Abstract
Bacteroides fragilis is found among the normal intestinal flora and is involved in host immunostimulation via TLR2. Its cell surface components, such as LPS and capsular polysaccharides, were reported to participate in host immunostimulation. In this study, we report on the existence of a lipoprotein that acts as a TLR2 stimulant in B. fragilis. The TLR2-stimulating lipoprotein was obtained using Triton X-114-water phase partitioning followed by preparative SDS-PAGE. Its N-terminal hydrophobic peptide, which was separated from a tryptic digest, was characterized as a triacylated lipopeptide, and the lipoprotein was identified as BF1333 by mass spectrometry of Asp-N-digested peptides. These results showed that the lipoprotein acts as a TLR2-stimulating component in B. fragilis.
Introduction
Bacteroides fragilis is an anaerobic commensal bacterium that is present in the microflora in the human gastrointestinal tract and is associated with intra-abdominal infections, including abscess formation and bacteremia. The cell surface components of B. fragilis are known to modulate the host immune system. Capsular polysaccharides such as polysaccharide A (PSA), which is produced by B. fragilis, are reported to stimulate T-cell lineage differentiation of the mammalian immune system 1 and cytokine production by macrophages and dendritic cells. 2 LPS preparations have also been shown to activate macrophages and induce them to secrete cytokines. 3
TLRs play a critical role in the innate immune system as they initiate the host response to bacterial invasion by recognizing their structural motifs. 4 In common Gram-negative bacteria, TLR4 is an important receptor for innate immune recognition of bacterial LPS. 5 In contrast, TLR2 signaling is reported to be critical for the immune response to B. fragilis components.2,6–8 TLR2- but not TLR4-transfected cells recognized an LPS preparation from B. fragilis.6,7 In addition, wild type and TLR4-hyporesponsive mice formed abscesses in response to challenge with B. fragilis PSA or bacteria, but TLR2-/- mice exhibited a reduced ability to form abscesses. 2 Bacteroides fragilis PSA induces signaling through TLR2 on regulatory T cells to promote immunologic tolerance. 8
The main role of TLR2 is as the predominant receptor for lipoproteins derived from various bacteria. In Gram-positive bacteria, a contribution of lipoproteins to TLR2-mediated inflammation has been reported.9–12 In Gram-negative bacteria, the inflammatory responses to membrane lipoproteins from Borrelia burgdorferi, Escherichia coli, Neisseria gonorrhoeae and Porphyromonas gingivalis are mediated by TLR2.13–16 However, the contribution of B. fragilis lipoproteins to immune responses has not been demonstrated. In the present study, we identified a lipoprotein from B. fragilis bacterial cells and elucidated its N-terminal TLR2-activating structure.
Materials and methods
Bacterial strain and bacterial components
Bacteroides fragilis JCM 11019 (NCTC 9343) was provided by RIKEN BRC through the National Bio-Resource Project of MEXT, Japan. The bacteria were cultured in GAM broth (Nissui, Tokyo, Japan) under anaerobic conditions at 37°C for 6 h. The bacterial cells were harvested by centrifugation and then washed three times with saline. To extract the lipoprotein fraction, B. fragilis cells were digested with lysozyme and then with DNase and RNase, and the digested cells were subjected to Triton X-114 (TX-114)-water phase partitioning according to the method described previously. 17 The lipoprotein fraction was designated as Bf-TX (B. fragilis TX-114 extract).
FSL-1, a synthetic diacylated lipopeptide, was purchased from EMC microcollections (Tübingen, Germany). Escherichia coli O55:B5 LPS (Sigma-Aldrich, St Louis, MO, USA) was purified by sodium deoxycholate reextraction, as described previously. 18
Mass spectra
Mass spectra (MS) were obtained by MALDI-TOF-MS with an Axima QIT TOF mass spectrometer (Shimadzu, Kyoto, Japan) and an Autoflex speed mass spectrometer (Bruker Daltonics, Billerica, MA, USA). 2,5-Dihydroxybenzoic acid was used as a matrix at a concentration of 10 mg/ml in aqueous 33% acetonitrile containing 0.1% trifluoroacetic acid for peptides or in chloroform-methanol (1:1) for lipopeptides. The peptides were analyzed in positive ion reflectron or reflector mode. Tandem MS (MS/MS) spectra were obtained in collision-induced dissociation (CID) mode using argon as the collision gas in the Axima QIT spectrometer or in post-source decay (PSD) mode in the Autoflex speed spectrometer. On-target oxidation was performed as described previously. 19 Briefly, 30% hydrogen peroxide was spotted onto the sample–matrix co-crystal on a MALDI sample target and the target was kept at ambient temperature until dry.
Separation and identification of lipoproteins and lipopeptides
Analytical SDS-PAGE was performed using the Tris–glycine method. Proteins were visualized with Coomassie brilliant blue (CBB) staining. The separated proteins were digested in gel with trypsin or Asp-N (Promega, Madison, WI, USA), as described previously. 20 The peptides were then analyzed by MALDI-TOF-MS and identified by a database search using the MASCOT software from Matrix Science. 21 Lipoproteins were identified by a DOLOP database search (http://www.mrc-lmb.cam.ac.uk/genomes/dolop/). The proteins were designated according to the locus tags for the B. fragilis NCTC9343 genome.
Preparative SDS-PAGE was carried out using an AE-6750 preparative electrophoresis apparatus (ATTO, Tokyo, Japan), according to the manufacturer's instructions. The eluates were analyzed by SDS-PAGE and subjected to acetone precipitation to remove any contaminating SDS and obtain the lipoprotein fraction. The concentration of lipoproteins relative to the concentration of BSA was estimated by SDS-PAGE and CBB staining. To isolate N-terminal lipopeptides, the lipoproteins were digested with trypsin, and the digests were subjected to phase partitioning using ethyl acetate and water. The N-terminal lipopeptides were detected in the organic phase and characterized by MALDI-TOF-MS. Fatty acids were analyzed by the methyl ester method, as described previously. 22
In-gel TLR-stimulation assay
The Bf-TX was separated by SDS-PAGE and visualized by CBB staining. The resolved proteins in the gel were cut into strips and each strip was separately broken down mechanically in RPMI-1640 medium. The resultant suspensions were applied to a luciferase assay using Ba/mTLR2 cells.
Luciferase and cytokine assay
Ba/F3 cells that stably expressed p55IgκLuc, an NF-κB/DNA binding activity-dependent luciferase reporter construct (Ba/κB), a murine TLR2 and p55IgκLuc reporter construct (Ba/mTLR2), and a murine TLR4/MD-2 and p55IgκLuc reporter construct (Ba/mTLR4/mMD-2) were kindly provided by Prof. K. Miyake (Institute of Medical Science, University of Tokyo, Japan). The NF-κB-dependent luciferase activity in these cells was determined as described previously. 9 The results are shown as relative luciferase activity, which was determined as the ratio of stimulated to non-stimulated activity. The concentration of stimuli giving half maximal activity was estimated by reading a scatter plot with logarithm scale for the x-axis.
Eight-week-old male BALB/c mice were obtained from Kyudo (Kumamoto, Japan). The animals received humane care in accordance with our institutional guidelines and the legal requirements of Japan. The stimulation of thioglycolate-elicited peritoneal exudate cells (PECs) and the TNF-α secretion assay were performed as described previously. 10 The data are shown as the mean ± SEM values obtained from three independent experiments.
Results
Separation of the lipoprotein fraction from B. fragilis
The lipoprotein fraction from B. fragilis cells was obtained by TX-114-water phase partitioning. Accordingly, 54.1 mg (ca 4.5%) of the lipoprotein fraction (Bf-TX) was obtained from about 7 ml wet bacterial cells (correspond to ca 1.2 g dry cells). The Bf-TX activated Ba/mTLR2 cells at 100 ng/ml but not Ba/mTLR4/mMD-2 or Ba/κB cells (Figure 1A). The SDS-PAGE profile of Bf-TX showed that it contained CBB-positive proteins (Figure 1B). An in-gel TLR stimulation assay of the proteins in the Bf-TX showed that a 33-kDa protein displayed strong TLR2-mediated NF-κB activation (Figure 1B). The proteins in the Bf-TX were identified by in-gel tryptic digestion followed by peptide mass fingerprinting or MS/MS spectro-metry. However, we failed to identify the 33-kDa protein by the triptic digestion method. No lipoproteins were detected in the other CBB-positive major bands.
TLR-stimulating activity in the Bf-TX. (A) The NF-κB activation induced by Bf-TX in Ba/κB, Ba/mTLR2 and Ba/mTLR4/mMD-2 cells. The cells were incubated for 4 h and their NF-κB activation levels were measured in a luciferase assay. (B) SDS-PAGE profile of the Bf-TX and the in-gel TLR2-stimulating activity of each CBB-positive band. The Bf-TX was separated with 10% gel and visualized by CBB staining. The NF-κB activation of each gel band from the Ba/mTLR2 cells was assessed using a luciferase assay.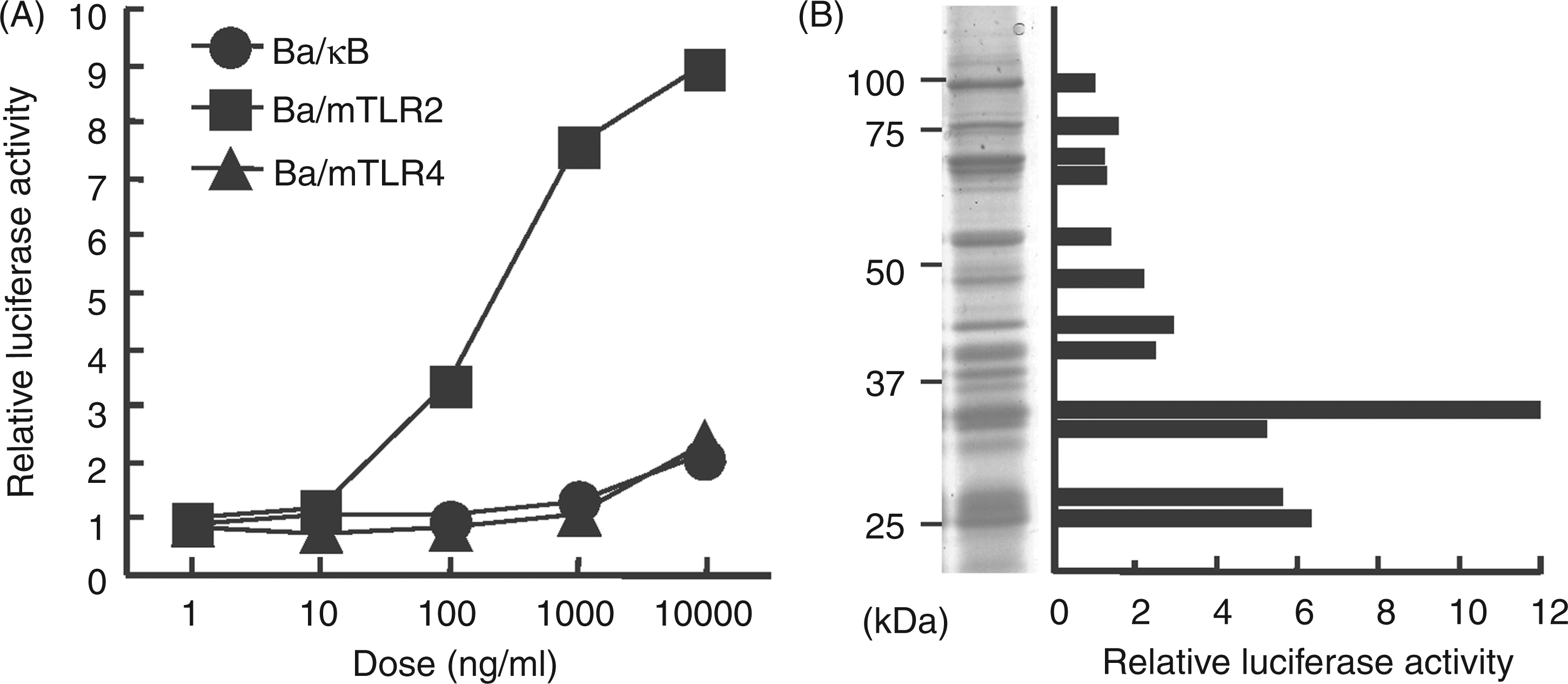
Immunological activity of the 33-kDa protein
Thus, the protein in the 33-kDa region was separated as a single band by preparative SDS-PAGE using 10% gel (Figure 2A). About 30 µg (0.6%) of the protein was obtained from 5 mg Bf-TX. The 33-kDa protein induced NF-κB activation in Ba/mTLR2 cells, but not in Ba/mTLR4/mMD-2 or Ba/κB cells (Figure 2B, C). In addition, the concentration of 33-kDa protein giving half maximal activity (ca 2 ng/ml, Figure 2C) displayed 100-fold lower activity than that of Bf-TX (ca 200 ng/ml, Figure 1A). It also stimulated murine PECs to produce TNF-α in a dose-dependent manner (Figure 2D). These results suggest that the 33-kDa protein is mainly responsible for the activity of Bf-TX.
Immunological activities of a 33-kDa protein isolated by preparative SDS-PAGE. (A) SDS-PAGE profile of the 33-kDa protein isolated using a 10% preparative gel. The proteins separated in 10% analytical gel were visualized by CBB. (B) The NF-κB activation induced by 10 ng/ml of the 33-kDa protein, 1 ng/ml FSL-1 and 10 ng/ml LPS in Ba/κB, Ba/mTLR2 or Ba/mTLR4/mMD-2 cells. (C) Dose-dependent NF-κB activation induced by the 33-kDa protein in Ba/mTLR2 cells. The cells were incubated with the above stimuli for 4 h. NF-κB activation was measured with a luciferase assay. (D) TNF-α production induced by the 33-kDa protein in murine PEC. The levels of TNF-α in the culture supernatants of the cells incubated for 4 h were measured by ELISA.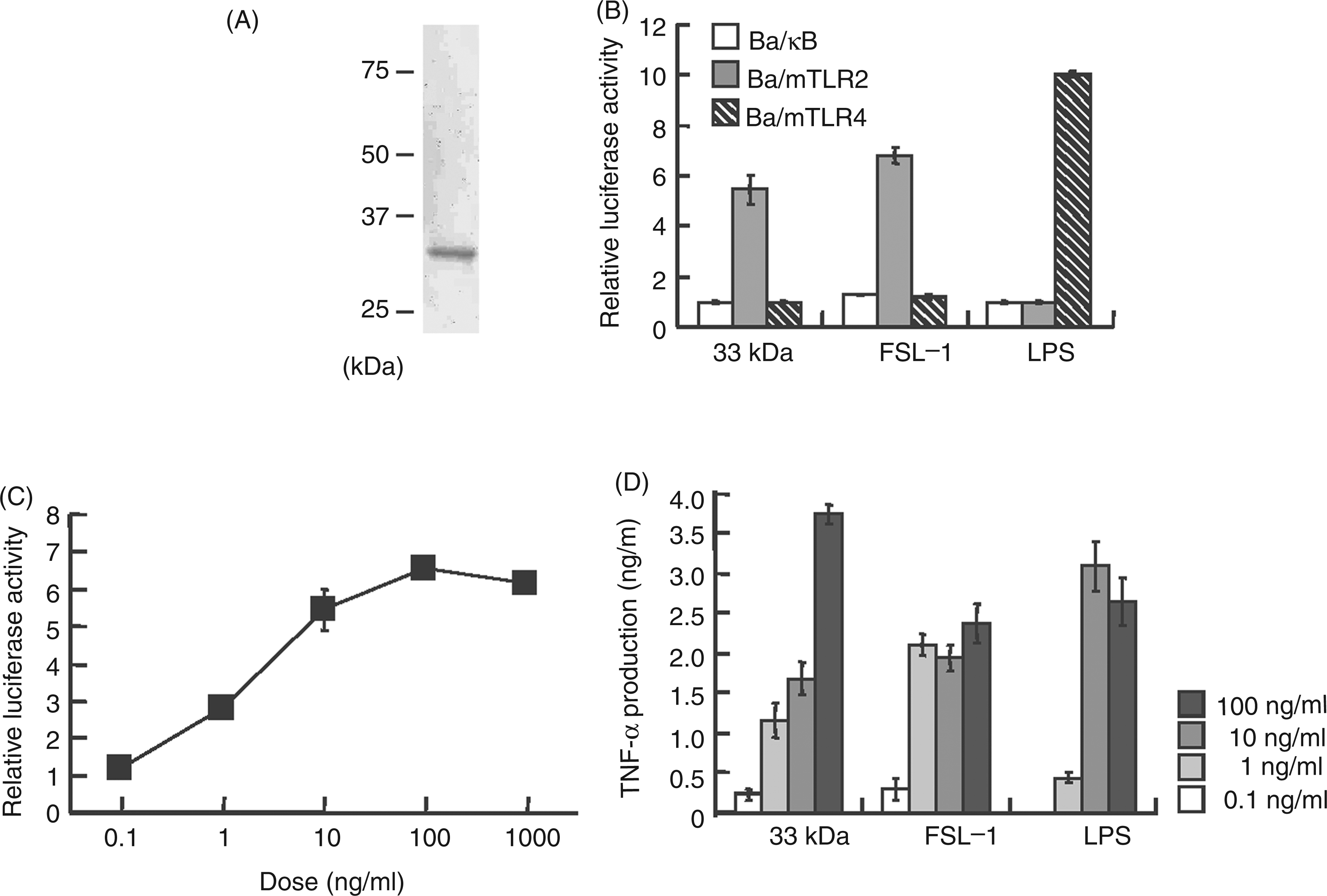
Evidence for the presence of N-terminal acyl groups in the 33-kDa protein
Bacterial lipoproteins have been shown to possess hydrophobic N-terminal moieties composed of acylated S-(2,3-dihydroxypropyl)cysteine.
23
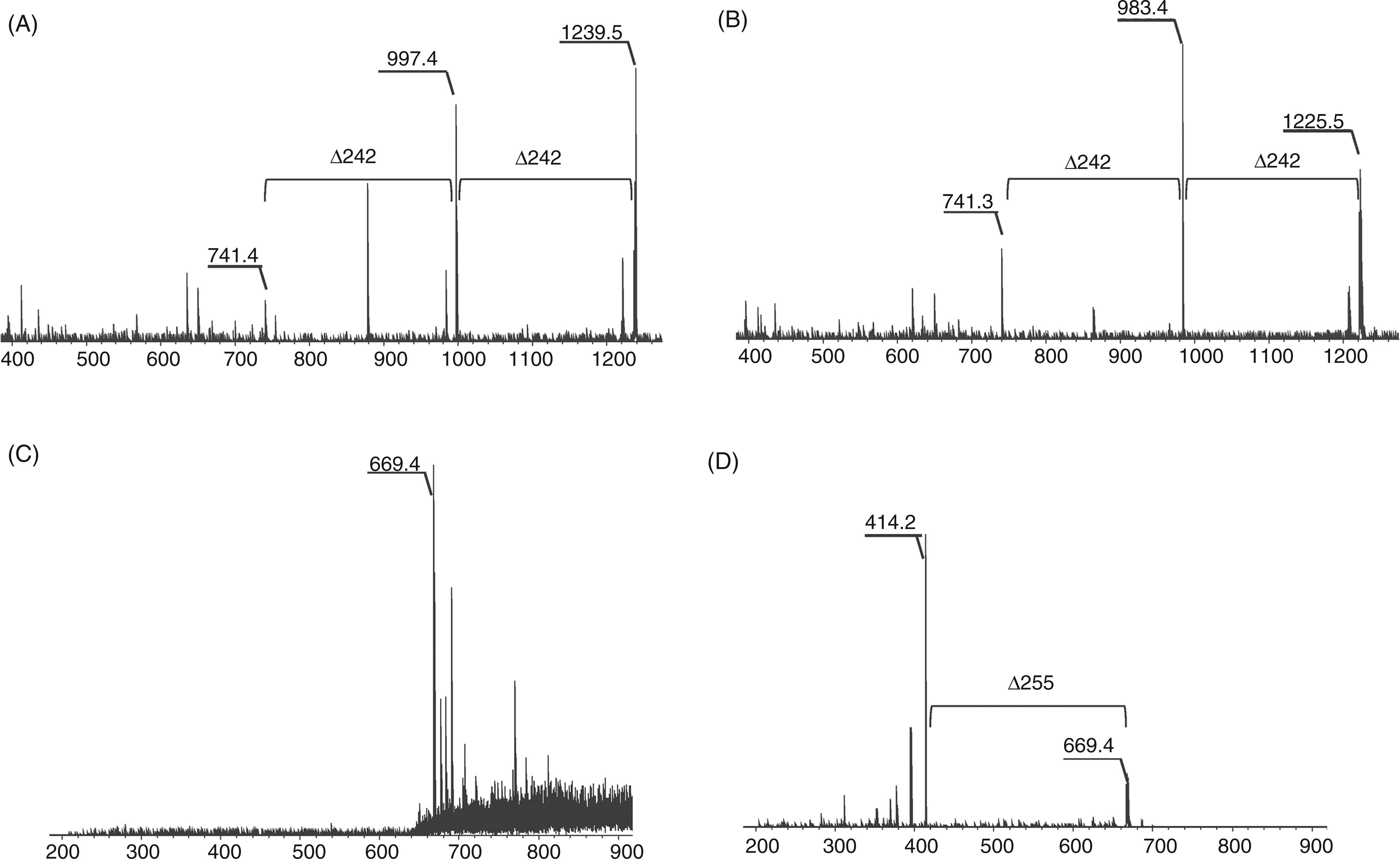
To clarify whether the 33-kDa protein obtained by preparative SDS-PAGE was a lipoprotein, its N-terminal structure was characterized. The 33-kDa protein was subjected to tryptic digestion followed by phase separation using ethyl acetate and water, and TLR2-stimulating activity was detected in the organic phase (Figure 3A), suggesting that the active compound was a hydrophobic component like lipopeptide. Thus, the structure of the active compound in the organic extract was analyzed by MS spectrometry using an Axima QIT spectrometer. The MALDI-TOF-MS spectrum of the active compound displayed two major pseudomolecular ions [M + H]+ at m/z 1225.5 and m/z 1239.5 (Figure 3B). The mass difference of each ion was m/z 14, which corresponded to a methylene (CH2) group. In the MS/MS spectrum of the precursor ion at m/z 1239.5, daughter ions were observed at m/z 997.4 and m/z 741.4, which corresponded to the neutral loss of the fatty acids 15:0 (Δm/z = 242) and 16:0 (Δm/z = 256) respectively (Figure 4A). In the MS/MS spectrum of the precursor ion at m/z 1225.5, daughter ions were observed at m/z 983.4 and m/z 741.3, which corresponded to the neutral loss of the fatty acid 15:0 (Δm/z = 242) (Figure 4B). When the active compound was subjected to on-target oxidation, which oxidized the sulfur atom of the dihydroxypropyl cysteine moiety of the lipopeptide, its MS spectrum showed an ion at m/z 669.4 (Figure 4C). The mass difference was consistent with the elimination of a 2,3-diacycloxypropane-1-thiol moiety, which contain two 15:0 (Δm/z = 556) O-acyl groups, or one 15:0 O-acyl group and one 16:0 (Δm/z = 570) O-acyl group. In the MS/MS spectrum of the precursor ion at m/z 669.4, daughter ions were observed at 414.2 u, which corresponded to the neutral loss of the fatty acid amide 15:0 (Δm/z = 255) (Figure 4D). These results suggested that the active compound was a triacyl type lipopeptide. Fatty acid analysis showed that the compound contained iso-15:0, anteiso-15:0 and 16:0 (2:2:2.5). These results agreed with the above MS data.
Characterization of active components in the tryptic digest of the 33-kDa protein. (A) NF-κB activation induced in Ba/mTLR2 cells by tryptic peptide extracts. The 33-kDa protein was digested with trypsin and partitioned with ethyl acetate and water. The cells were incubated with the extracts for 4 h. NF-κB activation was measured with a luciferase assay. (B) MS spectrum of the active compound in the organic extract. The spectrum was obtained with an Axima QIT spectrometer in positive ion mode. Determination of the N-terminal acyl group of the active component. (A) MS/MS spectrum of the ion at m/z 1239.5 in the organic extract. (B) MS/MS spectrum of the ion at m/z 1239.5. The precursor ion was decomposed using CID. (C) MS spectrum of the active component after on-target oxidation. The sample was oxidized with H2O2 on a MALDI target plate. (D) MS/MS spectrum of the ion at m/z 669.4. All spectra were obtained with an Axima QIT spectrometer in positive ion mode.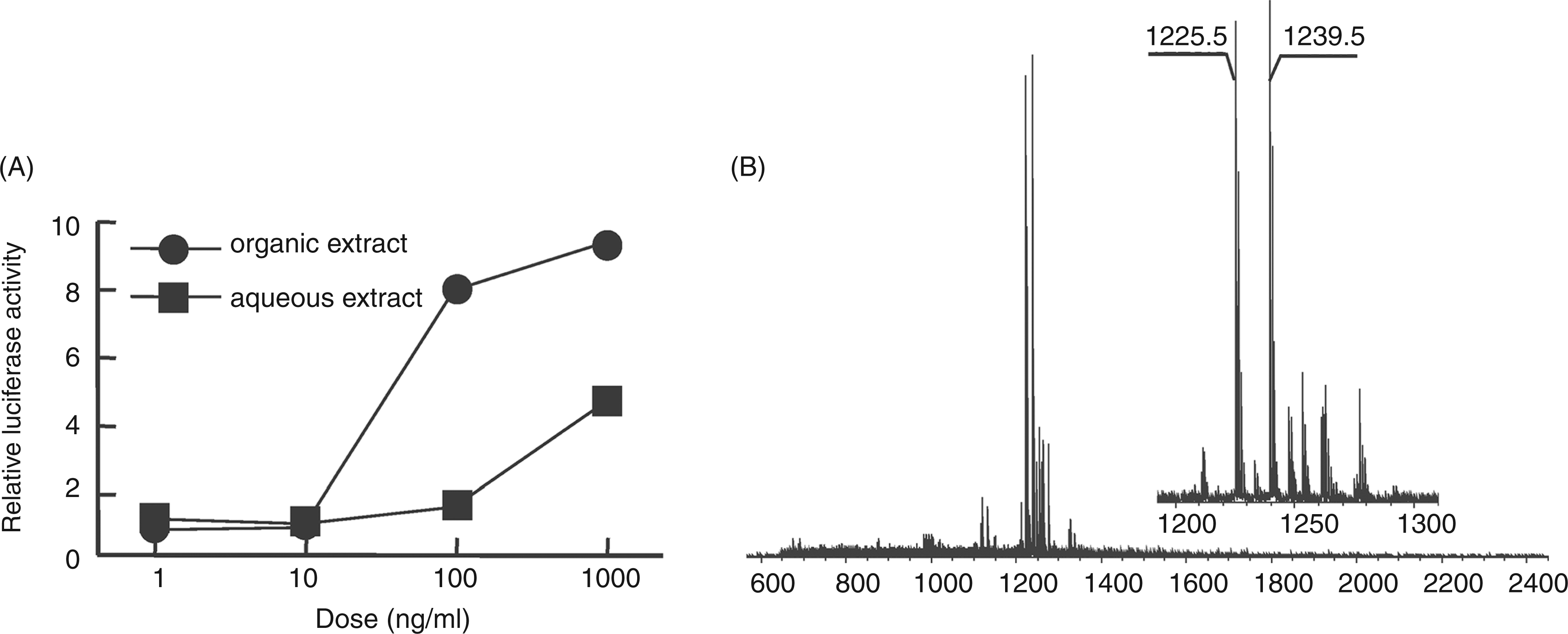
Identification of the 33-kDa B. fragilis protein
To elucidate the sequence of the 33-kDa protein, the active compound was subjected to MS/MS analysis with an Autoflex speed spectrometer (Figure 5A). C-terminal y ions were observed at m/z 362.1, m/z 234.1 and m/z 147.1, which suggested a sequence of QSK. Thus, the structure of the lipopeptide was considered to be as shown in Figure 5B. A DOLOP database search for the CQSK sequence indicated that the 33-kDa protein is BF1333, a B. fragilis NCTC 9343 protein. BF1333 possesses a lipobox sequence (LVSC) and is considered to be a lipoprotein (Figure 5C). BF1333 is considered to be difficult to identify by peptide mass fingerprinting of its tryptic digest because its K and R residues are too close together to obtain peptides of appropriate lengths. This might have contributed to our failure to identify the 33-kDa protein. In contrast, the location of the D residue made peptide fingerprinting possible. Thus, the 33-kDa protein was digested in gel with Asp-N and analyzed by MS (Figure 5D). Using a MASCOT database search, the protein was identified as BF1333. These results show that the TLR-2 stimulating lipoprotein in B. fragilis is BF1333.
Identification of the 33-kDa protein. (A) MS/MS spectrum of the ion at m/z 1239.5. The precursor ion was decomposed using PSD. (B) Structure of the N-terminal lipopeptide. (C) Amino acid sequence of BF1333 in B. fragilis NCTC9343. The isolated N-terminal sequence has been surrounded with a box. (D) MS spectrum of the 33-kDa protein digested with Asp-N. The underlined sequences in C were determined by a MASCOT search. All spectra were obtained with an Autoflex speed spectrometer in positive ion mode.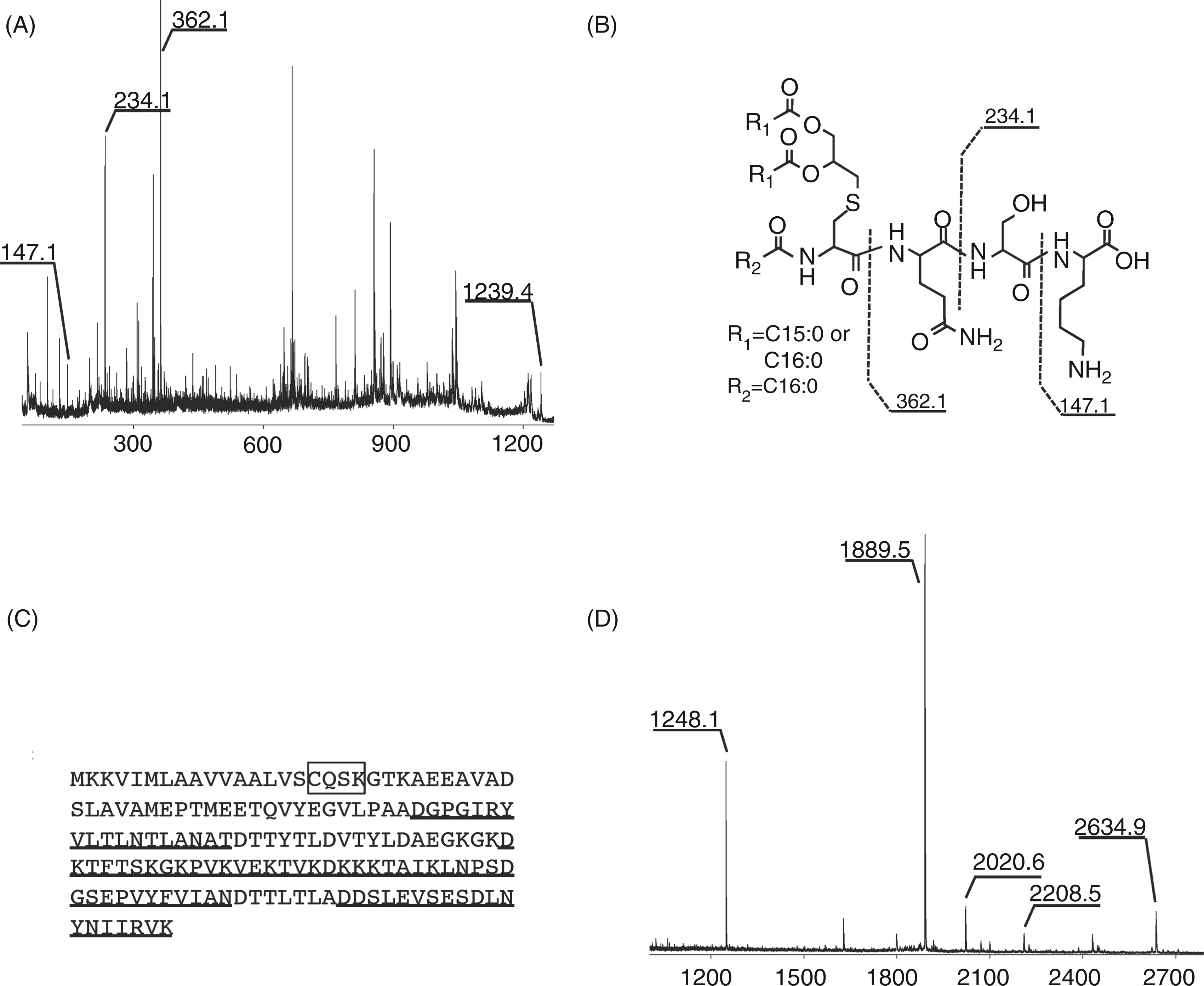
Discussion
The cell surface components of B. fragilis, such as PSA preparation, have been reported to modulate the immune system via TLR2.2,8 In this study, we detected a B. fragilis lipoprotein that also stimulates TLR2. BF1333 is classified as a hypothetical protein and no function has been assigned to it at present. However, the recovery of BF1333 from Bf-TX (ca 0.6%) and the activity enrichment ratio of BF1333 to Bf-TX (ca 100 times, estimated from Figures 1A and 2C) suggests that BF1333 is a relatively major constituent in Bf-TX and will significantly contribute to immune modulation by B. fragilis cells. We also observed slight TLR2-stimulating activity in the lower molecular mass range, e.g. 25 kDa (Figure 1B). In the B. fragilis NCTC9343 and YCH46 genomes, 177 and 184 lipoproteins have been identified respectively (predicted list of lipoproteins from bacterial genomes in DOLOP database, http://www.mrc-lmb.cam.ac.uk/genomes/dolop/compgen.shtml). Although we did not identify any additional lipoproteins in this study, some other lipoproteins might be expressed in B. fragilis. Furthermore, the bacterial cells derived from the present cultural conditions might not correspond to the in vivo pathogenic conditions of the organism. Thus, B. fragilis might display a different in vivo protein expression pattern to that detected in this study. A further study aimed at detecting lipoproteins that act as virulence factors in vivo might be required.
In a previous study, a B. fragilis LPS preparation was reported to stimulate TLR2;6,7 however, a contradictory report found that a highly purified LPS preparation produced using a modified phenol–water re-extraction method activated cells via TLR4 and its activity was independent of TLR2. 24 At present, the latter observation is considered to be reasonable. In LPS molecules, the lipid A moiety is proved as the structural motif responsible for by TLR4 recognition. 25 Escherichia coli-type lipid A which consists of diglucosamine backbone, 1,4'-bisphosphate and six fatty acids composed of four 14:0(3-OH), 12:0 and 14:0, is one of the most powerful stimuli. 26 In contrast, B. fragilis lipid A was identified as a penta-acylated monophospho-lipid A which consists of relatively long-branched chain fatty acids and lacks 4'-phosphate. 27 Such structurally-truncated lipid A is expected to have a significant reduced TLR4 activity compared with the E. coli-type one. 26 Recently, Park et al. showed the crystal complex of LPS/MD-2/TLR4 in which the number of acyl chains and the two phosphates at the lipid A backbone play a crucial role in the TLR4 recognition. 28 In contrast, no study has reported that TLR2 recognizes any synthetic lipid A molecule. Therefore, B. fragilis lipid A activates TLR4 even weakly, but does not activate TLR2 at all. Furthermore, the present observations suggest that contaminating lipoproteins in the LPS preparation might have been responsible for the observed TLR2-mediated cellular activation. Contaminating lipoproteins in LPS preparations are often removed by re-extraction;14,18 however, it is sometimes not possible to completely remove all lipoproteins. We demonstrated previously that contaminating lipoproteins in the LPS fraction of P. gingivalis were responsible for its activation of the innate immune system through TLR2. 16 We also found that lipoproteins could not be completely removed from the LPS fraction by re-extraction. Mancuso et al. also noted that TLR2-stimilating contaminants in B. fragilis LPS are unlikely to be proteins. 24 However, BF1333 is difficult to be identified by commonly-used peptide mass fingerprinting of tryptic digest. Therefore, the immunomodulatory activity of B. fragilis LPS preparations should be re-investigated.
In conclusion, we identified a TLR2-stimulating lipoprotein in B. fragilis cells and found that it possessed a triacylated N-terminal lipopeptide structure. Although immunostimulatory lipoproteins are known to be expressed in many bacteria, the lipoproteins in B. fragilis have not previously been considered to be pathogenic. Our findings might contribute to preventing a symptom of B. fragilis infection and elucidating the mechanism underlying immune homeostasis in the intestinal tract.
Footnotes
Acknowledgements
We thank Mr Atsushi Saito and Ms Maiko Furuyashiki for providing technical assistance.
Funding
This study was supported in part by a SUNBOR grant from the Suntory Institute for Bioorganic Research.
Conflict of interest
The authors declare that there is no conflict of interest.
