Figure 8B). The results showed that while recombinant IL-1Fm2 had a very weak effect, if any, on the expression of the gene encoding IL-1β, IL-1Fm2, TNF-α, IL-8 and IL-10 when used alone, it showed a powerful synergistic effect with bacterial DNA by increasing the mRNA levels of IL-1β, IL-1Fm2, IL-8 and IL-10 in macrophages and IL-1β and IL-1Fm2 in acidophilic granulocytes (P < 0.0001 for the interaction according to a two-way ANOVA). Collectively, these results indicate that the crosstalk between TLR and IL-1Fm2 signaling regulates the fish inflammatory response.
Recombinant gilthead seabream IL-1Fm2 regulates the expression of pro- and anti-inflammatory genes in phagocytes. The mRNA levels of IL-1β, IL-1Fm2, TNF-α, IL-8 and IL-10 were determined by RT-qPCR in macrophages (A) and acidophilic granulocytes (B) incubated for 3 (macrophages) or 24 h (acidophilic granulocytes) with 50 µg/ml VaDNA and/or 10 ng/ml recombinant seabream IL-1Fm2 (rsbIL-1Fm2). The results are expressed as the mean ± SEM of triplicate samples. Different letters denote statistically significant differences among the groups according to a Tukey test. The groups marked with ‘a’ did not show statistically significant differences from control cells.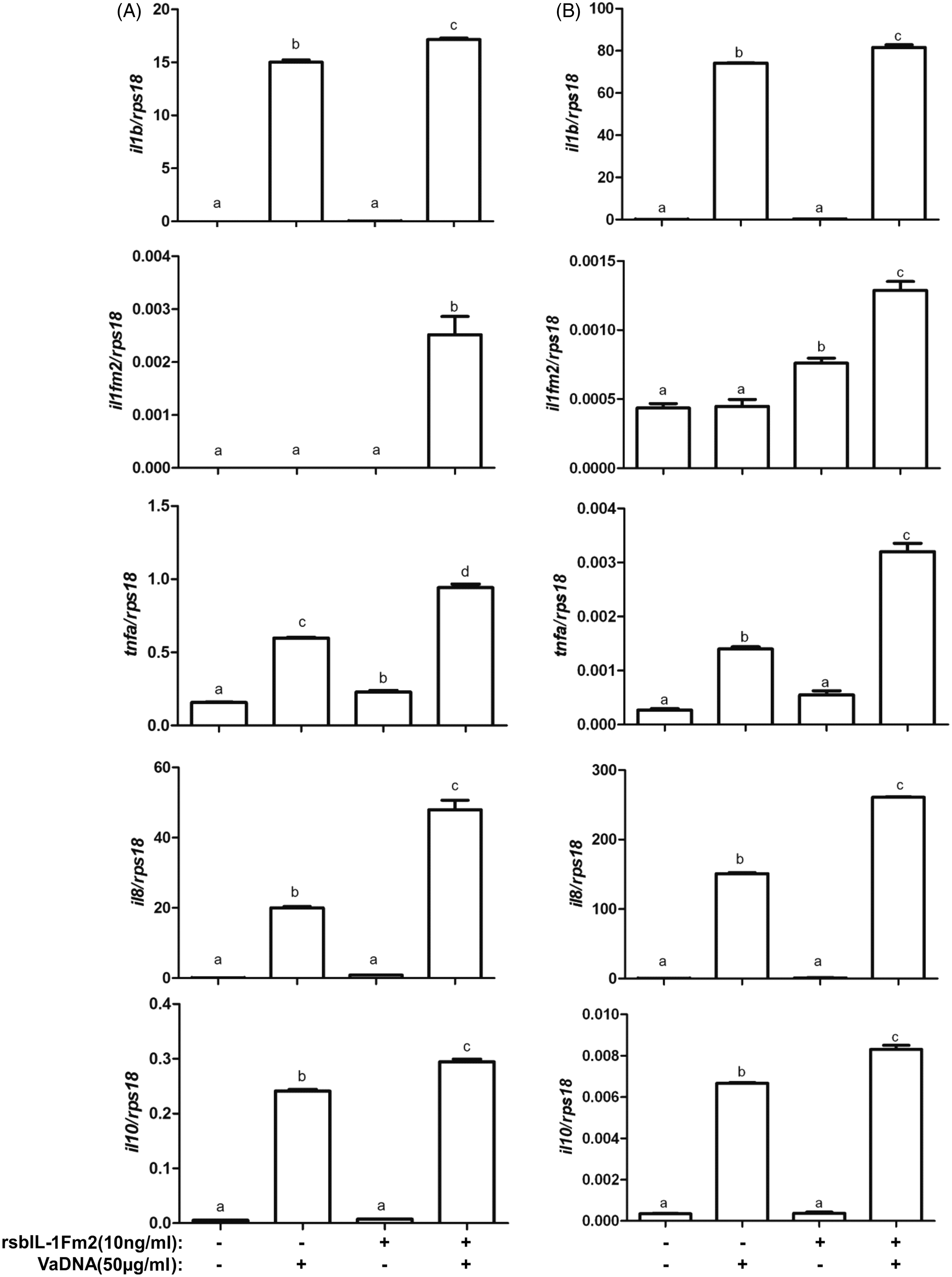
The strong induction of IL-1Fm2 expression by the combination of bacterial DNA and recombinant IL-1Fm2 led us to examine IL-1Fm2 protein levels in cell extracts and supernatants from macrophages co-stimulated with this combination. It was found that bacterial DNA and recombinant IL-1Fm2 strongly increased the intracellular accumulation of IL-1β (Figure 9A), whereas it moderately increased a polypeptide of the expected size for the precursor form of IL-1Fm2 (approximately 30 ku) (Figure 9B), confirming the gene expression studies. Strikingly, the 6D1E4 mAbs also reacted with a polypeptide of about 18 ku present in the supernatants of bacterial DNA/IL-1Fm2-stimulated macrophages (Figure 9B), which might represent the matured form of IL-1Fm2. This polypeptide was synthesized by macrophages, as recombinant IL-Fm2 showed a faster electrophoretic mobility (Figure 9B). Interestingly, the pharmacological inhibition of all caspases, using the pan-caspase inhibitor Q-VD-OPh, or caspase-1, using the specific caspase-1 inhibitor Ac-YVAD-CMK, both failed to inhibit the processing and release of IL-1Fm2 (Figure 9C), as it has already been shown for gilthead seabream IL-1β.
10
Gilthead seabream IL-1Fm2 is processed by a caspase-independent mechanism. Macrophages pre-treated for 1 h with 100 µM of the caspase-1 inhibitor Ac-YVAD-CMK or 50 µM of the pan-caspase inhibitor Q-VD-OPh were stimulated for 16 h with 50 µg/ml VaDNA and 10 ng/ml rsbIL-1Fm2. Cell lysates from unstimulated and VaDNA/rsbIL-1Fm2-activated macrophages (30 µg protein) and concentrated supernatants (corresponding to 100 µg protein from cell extracts) were probed with a monospecific polyclonal Ab to sbIL-1β (A) or the mAb 6D1E4 against rsbIL-1Fm2 (B, C). Forty nanograms of rsbIL-1Fm2 were also probed with the 6D1E4 mAb (B, lower panel, C+). Migration position for the pro-cytokine and mature forms are indicated. The results are representative of two independent experiments.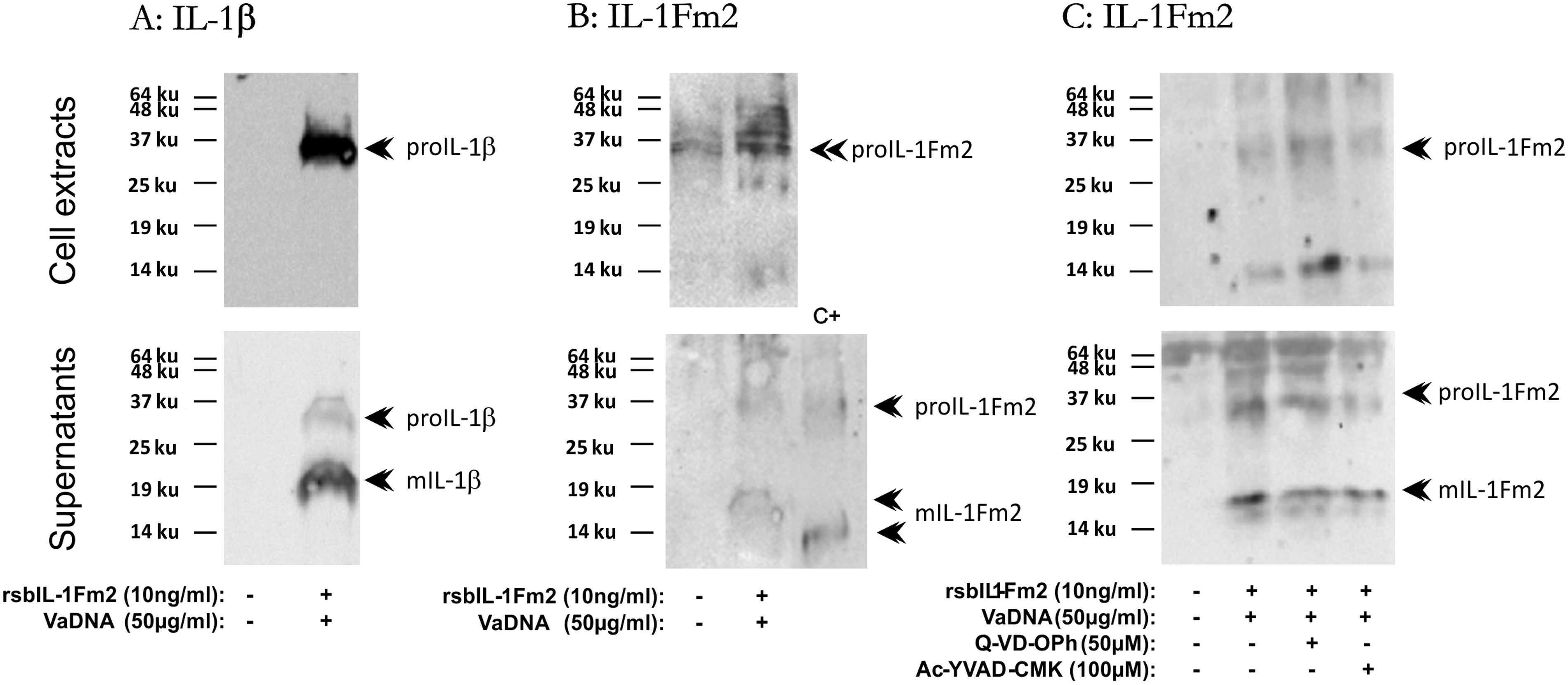
Discussion
In this study, we have identified a new member of the IL-1 family, IL-1Fm2, which is present in most evolutionarily advanced teleost fish, the Series Percomorpha. Surprisingly, after searching their genomes it appears that IL-1Fm2 is absent in pufferfishes, Order Tetraodontiformes. The conservation of synteny of several genes across teleost IL-1β and IL-1Fm2 loci supports the notion that both IL-1 members arose from the FSGD that occurred approximately 350 million years ago. 29 The secondary loss of many genes after this event 29 might explain the absence of IL-1Fm2 in pufferfishes. The present study clarifies the evolution of the IL-1 family in teleost fish, as the recent discovery of the IL-1β3 gene and IL-1β4 pseudogene in salmonids (rainbow trout, Oncorhynchus mykiss, and Atlantic salmon, Salmo salar) led to the proposal of two types of il1b gene in all teleost fish analyzed, including Perciformes (tilapia) and Beloniformes (medaka), 30 both belonging to the Percomorpha. The new IL-1 member identified in tilapia and medaka were named IL-1β2 and were classified as type I teleost IL-1β. 30 However, the phylogenetic analysis of these two new members together with the ones identified in gilthead seabream and European sea bass (this study) and Japanese flounder 28 demonstrates that all of them form a separated clade from type I teleost IL-1β and constitute a subclade together with the nIL-1Fm, which functions as an IL-1 antagonist in rainbow trout. 31 Therefore, we propose the name il1fm2 for the genes identified in the present study. Furthermore, our phylogenetic analysis also shows that the two new IL-1 members described in the Japanese flounder as IL-1β-like1 and IL-1β-like2 28 are, indeed, the IL-1Fm2 and nIL-1Fm orthologs, respectively, of this species.
The weak inducibility of gilthead seabream (this study) and Japanese flounder 28 il1fm2 genes in infected animals and in leukocytes stimulated in vitro also supports that IL-1Fm2 is a new member of the IL-1 family rather than an IL-1β paralogue belonging to the type I teleost IL-1β. In fact, although rainbow trout IL-1β3 gene (type I) was induced at lower levels than IL-1β2, and to some extent IL-1β1 (both type II), upon macrophage stimulation with phytohemagglutinin A (PHA), 30 all type I il1b genes reported so far, including zebrafish32–34 and common carp (Cyprinus carpio) 35 (Cypriniformes), catfish (Ictalurus punctatus, Siluriformes) 36 and conger eel (Conger myriaster, Anguilliformes), 37 are heavily induced in infected animals and/or stimulated leukocytes.
In mammals, IL-1β needs to be processed to exert biological activity by caspase-1, which is activated in a cytosolic molecular platform called the inflammasome.6,7 However, all non-mammalian il1b genes reported so far lack a conserved caspase-1 recognition site. 5 Although gilthead seabream IL-1Fm2 does not show a conserved caspase-1 processing site, gilthead seabream macrophages processed IL-1Fm2 before being released. However, both pan-caspase and caspase-1-specific inhibitors fail to inhibit its processing; despite this, they are able to inhibit apoptosis in fish38–40 and the activity of recombinant gilthead seabream caspase-1, 24 respectively. Similarly, the processing and release of gilthead seabream IL-1β is caspase-1-independent. 10 In sharp contrast, it has been found that zebrafish inflammatory caspase A and caspase B 8 and European sea bass caspase-1 9 can process IL-1β in these species. Nevertheless, classical NLRP3 and NLRC4 inflammasome activators also fail to trigger the processing and secretion of IL-1β in the gilthead seabream.10,12 Therefore, the mechanism of processing and release of fish IL-1 family members deserves further investigation.
Although information on the biological activity of bony fish cytokines is scant, IL-1β was one the best cytokines characterized in these animals. In rainbow trout, recombinant IL-1β and derived peptides were found to promote leukocyte recruitment in vitro 41 and induce potent inflammatory response when injected i.p., which results in enhanced resistance to a experimental bacterial 42 and viral 43 infections. Similarly, recombinant IL-1β from the phylogenetically distant European sea bass also shows pro-inflammatory effects and induced the proliferation of thymocytes. 44 Although il1fm2 genes seem to be phylogenetically close to nil1fm genes, which at least in rainbow trout acts as an IL-1 antagonist, recombinant gilthead seabream IL-1Fm2 seems to be pro-inflammatory, as it is able to activate the respiratory burst of phagocytes and to induce the expression of several genes encoding pro-inflammatory mediators, such as IL-1β, TNF-α and IL-8. Strikingly, IL-1Fm2 and PAMPs synergistically induce its own expression and that of il10 gene, suggesting a key role for this cytokine in the resolution of inflammation and the polarization of the fish adaptive immune responses.
In conclusion, we have identified a new IL-1 family member that is present in most phylogenetically advanced teleost fish and which plays an important role in the regulation of the immune responses of this group of animals. The identification of IL-1Fm2 sheds light on the evolution of the IL-1 family in vertebrates and points to the complexity of this cytokine family.
Footnotes
Funding
This work was supported by the Spanish Ministry of Economy and Competence (grants BIO2008-01379, BIO2011-23400 and CSD2007-00002 to VM, Juan de la Cierva contract to JM and fellowship to DA, all co-funded with Fondos Europeos de Desarrollo Regional/European Regional Development Funds) and the Fundación Séneca-Murcia (grant numbers 04538/GERM/06 to VM).
Acknowledgements
We thank I. Fuentes and P. Martínez for excellent technical assistance, and Professors A.E. Toranzo and J.L. Barja for the R82 strain of V. anguillarum.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
