Abstract
The innate response interferon-inducible transmembrane (Ifitm) proteins have been characterized as influencing proliferation, signaling complexes and restricting virus infections. Treatment of cells lacking these proteins (IfitmDel) with IFN-β resulted in the loss of clathrin from membrane compartments and the inhibition of clathrin-mediated phagocytosis, suggesting a molecular interaction between clathrin and Ifitm proteins. The pH of endosomes of IfitmDel cells, with or without IFN activation, was neutralized, suggesting the function of the vacular ATPase proton pumps in such cells was compromised. Co-immunoprecipitation of Ifitm3 with Atp6v0b demonstrated a direct interaction between the Ifitm proteins and the v-ATPase. These data suggest that the Ifitm proteins help stabilize v-ATPase complexes in cellular membranes which, in turn, facilitates the appropriate subcellular localization of clathrin.
Introduction
The interferon-inducible transmembrane (Ifitm) proteins are encoded by a set of highly conserved duplicated genes.1,2 The Ifitm proteins are about 130 amino acids long and possess gene/protein specific N-terminal sequences followed by conserved transmembrane (TM) and cytoplasmic domains (the latter about 15 amino acids long) followed by a third, less conserved domain with TM-like, alpha-helical characteristics.
The Ifitm protein family has been studied in humans, cows, rats and mice. There are four functional members of this gene family in the human: IFITM1 (Leu-13), IFITM2 (1-8D), Ifitm3 (1-8U) and IFITM5 (BRIL).2,3 Ifitm3 is highly inducible with type I and type II IFNs, IFITM1 is less responsive and the expression of IFITM2 is unchanged.4,5 Human IFITM1 (Leu-13) was first identified, with a monoclonal Ab, as a 16-ku protein expressed on the surface of human cells. 6 Treatment of cells with the anti-Leu13 Ab inhibited the proliferation of resting and activated B cell lines, induced lymphocyte aggregation and depressed proliferation in T cells following stimulation. 7 Anti-Leu-13 treatment also led to the rapid release of L-selectin, 8 suggesting that IFITM1 may play a role in membrane organization. IFITM1 serves as a negative regulator of MAPK signaling that is induced by IFN-γ; over-expression of IFITM1 resulted in the arrest of cell cycle progression in the G1 phase in a p53-dependent fashion. 9 IFITM1 was also characterized as interacting with caveolin-1 in the caveola via the hydrophobic IFITM TM domain. This interaction inhibits ERK activation by enhancing the inhibitory function of CAV-1. 10 IFITM1 is associated with CD81, a tetraspanin protein, on the cell surface of B cells. 11 The best-defined signaling complex that includes IFITM1 is that of CD81, CD19 and the CR2 (CD21) complement receptor. 12 A report linked specific alleles of Ifitm3 with ulcerative colitis, suggesting that altered expression of the protein may be instrumental in leading to susceptibility to colitis and inflammatory bowel disease. 13 Analysis of the rat IFITM/Rat8 protein, using a rat breast adenocarcinoma model system, demonstrated that the protein was held within vesicular sites in the cytoplasm of the cell prior to induction. After activation, the protein moved to the cell surface and was found in lipid rafts associated with the Lyn and Fyn protein kinases.14,15
The mouse genome possesses 5 functional Ifitm genes clustered within 68,000 base pairs (bps) on mouse chromosome 7. 1 A series of studies proposed that the Ifitm proteins were critical for the appropriate homing and repulsion of primordial germ cells and were thus essential for the appropriate organization of the developing mouse embryo.1,16,17 However, mice lacking all Ifitm proteins (IfitmDel) developed normally without any embryonic complications. 18 We have shown previously that the Ifitm2 and Ifitm3 proteins associate with the tetraspanin proteins CD81 and CD9 on the surface of activated B cells and also entered lipid rafts following anti-IgM/anti-CD19 crosslinking. 19 Activation of mouse fibroblasts and macrophages with IFN-β leads to the movement of the Ifitm proteins from a cytoplasmic location to the membrane.
A number of recent studies have illustrated the importance of the Ifitm proteins in the anti-viral, innate immune IFN response. Using a siRNA screen, the Ifitm3 protein was defined as restricting the infection of cells with influenza A. 20 Over-expression of IFITM1, 2 or 3 could also block influenza infection in the absence of IFN signaling. Cells obtained from the IfitmDel animal were also more susceptible to influenza infection than IFN-activated wild type (WT) controls were: forced expression of Ifitm2 or Ifitm3 in the IfitmDel cells restored the relative resistance of such cells to infection. The IFITM proteins were shown to inhibit infection of a number of viruses, including HIV, West Nile flavivirus, severe acute respiratory syndrome (SARS) coronavirus, Marburg and Ebola filoviruses, Dengue virus and vesicular stomatitis virus.21–24 Furthermore, the over-expression of Ifitm3 led to the expansion of acidified compartments within host cells, potentially providing a means for blocking cytosolic entry of the virus. 25 These data suggested that the IFITM proteins influenced viral infections by binding to cellular or viral partners, and blocking the entry of the virus into the cytoplasm, required for replication. Many of the viruses blocked from infection by the IFITM proteins utilize a clathrin dependent pathway, 26 suggesting that one anti-viral function of the IFITM proteins may be in altering normal clathrin functions.
In this study, we have further defined the roles of the Ifitm proteins in normal signaling complexes and IFN responses by analyzing the localization of membrane associated proteins in unstimulated and IFN-β activated cells from WT and IfitmDel animals. Intriguingly, we have found that IFN-treated mouse cells lacking Ifitm proteins show a dramatic loss of clathrin heavy chain (CHC) in the membrane that is coincident with the loss of endocytosis via the clathrin-dependent pathway. We also demonstrate that the pH of the endosomes from resting or IFN-treated cells from the IfitmDel animal is unusually high and that the Ifitm3 protein has direct physical contact with a multi-membrane spanning protein of the vacuolar ATPase (v-ATPase) complex required for normal lysosome acidification.
Materials and methods
Mice and primary cell culture
Mice lacking the Ifitm genes (IfitmDel) 18 were obtained from Dr M. Azim Surani (University of Cambridge, UK). Animals used in these experiments were females aged 6–10 wks. The genotype of the IfitmDel mouse was determined by PCR using two oligonucleotides whose product is specific for the genetic deletion. Oligonucleotide sets were used to define the WT and the IfitmDel alleles: WT allele: forward, 5′-AACATGCCTTGCATCCCTGGAGTTCCTTCTAAAGGA-3′; reverse, 5′-CCCTAAAACACTTAGCAGTGACCCCTCACAAGCC-3′. IfitmDel allele: forward, 5′-ACTCTAGCCAGAGTCTTGCATTTCTCAGTCCTAAAC-3′; reverse, 5′-TCTAGTACAGTCGGTAAGAACAAAATAGTGTCTATCA-3′.
Cells from the IfitmDel mouse used in these experiments were obtained from mixed C57BL/6/129/sv heritage and WT cells were obtained from Ifitm sufficient littermates. The experiments detailed in this study were approved by the University of Utah Institutional Animal Use and Care Committee; Protocol 10-06003.
Cell isolation and primary cell culture
Bone marrow-derived macrophages (BMDM) were obtained by flushing femurs of mice described with serum-free RPMI and culturing with supernatant from L929 cells for 7 d. Spleens were disrupted through 100 µm cell strainers and washed with PBS, followed by RBC lysis. Spleen cells were plated in 10-cm dishes in RPMI with
Sucrose gradient lipid raft fractionation and Western blot analysis
Cells were prepared from spleens of WT and IfitmDel animals (two spleens from each genotype for a total of about 3 × 107 cells per gradient). After 18 h of IFN treatment (500 units/ml of Universal Type 1 IFN), cells were lysed in 1 ml TNEV (10 mM Tris-HCl, 5 mM EDTA, 150 mM sodium chloride, 1 mM Na orthavanadate and protease inhibitors) for 30 min on ice with 10 strokes of a dounce homogenizer. The nuclear fraction and cell debris were removed by centrifugation at 15,000 g for 5 min. The supernatant was diluted with an equivalent volume of 80% sucrose over which was loaded a 5–30% sucrose step gradient. The gradients were centrifuged at 200,000 g in a SW41Ti rotor (Beckman Coulter, Fullerton, CA, USA) for 20 h. After centrifugation, twelve 1-ml fractions were collected from the top, precipitated with 30% (v/v) trichloroacetic acid in acetone at −20°C for 18 h and separated by SDS-PAGE.
For Western blot analysis, proteins were transferred onto Immobilon-P Transfer membrane (Millipore, Bedford, MA, USA) after resolution in a 4–20% protein SDS-PAGE. Membranes were blocked with 5% skimmed milk and 0.1% Tween 20 in TBS (25 mM Tris-HCl, pH 7.5, 150 mM NaCl, 2 mM KCl), and then blotted with primary Ab for 18 h: anti-clathrin monoclonal Ab (BD. Transduction Laboratories, San Jose, CA, USA) (1:2500 dilution), anti-Lyn polyclonal Ab (Santa Cruz, Santa Cruz, CA, USA) (1:2500 dilution) and anti-Ifitm3 rabbit sera (1:2500 dilution). 19 After incubation with HRP-conjugated secondary Abs (1:5000 dilution), signals were enhanced with enhanced-chemiluminescent substrate (ECL) (Thermo Scientific, Rockford, IL, USA).
Retroviral infection of cells
Plat-E cells (1 × 106) were plated in 10-cm dishes in DMEM with 10% FBS containing 10 µg/ml blasticidin and 1 µg/ml puromycin before transfection. Cells were transfected with pBMN-I-GFP plasmid vector containing Ifitm3 with FLAG tagged at the 3′ end or empty pBMN-I-GFP plasmid vector by using a Fugene HD transfection reagent (Roche). Retrovirus was applied at appropriate dilution to the MEF cells plated in 12-well dishes and 8 µg/ml Polybrene was added to each well. The efficacy of infection was obtained by FACS.
Uptake assays, immunofluorescence staining and confocal microscopy
For co-localization studies, BMDM and MEFs from WT and IfitmDel animals were grown on coverslips with or without IFN-β (500 units/ml of Universal Type 1 IFN) for 18 h. Cells were then fixed using 4% paraformaldehyde for 15 min at 20–22°C. After permeabilization with 0.25% Triton X-100 for 10 min, the samples were blocked with 1% BSA for 30 min. Primary Abs, including anti-clathrin monoclonal Ab (BD Transduction Laboratories) and anti-Lyn polyclonal Ab (Santa Cruz), and appropriate secondary Abs were used. Cells were counterstained with DAPI (Molecular Probes/ Invitrogen, Carlsbad, CA, USA) and mounted with Fluoro Save reagent (Calbiochem, San Diego, CA, USA).
For transferrin (25 µg/ml Alexa 594-Tf, Invitrogen, Carlsbad, CA, USA) and endothelial growth factor (EGF) (1.5 ng/ml Alexa 488-EGF, Invitrogen) uptake assays, cells were serum starved for 1 h and uptake assays performed by incubating cells for 1 h at 37°C in RPMI (Invitrogen). The dextran (0.5 mg/ml 10,000 MM, Invitrogen) uptake assay was performed by incubating cells for 30 min at 37°C in RPMI. After incubation with the ligands, cells were washed and fixed.
MEFs were incubated with 50 nM Lysotracker Red DND-99 (Invitrogen) for 5 min at 20–22°C in Hanks’ Balanced Salt Solution (HBSS, Invitrogen). After incubation, the cells were washed with HBSS, fixed and mounted as described previously. All fluorescent images were obtained by using a confocal microscopy (Olympus FV-1000, Olympus, San Diego, CA, USA).
Statistical analysis
For co-localization, transferrin, dextran uptake assays and Lysotracker Red DND-99, images were obtained using a 60 × oil immersion objective (Olympus) and the z-stack images were scanned at ˜3 µm intervals throughout the depth of the cells on the cover slips. Three independent experiments were carried out in order to obtain 4–5 60 × field images that contained 10–15 cells per field (obtained from groups of 40–50 cells). For EGF uptake assays, images were obtained using 40 × oil immersion objective (Nikon, Tokyo, Japan). Five independent experiments were carried out and a total 500 cells were analyzed in 5 groups of 100 cells. Volocity software (PerkinElmer, Waltham, MA, USA) was used for quantification and image processing was achieved by using Photoshop CS3 (Adobe Systems Inc, San Jose, CA, USA). Statistical differences were determined with one-way ANOVA.
Immunoprecipitation analysis
J774 macrophages (1 × 108 cells per immunoprecipitation) were lysed in Nonident P-40 [0.5% (v/v) NP-40, 150 mM sodium chloride, 50 mM Tris-HCl pH 7.5, 0.05 mM EDTA and Roche protease inhibitors] or RIPA [50 mM NaCl, 25 mM Tris-HCL, pH 7.5, 1 mM EDTA, 0.1% SDS, 1% sodium deoxycholate (DOC), 1% NP40 and Roche protease inhibitors). Lysates were incubated with rabbit serum that was raised against the conserved intracellular domain of Ifitm proteins for 18 h at 4°C. Then, a 50% slurry of Protein G–agarose beads (Sigma, St Louis, MO, USA) was added and the reaction mixtures were further incubated for 2 h at 4°C. The beads were separated from the supernatant by centrifugation and washed three times. The agarose beads were boiled in 2 × SDS sample buffer and resolved by 4–20% gradient protein gel (Thermo Scientific). The Western blot was probed with a goat Ab against ATP6v0b (sc-241867, Santa Cruz) (1:200 dilution) followed by anti-goat HRP secondary Ab (1:10,000).
Results
Cells lacking Ifitm proteins remain responsive to type I IFN activation
Members of the mouse Ifitm gene family demonstrate enhanced expression in the presence of type I and II IFNs, and the Ifitm1,3,6 genes possess canonical IFN response elements.
27
Using quantitative RT-PCR with primer sequences specific for each of the five Ifitm family members, we analyzed BMDM, mouse embryonic fibroblasts MEFs and total mouse splenocytes for IFN-β-induced transcriptional responses of the five Ifitm genes. As demonstrated in Supplementary Figure 1, the expression of Ifitm3 was most pronounced with the addition of IFN-β in these cell samples. The expression of Ifitm2 and Ifitm6 in the BMDM and Ifitm2 in MEFs remained constant, regardless of IFN addition. Therefore, for the following studies, the primary target for IFN-β responses in the Ifitm family is Ifitm3.
Clathrin heavy chain is lost from membrane fractions in activated IfitmDel cells. Sucrose gradient sedimentation analysis of total spleen cells from WT and IfitmDel animals was performed following with no treatment (mock) (A) or treated with IFN-β for 18 h (B). After fractionation, samples were analyzed by Western blot. The more buoyant fractions are to the left (indicated with LR for lipid raft) whilst the more dense cytoplasmic fractions are to the right. This experiment is representative of five similar analyses.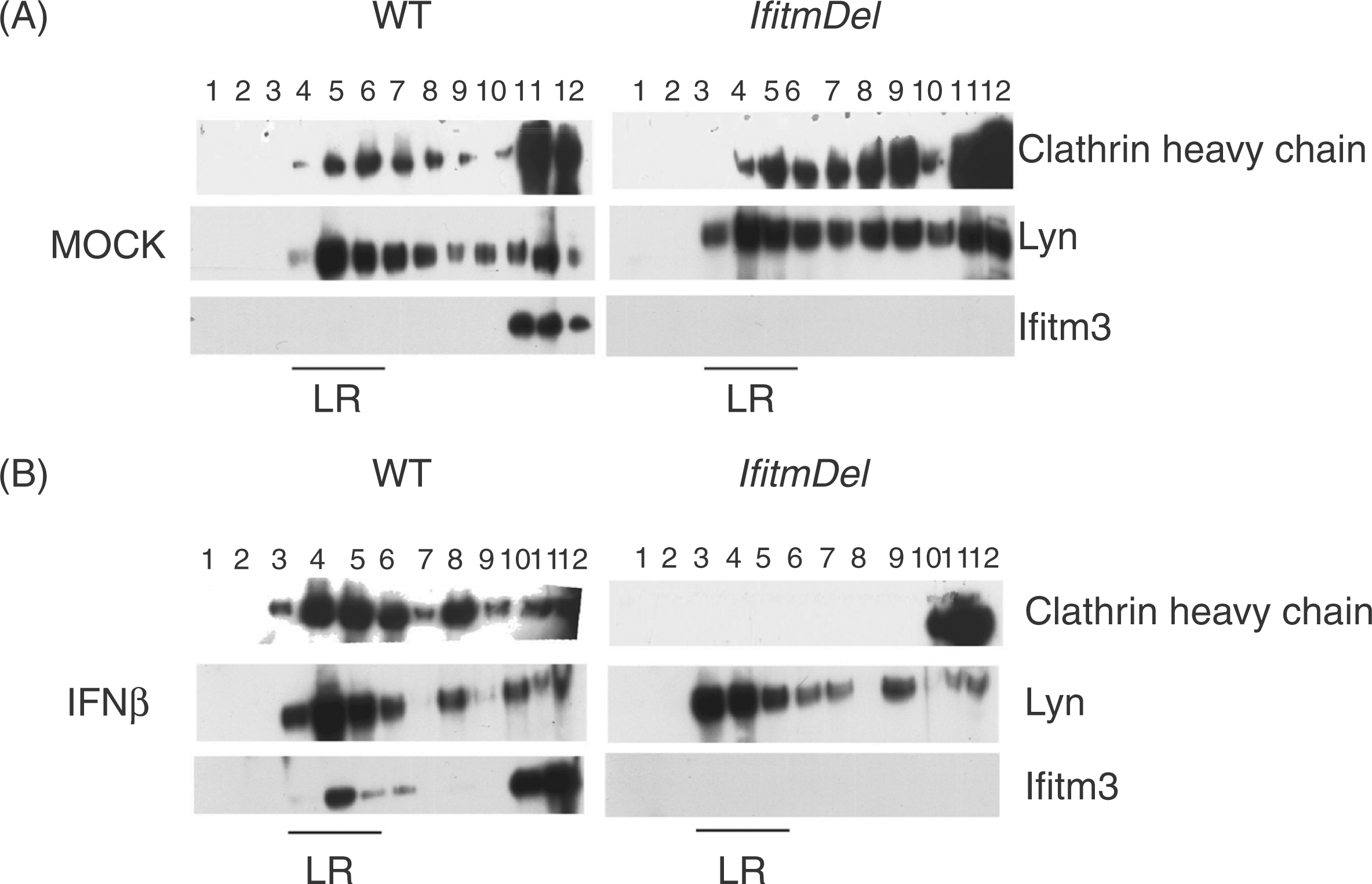
The IfitmDel animal lacks all five of the Ifitm genes. Before probing the intracellular pathways utilized by the Ifitm proteins in normal and IFN-activated cells, we first wanted to demonstrate that cells obtained from the IfitmDel animal were still fully responsive to activation with IFN-β. Accordingly, BMDM and MEFs obtained from WT and IfitmDel animals were treated with IFN-β and analyzed for the expression of the type I IFN-inducible Cxcl10 and Ifit1 genes. As shown in Supplementary Figure 2, both the BMDM (panel A) and MEFs (panel B) showed elevated expression of Cxcl10 and Ifit1 2 hrs after treatment with IFN-β. Such elevated expression was largely lost 18 h after treatment except for the Ifit1 transcript in the fibroblast lines. Similar induction of IFN response genes was seen using total splenocytes from WT and IfitmDel animals (data not shown). Thus, the primary responsiveness of the IfitmDel cells to type I IFN treatment was similar to that of the WT cells.
Cells lacking the Ifitm proteins lose clathrin localization at the cell surface following IFN activation
We, and others, have shown that the Ifitm proteins co-associate with tetraspanin proteins within lipid rafts following cellular activation/IFN induction,12,19,28,29 which led us to question if the absence of the Ifitm proteins could alter the localization of the normal membrane constituents. One method for defining lipid raft constituents is discontinuous sucrose gradient centrifugation in which the lighter, cholesterol-containing fractions are separated from the denser, cytoplasmic and membrane fractions. Accordingly, total splenocytes from WT and IfitmDel animals were cultured with or without IFN-β for 18 h and analyzed by discontinuous sucrose centrifugation (Figure 1). In the absence of IFN-β, the presence of the Lyn kinase and CHC proteins were both evident in the low-density region of the gradient (associated with clathrin-coated pits and lipid rafts), while the Ifitm3 protein was present in the cytoplasmic, dense membrane fraction. There was no difference in the localization of clathrin or Lyn proteins in cells derived from the IfitmDel animal. After treatment of the WT cells with IFN-β for 18 h, clathrin and Lyn remained within the lipid raft region of the gradient and were joined with Ifitm3. Strikingly, the IFN-β treated IfitmDel cells showed a complete loss of clathrin within the low-density region of the gradient while the localization of Lyn was left undisturbed. These data suggest that Ifitm protein(s) play a role in maintaining the position of clathrin within the clathrin-coated pits of the membrane following activation with IFN-β.
We further evaluated the localization of clathrin in the IfitmDel cells by direct visualization of the localization of the protein following IFN-β activation. Using BMDM derived from WT and IfitmDel bone marrow (untreated or treated with IFN-β), cells were fixed and stained for Lyn and CHC, and analyzed by confocal microscopy. As shown in Supplementary Figure 3, treatment of WT cells with IFN-β maintained the co-localization of the CHC protein (green) with the Lyn protein (red), resulting in a large swath of merged yellow staining. However, the IFN-β treated IfitmDel cells showed a distinct segregation of the green and red fluorescence, indicating that the Lyn proteins remained associated with the membrane while the clathrin was largely present in the cytoplasm. Control staining of the BMDM with non-specific secondary Abs failed to identify Lyn or CHC within the cells (Supplementary Figure 4).
To quantify the confocal staining data described above, BMDM obtained from the WT and IfitmDel animals (plus or minus IFN-β activation) were stained and examined by Z-stacking confocal analysis. Figure 2A shows the Z-stack images from single cell samples demonstrating the green cytoplasmic staining of clathrin in the IFN-β treated IfitmDel cells. Dozens of such cells from multiple experiments were similarly analyzed and the co-localization of clathrin and Lyn was quantified (Figure 2B). The IfitmDel cells treated with IFN-β demonstrated a significant decrease in the co-localization of Lyn and CHC compared with the other samples. Thus, these data support the sucrose density centrifugation data described above, indicating that, following activation with IFN-β, Ifitm proteins (presumably Ifitm3) are required to maintain the appropriate localization of clathrin within the cell.
Decrease in the co-localization of clathrin heavy chain (CHC) and Lyn in IfitmDel cells following IFN-β activation. (A) After 18 h with or without IFN-β treatment, BMDM cells from WT and IfitmDel animals were fixed and co-stained with CHC (green) and Lyn (red) Abs, followed by counterstaining with DAPI. Co-localization of these two proteins in WT and IfitmDel cells was revealed by z-series stack images. Scale bar = 21µm. (B) Quantitative analysis of co-localization of CHC with Lyn based on 50 cells counted for each experimental group. The percentage of CHC co-localized with Lyn was calculated as the voxel count of the green fluorescence that co-localized with the red fluorescence divided by the total voxel count of the green fluorescence. Error bars indicate SEM. (**P ≤ 0.01 or ***P ≤ 0.001 of samples compared to the IFN-β treated IfitmDel cells.).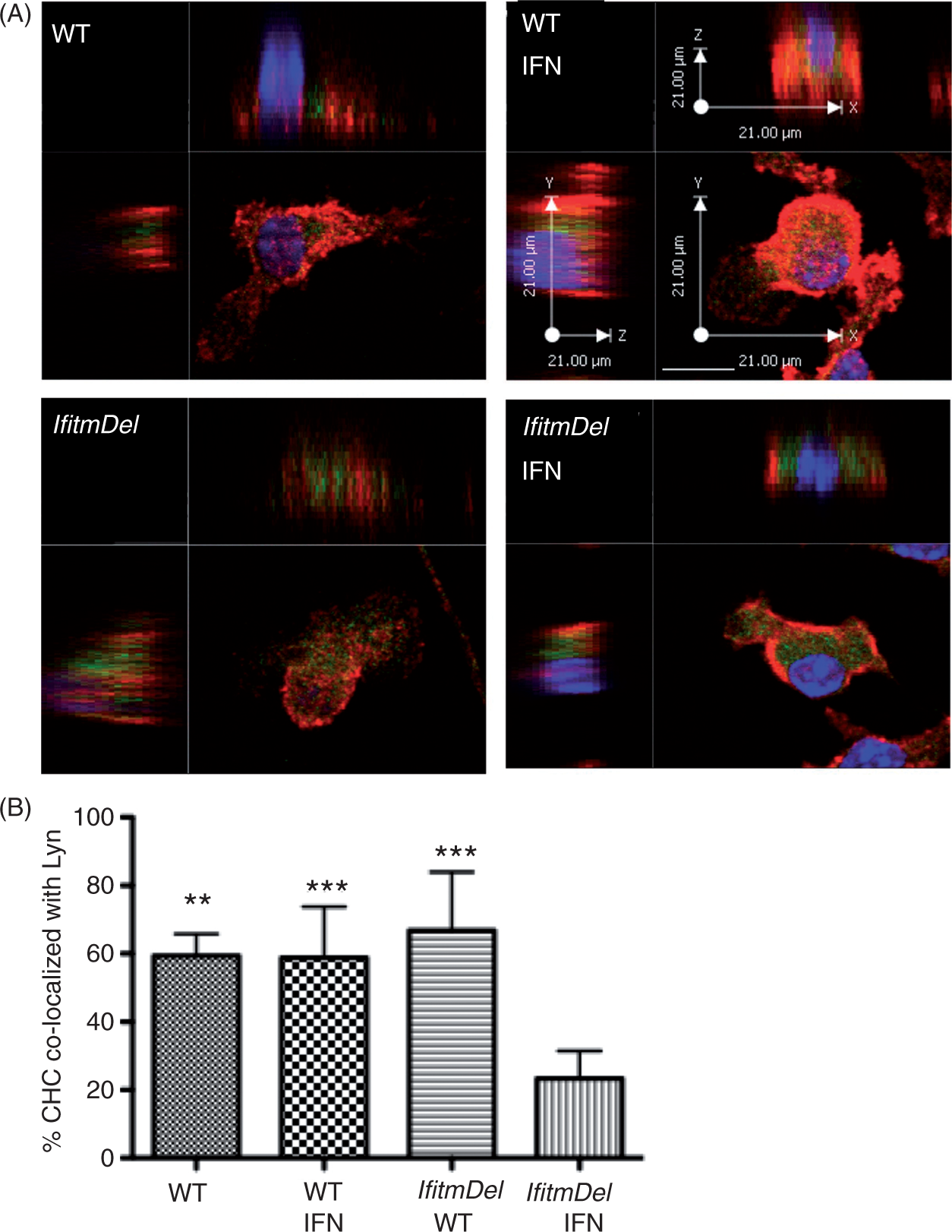
The data in the preceding figure were obtained from the analysis of BMDM yet the role of clathrin in endocytosis is critical for all cell types. The analysis of MEF cells derived from the IfitmDel animals also showed a selective loss of clathrin at the cell membrane following IFN-β treatment (Supplementary Figure 5). As Ifitm3 is the most highly IFN-β-induced gene of the Ifitm gene family, we chose to attempt to rescue the WT localization of clathrin in the IfitmDel/IFNβ-treated MEFs by introducing a vector expressing both GFP and Ifitm3. The construct of the Ifitm3-expressing retrovirus and demonstration of it encoding both GFP and Ifitm3 is shown in Supplementary Figure 6. As shown in Figure 3, IfitmDel MEF cells were infected with either an empty virus vector expressing GFP (top four panels) or a retrovirus vector co-expressing Ifitm3 and GFP (bottom four panels). Such cells were treated with IFN-β for 18 h and analyzed by confocal imaging. Green arrows indicate cells that were not infected and pink arrows indicate infected cells. The introduction of the Ifitm3 protein in these cells clearly re-constituted the appropriate co-localization of clathrin and Lyn (indicative of the yellow overlap), whilst the vector-alone samples (or the uninfected cells) still demonstrated the mutant phenotype. Control staining (using non-specific secondary Abs) of the IfitmDel MEF cells infected with the empty vector control or with the Ifitm-expressing retrovirus is shown in Supplementary Figure 7. In total, the data indicate that Ifitm3 is critical for the appropriate localization of clathrin in IFN-β-treated cells.
Reconstitution of Ifitm3 restores appropriate clathrin localization. IfitmDel MEFs were infected with empty vector retrovirus (EV) expressing eGFP (cyan) or with the same retrovirus also expressing Ifitm3. After IFN-β treatment for 18 h, cells were stained for Lyn (pseudo red) and clathrin heavy chain (CHC) (green). Pink arrows show infected cells and green arrows indicate uninfected cells. Scale bars = 10 µm.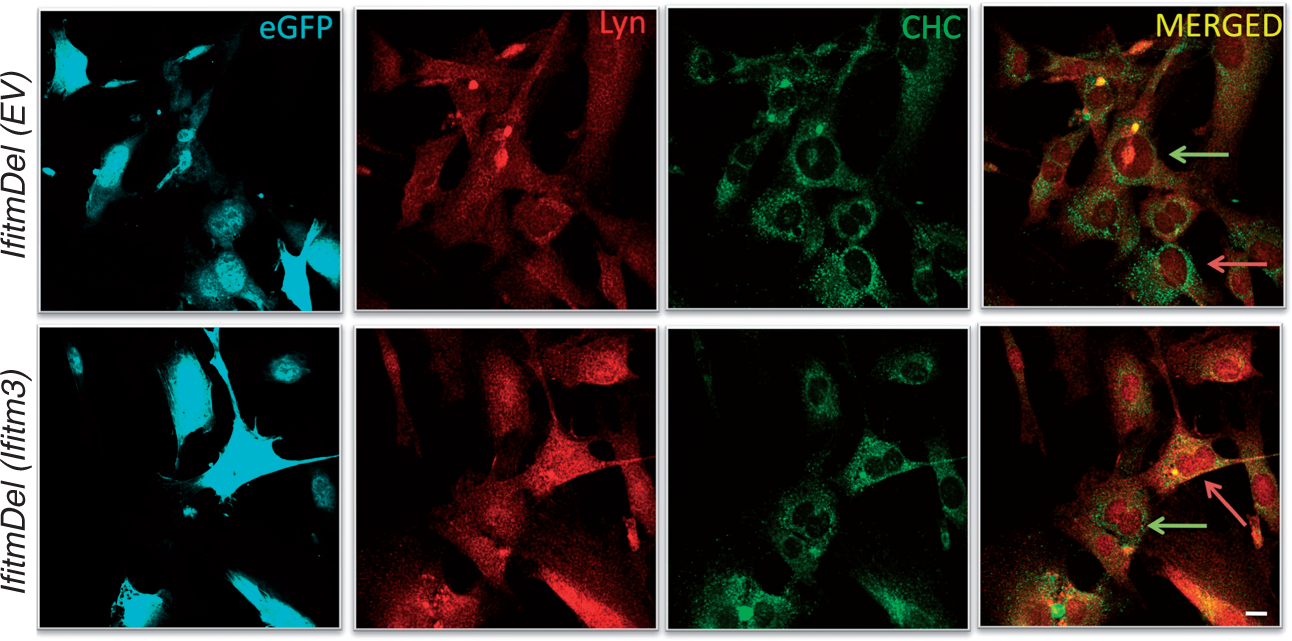
Cells lacking the Ifitm proteins demonstrate depressed clathrin-dependent endocytosis but enhanced macropinocytosis
The transferrin and EGF receptors utilize the clathrin pathway for the internalization of ligand whilst the entry of dextran into the cells is clathrin-independent and primarily accomplished by macropinocytosis. Based upon the clathrin localization data described above, we decided to test the efficiency of IFN-β-activated IfitmDel cells for clathrin-dependent and clathrin-independent uptake. To quantify the internalization of transferrin, BMDM from WT and IfitmDel animals were grown in culture, treated with IFN-β for 18 h, incubated with labeled transferrin for 1 h and analyzed for ligand uptake. As shown in Figure 4A, the activation of WT cells with IFN-β greatly increased their phagocytic potential for transferrin relative to untreated WT BMDM. The IfitmDel BMDM treated with IFN-β, however, showed little uptake. Transferrin uptake by MEFs derived from the IfitmDel animal is similarly inhibited as the IfitmDel BMDM (Supplementary Figure 8).
Transferrin and EGF uptake is inhibited in IFN-β treated IfitmDel cells while macropinocytosis of dextran is elevated. (A) Endocytosis of labeled transferrin by BMDM derived from WT and IfitmDel mice plus or minus IFN-β activation (18 h). Quantification of transferrin endocytosis was calculated as uptake in each cell sample divided by the total transferrin uptake of the four experimental groups in a single individual experiment. Averages are from five independent experiments. (B) Endocytosis of low dose EGF (1.5 ng/ml) by BMDM derived from WT and IfitmDel mice plus or minus IFN-β activation (18 h). Quantification of EGF endocytosis was calculated as uptake in each cell sample divided by the total EGF uptake of the four experimental groups in a single individual experiment. Averages are from five independent experiments. (C) Macropinocytosis of dextran by BMDM derived from WT and IfitmDel mice plus or minus IFN-β activation (18 h). Quantification of dextran endocytosis was calculated as uptake in each cell sample divided by the total dextran uptake of the four experimental groups in a single individual experiment. Averages are from five independent experiments. Error bars indicate SEM. Statistical significance was achieved (*P ≤ 0.05 or **P ≤ 0.005) of the samples indicated with the bar.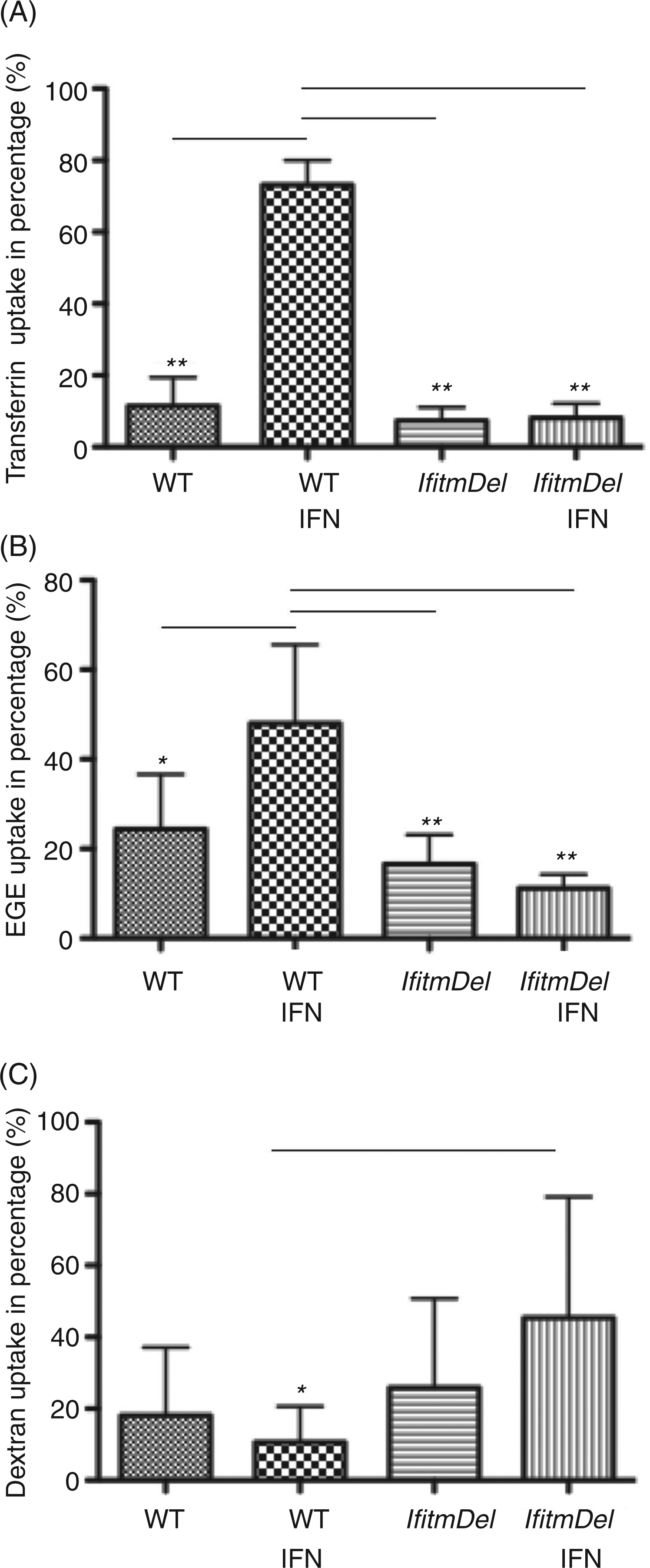
To determine if this effect was unique to the transferrin receptor, we also analyzed the ligand-dependent uptake of the EGF receptor, which also utilizes clathrin-coated pits. Accordingly, BMDM from WT and IfitmDel animals were treated with IFN-β and tested for their uptake of labeled EGF. WT cells treated with INF-β demonstrated an elevated uptake of EGF that was lost in the cells from the IfitmDel animals (Figure 4B). These experiments were done with low levels of EGF binding to the receptor which preferentially leads to receptor internalization via the clathrin pathway. Using high levels of the ligand, however, the receptor can enter the cell in a clathrin-independent fashion. 30 When the WT and the IfitmDel cells were treated with IFN-β and then incubated with high levels of EGF (100-fold greater than levels used in Figure 4B) there was no difference in uptake between these treated and untreated cells (data not shown).
Macropinocytosis of fluid phase markers is a clathrin-independent means of uptake. 31 We chose to assay the rate of uptake of dextran by macropinocytosis in the IfitmDel cells plus or minus IFN-β activation. As shown in Figure 4C, the ability of the IfitmDel cells treated with IFN-β to ingest dextran by macropinocytosis was significantly greater than untreated IfitmDel cells, or IFN-β-treated or IFN-β-naive WT cells. Thus, the absence of clathrin in the membrane of the IFN-β-treated IfitmDel cells does not impair macropinocytosis by such cells.
The Ifitm proteins directly bind to the v-ATPase required for appropriate endosome acidification
The Ifitm proteins were defined previously as binding partners to tetraspanin proteins such as CD81 and CD9. 19 The tetraspanin proteins are critical in the formation of membrane activation complexes (forming what is referred to as the tetraspanin web), allowing for appropriate signal transduction from receptor signaling. 32 In two unpublished yeast hybrid studies we found that Ifitm proteins can bind to known plasma membrane-bound tetraspanin proteins such as CD81 and CD9, as well as those found associated with internal membranes such as lysosomal-associated transmembrane protein 4B (Laptm4b) which is preferentially found associated with lysosomes and the multi-membrane spanning Atp6v0b subunit of the v-ATPase (data not shown).
The primary determinant of pH regulation in compartments of the endocytic compartment is the vacuolar ATPase (v-ATPase).33–35 The v-ATPase is a multi-subunit protein complex comprising a number of proteins, including multiple membrane bound V0 subunits.
36
Two of these ATP6V0 subunits (B and E2) were implicated in providing resistance to virus infection following IFN-β treatment.
20
The mouse Atp6v0b protein is 21 kDa in size and is predicted to span the membrane with five transmembrane domains. Owing to its structure and localization in the membrane, it is possible that the Ifitm proteins are required for the appropriate organization and/or stabilization of the ATPase in the lysosomal membrane. To investigate the potential of altered pH of lysosomes in the IfitmDel cells and to potentially link a change in pH to possible Ifitm functions, we utilized Lysotracker Red DND 99 to label acidic vacuoles within cells. DND-99 is a red-fluorescent dye linked to a weak base that can freely permeate cells but is retained and concentrated in acidic organelles. MEFs from WT and IfitmDel animals were treated (plus or minus) with IFN-β for 18 h and incubated with the dye for 60 min. Live cells were then fixed and analyzed for retention and concentration of the dye. As shown in Figure 5A, the intensity of staining of the WT cells treated with IFN-β treatment was higher than the staining of similarly activated IfitmDel cells. The staining of such cells was quantified in Figure 5B and demonstrated significant loss of acidic pH-dependent retention of the DND-99 dye in the IFN-β-treated IfitmDel cell samples, confirming that the Ifitm proteins are required for this response.
Ifitm proteins are required for endosomal acidification and directly interact with the v-ATPase. (A) MEFs cells derived from WT and IfitmDel animals were stained with Lysotracker Red DND-99 in the presence or absence of IFN-β for 18 h. Scale bars = 10 µm. (B) Quantification of Lysotracker Red DND-99 staining in percentages was calculated as the Lysotracker Red DND-99 staining in each group divided by the total Lysotracker Red DND-99 staining of the four experimental groups in an individual experiment. Error bars indicate SEM. *P ≤ 0.05 or **P ≤ 0.005 compared with the WT IFN sample. These data are representative of three identical experiments. (C) J774 macrophages were lysed with either RIPA (R) or NP-40 (N). Rabbit anti-Ifitm3 (intracellular domain: int24)
19
or rabbit polyclonal control was used to precipitate protein complexes that were then resolved by SDS-PAGE and immunoblotted with anti-ATP6v0b (15,000 mr). Results from a single experiment are shown but are representative of three similar analyses.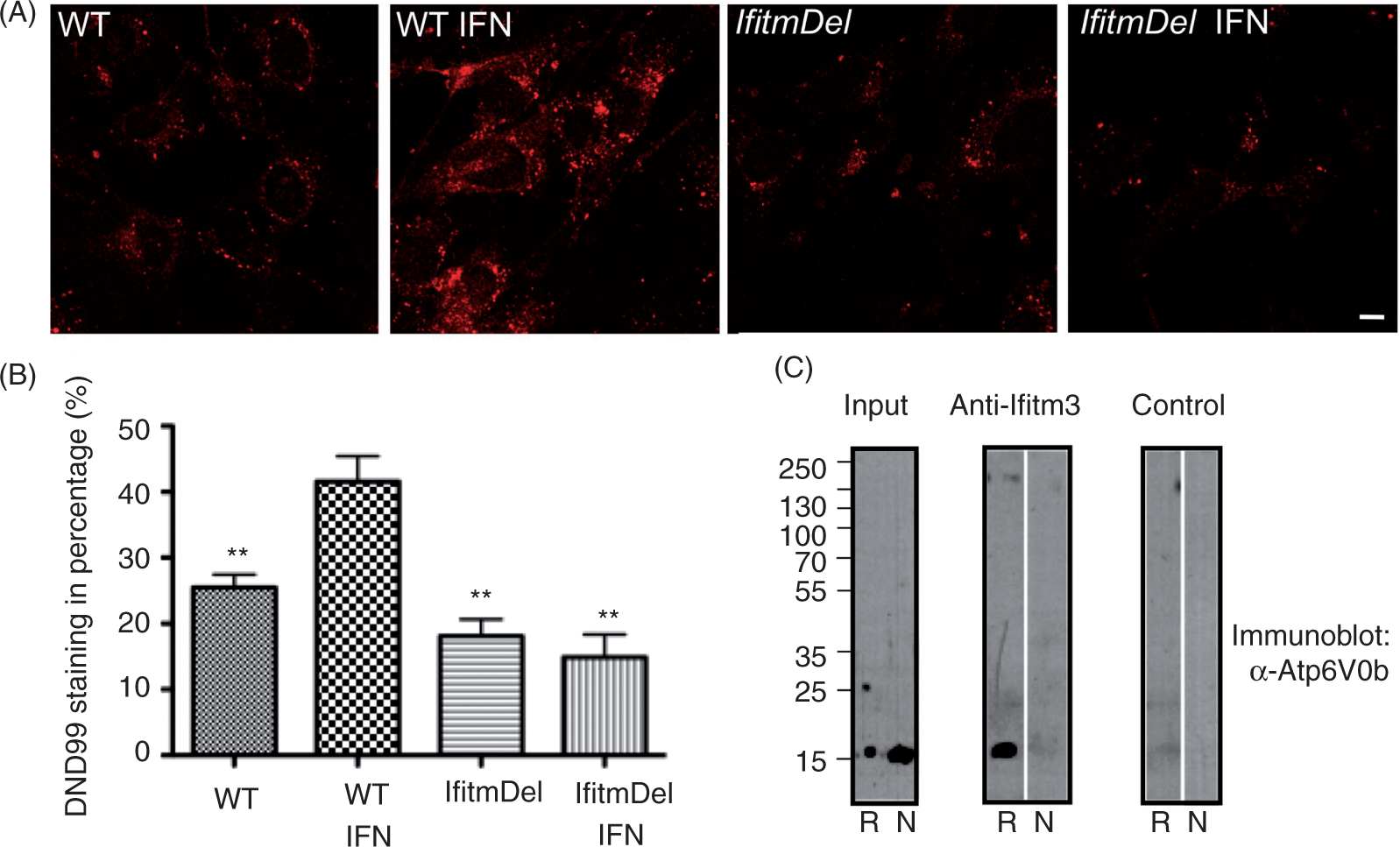
To determine if the Ifitm proteins are, indeed, associated with the Atp6v0b v-ATPase subunit, J774 murine macrophages (that express both Ifitm3 and Atp6v0b) were lysed in the presence of either RIPA (0.1% SDS, 1% DOC, 1% NP40) or NP-40 alone (0.5%), and precipitated with the anti-Ifitm3 Ab raised against the conserved cytoplasmic domain sequence.
19
The immunoprecipitates were resolved by SDS-PAGE and immunoblotted with an anti-Atp6v0b polyclonal Ab. As shown in Figure 5C, the co-association of Ifitm3 with Atp6v0b was evident in the RIPA but not NP-40 lysates, suggesting that the higher detergent stringency is required to expose the Ifitm epitope for Ab recognition or to solubilize the lipid compartment in which this complex is found. Interestingly, an Ab specific for the N-terminal region of the Ifitm3 protein
19
did not co-precipitate the Atp6v0b protein in either RIPA or NP-40 lysates (data not shown) suggesting that the N-terminus of the protein is masked when these two protein co-associate. The data in this figure indicate that Ifitm3 and the Atp6v0b subunit of the v-ATPases have direct physical contact and that the absence of Ifitm3 alters the function of the v-ATPases in regulating the pH of phagocytic endosomes (Figure 6).
Model of the Ifitm3, v-ATPase and clathrin complex. (A) The multi-subunit v-ATPase is shown in a WT cell localized within a lipid raft microdomain in association with Ifitm3. This complex is linked to clathrin heavy chain trimers via the AP-2 adaptor protein; appropriate acidification of the phagocytic endosome occurs following IFN-β induction. (B) The lack of Ifitm proteins in cells from the IfitmDel animal results in a functional disruption of the activity of the v-ATPase following IFN-β induction. This disruption interrupts the clathrin/v-APTase interaction leading to decreased phagocytic activity, a neutral pH within the phagocytic endosomes and the cytoplasmic localization of untethered clathrin.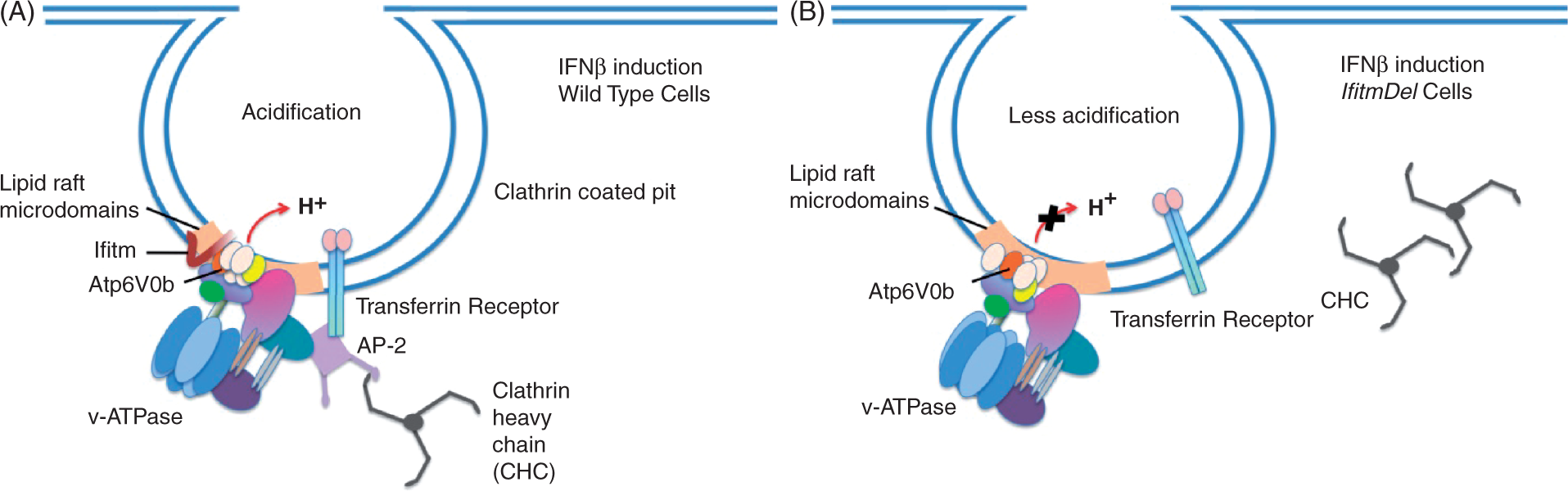
Discussion
The mammalian Ifitm gene family consists of multiple genes encoding closely related proteins. Members of the family were first described as demonstrating induced expression after treatment with type I and/or type II IFNs,4,5 although not all members of the family are responsive to IFN treatment (Supplementary Figure 1). 19 Therefore, members of this family may have roles in normal cell signaling (such as B cell receptor signaling) independent of those utilized in restricting viral replication. The Ifitm proteins are structurally simple, consisting of only two highly conserved domains: a TM-like domain and a conserved cytoplasmic domain. The Ifitm proteins are found associated with intracellular endosomal compartments, as well as the plasma membrane. Cellular activation can promote the movement of the Ifitm proteins from their intracellular location to the surface membrane where they are known to associate with several members of the tetraspanin family.14,17,20 The localization of the Ifitm proteins to lipid raft signaling domains of the plasma membrane of cells following activation is likely a result of their association with the tetraspanins, although both the Ifitm proteins and tetraspanins obtain palmitoylation modifications following cell activation which can dictate the movement of such proteins into raft-like structures of cells.37,38
The function(s) of constitutively expressed, as well as IFN-induced Ifitm proteins is not clear. A number of investigators have demonstrated their role in blocking virus infections following IFN treatment. siRNA screens, as well as the analysis of cells derived from the IfitmDel mouse strain, demonstrated that the Ifitm proteins help limit the release of viruses from the endosomal compartments into the cytoplasm and, accordingly, suppress infection.21,23,24 A study utilizing SARS coronavirus pseudotypes concluded that Ifitm-mediated restriction occurred at a late stage of the endocytic pathway and did not block the entry of the virions from entry into acidic cellular compartments. 22 Thus, viruses that require late endosomal processing for infection are sensitive to the effects of the Ifitm proteins whilst those that enter the cell directly from the plasma membrane are not. However, the specific function of the Ifitm proteins in controlling this process is unknown.
A number of groups have performed genome-wide siRNA silencing experiments to define the host factors that limit viral infections.20,39,40 One group of proteins that are consistently identified in these assays are components of the v-ATPase complex, required for the acidification of endocytic compartments. 33 Members of this family of proteins have also induced expression following IFN treatment. 20 Reducing the effectiveness of this complex provides the same phenotype as reducing the expression of the Ifitm proteins. As shown in Figure 5 (and by others 22 ), the pH of the endosomes of cells derived from the IfitmDel mice is higher than that of WT cells, with or without IFN induction, indicating that a more neutral endosomal pH increases the cell’s permissiveness to virus infection. Thus, these data suggest that the v-ATPase is functionally compromised in the absence of the Ifitm proteins leading to an inability to appropriately acidify the endosomal compartments. The observation that Ifitm3 forms a co-precipitating complex with the multi-spanning v-ATPase component Atp6v0b (Figure 6) implies that these two proteins interact within the cell to form a functional v-ATPase complex.
There are other examples of how the Ifitm proteins and the v-ATPase may interact. One of the Ifitm proteins, Ifitm5, is expressed primarily in bone tissues. Animals lacking Ifitm5 possess smaller bones but maintain equivalent bone mass and normal levels of osteocalcin, suggesting osteoblast maturation is normal. 41 Osteoclasts, however, utilize a membrane-bound v-ATPase to acidify the extracellular space to help remodel forming bone. 42 The lack of Ifitm5 may compromise osteoclast function by inhibiting the function of the v-ATPase, thus impairing bone formation. Similarly, v-ATPases are linked to the acidity of tumors; the extracellular release of protons by the tumor cells is implicated in tissue damage and metastasis of the cells. 43 The expression of the Ifitm protein varies widely in tumor cells. Induction of the rat Ifitm3 protein has been shown to promote tumor formation and differentiation in a rat breast cancer model. 15 Thus, the elevated expression of the Ifitm proteins may promote the expansion and subsequent metastasis of tumor cells. Additionally, the activation of B cells via the B cell antigen receptor (BCR) leads to the transcriptional up-regulation of multi-spanning components of the v-ATPase. 44 We have demonstrated the translocation of the Ifitm3 into the lipid raft constituents of the B cell membranes following BCR activation; 19 these proteins have been previously documented as constituents of the CD21-CD19-CD81 co-receptor complex. 12 The translocation of the Ifitm proteins into defined membrane compartments following BCR activation may lead to enhanced endosomal acidification required for the appropriate processing of the internalized antigen.
IFN-β treatment of WT cells resulted in the enhanced phagocytosis of clathrin-dependent ligands (Figure 4). 45 This elevated uptake was lost in the IFN-β-activated IfitmDel cells, although macropinocytosis was enhanced. Resting WT and IfitmDel cells possess membrane bound clathrin but, following IFN-β activation, clathrin is lost from the membrane in the IfitmDel cells and found in the cytoplasm of the cell (Figures 1 and 2). The lack of clathrin at the membrane and within the lipid rafts of the IfitmDel IFN-β-treated cells was not, however, because of generalized destruction of membrane structure as the Src kinase family member Lyn remained in the membrane in such low-density, cholesterol-rich compartments. These data suggest that IFN-β activation alters membrane constituents and requires the presence of the Ifitm proteins to maintain appropriate membrane organization. Clathrin and the v-ATPases are linked together in endosomal and plasma membrane sites via the clathrin assembly protein AP-2 (Figure 6). 46 The AP-2 adaptor protein directly binds to receptor cargo proteins such as the transferrin receptor (Tfr), as well as clathrin subunits and the v-ATPase.47,48 Thus, if the localization of the v-ATPase is destabilized via the loss of the Ifitm proteins then the stability of the clathrin cage required for phagocytosis and endosome routing could also be compromised.
The model of Ifitm function proposed in this article may appear paradoxical in that the virus-resistant IFN-β-treated WT cells maintained clathrin within the membrane and showed enhanced clathrin-mediated endocytosis whilst the virus-sensitive cells lacking the Ifitm proteins did not. Indeed, the opposite findings might have been predicted. The redistribution of clathrin during cell activation has been reported previously. EGF binding to its receptor triggers the phosphorylation of CHC, which results in its redistribution at the cell membrane. 49 Additionally, the creation of non-productive complexes of clathrin and adaptor proteins can be influenced by the phosphorylation of these adaptor proteins in the cytosol. 50 We have not, however, found any report in the literature of CHC sequestration in the cytoplasm similar to that described in this report. Instead, the anti-viral effect of the Ifitm proteins may be to maintain clathrin within the membrane of the IFN-β-activated cell by through stabilization of the IFN-induced, newly synthesized v-ATPases. Defining the specific interactions of the Atp6v0b and Ifitm proteins in resting and in IFN-activated cells will help us to define the molecular basis of v-ATPase and clathrin functions in the anti-viral response.
Footnotes
Funding
This work was supported by grants from the National Institute of Allergy and Infectious Diseases (AI-24158, AI-088451, JHW: AI-32223, JJW).
Acknowledgements
We thank Drs M. Azim Surani, Louise van der Weyden and David J. Adams for generously providing us with the IfitmDel animal. We also thank Dr Chris Rodesch of the University of Utah imaging core facility for his help with the confocal analysis and Dr Diane Ward for her assistance in the transferrin and dextran uptake experiments. As always, we are indebted to the members of the Weis’ laboratories for their critique of the research and many valuable contributions. The content is solely the responsibility of the authors and does not necessarily represent the official views of the Institute of Allergy and Infectious Diseases or the National Institutes of Health.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
