Abstract
Phosphodiesterase
Keywords

Introduction
It is well known that cAMP and cGMP, as important secondary messengers, affect the physiological activities of the human body, and, importantly, their imbalance in cell concentration in the body may lead to a variety of diseases.1–3 Previous studies have shown that the activity of phosphodiesterases (PDEs) directly affects the expression concentration of cAMP and cGMP in vivo. 4 PDEs are a large family of polygenes, including 11 different subtype structures, which have different substrate specificities, enzyme kinetics, and cell distribution. They also have similar cell structures, show their own homology, and determine the specificity of the substrate or inhibitor.5–7 Among them, PDE2 is a dual-substrate enzyme that can hydrolyze cAMP and cGMP. Although cAMP and cGMP are expressed in the brain, heart, liver, platelets, and T cells, the expression level is the highest in the brain regions,8–10 which indicates that PED2 plays an important role in the emotional, cognitive, and other behavior of humans.7,10,11 Therefore, PDE2 inhibitors have the potential to treat a variety of diseases, such as Alzheimer’s disease, angina pectoris, vasodilatation, heart failure, hypertension, and arrhythmia.12–22 However, due to poor pharmacological activity, low blood–brain barrier permeability, and poor metabolic stability, no PDE2 inhibitor has been approved for clinical use. Therefore, the development of novel PDE2 inhibitors is an important research goal.
BAY 60-7550 (Figure 1) is an imidazole–triazine compound with the high selectivity and activity toward PDE2. Moreover, it is often used as an important standard in PDE2 bioactivity tests to investigate other PDE2 inhibitors.23,24 Considering the structural–activity relationship between BAY60-7550 and PDE2 protein (PDB ID: 4HTX, http://www.rcsb.org), a series of phthalimide and phenylpyrazolo[1,5-a]pyrimidine PDE2 inhibitors have been designed and prepared. Furthermore, the IC50 values were determined by the biological activity detection method. The structure–activity relationship between the optimum product and 4HTX was simulated by the Discovery Studio software to illustrate the possible binding modes and for the further development of effective PDE2 inhibitors.

The structure of BAY 60-7550.
Results and discussion
Synthetic pathways
The synthetic strategy toward phthalimide PDE2 inhibitors

The synthetic pathway toward phthalimide and phenylpyrazolo[1,5-a]pyrimidine PDE2 inhibitors
Bioassays
Protein expression and purification
In order to express the catalytic domain (residues 580–941), the recombinant pET15b-PDE2A plasmid was subcloned and purified by the previously reported method.
26
Subsequently, the plasmid was transferred into Escherichia coli strain BL 21 (CodonPlus) to grow in 2XYT medium at 37 °C until the absorbance (A) at 600 nm was 0.6–0.8. Next, 0.1 mmol L−1 of isopropyl-β-
Enzymatic assays
The enzymatic activities of 9 compounds were assayed using Bio-cAMP as the substrate.1,26 Briefly, a reaction mixture of 20 mmol L−1 of Tris HCl, 10 mmol L−1 of MgCl2, 0.5 mmol L−1 of DTT, 3H-cAMP or 3H-cGMP (20000-40000 cpm 3H-cAMP or 3H-cGMP per assay ). and proteins were incubated at 25 °C for 15 min. The reaction was then terminated by the addition of ZnSO4 and precipitated out by Ba(OH)2. The compound was dissolved in with DMSO, and the possible systematic errors of the method were evaluated through the different contents of DMSO. Finally, BAY60-7550, a typical PDE2 positive inhibitor, was selected to determine its activity and to calculate the accuracy of the IC50 verification method. The hydrolysis rate of H-cAMP or H-cGMP expressed the inhibitory activity of the compound. Seven concentrations of inhibitors were used for measuring the IC50 values. Each measurement was repeated three times, and the IC50 values were calculated by nonlinear regression. BAY 60-7550 was used as the reference compound for PDE2 enzyme activity detection.
Analysis of biological activity results
The AlphaScreen kit method was used to detect the activity of the nine synthesized compounds, which were diluted at seven concentrations, and three sets of parallel experiments were performed. The IC50 values were obtained by fitting calculations using the software GraphPad Prism 7. The results are given in Table 1.
BAY60-7550 was used as the reference compound with an IC50 of 8.4 nM.
IC50 values determined against 3H-cAMP.
First, according to the IC50 values of phthalimide compounds
Molecular docking
Discovery Studio software is a comprehensive molecular simulation platform. The working principle is based geometric complementarity, chemical environment complementarity, and energy complementarity. If there is an interaction between the ligand and the receptor, then molecular docking can be evaluated and we can obtain the structure–activity relationship and scoring results of the interaction between the molecule and the protein. According to the results, we can judge whether the binding between the molecule and the protein is good or not, and further guide modifications of the compound structure. In this paper, a semi-flexible CDOCKER docking is used to simulate the binding mode of the synthesized compound and the PDE2 protein 4HTX.
According to the analysis of the structure–activity relationship of BAY60-7550 and PDE2 protein 4HTX, we found that the carbonyl oxygen on the pyrimidinone ring forms a hydrophilic region with GLN812 and GLN859. The hydrophobic region is mainly due to the imidazotriazine skeleton, while the amino acids of ILe826, Phe862, Leu770, Leu809, Tyr655, and Met847 can form π–σ, π–π and π–alkyl stacking interactions with this core skeleton, and promote the PDE2 effects of BAY60-7550 (Figure 2).
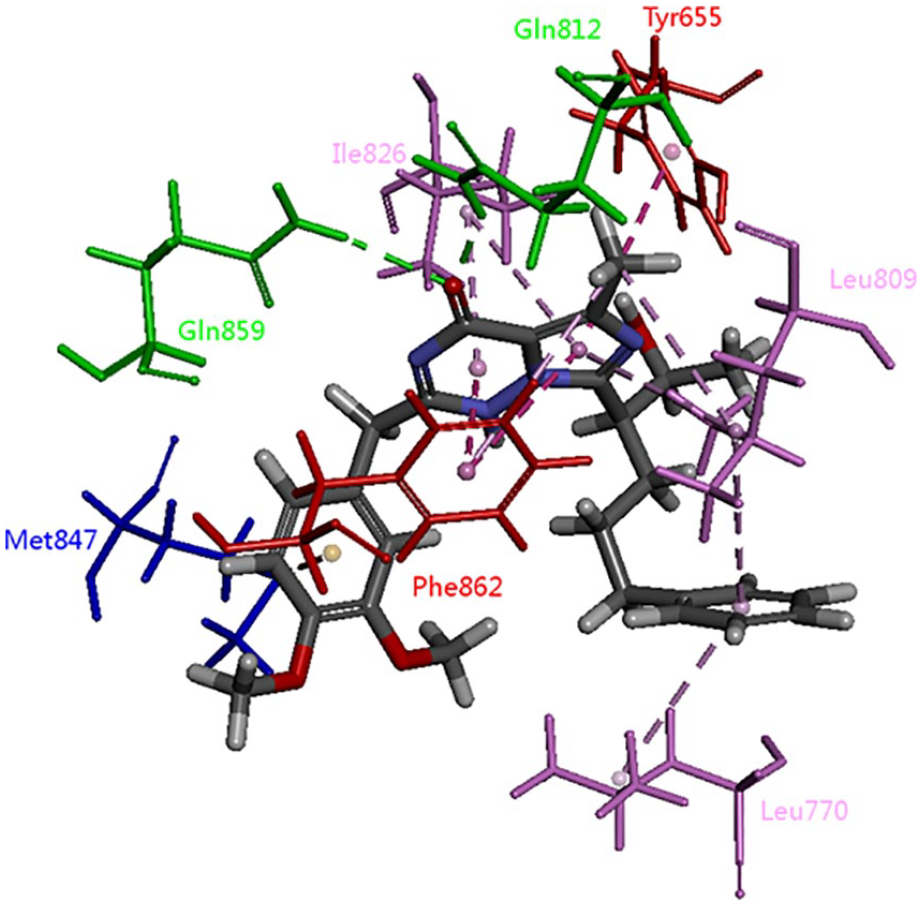
A three-dimensional plan view of the BAY 60-7550–PDE2 protein interaction active site.
To further explain the inhibitory activity of compound

A three-dimensional plan view of the
Conclusion
In this study, two series of phthalimides (
Experimental
General
1 H NMR spectra were recorded on a BrukerBioSpin GmbH spectrometer at 300 and 400 MHz, 13C NMR spectra were recorded on a BrukerBioSpin GmbH spectrometer at 75 MHz; the coupling constants are given in Hz. High-resolution mass spectrometry (HRMS) was performed using an Agilent 6200 accurate-mass time-of-flight (TOF) liquid chromatography (LC)/mass spectrometry (MS) system with electrospray ionization (ESI). Thin-layer chromatography (TLC) was performed on precoated silica gel F-254 plates (25 mm × 75 mm, Shanghai Pinjia Chemical Co., Ltd), and the samples were visualized with UV light. High-performance liquid chromatography (HPLC) analysis was performed using a SHIMADZU LC-20AB instrument. Melting points were measured with an X4-A microscopic melting point apparatus. All the starting materials and reagents were purchased from commercial suppliers.
Synthesis of compounds (1a–c ); general procedures
To a solution of maleic anhydride (30 mmol, 2.94 g) in ether (30 mL) was added ethyl furan (30 mmol, 2.88 g) in a dropwise manner, and the resulting mixture was stirred overnight at room temperature in the dark. The progress of reaction was monitored by TLC until all the starting materials had been consumed. The solution was filtered, and the filter cake was washed with ether (60 mL) to afford
2-(3,4-Dimethoxybenzyl)-4-ethylisoindoline-1,3-dione (1a )
Using 3,4-dimethoxybenzylamine as the starting martial, the desired product
4-Ethyl-2-(1H-pyrazol-3-yl)isoindoline-1,3-dione (1b )
Using 3-aminopyrazole as the starting martial, the desired product
4-Ethyl-2-[3-(benzyloxy)phenyl]-isoindoline-1,3-dione (
1c
)
Using 3-(benzyloxy)aniline as the starting martial, the desired product
Synthesis of compounds (2a–f ); general procedures
A mixture of 3-amino-4-ethoxycarbonylpyrazole (25 mmol, 3.82 g) and ethyl benzoyl acetate (25 mmol, 4.74 g) was heated in acetic acid (15 mL) at 120 °C for 12 h. After the completion of the reaction as evident by TLC, the solution was cooled and filtered. The residue was washed with acetic acid, filtered, and dried to give
7-Oxo-N-phenethyl-5-phenyl-4,7-dihydropyrazolo[1,5-a]pyrimidine-3-carboxamide (2a )
Using phenethylamine as the starting martial, the desired product
N-Benzyl-7-oxo-5-phenyl-4,7-dihydropyrazolo[1,5-a]pyrimidine-3-carboxamide (2b )
Using benzylamine as the starting martial, the desired product
N-Butyl-7-oxo-5-phenyl-4,7-dihydropyrazolo[1,5-a]pyrimidine-3-carboxamide (2c )
Using n-butylamine as the starting martial, the desired product
N-Hexyl-7-oxo-5-phenyl-4,7-dihydropyrazolo[1,5-a]pyrimidine-3-carboxamide (2d )
Using n-hexylamine as the starting martial, the desired product
7-Oxo-5-phenyl-N-[(tetrahydrofuran-2-yl)methyl]-4,7-dihydropyrazolo[1,5-a]pyrimidine-3-carboxamide (2e )
Using (tetrahydrofuran-2-yl)methanamine as the starting martial, the desired product
N-(Furan-2-ylmethyl)-7-oxo-5-phenyl-4,7-dihydropyrazolo[1,5-a]pyrimidine-3-carboxamide (2f )
Using furfurylamine as the starting martial, the desired product
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (no. 81803471).

