Abstract
We report the supramolecular self-assembly of cucurbit[8]uril(Q[8]) and 1-(4-methoxyphenyl)piperazine inclusion complex. The complex is cultured in 3 M hydrochloric acid solution under CdCl2 induction conditions to obtain a new crystal, the structure of which was characterized by X-ray single-crystal diffraction. The results show that the crystal belongs to the monoclinic crystal system with a C2/m space group. The host–guest interaction ratio of Q[8] and 1-(4-methoxyphenyl) piperazine is 1:2.
Keywords
We report the supramolecular self-assembly of an cucurbit[8]uril(Q[8]) and 1-(4-methoxyphenyl)piperazine, and it was cultured in 3 M hydrochloric acid solution and CdCl2 induction conditions to obtain a new crystal, whose structure was characterized using X-ray single-crystal diffraction. The results show that the crystal belongs to the monoclinic crystal system with a C2/m space group and a = 25.663(4), b = 38.692(7), c = 21.286(4), α = 90°, β = 120.818(4)°, γ = 90°, and Z = 2. Obtain the host–guest interaction ratio of Q[8] and 1-(4-methoxyphenyl) piperazine is 1:2.

Introduction
Supramolecular self-assembly has attracted much attention because it covers various fields, such as chemistry, materials, and life sciences.1–5 To construct inorganic, organic, or supramolecular frameworks, the key is to select the appropriate building blocks to generate periodic and porous frameworks through coordination, covalent, and supramolecular interactions. These frameworks have different functions and applications, such as adsorption, release, sensing, and catalysis.6–14 Cucurbit[n]urils (Q[n]s or CB[n]s)15–22 (Figure 1) are noved macrocyclic compounds that were discovered after cyclodextrins, crown ethers, and calixarenes. They have an electrically neutral hydrophobic cavity, two negative potential carbonyl portals, and an outer surface with a positive potential. Because of this unique structure, three major research directions are based on cucurbit[n]uril host–guest chemistry,23–26 coordination chemistry,27,28 and non-covalent supramolecular frameworks.29,30

The structure of Q[5–8], Q[10], and Q[13–15].
For our study, we selected 1-(4-methoxyphenyl)piperazine, an antihypertensive intermediate of phenylpiperazine, as the guest, while cucurbit[8]uril(Q[8]), which has a large hydrophobic cavity and two polar ports, was chosen as the host (Figure 2). The supramolecular self-assemblies formed between the guest molecule and Q[8] have been studied by means of single-crystal X-ray diffraction analysis. The results show that the self-assemblies formed by the guests with Q[8] are exclusion complexes, involving interactions with the electronegative carbonyl groups around the portal of Q[n], as well as host–guest interactions with Q[n] and anionic [Cd6Cl19]7− to form a multi-dimensional, multi-level, self-assembled supramolecular framework.
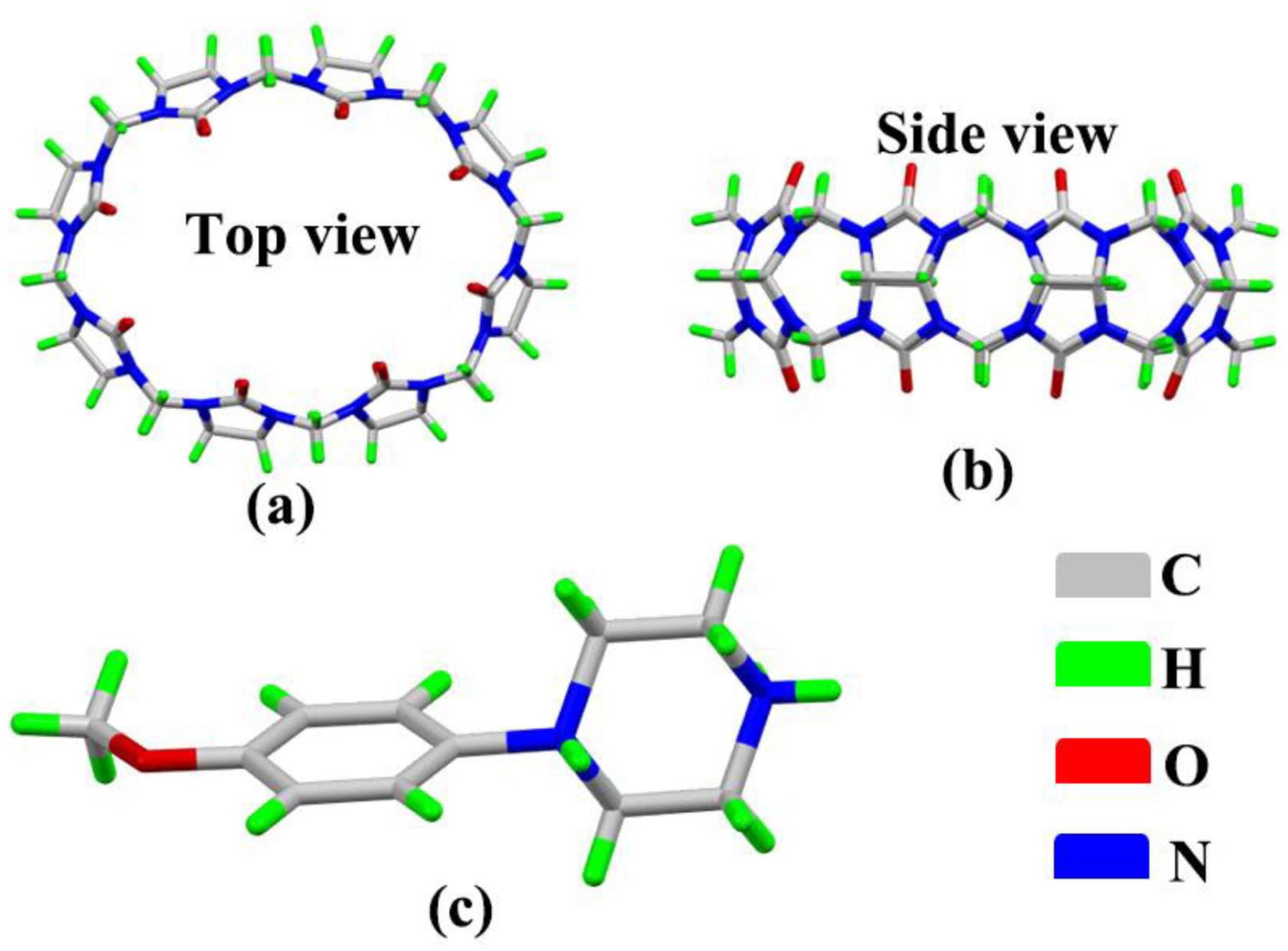
Structure of Q[8] (top view (a) and side view (b)) and 1-(4-methoxyphenyl)piperazine (c).
Results and discussion
Description of the crystal structures of complex
Figure 3(a) shows the host–guest interaction ratio of cucurbit[8]uril (Q[8]) and 1-(4-methoxyphenyl)piperazine, which contains one Q[8] and two 1-(4-methoxyphenyl)piperazine. Two 1-(4-methoxyphenyl)piperazine molecules enter the cavity via the two ports of Q[8], respectively, while the benzene ring of the 1-(4-methoxyphenyl) piperazine molecule entering the cavity of cucurbit[n]uril. As shown in Figure 3(b), there are two free water molecules at each port of the cucurbit[n]uril. The water molecule interacts with the cucurbit[n]uril and the 1-(4-methoxyphenyl)piperazine molecule through hydrogen bonds. [Cd6Cl19]7− participates in an ionic dipole interaction (C–H···Cl interaction) with the cucurbit[n]uril. It also serves as a bridge between cucurbit[n]urils.
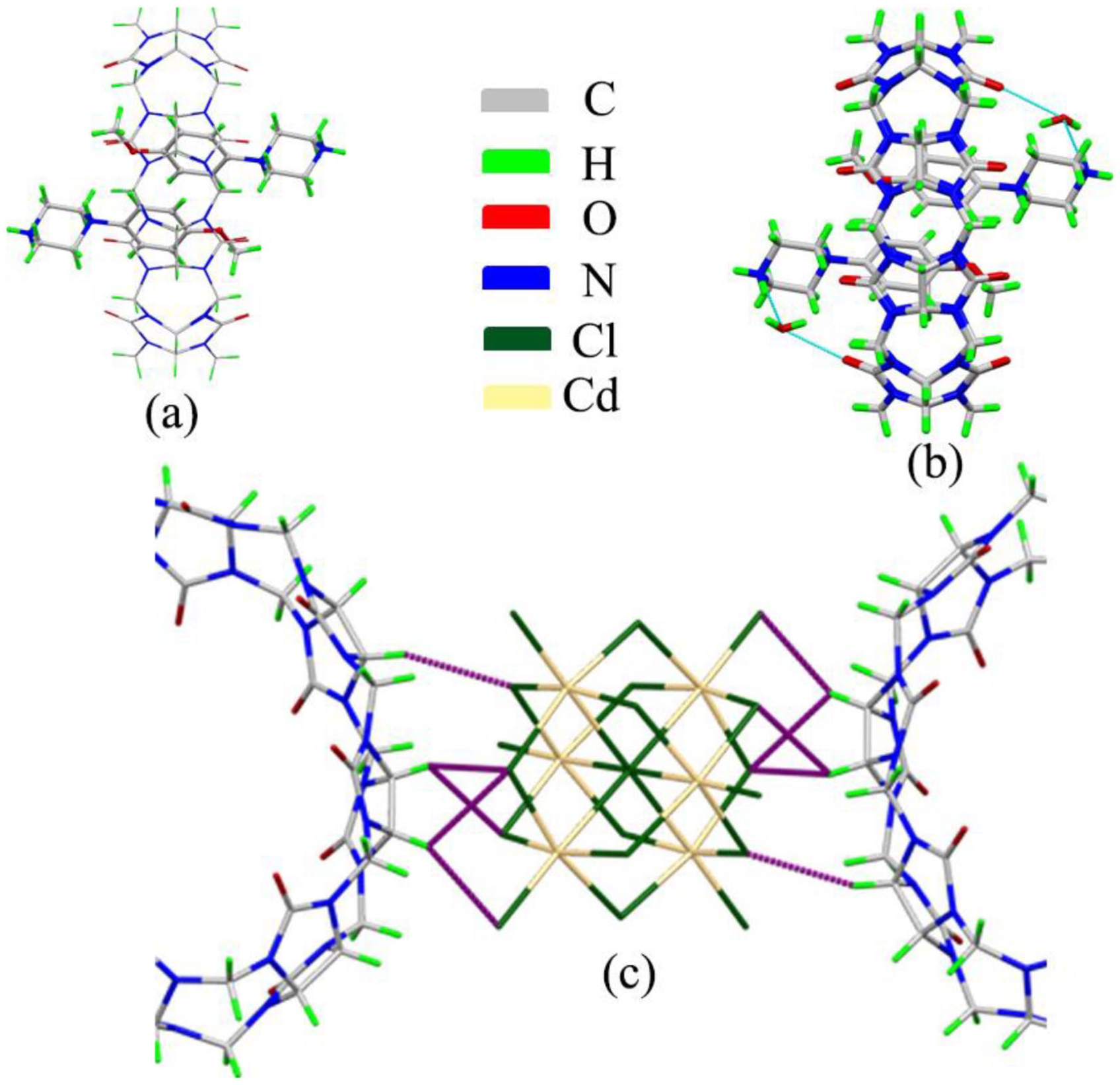
Crystal structure of complex (Q[8] and 1-(4-methoxyphenyl)piperazine): (a) host–guest interaction, (b) hydrogen bond interaction, and (c) ion–dipole interaction (C–H···Cl interaction).
Figure 4(a) shows the stacked diagram along the a-axis view direction and Figure 4(b) shows the stacked diagram along the c-axis view direction. We can clearly see that there are six cucurbit[n]uril molecules around each [Cd6Cl19]7− unit. Interestingly, there are four Q[8] units that interact with the 1-(4-methoxyphenyl)piperazine molecule, and the interaction ratio is 1:2. However, the other two Q[8] molecules do not interact with the guest molecule. Whether along the a-axis view direction and along the c-axis view direction, we find that the stacked diagram structure is highly symmetrical. Figure 5(b) shows the stacked diagram along the c-axis view direction. The [Cd6Cl19]7− anion enters the holes formed between Q[8]. Q[8]@1-(4-methoxyphenyl)piperazine and anionic [Cd6Cl19]7− form a multi-dimensional, multi-level, self-assembled supramolecular framework.

Stacked diagram with one [Cd6Cl19]7−: (a) along the a-axis view direction and (b) along the c-axis view direction.

Stacked diagram along the c-axis view direction.
Conclusion
The supramolecular self-assemblies formed by 1-(4-methoxyphenyl)piperazine molecules with Q[8] have been studied by means of single-crystal X-ray diffraction analysis. The results show that the self-assemblies formed by 1-(4-methoxyphenyl)piperazine with Q[8] are exclusion complexes and involve their interaction with the electronegative carbonyl groups around the portal of Q[8]. There are six cucurbit[n]uril molecules around each [Cd6Cl19]7− anion. Of note is the fact that there are four Q[8] units that interact with the 1-(4-methoxyphenyl)piperazine molecule, and the interaction ratio is 1:2. However, the other two Q[8] molecules do not interact with the guest molecule. As well as host–guest interactions, Q[8] and anionic [Cd6Cl19]7− form a multi-dimensional, multi-level, self-assembled supramolecular framework.
Experimental
General materials
All materials were reagent grade and were used without any further purification. Q[8] (purity ⩾97%) was prepared in the Key Laboratory of Macrocyclic and Supramolecular Chemistry of Guizhou Province, China.
Preparation and testing of the crystalline complex
Q[8] (10 mg, 7.52 μmol) and 1-(4-methoxyphenyl)piperazine (5 mg, 25.74 μmol) were dissolved in a 10-mL round-bottom flask containing HCl (5 mL, 3 mol/L), CdCl2 (5 mg, 27.26 μmol) was added, and the mixture was shaken for 3 min at room temperature and then heated to reflux for 5 min with stirring. The resulting mixture was cooled and left to stand for about 1 week to afford colorless crystals. A regular and transparent complex crystal was selected and fixed on a glass filament using petroleum jelly. X-ray single-crystal diffraction analysis was carried out on a Bruker D8 Venture X-ray single-crystal diffractometer in ω-scan mode using graphite to monochromatize the Mo-Kα rays (λ = 0.71073 Å, μ = 0.828 mm−1). The crystal data were collected, and the Lorentz polarization and absorption corrections were carried out. The SHELXT-14 and SHELXL-14 program packages were used for structural analysis and full-matrix least squares refinement. All non-hydrogen atoms were anisotropically refined using the analytical expression of the neutral atom scattering factor and combined with anomalous dispersion correction. The SQUEEZE program in the PLATON package was used to delete some of the disordered solvent molecules. The X-ray crystallographic data for the structure reported in this study have been deposited at the Cambridge Crystallographic Data Centre under accession number CCDC: 2101349. These data can be obtained free of charge via http://www.ccdc.cam.ac.uk/data request/cif. The crystal data and structure modification parameters are shown in Table 1.
The crystallographic parameters of the complex.
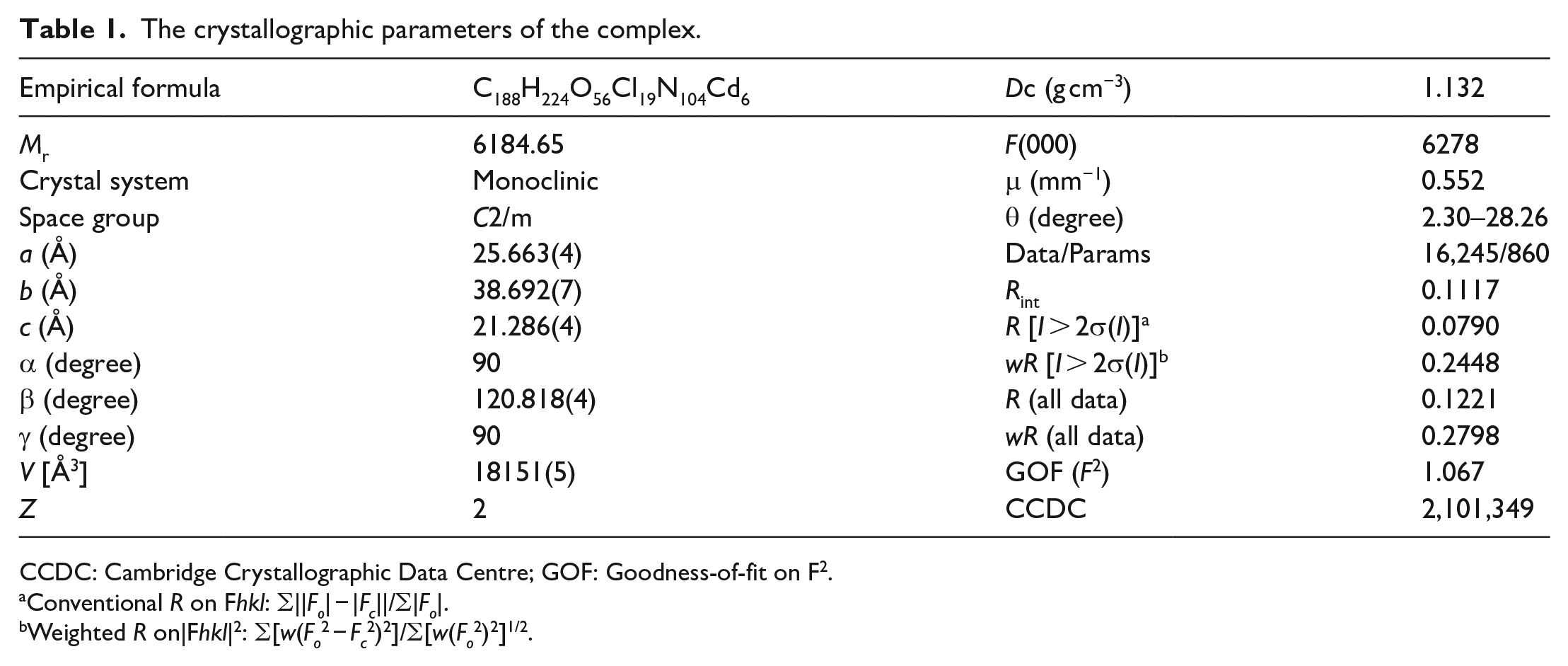
CCDC: Cambridge Crystallographic Data Centre; GOF: Goodness-of-fit on F2.
Conventional R on Fhkl: ∑||Fo| − |Fc||/∑|Fo|.
Weighted R on|Fhkl|2: ∑[w(Fo2 − Fc2)2]/∑[w(Fo2)2]1/2.
Footnotes
Author contributions
All authors contributed to the study, conception, and design. Material preparation, data collection, and analysis were performed by J.Z., L.Z., Y.C., Y.M., X.D., L.S., Y.L., X.Y., and P.M. The first draft of the manuscript was written by J.Z., and all authors commented on previous versions of the manuscript. All authors have read and approved the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (grant no. 22161010).
