Abstract
Alterations in innate immunity that predispose to chronic obstructive pulmonary disease (COPD) exacerbations are poorly understood. We examined innate immunity gene expression in peripheral blood polymorphonuclear leukocytes (PMN) and monocytes stimulated by Haemophilus influenzae and Streptococcus pneumoniae. Thirty COPD patients (15 rapid and 15 non-rapid lung function decliners) and 15 smokers without COPD were studied. Protein expression of IL-8, IL-6, TNF-α and IFN-γ (especially monocytes) increased with bacterial challenge. In monocytes stimulated with S. pneumoniae, TNF-α protein expression was higher in COPD (non-rapid decliners) than in smokers. In co-cultures of monocytes and PMN, mRNA expression of TGF-β1 and MYD88 was up-regulated, and CD14, TLR2 and IFN-γ down-regulated with H. influenzae challenge. TNF-α mRNA expression was increased with H. influenzae challenge in COPD. Cytokine responses were similar between rapid and non-rapid decliners. TNF-α expression was up-regulated in non-rapid decliners in response to H. influenzae (monocytes) and S. pneumoniae (co-culture of monocytes and PMN). Exposure to bacterial pathogens causes characteristic innate immune responses in peripheral blood monocytes and PMN in COPD. Bacterial exposure significantly alters the expression of TNF-α in COPD patients, although not consistently. There did not appear to be major differences in innate immune responses between rapid and non-rapid decliners.
Introduction
Acute exacerbations are common and potentially serious events in patients with chronic obstructive pulmonary disease (COPD).1,2 Acute exacerbations of COPD (AECOPD) increase health care costs associated with COPD management. 3 Frequent exacerbations are associated with progression of COPD4,5 and predict risk of future exacerbations. 1 Understanding the pathogenesis of AECOPD is vital in order to develop more effective preventive and therapeutic strategies.
Bacteria and viruses are the main causes of AECOPD,6,7 and airway inflammation correlates directly with bacterial load and pathogenicity. 8 Detailed profiling of COPD biospecimens (bronchial lavage, sputum and tissues) identifies a bacterial microbiome consisting mainly of Haemophilus influenzae, Streptococcus pneumoniae, Pseudomonas aeruginosa, Moraxella catarrhalis and Staphylococcus aureus. 6 Lung colonisation with H. influenzae is associated with increased frequency of AECOPD and decreased forced expiratory volume in one second (FEV1). 9 Pulmonary and systemic inflammatory markers are increased during AECOPD, including C-reactive protein (CRP), serum amyloid A, IL-8, IL-1β, IL-10 and NF-κB. Upon recovery, lung function may return to baseline, but excessive inflammation and oxidative stress often persist, predisposing to future exacerbations. 10
Many immune cells, including monocytes, macrophages, polymorphonuclear leukocytes (PMN), lymphocytes, NK cells, dendritic cells and eosinophils, as well as airway epithelial cells, are involved in the detection and removal of microbial pathogens. 11 Monocytes/macrophages and PMN promote bacterial clearance, but also contribute to tissue destruction through the release of free radicals and proteases that cause parenchymal destruction. 12 Innate immunity is the first line of host defence against pathogens. Smoking impairs the immune response in COPD patients, thereby reducing bacterial clearance and predisposing patients to recurrent infections and AECOPD.13,14 Dysregulated innate immunity may, therefore, predispose to risk of exacerbations and excessive inflammation, and needs to be better understood in patients with COPD.
We hypothesised that COPD patients have an altered innate immune response to bacteria in peripheral blood immune cells, which may then predispose them to frequent exacerbations and, consequently, accelerated lung function decline. To understand this aspect of COPD pathogenesis better, we aimed to characterise the molecular response of immune cells (PMN and monocytes) to bacteria in patients with COPD. We used quantitative real-time PCR (qRT-PCR) and ELISA to examine expression of selected innate immune genes in peripheral blood PMN and monocytes after in vitro stimulation with H. influenzae and S. pneumoniae. To characterise the function of the innate immune genes in driving COPD severity we compared COPD patients with rapid and non-rapid trajectories of lung function decline against healthy smoker controls.
Materials and methods
Participants
Patients with COPD were identified from lung function data recorded in hospital medical records and recruited to donate blood for this study. COPD was defined according to the GOLD guidelines. 15 Spirometry had been measured according to standard guidelines, 16 as described previously, 17 at routine clinic visits to The Prince Charles Hospital. To calculate the rate of decline of FEV1 as a percent of the predicted value based on age and height, we analysed at least three FEV1 measurements obtained over at least a three-year period using multiple linear regression analysis (SPSS Version 20; SPSS, Chicago, IL, USA). ‘Rapid’ decline was defined arbitrarily as decline of greater than 1.5% of the predicted value per year, whereas ‘non-rapid’ was decline at a rate of <0.16% of the predicted value per year). Fifteen cases of COPD with rapid rates of decline were compared with 15 cases of COPD with non-rapid decline and 15 cases of smokers without airflow obstruction (non-COPD controls). All participants were former smokers and stable with no infective exacerbations or respiratory infections in the 6 wk preceding blood donation. This study was approved by the institutional ethics committees of The Prince Charles Hospital and The University of Queensland. Written informed consent was obtained from all participants.
Inflammatory markers
Full blood count, including total white cell count (WCC) and differential cell count was measured at recruitment.
Isolation and culture of monocytes and PMN
Forty ml of peripheral blood collected in lithium heparin tubes was diluted 1:1 with Hanks balanced salt solution. Twenty ml of this mixture was overlayed on 10 ml Ficoll-Hypaque density gradient reagent and centrifuged. White blood cells were isolated from the interface and PMN from the red blood cell (RBC)/Ficoll mixture.
To isolate monocytes, 1 × 107 cells from the WBC pool containing lymphocytes and monocytes were labelled with 20 µl of CD14 magnetic microbeads (Miltenyi Biotech, Bergisch Gladbach, Germany) and incubated for 15 min at 4℃. The cell/bead mixture was then passed through a column attached to a magnet to capture monocytes. After a series of PBS washes to remove CD14 negative cells, the column was removed from the magnet and flushed with RPMI cell culture medium to elute monocytes into a fresh collection tube. Monocytes (0.25 × 106 cells) were then distributed into each well of a 24-well plate and incubated overnight at 37℃ before challenge.
To isolate PMN, the RBC/Ficoll mixture was washed three times with red cell lysis buffer to remove RBCs. 18 The remaining PMN were resuspended in RPMI, counted and diluted to the desired concentration. Experiments on three control subjects were performed to assess monocyte and neutrophil morphology by cytospin and differential staining (DiffQuik) independently. A cytologist scored these slides and confirmed that more than 99% and 95% of the cell population were monocytes and PMN respectively.
To test the interaction of monocytes and PMN in response to bacterial challenge (modelling the situation in COPD airways), PMN were co-cultured with monocytes at a ratio of 5:1 respectively (1.25 × 106 PMN to 0.25 × 106 monocytes per well) for 24 h. This ratio was based on the observed ratio of PMN to macrophages in COPD airways19,20 and previous model systems in other diseases. 21
Bacterial culture
Gram-positive bacteria S. pneumoniae (ATCC strain 49619) and Gram-negative bacteria H. influenzae (ATCC strain 49247) were used for live, in vitro bacterial challenges because of their relevance as pathogenic bacteria in COPD infective exacerbations. Broth culture of S. pneumoniae was grown for 18 h in brain heart infusion broth supplemented with 1% fetal bovine serum. Next, S. pneumoniae was diluted 1:10 in broth culture and incubated for a further 2 h to achieve log phase bacterial growth. Haemophilus influenzae were similarly prepared, except in Mueller–Hinton broth supplemented with 10 mg of β-NAD and hemin.
Bacteria were then pelleted by centrifugation and resuspended in RPMI without antibiotics. The concentration of bacteria was measured using a spectrophotometer set at absorbance wavelength of 600 nm. The optimal cell (monocytes/PMN) to bacteria ratio was determined to be 1:25 by experiments measuring TNF-α response in the supernatant (ELISA; R&D Systems, Minneapolis, USA) (data not shown).
Bacterial challenges to cells
For live bacterial challenges, published methods were used.22,23 Briefly, RPMI containing H. influenzae or S. pneumoniae (6.25 × 106 bacteria/ml) was added to wells containing (i) monocytes only, (ii) PMN only, or (iii) monocytes and PMN in co-culture, and incubated for 4 h at 37℃. Supernatant was harvested and stored at −80℃ for cytokine analysis. To avoid monocyte or neutrophil cell death from prolonged bacterial infection, fresh RPMI containing antibiotics (penicillin and streptomycin) was then added to the wells, and incubation was continued for a further 20 h to a total of 24 h. The supernatant at 24 h was harvested and stored at −80℃ for cytokine analysis. Monocytes and PMN were then harvested in TRIzol (Invitrogen, Carlsbad, CA, USA) for RNA extraction. A non-bacterial negative control containing medium only (without bacteria) was cultured for 24 h. All experiments were performed in triplicate.
Protein expression of cells challenged with bacteria
The expression of four inflammatory cytokines (TNF-α, IFN-γ, IL-8, IL-6) was measured using Bioplex beads (Biorad Laboratories, Hercules, CA, USA), following the manufacturer’s instructions. Briefly, 50 μl of undiluted sample was sandwiched between 50 μl of premade multiplex bead working solution and 25 μl of detection Ab. The labelled Abs were conjugated with streptavidin-phycoerythrin and detected using Luminex-100 system (Luminex Corporation, Austin, TX, USA). A broad range standard curve prepared as a fourfold dilution from the stock (32,000 pg/ml) was included. All samples were performed in duplicate. Coefficient of variation (CV) was calculated for all samples. Those with CV <20% were included, and the other samples were repeated or excluded.
mRNA expression of cells challenged with bacteria
Total RNA was extracted from cells using TRIzol. Contaminating DNA was removed using a DNA-free™ DNAse treatment kit (Ambion, Austin, TX, USA). Whole genome mRNA amplification using SuperScript™ RNA Amplification System (Invitrogen) was performed following the manufacturer’s instructions. mRNA was reverse transcribed by its first strand to cDNA followed by second strand synthesis and amplification. The amplified DNA was then transcribed back to RNA and purified. The amplified RNA underwent a second round of first strand cDNA synthesis to prepare for qRT-PCR using TaqMan® gene expression assays (Applied Biosystems, Foster City, CA, USA) for 13 innate immunity genes. The expression of target genes was normalised to the geometric mean of two housekeepers: β-2 microglobulin (B2M) and hypoxanthine phosphoribosyltransferase 1 (HPRT1). Relative expression of the genes was calculated using Pfaffl’s method, 24 and the ratio of expression in unstimulated to stimulated cells was used for class comparison and fold-change analysis.
Statistical analysis
To identify significant differences in WCC, gene expression and protein expression the following comparisons involving the three participant groups were made: (i) rapid decliners versus non-rapid decliners, (ii) rapid decliners versus non-COPD controls, (iii) non-rapid decliners versus non-COPD and (iv) COPD (rapid and non-rapid decliners) versus non-COPD controls. The differences between groups for WCC and erythrocyte sedimentation rate were calculated using t-tests. For protein expression, the difference between stimulated and unstimulated levels measured 4 h and 24 h after bacterial exposure in macrophage, neutrophil and macrophage/neutrophil co-cultures were calculated. A P-value of <0.05 (two-tailed) was considered statistically significant in all analyses. Unsupervised hierarchical clustering was performed in BRB Array Tools V4.2.1 (developed by Dr Richard Simon and Amy Peng Lam, freely accessible at http://linus.nci.nih.gov/BRB-ArrayTools.html) to determine whether inflammatory cytokine protein expression classified H. influenzae and S. pneumoniae exposure or the immune cell types. Finally, the ratios of unstimulated to stimulated gene expression were calculated for each of the 13 innate immunity genes in macrophage/neutrophil co-cultures exposed to bacteria for 24 h. Class comparison analysis of log fold-change using t-test was performed to identify genes significantly associated with each participant group. Unsupervised hierarchical clustering was also performed in BRB Array Tools V4.2.1 to determine whether gene expression classified H. influenzae and S. pneumoniae exposure.
Results
Demographics
Patient demographics of innate immunity study.
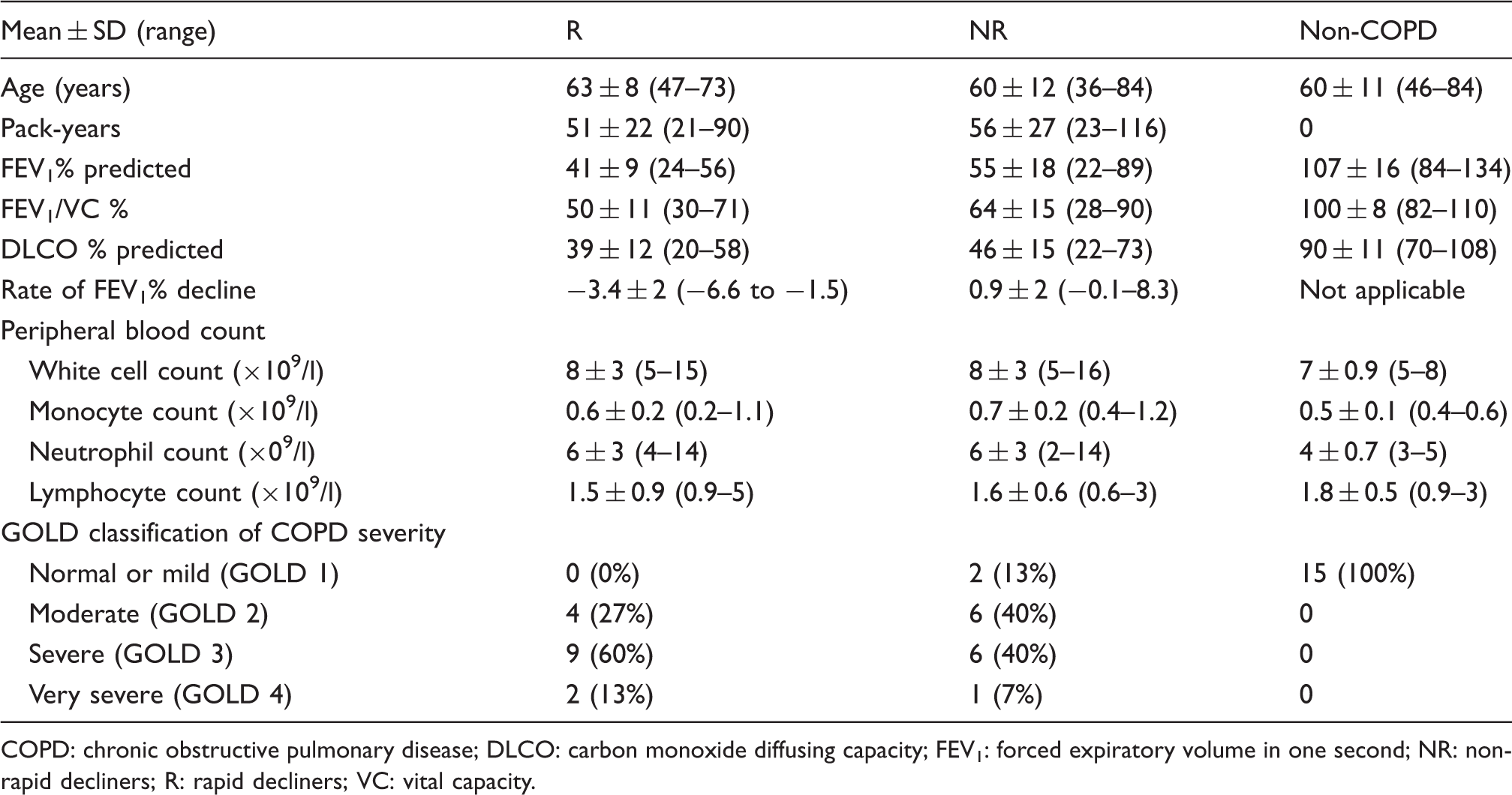
COPD: chronic obstructive pulmonary disease; DLCO: carbon monoxide diffusing capacity; FEV1: forced expiratory volume in one second; NR: non-rapid decliners; R: rapid decliners; VC: vital capacity.
WCC
Mean peripheral blood total white blood cell count and neutrophil counts were significantly higher in COPD patients than in non-COPD controls (P < 0.01) (Supplementary Table 1), whereas lymphocyte and monocyte counts were not statistically different between these groups. COPD rapid decliners had significantly lower mean monocyte counts than non-rapid decliners.
Protein expression of cytokines from immune cells challenged with bacteria
Response to bacteria
The protein expression of four cytokines (IL-8, IL-6, TNF-α and IFN-γ) from the immune cells (monocytes, PMN and monocyte/neutrophil co-culture) was measured at 4 h and 24 h following exposure to bacteria and compared with control conditions that had no bacterial exposure (Supplementary Table 2). At 24 h after H. influenzae challenge, levels of IFN-γ and TNF-α protein in monocyte, neutrophil and monocyte/neutrophil cultures were significantly increased, as was IL-6 in monocytes and co-cultures, but not PMN. IL-8 was increased 24 h after commencement of exposure, but 4 h levels were not higher than baseline. The pattern of results was similar for S. pneumoniae challenges, although the magnitude of the increases in IL-6, TNF-α and IFN-γ were generally not as high as with H. influenzae.
Disease groups and response to bacteria
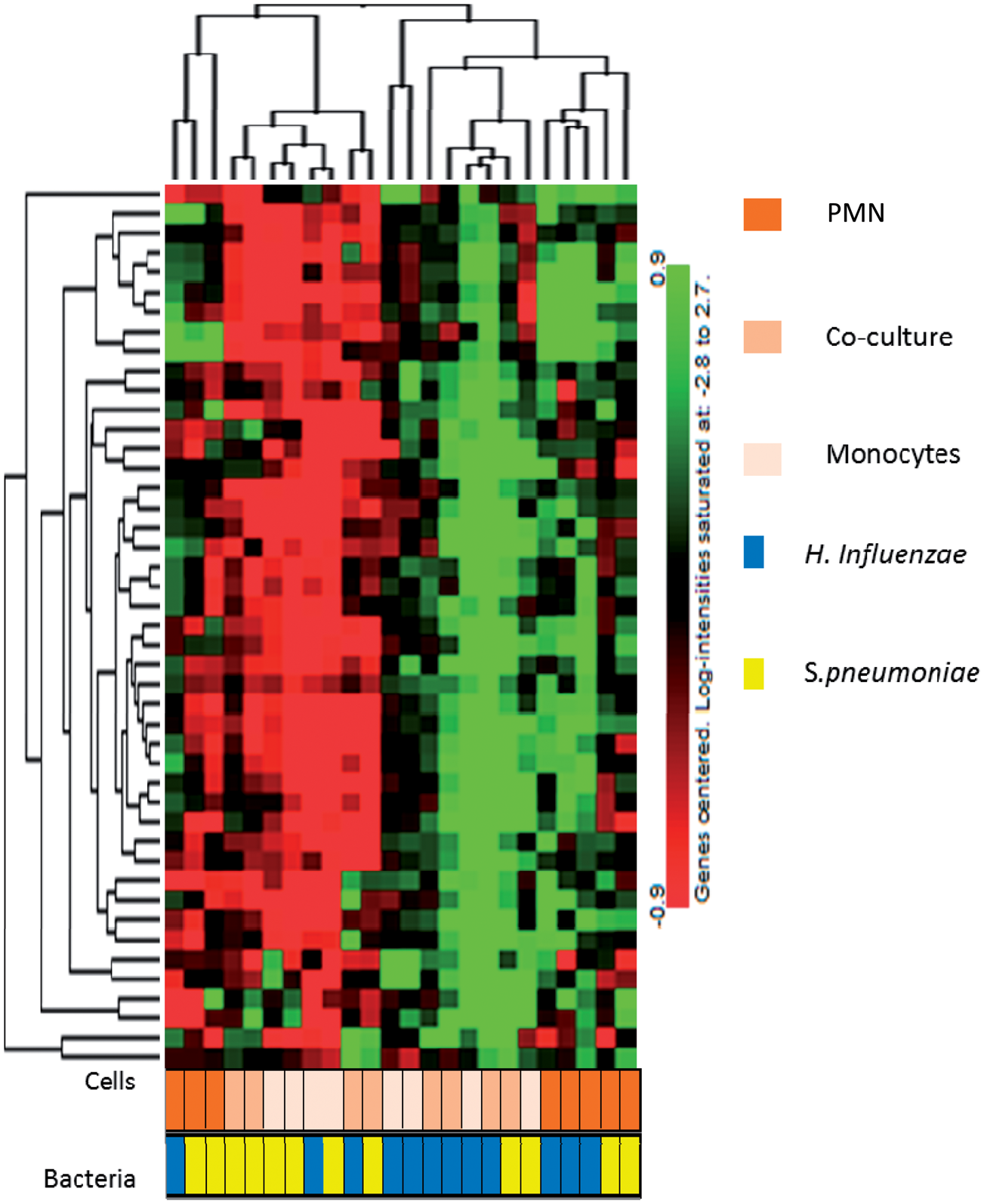
There were no differences between COPD and non-COPD controls in cytokine levels 24 h after in vitro challenge with H. influenzae or S. pneumoniae (Supplementary Tables 3 and 4), except for higher TNF-α in monocyte cultures stimulated with S. pneumoniae in the COPD group with non-rapid decline compared with the non-COPD group (P = 0.03) (Figure 1 and Supplementary Table 4). There were no statistically significant differences in cytokine levels between COPD patients with rapid decline and non-rapid decline in lung function. Unsupervised clustering analysis showed a separate pattern of expression of these four cytokines in PMN from that of either monocytes or co-cultures (Figure 2).
TNF-α protein expression in monocytes cells exposed to S. pneumoniae for 24 h. A significant difference was observed in TNF-α cytokine expression in monocytes from COPD and non-COPD patients after S. pneumoniae exposure. *P < 0.05. Unsupervised clustering analysis based on protein expression of cytokines in monocytes, PMN and co-culture. In the dendrogram red and green spots indicates genes that are over-and under-expressed respectively. Protein expression of samples exposed to H. influenzae samples and S. pneumoniae are represented as blue and yellow bars respectively. The cell types are represented as gradients of orange.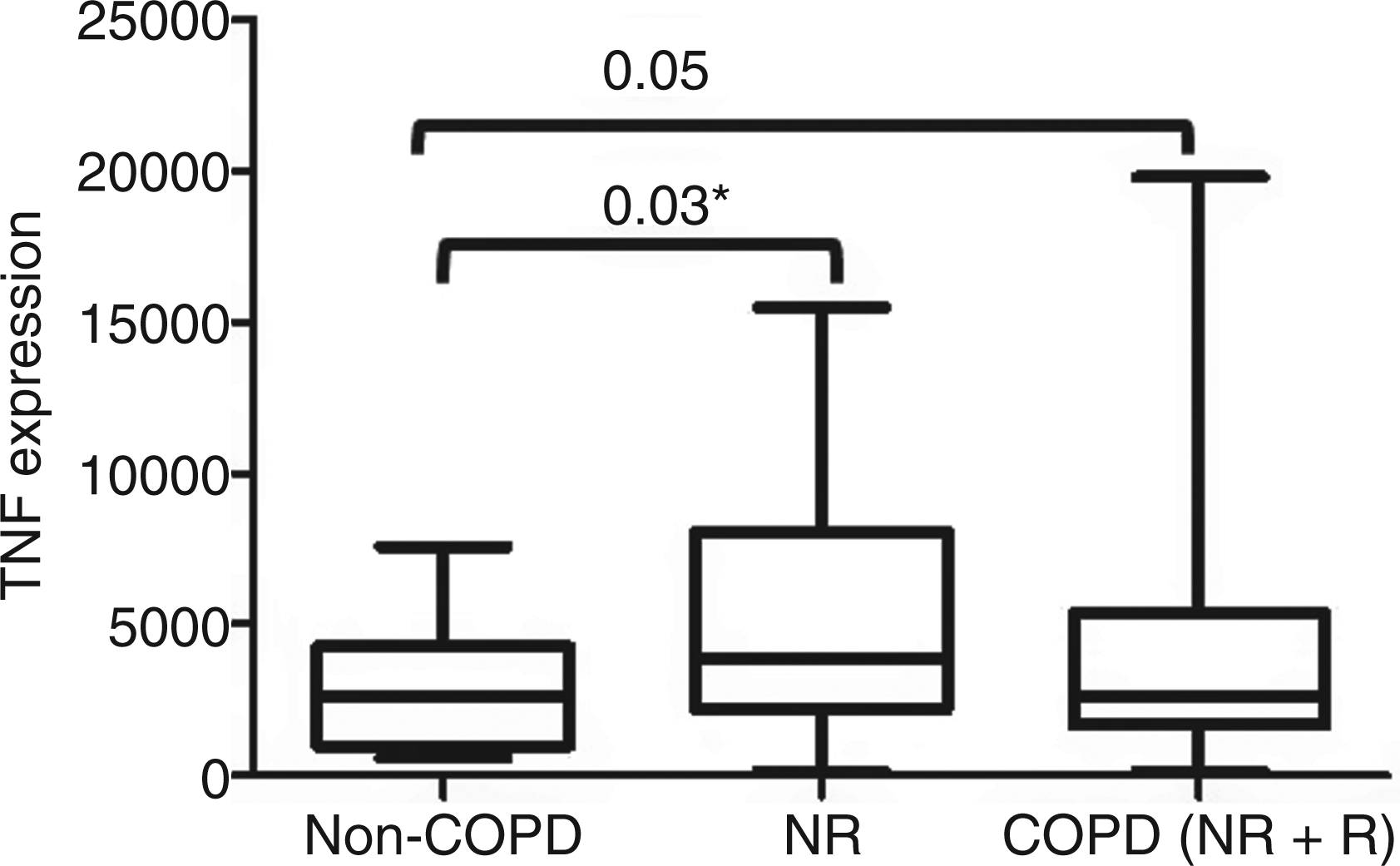
mRNA expression of cytokines from immune cells challenged with bacteria
Response to bacteria
The expression of 13 innate immunity pathway genes (Supplementary Table 5) was measured by qRT-PCR on mRNA extracted from monocyte/neutrophil co-cultures 24 h after an initial 4 h exposure to live bacteria (Supplementary Table 6). TLR5 expression did not pass quality control filtering in BRB Array Tools and was excluded from analysis. When controlled for housekeeper gene expression there was significantly higher TGF-β1 and MYD88 expression, and lower CD14, TLR2 and IFN-γ expression following H. influenzae challenge than following S. pneumoniae challenge (P < 0.01).
Disease groups and response to bacteria
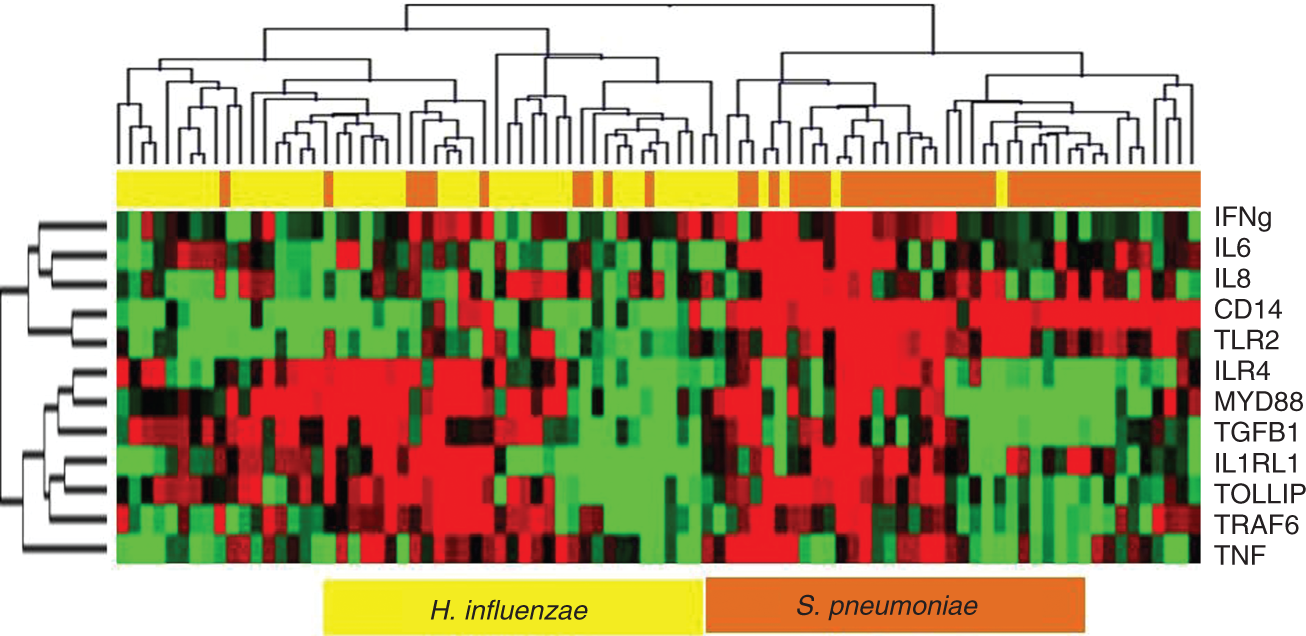
In COPD patients, only TNF-α mRNA expression demonstrated statistically higher up-regulation with H. influenzae challenge than non-COPD controls (Figure 3 and Supplementary Table 7). Although some genes showed more than twofold increase or decrease in expression in monocyte/neutrophil co-cultures challenged with S. pneumoniae, these did not reach statistical significance. Unsupervised clustering of gene expression separated the samples exposed to H. influenzae from those exposed to S. pneumoniae (Figure 4), and the expression of innate immunity genes was capable of classifying H. influenzae and S. pneumoniae infected cells with more than 90% accuracy.
TNF-α mRNA expression in monocyte/PMN co-culture cells exposed to H. influenzae for 24 h. TNF-α expression was significantly altered in monocyte/PMN co-culture cells from non-rapid decliners and COPD patients (vs non-COPD) 24 h after H. influenzae exposure. Unsupervised clustering of all samples exposed based on mRNA gene expression in co-culture only. In the dendrogram red and green spots indicate genes that are over- and under-expressed respectively. mRNA expression of samples exposed to H. influenzae and S. pneumoniae are represented as yellow and orange bars at the top.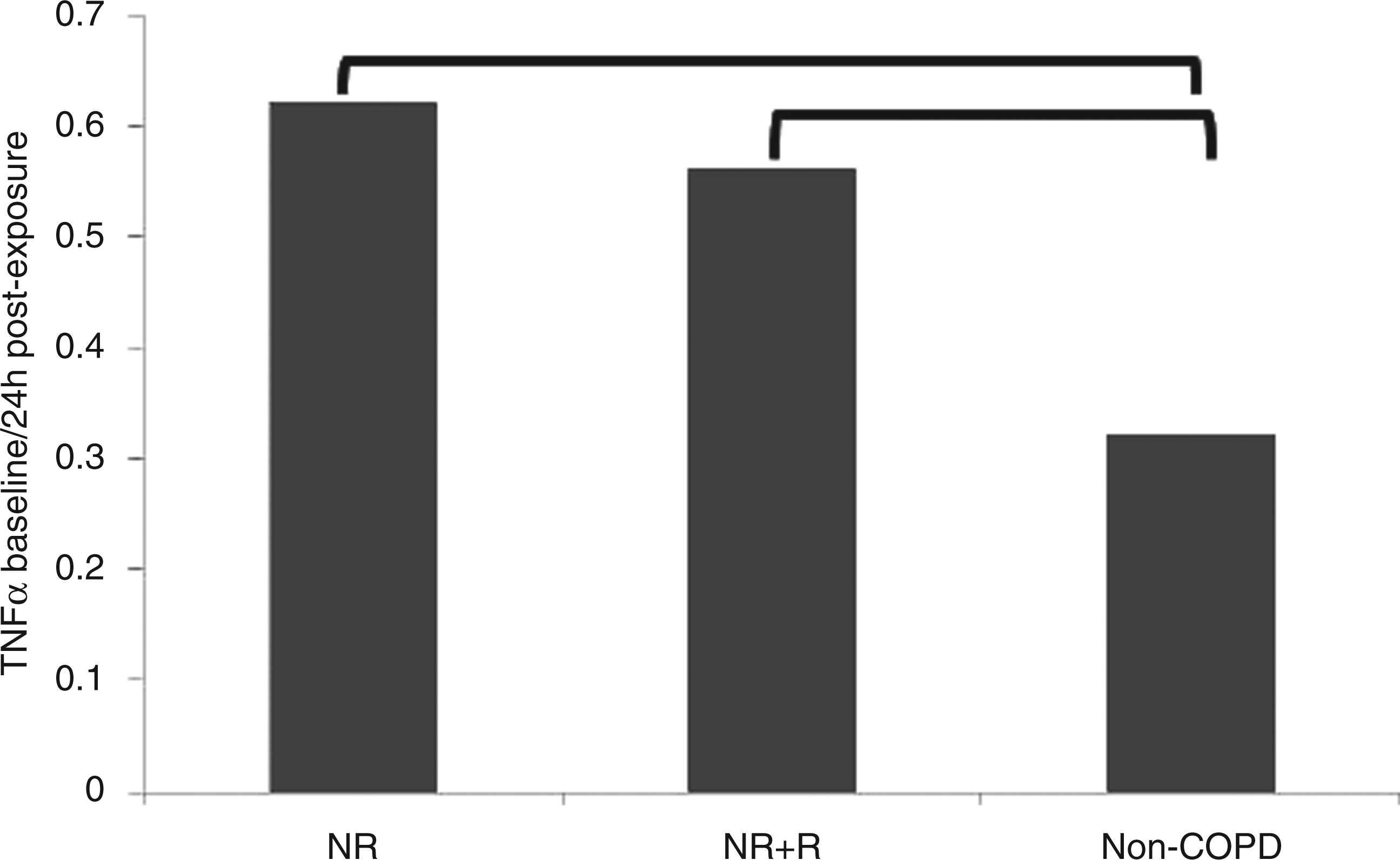
Discussion
Innate immunity plays an important role in host defence against infection in COPD. 12 Furthermore, exacerbations of COPD are a risk factor for accelerated decline in lung function. 5 Here we endeavoured to link these concepts by identifying an altered innate immune response in COPD patients with rapidly declining lung function.
In this study of the in vitro response of COPD patients’ peripheral compartment cells to bacterial challenge, we found evidence that different pathogens elicit different innate immune responses and that the various different cell types represented in the peripheral blood compartment mount different cytokine and innate immunity gene responses to bacterial challenges. However, we found no major differences in innate immune responses to in vitro bacterial challenge between COPD patients with and without rapidly declining lung function. Although this work does not exclude that a difference in innate immune response to bacterial pathogens might contribute to rapid decline in lung function in COPD patients with frequent acute exacerbations, it provides no significant additional support for this theory. Other possible drivers still need to be considered in the pathogenesis of rapidly declining lung function in COPD, such as inflammation, oxidative stress markers, phagocytosis 25 and genetic variation.
This study identifies innate gene expression patterns capable of classifying samples exposed to H. influenzae and to S. pneumoniae. This was reported previously in respect of challenge by killed pathogens.26,27 Heat-killed bacteria trigger different host responses to live bacteria 28 and are phagocytosed slower than live bacteria. 29 Therefore, we used live bacterial cultures to test a biologically realistic model of innate immune response against infection. Whereas previous whole genome profiling of PBMCs exposed to Bordetella and Staphylococcus spp. have been reported, 28 in this study we focused on bacterial strains of direct relevance to COPD. We found a novel profile, based on the expression of 13 genes, which distinguished H. influenzae from S. pneumoniae challenge. Future whole genome profiling studies will, no doubt, provide more detailed insight into host mechanisms in response to bacterial infection in COPD patients. Identification of the host response pattern specific to different bacterial species could be useful in early diagnosis and treatment of exacerbations.
Both protein and mRNA expression of the pro-inflammatory cytokine TNF-α was significantly increased in non-rapid decliners than in non-COPD controls exposed to H. influenzae (monocytes) and S. pneumoniae (co-culture) respectively. Although the TNF-α responses were initiated by different cell types in response to different bacteria, the trend we noticed is consistent with previous work showing that levels of TNF-α expression increase in the sputum of COPD patients during exacerbations.30–32 In fact, TNF-α antagonists are currently being tested for clinical use in COPD and asthma patients. 33 The fact that TNF-α expression in response to pathogens was not significantly different between rapid and non-rapid decliners lends no support to the concept of TNF mediated disease progression.
mRNA expression of innate immunity mediators and cytokines, measured from co-cultured cells exposed to bacteria, showed statistically significant up-regulation of TGF-β1 and MYD88 following H. influenzae challenge (compared with the S. pneumoniae challenge), and down-regulation of CD14, TLR2 and IFN-γ. Few other studies have measured gene expression of pattern recognition receptors (PRRs) in response to bacteria in COPD patients. However, it was shown that COPD patients, either when stable or in an acute exacerbation, had higher TLR2 expression in peripheral blood monocytes, than healthy smokers or controls. 34 Also, farmers chronically exposed to respirable organic material in the workplace showed reduced monocytic TLR2 response to LPS challenge. 35 Another study of immune responses in COPD patients found increased IL-8 and TLR2 gene expression in sputum, and increased IL-8, TLR2 and TLR4 expression from peripheral blood PMN challenged with LPS. 36 Our finding that S. pneumoniae stimulated TLR2 in co-cultured cells is consistent with the known function of TLR2 as a PRR for Gram-positive bacteria.
There have been few studies relating rate of decline of lung function in COPD to innate immunity. Although Donaldson et al. found that higher levels of sputum IL-6 were associated with more rapid decline in FEV1, 37 we did not detect major differences in peripheral compartment cytokine levels or innate immunity gene expression, including IL-6 in rapid decliners compared with non-rapid decliners. The disparity could be a result of the different compartments tested (sputum vs peripheral blood).
Other studies have shown enhanced cytokine responses to bacteria or bacterial antigens in COPD patients. LPS challenge stimulated release of IL-6 from peripheral blood monocytes in COPD patients, 38 and (although highly variable) plasma levels of IL-6 were found elevated in COPD patients. 39 LPS stimulated release of TNF-α by monocytes was higher in COPD patients with weight loss compared with those without. 40 H. influenzae and Moraxella spp. induced higher levels of cytokines (IL-23, IL-12p70, IL-10) from blood monocyte-derived dendritic cells than other commensal airway bacteria, 41 indicating an augmented response to bacteria commonly found in COPD patients.
Strengths
To our knowledge, this is the first study to profile the innate immune response of peripheral blood monocytes and PMN in COPD patients with and without rapid lung function decline. This COPD phenotype of accelerated decline in FEV1 was selected for study because of its importance as a marker of progression of COPD, as used in recent large epidemiological and clinical studies.5,42,43 We recruited polarised groups of patients (rapid versus non-rapid lung function decline) to maximise the power of detecting differences in innate immunity responses between these disease progression phenotypes of COPD. The decline in lung function in the COPD patients was determined for at least three years or longer. Current smokers were excluded in order to remove the confounding effect of current smoking on gene expression and immune responses.
Peripheral blood monocytes and PMN were studied, as these are biologically relevant and clinically accessible cells for demonstrating the innate immune response against bacteria.12,44 Monocytes, as peripheral immune system cells, are precursors to lung macrophages, and peripheral blood PMN traffic to the lungs in the normal course of the host response to bacteria.
The bacteria studied—H. influenzae and S. pneumoniae—are two of the most common organisms causing an acute infective exacerbation of COPD 15 and highly relevant to characterising immune responses in COPD. The dose for bacterial challenge was chosen based on the IL-8 and TNF-α response of the cells. We used commercially available ATCC strains in order to ensure standardised challenges, with virulence that would be expected to be similar to clinically isolated strains. The biological outcomes measured were direct responses of mRNA and protein expression of important PRRs and innate immune pathway mediators (CD14, TLR2, TLR4, TLR5, TLR9, MYD88, TOLLIP) and inflammatory cytokines (IL-8, IL-6, TNF-α, IFN-γ, TGF-β1, TRAF6). We challenged our patient cells with live bacteria as opposed to killed bacteria or bacterial antigens in order to model biologically relevant cellular conditions.
Limitations
Owing to the relatively small number of samples, the subtle differences in protein and gene expression between patient groups did not reach significance statistically. Nevertheless, statistically significant changes in cytokine responses were demonstrated with bacterial challenge when all subjects were analysed. The source of monocytes and PMN were limited to peripheral blood, and immune cells of lung origin were not studied. Previous studies have shown that alveolar macrophages, but not peripheral blood macrophages from COPD patients have defective cytokine responses to H. influenzae. 45 However, alveolar macrophages and PBMCs show similar expression patterns in COPD patients. 46 Furthermore, peripheral blood monocytes 38 and PMN 44 have been studied previously by others to characterise immune responses in stable COPD. In the future, studying immune system response during exacerbations would be useful. Because we aimed to analyse immune response regulated by the interaction of immune cells from the blood compartment in host defence we measured gene expression in monocyte/neutrophil co-cultures; however, it may be useful in future studies to examine the separate responses of each immune cell type. Because innate immune function is complex, in this discovery investigation we studied numerous variables of relevance to our hypothesis that innate immune response may relate to rate of decline in lung function. In doing so, we did not explicitly correct for the multiple comparisons involved—this would require a much larger study.
Conclusions
This study aimed to identify differences in host innate immune responses of monocytes and PMN to bacterial infections in COPD patients with various degrees of lung function decline. We found that protein expression of cytokines increased in immune cells challenged with H. influenzae and S. pneumoniae, particularly expression from monocytes. There were no consistent differences in protein expression of cytokines across disease groups or bacteria. In co-cultures of monocytes and PMN, there was statistically significant up-regulation of mRNA expression of TGF-β1 and MYD88, and down-regulation of CD14, TLR2 and IFN-γ in H. influenzae-challenged cells compared with S. pneumoniae. The protein and mRNA expression of TNF-α was significantly up-regulated in non-rapid decliners than in non-COPD controls in response to H. influenzae (monocytes) and S. pneumoniae (co-culture) exposure respectively. Unsupervised clustering of gene expression separated the samples exposed to H. influenzae from those exposed to S. pneumoniae. We conclude that exposure to bacterial pathogens causes characteristic innate immune responses in peripheral blood monocytes and PMN in COPD. Under the given experimental conditions and the limited genes tested, there do not appear to be consistent differences in peripheral blood innate immune responses between COPD patients (whether rapid or non-rapid decliners) and controls, with the exception of TNF-α. Future studies could undertake global gene expression profiling of immune cells challenged with bacteria in order to better understand dysregulation of immune responses in COPD.
Footnotes
Acknowledgements
We sincerely thank the patients and staff of The Prince Charles Hospital for their participation in this study. Bacteria were kindly supplied by Dr Chris Coulter and Sonali Coulter, Microbiology Department, The Prince Charles Hospital. We thank John Cardinal, Cardinal Biosciences, Brisbane for assistance with the protein expression assays.
Funding
This work was supported by NHMRC project grant 401500, NHMRC Biomedical Scholarship (SSF), NHMRC Career Development Fellowship (IY), NHMRC Practitioner Fellowship (KF), Australian Lung Foundation/BoehringerIngelheim COPD Research Fellowship (IY), The Prince Charles Hospital Foundation and Asthma Foundation Queensland.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
