Abstract
Upconversion nanoparticles, Yb,Tm,Fe-doped NaYF4 nanoparticles, are synthesized and modified with polymyxin B for the selective detection of Gram-negative bacteria. Polymyxin B, a cyclic cationic antimicrobial peptide which can specifically bind to the lipopolysaccharides of cell wall of Gram-negative bacteria, such as Escherichia coli, is used to target and bind Gram-negative bacteria. The bacteria are then quantified by measuring the fluorescence intensity of the upconversion nanoparticle–bacteria complexes at 801 nm under 980 nm excitation. A limit of detection of 36 CFU/mL is achieved in the detection of Escherichia coli, and Escherichia coli in soybean milk is successfully detected. The limited autofluorescence and photobleaching properties of the upconversion nanoparticles make the proposed method useful for in vivo fluorescence imaging of Gram-negative bacteria.

Introduction
Foodborne pathogens are a threat to public health and food safety all over the world. 1 Although many efforts have been made to combat foodborne pathogens, the number of diseases caused by foodborne pathogenic bacteria remains high across many countries. An European Union (EU) report showed that more than 300,000 cases of foodborne zoonotic infections happened in 2014. 2 According to statistics, the estimated cost of foodborne diseases is more than US$15.6 billion each year in the United States, with US$300 million for the diseases caused by Escherichia coli (E. coli). 3 Therefore, it is crucial to human health and food safety to develop effective bacterial detection systems. 4
Plate colony counting is the most traditional method for bacteria detection, 5 but it suffers from low sensitivity and requires a long time to obtain results. Other detection methods including the polymerase chain reaction (PCR), 6 enzyme-linked immunosorbent assays (ELISA), 7 electrochemical detection,8,9 and flow cytometry 10 have been developed to replace plate colony counting due to their high sensitivity and effectiveness in bacteria detection. However, there are still various shortcomings, such as high cost, shortage of antibodies, time-consuming processes, or the need for expensive equipment. Recently, fluorescence methods with ultra-high sensitivity have shown great potential for bacteria detection.11,12 However, photobleaching and autofluorescence of organic dyes 13 or the cytotoxicity of quantum dots, due to their heavy metal components, 14 limits their biological applications. Rare earth–doped upconversion nanomaterials have attracted a lot of attention due to the advantages of innocuity, narrow emission width, low background interference, and no autofluorescence. 15 Therefore, upconversion nanomaterials, which can convert long-wave emission into short-wave emission, 16 have been widely used in biological detection, imaging, marking, and disease treatment.16–18 Chen et al. synthesized a new core-shell LiLuF4:Ln3+ for the detection of the disease marker beta-hCG17, 17 while Chen and co-workers prepared NaYF4:Yb,Er upconversion nanoparticle (UCNP) binding antibodies for the specific detection of E. coli. 18 Cai et al. synthesized new alkaline earth sulfide–based materials, such as CaS:Eu,Sm,Mn, as upconversion fluorescent materials. Furthermore, the electron-trapping mechanism can greatly improve the fluorescence efficiency of the upconversion emission, 19 and the emitting spectrum can be tuned by changing the doped elements and their content. 20 These efforts show that upconversion fluorescent materials have great potential in biological fields.
In this work, UCNPs were prepared and modified with polymyxin B (PMB) for the selective detection of Gram-negative bacteria, with E. coli as a model as shown in Scheme 1. Polymyxin sulfate B is a cyclic cationic antimicrobial peptide which can specifically bind to the lipopolysaccharides (LPS) of the cell wall of Gram-negative bacteria via its alkaline hydrophilic groups.21,22 Therefore, polymyxin sulfate B–modified UCNPs can selectively target Gram-negative bacteria, forming bacteria–PMB–UCNP complexes which can be fluorescently detected.
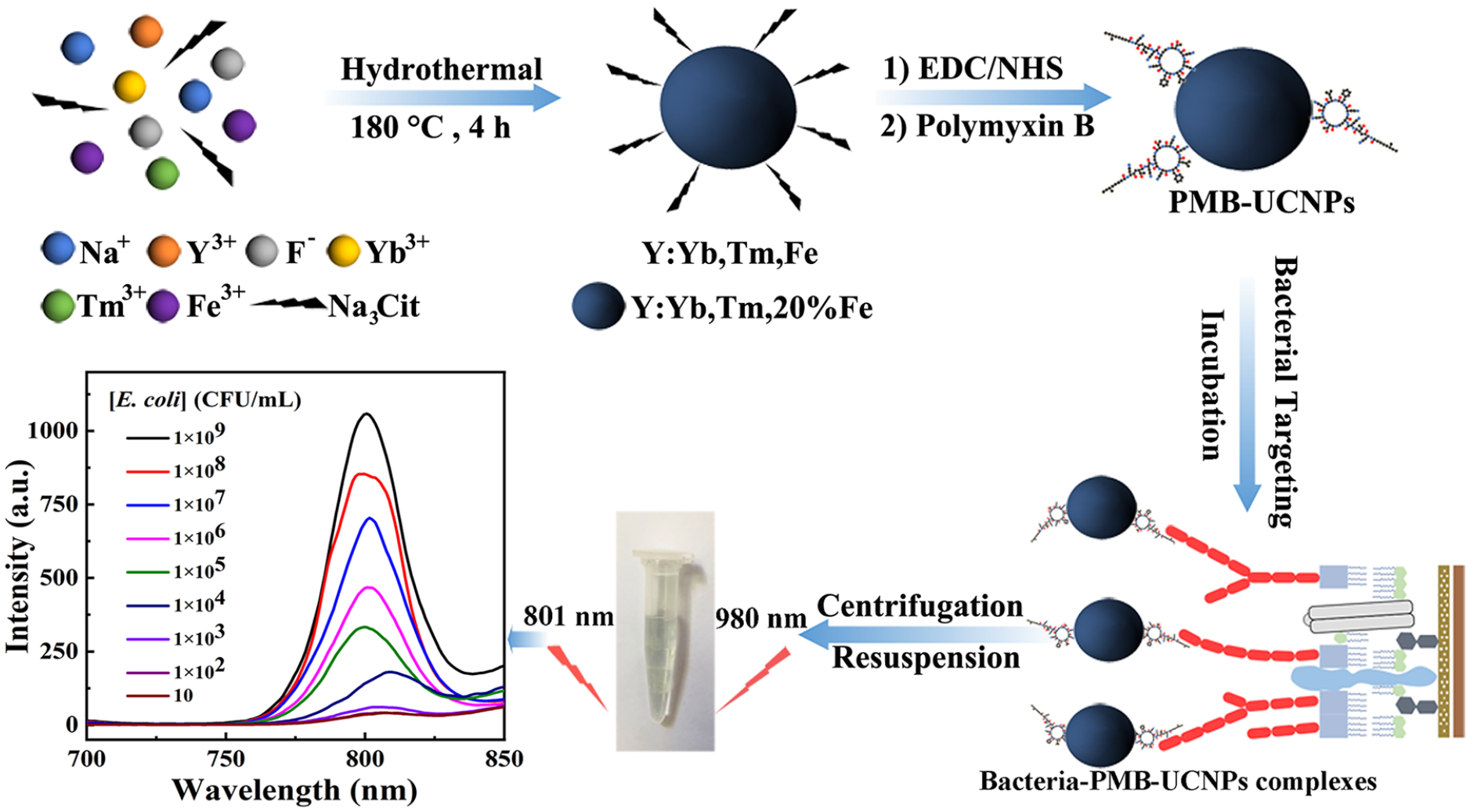
Preparation of polymyxin B–modified upconversion nanoparticles (PMB-UCNPs) and the specific detection of Gram-negative bacteria.
Results and discussion
Characterization of the UCNPs and the PMB-UCNPs
The NaYF4:Yb,Tm,Fe nanoparticles (NPs) were prepared based on our previous work, in which the doped elements had been carefully optimized. 23 The introduction of Fe3+ (20 mol%) resulted in an obvious enhancement of fluorescence intensity by 20 times, achieved by changing the molar content of Fe3+ relative to that of the Y3+ ions (Fe/Y = 0–1.32 mol).23,24 X-ray diffraction (XRD) (Figure 1(a)) shows that the synthesized UCNPs are in a mixture of standard cubic (JCPDS No. 06-0342) and hexagonal (JCPDS No. 16-0334) phases. No impurity peaks are observed. The mixed phase is probably due to the substitution of yttrium ions by iron ions forming a Yb3+–Fe3+ dimer complex. The transmission electron microscopy (TEM) image (Figure 1(b)) shows that the particles are spherical with a diameter of about 70 nm. Figure 1(c) shows the Fourier transform infrared (FT-IR) spectrum of the UCNPs. The broad peak at 3446 cm−1 corresponds to the stretching vibration peak of the hydroxy groups. The single characteristic peak at 1639 cm−1 is correlated to the asymmetric stretching vibration of the carboxyl groups (–COOH) on the surface of the UCNPs. The peak at 1392 cm−1 is the symmetrical stretching vibration, and that at 1068 cm−1 is the bending vibration absorption peak of the oxygen–hydrogen bond, which proves the existence of carboxyl groups on the surface of the UCNPs. The surface carboxyl groups make the UCNPs water-soluble. The energy-dispersive X-ray spectroscopy (EDS) spectrum (Figure 1(d)) revealed the existence of Fe, O, F, Na, Yb, Tm, C, and Y elements, confirming the successful preparation of the UCNPs.

XRD spectrum of (a) NaYF4:Yb,Tm,Fe, (b) TEM image, (c) FTIR spectrum, and (d) EDS spectrum of the NaYF4:Yb,Tm,Fe NPs.
The modification of UCNPs with PMB was characterized by ultraviolet–visible spectroscopy (UV-Vis) (Figure 2(a)) and FT-IR spectroscopy (Figure 2(c)). The modification of PMB results in new characteristic peaks at 3443, 1631, and 1230 cm−1. The peak at 3443 cm−1 corresponds to the stretching vibration of hydroxide radicals, while the peaks at 1631 and 1230 cm−1 correspond to the distinct amide I and amide II vibration modes. These results indicate the success of the immobilization of PMB on the UCNPs. The upconversion luminescence (UCL) spectrum (Figure 2(b)) shows that the covalent linkage with PMB has little effect on the emission spectrum of the UCNPs, with the maximum emission wavelength of either UCNPs or PMB-UCNPs being 801 nm. This successful modification makes possible the specific targeting of Gram-negative bacteria.

(a) UV spectra of PMB, UCNPs, and PMB-UCNPs; (b) UCL spectra of UCNPs (red) and PMB-UCNPs (black); and (c) FTIR spectra of PMB and PMB-UCNPs.
Detection of E. coli
The NaYF4:Yb,Tm,Fe NPs were synthesized with carboxylic acid ligands on the surface by the hydrothermal method. The surface carboxylic acid ligands can covalently bind with PMB to form amide bonds by the 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide hydrochloride (EDC) coupling method. 25 The detection of E. coli is based on the specific binding of PMB to LPS on the outer wall of E. coli. 26 LPS are about 8 nm in thickness, are located on the outer layer of cell walls, and are covered with mucopeptides, which are composed of core polysaccharides, O-polysaccharide side chains, and lipoid A groups. The specific binding of polymyxin and LPS occurs via a long-recognized model called the self-promoting uptake mechanism. The model considers that the free amino groups in the polymyxin Dab residue undergo protonation under physiological conditions and electrostatic adsorption with proton-like phosphate anions. 22
Incubation of the PMB-UCNPs and the bacterium solution formed bacteria–PMB–UCNP complexes which were centrifugally separated, with the unbound PMB-UCNPs leaving in the supernatant and the bacteria–PMB–UCNP complexes remaining in the precipitate. The bacteria–PMB–UCNP complexes were re-suspended in 800 μL of phosphate-buffered saline (PBS) to measure the fluorescence at 801 nm under 980 nm light excitation. The fluorescence intensity increases as the E. coli concentration increases from 10 to 1 × 109 CFU/mL, as shown in Figure 3(a). A good linear correlation (R2 = 0.9902) was obtained between the UCL intensity and the logarithm of the E. coli concentration (Figure 3(b)). The limit of detection (LOD) was calculated as 36 CFU/mL based on being three times the noise of the signal, where the noise is the standard deviation of eight control measurements. The achieved sensitivity is superior or comparable to the reported fluorescence methods18,27–35 as shown in Table 1.

(a) Fluorescence spectra of PMB-UCNPs in the presence of different concentrations of E. coli. (b) Calibration graph between the changed UCL intensity (ΔI = I − I0) and E. coli concentration.
Comparison of the fluorescence-capturing methods aimed at detection of bacteria.

AuNC: gold nanocluster; LOD: limit of detection; UCNP: upconversion nanoparticle; QD: quantum dot.
The specificity of the PMB-UCNP probe toward Gram-negative bacteria was evaluated by detection of 1.0 × 105 E. coli in the presence of other Gram-positive bacteria, Staphylococcus aureus and Bacillus subtilis, at concentrations of 1.0 × 105 CFU/mL. As shown in Figure 4, the coexisting S. aureus and B. subtilis resulted in a positive deviation of 23.8% and 31.6%, respectively, which can be ascribed to the nonspecific adsorption of these interfering bacteria.
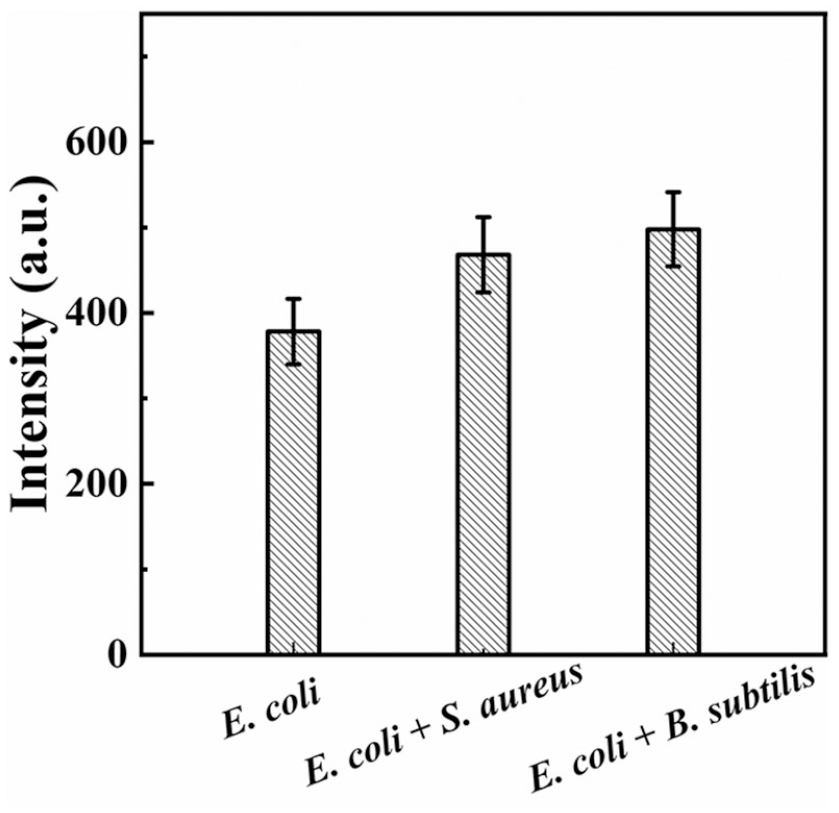
UCL intensity in the presence of 1.0 × 105 CFU/mL of E. coli and together with 1.0 × 105 CFU/mL of S. aureus or B. subtilis.
Detection of E. coli in soybean milk
In order to study the applications of the probe in the analysis of practical samples, soybean milk purchased at a local supermarket was centrifuged at 10,000 rpm for 10 min. The yellow precipitate and the upper emulsion were removed. The suspension was centrifuged once again at 4°C. The clean soybean milk solution was preserved and spiked with E. coli at different concentrations. Figure 5 is the calibration curve determined in the soybean milk medium. A linear relationship between the fluorescence intensity and the logarithmic concentration of E. coli was obtained in the range of 103–109 CFU/mL. The calibration curve determined in soybean milk medium is a little different from that determined in PBS. Based on this calibration curve, the E. coli concentration in soybean milk was measured. Table 2 lists the results determined by the proposed method and the plate-counting method for comparison. The E. coli concentration determined by the proposed method is basically consistent with the plate-counting results, with a satisfying relative standard deviation (RSD ⩽5.22%). The results showed that the bacteria count recovered using the plate-counting method was slightly lower than the added cells, which may be due to the complexity of the practical matrixes and antibacterial activities of PMB. The whole detection process is less than 2.5 h using the sensitive bacterium assay method, the result of which is acceptable.18,29,30,34 Hence, the proposed fluorescent probe can be applied for the detection of Gram-negative bacteria in real samples.
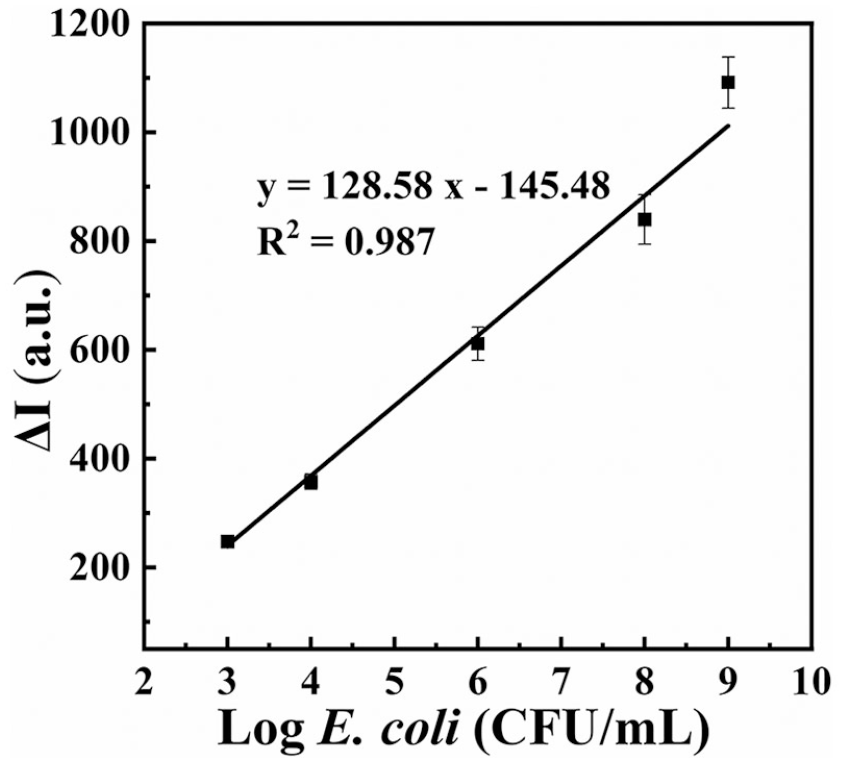
Calibration graph for detecting E. coli in soybean milk samples.
Assay of E. coli in practical samples using the proposed system and plate-counting methods.
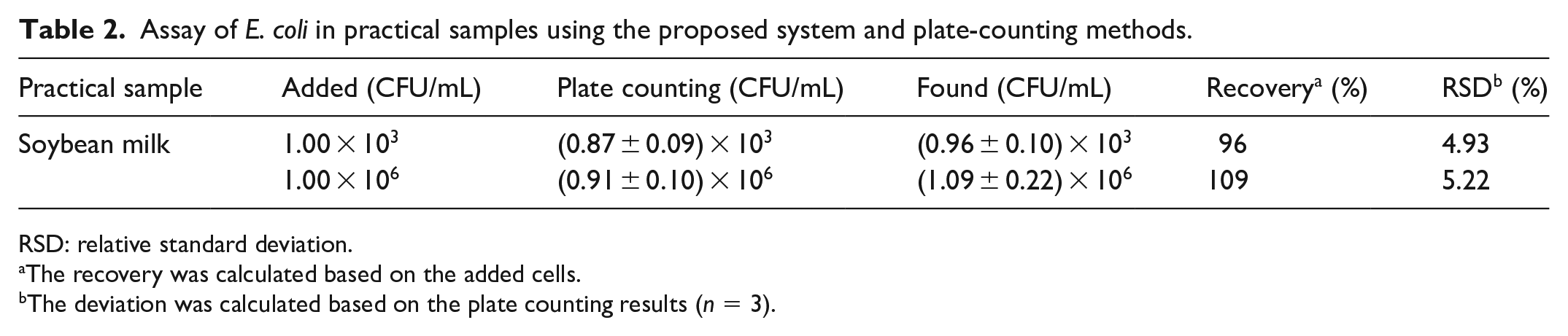

RSD: relative standard deviation.
The recovery was calculated based on the added cells.
The deviation was calculated based on the plate counting results (n = 3).
Conclusion
In summary, a fast, simple, and sensitive UCL nano-platform for the detection of Gram-negative bacteria has been constructed and applied for the detection of E. coli in soybean milk with an LOD of 36 E. coli cells per milliliter. The proposed UCL nano-platform has little background fluorescence interference, a stable fluorescence, a wide detection range, and high sensitivity. Thus, it is a good platform for the fluorescence imaging of Gram-negative bacteria.
Experimental
Reagents and materials
YCl3·6H2O (99.99%), TmCl3·6H2O (99.99%), and YbCl3·6H2O (99.99%) were purchased from Alfa Aesar. NaF, cetyltrimethyl ammonium bromide (CTAB), FeCl3·6H2O, sodium citrate, and concentrated nitric acid were of analytical reagent grade and were obtained from Sinopharm Chemical Reagent Co., Ltd. (Shanghai, China). Poly-myxin B sulfate was purchased from Sigma Aldrich. EDC was purchased from Adamas Reagent Co., Ltd. 4-N-Hydroxysuccinimide sodium (NHS) was acquired from Dalian Meilun Biological Technology Co., Ltd. Tryptone and agar were purchased from Sangon Biotech, and bacto-yeast extract powder was purchased from Aladdin. Sodium chloride was purchased from Sinopharm Chemical Reagent Co., Ltd. Normal saline was purchased from Chenxin Pharmaceutical Co., Ltd., and PBS (pH = 7.4) was purchased from Beijing Solarbio Science & Technology Co., Ltd. All reagents were used as received without further purification. Ultrapure water was obtained from a Millipore water purification system (18.2 MΩ cm, Milli-Q; Millipore, USA) and used in all assays.
Synthesis of water-soluble UCNPs
The water-soluble NaYF4:Yb,Tm,Fe UCNPs were prepared according to our reported work.23,24 YbCl3·6H2O (0.1 M, 0.5 mL), TmCl3·6H2O (0.1 M, 0.1 mL), YCl3·6H2O (0.2 M, 1.7 mL), FeCl3·6H2O (0.1 M, 1.0 mL), CTAB (0.1 g), and citric acid (0.1 M, 1.75 mL) were dissolved in a mixed solvent consisting of 15 mL of ethanol and 2.2 mL of H2O. Then, NaF (0.1 M, 6.0 mL) was added dropwise, and 1.0 mL of concentrated nitric acid was subsequently added. After stirring for 2 h, the solution was transferred into a 50-mL autoclave flask which was then heated to 180°C at a heating rate of 5°C/min and kept at 180°C for 4 h. After cooling down to ambient temperature, the products were collected by centrifugation (10 000 rpm, 10 min), washed three times using ethanol and H2O, and dried in an oven at 60°C for 10 h. The final obtained NPs are white powders.
Modification of UCNPs with PMB
PMB was covalently bonded to cit-Y:Yb,Tm,20%Fe NPs via the EDC–NHS reaction. 25 Citric acid–capped UCNPs (4 mg) were dissolved in 5 mL of PBS buffer (pH = 7.4, 10 mM), and then 7 mg of EDC and 21 mg of NHS were added under stirring for 1 h at 37°C. The resulting precipitate was centrifugally separated, washed twice with water, and dissolved in 2 mL of PBS buffer. Then 2 mg of PMB was added. After keeping at ambient temperature for 6 h under slow stirring, 500 μL of 1% bovine serum albumin (BSA) was added, and 30 min later, the solution was stored at 4°C in a refrigerator overnight. The PMB-UCNP bioconjugates were centrifugally separated, washed twice with PBS buffer to remove the unbound PMB molecules, and dispersed in 2 mL of PBS buffer. The solution was stable for 1 week.
Bacterial culture and sample suspension preparation
Each bacterium was directly subjected to the following experiments in this contribution. E. coli, S. aureus, and B. subtilis were cultured in this work. Initially, the frozen bacteria were activated from a −20°C refrigerator onto Luria–Bertani (LB) broth (1.0 g of tryptone, 0.5 g of bacto-yeast extract powder, 1.0 g of NaCl) and incubated at 37°C for 17 h. Next, the bacteria were grown in LB suspended in 100 mL of distilled water and shaken in an incubator shaker at 37°C to reach the growing stationary phase. After overnight incubation, 5 mL of the original bacterial suspensions was centrifuged at 8000 rpm for 3 min to remove the supernatant, washed three times with normal saline, and re-suspended in 5 mL of PBS (pH = 7.4, 20 mM). By measuring the optical density (OD) value at 600 nm, the bacterium liquid concentration was adjusted to the correct level for future use. The E. coli bacterial suspensions were diluted 10-fold to different concentrations from 10 to 109 CFU/mL with sterile PBS. By plating the bacteria on LB plates, the amounts of bacteria per milliliter could be acquired by counting the related colony-forming units after incubation overnight at 37°C.
Fluorescence detection with PMB-UCNPs
Five hundred microliters of E. coli suspension and 100 μL of PMB-UCNP solution were mixed and incubated in a 1.5-mL tube with shaking at 37°C for 2 h to form the bacteria–PMB–UCNP complexes, which were then separated by centrifugation at 5000 rpm for 5 min to remove the unbound PMB-UCNPs. The precipitate was suspended in 800 μL of PBS buffer, and the fluorescence at 801 nm was measured under 980 nm excitation.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (No. 21874038).
