Abstract
A new asymmetrical Schiff base ligand, N1-(3-((pyridin-2-ylmethylene)amino)propyl)ethane-1,2-diamine, and its copper(II) complex, [Cu(PPEA)Br]Br·H2O (

Introduction
Schiff base compounds and their metal complexes have been extensively investigated due to their wide range of applications including catalysts,1–3 medicine, 4 anticorrosion agents, 5 photovoltaic materials, 6 chemosensors,7,8 anode materials in sodium ion batteries,9,10 organic light-emitting diode (OLEDs) 11 and polymer light-emitting diode (PLED) materials.12,13 These compounds serve as the photosensitizers, electrolytes or redox mediators in DSSCs (dye-sensitized solar cells). 12 Metal Schiff base complexes can be exploited to act as the antenna for lanthanide ion sensitization. 14 Schiff bases are known to be good ligands for metal ions.15,16 Schiff base ligands and their complexes show interesting pharmacological effects such as antimicrobial, 17 antitumor,18,19 antibacterial, 20 antifungal, 20 antioxidative 21 and urease inhibitory22,23 activities. The cleavage of plasmid DNA by Schiff base complexes has also been reported.24,25
In the following paper, we report the synthesis, characterization (Fourier-transform infrared spectroscopy (FT-IR), Raman, X-ray) and coordination of new pyridine-based asymmetrical Schiff base ligand, N1-(3-((pyridin-2-ylmethylene)amino)propyl)ethane-1,2-diamine (PPEA; Scheme 1), towards copper(II), [Cu(PPEA)Br]Br·H2O (
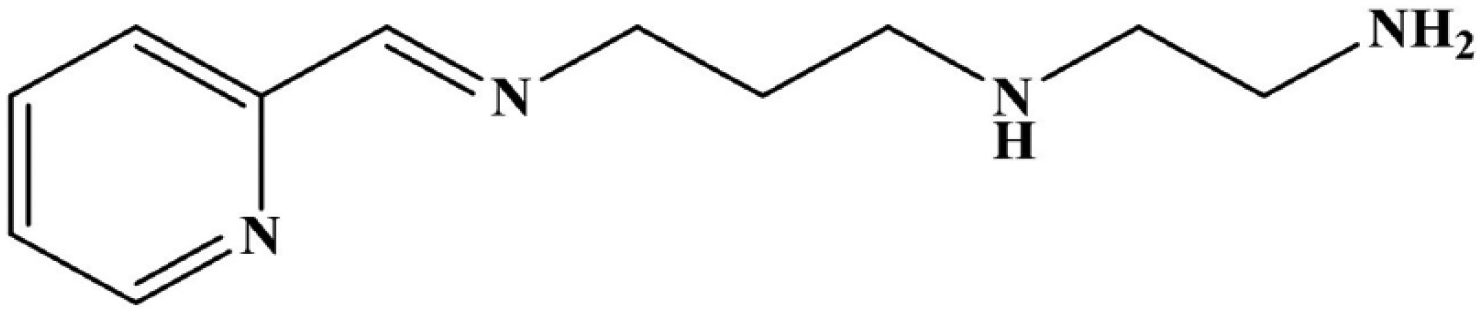
Structure of N1-(3-((pyridin-2-ylmethylene)amino)propyl)ethane-1,2-diamine (PPEA).
Schiff-base ligands are potential anticancer, antibacterial and antiviral agents, and their activities tend to increase in binding to metal centres.26,27 In addition to the expected biological properties of the PPEA ligand, binding it to the copper(II) ion makes the resulting complex a good choice as a biologically active compound. Many copper(II) complexes have been reported as potential antitumour agents and they have been found to be active both in vitro and in vivo.28–31
For the study of the biological activities of the ligand and its copper complex, docking calculations were run to investigate the possibility of an interaction between these compounds with 10 biomacromolecule targets:32–34 BRAF kinase, cathepsin B (CatB), DNA gyrase, histone deacetylase (HDAC7), recombinant human albumin (rHA), ribonucleotide reductases (RNR), thioredoxin reductase (TrxR), thymidylate synthase (TS), topoisomerase II (Top II) along with B-DNA. These proteins were selected either due to their reported roles in cancer growth or as transport agents that affect drug pharmacokinetic properties (e.g. rHA). The DNA gyrase was included to study the possibility of anticancer properties and their activity as antimalarial agents. 35 The knowledge gained from docking on the B-DNA should be useful for the development of potential probes for DNA structure and new therapeutic reagents for tumours and other diseases. 36
Experiment
Materials and instrumentation
All chemicals and solvents (Merck, Aldrich) were used as received without further purification. Infrared (IR) spectra (as KBr pellets) in the range 4000–400 cm−1 were recorded on a FT-IR 8400-Shimadzu spectrometer. The carbon, hydrogen and nitrogen contents were determined using a Thermo Finnigan Flash Elemental Analyzer 1112 EA. The melting points were measured with a Barnsted Electrothermal 9200 electrically heated apparatus. Raman spectra were obtained using a Nicolet Model 910 Fourier transform spectrometer. The microwave-assisted reactions for the synthesis of the ligand were carried out using a Microwave Laboratory Systems MicroSYNTH reactor from Milestone S.r.l.
PPEA
A mixture of N1-(2-aminoethyl)propane-1,3-diamine (0.11 g, 0.94 mmol) and pyridine-2-carboxaldehyde (0.10 g, 0.93 mmol) was irradiated inside a microwave reactor for 30 min with power up to 300 W. After microwave irradiation, an oily liquid was obtained. Excess precursors were removed using a rotary evaporator and a brown oil was obtained. Several attempts to crystallize the compound were unsuccessful. Yield: 0.14 g, 72%. Anal. calcd for C11H18N4: C, 64.05; H, 8.80; N, 27.16; found: C, 64.73; H, 8.43; N, 26.84%. IR (KBr disc, cm−1): 3400 m (νas NH2), 3306 m (νs NH2 and/or ν N−H), 3100 m (ν C−Har and/or C−Himine), 2936 s (νas CH2), 2858 m (νs CH2), 1660 m (ν C=N)imine, 1590 s (δ NH2), 1570 s (ν C=N)py, 1470 s (ν C=C)ar, 1435 s (δas CH2), 1338 m (δs CH2), 1049 m (ν C−N), 779 m and 621 m (γ py).
[Cu(PPEA)Br]Br H2O (1 )
A solution of PPEA (0.08 g, 0.39 mmol) in ethanol (15 mL) was added with stirring of CuBr2 (0.09 g, 0.39 mmol) in the same solvent (15 mL). The reaction mixture was then heated to reflux for 1 h. Blue crystals suitable for X-ray diffraction were obtained by slow evaporation of the solution for 3 days and collected by filtration. Yield: 0.09 g, 52%; m.p. 169 °C. Anal. calcd for C11H20Br2CuN4O: C, 29.51; H, 4.50; N, 12.52; found: C, 29.37; H, 4.66; N, 12.03%. IR (KBr disc, cm−1): 3383 s (νas NH2), 3213 s (νs NH2 and/or ν N−H), 3136 m (ν C−Har and/or C−Himine), 2936 s (νas CH2), 2862 m (νs CH2), 1643 m (ν C=Nimine and/or δ H2O), 1597 m (δ NH2 and/or C=Npy), 1470 m (ν C=C)ar, 1454 m (δas CH2), 1325 w (δs CH2), 1061 m (ν C−N), 783 m and 629 m (γ py and/or ρr H2O), ρw = 536 w (H2O). Raman (cm−1): 1601 m (δ NH2 and/or C=Npy), 1522 m (ν C=C)ar, 709 m and 616 m (γ py and/or ρr H2O), 461 m (ν Cu−N), 309 s (ν Cu−Br).
Crystal structure determination
Diffraction data were collected at 170 K on a Rigaku XtaLAb mini diffractometer. Data processing and absorption correction were carried out using CrystalClear-SM Expert. 37 The structures were solved with direct methods and refined with least squares using the OLEX-II package. 38 All hydrogen atoms were placed at their calculated positions. Diagrams of the molecular structure and unit cell were created using Ortep-III39,40 and Diamond. 41 Crystallographic data and details of the data collection and structure refinement are listed in Table 1, selected bond lengths and angles in Table 2 and hydrogen bond geometries in Table 3.
Crystal data and structure refinement for complex

Selected bond length (Å) and angles (°) for complex

Hydrogen bonds and short contacts dimensions (Å and °) in complex

Computational details
All structures were optimized with the Gaussian 09 software
42
and calculated for an isolated molecule using DFT
43
at the B3LYP
44
/6-31G
45
(d,p) level of theory for ligand and B3LYP/LanL2DZ
46
for complex as well as for NBO analysis. The X-ray structural data of complex
Docking studies
The pdb files 4r5y, 3ai8, 5cdn, 3c0z, 2bx8, 1peo, 3qfa, 1njb, 4gfh and 1bna for the 10 receptors, BRAF kinase, CatB, DNA gyrase, HDAC7, rHA, RNR, TrxR, TS, Top II and B-DNA, respectively, used in this research were obtained from the Protein Data Bank (pdb). 47 The full version of Genetic Optimisation for Ligand Docking (GOLD) 5.548 was used for the docking studies. The Hermes visualizer in the GOLD Suite was used to further prepare the ligand, complex and the receptors for docking. The cif file of the complex and optimized structure of ligand was used for the docking studies. The region of interest used for GOLD docking was defined as all the protein residues within 6 Å of the reference ligand ‘A’ that accompanied the downloaded protein. For B-DNA, the region of interest was defined on the DNA backbone within 10 Å of the O4, DT19 and O2, DT19 atoms for major and minor graves, respectively. All free water molecules in the structure of the proteins were deleted before docking. Default values of all other parameters were used and the compounds were submitted to 10 genetic algorithm runs using the GOLDScore fitness function.
Results and discussion
Spectroscopic characterization
In the IR spectra of PPEA and
The presence of the water molecule in
Information regarding the low frequency metal-ligand vibrations was obtained by Raman spectroscopy. In the Raman spectrum of
Crystal structure of [Cu(PPEA)Br]Br·H2O (1 )
X-ray analysis of
Ortep diagram of the molecular structure of complex
Studying the CSD database also revealed that the complexes with a Cu(N-sp3)2(N-sp2)2Br unit can be classified into two types (Scheme 2): the common (a) type (75%), in which the bromide ligand is located in the axial position and (b) type with the bromide ligand in the equatorial plane. The difference between the bond lengths of the axial and equatorial bromide ligands in (a) type complexes is 0.250 Å, confirming elongation along the z axis. Comparing the axial and equatorial Cu−N bond lengths in (b) type complexes also confirms this elongation; the average equatorial Cu−N bond length is 0.330 Å shorter than the axial one and is comparable with that of (a) type complexes (Scheme 2). Complex

CSD average for all complexes with Cu(N-sp3)2(N-sp2)2Br environment. The bromo ligand can locate in the axial position (a) or on the equatorial plane (b).
In this structure, the PPEA ligand acts as an N4-donor by forming two 5- and one 6-membered chelate rings. The five-membered chelate ring containing the imine group is planar (root mean square deviation: 0.055 for N1 atom) while the other rings are non-planar. The bond angle of the six-membered chelate ring (93.56(9)°) is larger than the two 5-membered ones (81.28(9) and 85.50(9)°). The PPEA ligand itself does not have chiral centre but a new one (N3) is formed upon coordination. Although
Crystal network interactions
In crystal structure of

Packing of complex
Theoretical studies
For comparing the structure of the complex

Ortep diagram of the optimized structures of PPEAopt and
For studying the charge distribution pattern of the ligand before and after coordination, an NBO analysis was performed (Table 4). The results reveal that the calculated charge on the carbon atom of the imine moiety is positive while other carbon atoms have a negative charge. Among the three nitrogen atoms, the amine nitrogen has higher electronegativity than the others. In
The NBO analysis results (e) for PPEA ligand and complex

PPEA: N1-(3-((pyridin-2-ylmethylene)amino)propyl)ethane-1,2-diamine; NBO: natural bond orbital.
The values are the average of charge on similar atoms.
In the optimized structure of ligand, the lowest unoccupied molecular orbital (LUMO) is delocalized on the aromatic and imine portions while the highest occupied molecular orbital (HOMO) is delocalized mainly on the aliphatic moiety (Table 5). In
HOMO and LUMO, total energy and LUMO/HOMO gap for optimized structures of PPEA ligand and complex

PPEA: N1-(3-((pyridin-2-ylmethylene)amino)propyl)ethane-1,2-diamine; LUMO: lowest unoccupied molecular orbital; HOMO: highest occupied molecular orbital.
Docking studies
For predicting the biological activities of the PPEA ligand and its copper complex, interactions of these compounds with 10 macromolecular receptors were studied using the GOLD 48 docking software. The GOLD docking results are reported in terms of the values of fitness, which means that the higher the fitness, the better the docking interaction of the compounds.32–35 The results of the docking presented in this work are the best binding results out of 10 favourites predicted by GOLD. Also for evaluation of the calculated fitness values, these scores were compared with those of the anticancer drug, doxorubicin (a cancer medication that interferes with the growth and spread of cancer cells in the body 62 ).
The general features from the GOLD docking prediction (Table 6) show that all studied structures can be considered as biologically active compounds.32–34 The best predicted protein targets for PPEA and
The calculated fitness values for PPEA ligand and complex

PPEA: N1-(3-((pyridin-2-ylmethylene)amino)propyl)ethane-1,2-diamine.

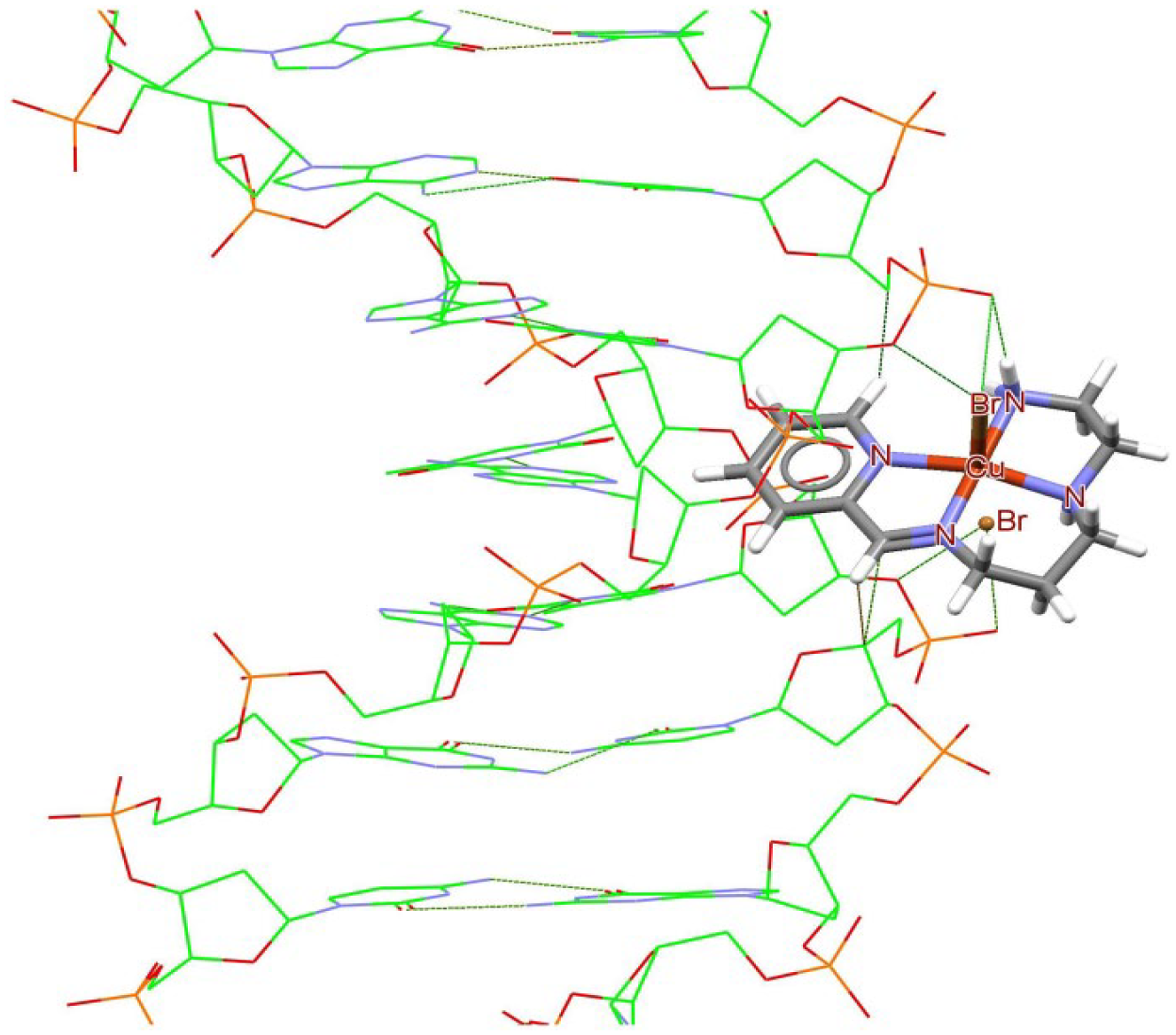
Docking study results, showing the interaction between the PPEA and B-DNA (minor grave).

Docking study results, showing the interaction between complex
Conclusion
In this work, a new asymmetrical Schiff base ligand, PPEA, and its copper(II) complex, [Cu(PPEA)Br]Br·H2O (
Footnotes
Acknowledgements
We are grateful to Payame Noor University and Urmia University of Iran for financial support.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
