Abstract
Background
Accumulating evidence has indicated that runt-related transcription factor 3 (RUNX3) gene polymorphism (rs7528484) is associated with an alimentary system cancer risk. However, the role of rs7528484 in colorectal cancer is still unclear. The present study aimed to explore the association between rs7528484 and colorectal cancer susceptibility in a Chinese Han population.
Material and methods
We firstly investigated the effect of the polymorphism rs7528484 in distal promoter of RUNX3 polymorphism on colorectal cancer risk in a Chinese Han population comprising 427 colorectal cancer patients and 503 controls. We then carried out a phenotype–genotype association analysis to validate its influence on the adjacent gene RUNX3.
Results
Logistic regression analysis demonstrated that the T allele of rs7528484 was significantly associated with an increased risk for colorectal cancer occurrence in our case-control study (odds ratio = 1.33; 95% confidence interval = 1.09–1.65; P = 0.005). In stratified analysis, the susceptibility of colorectal cancer in the T allele carriers increased among the smokers, III and IV tumor stage, and at the rectum. Furthermore, the T allele was significantly correlated with lower expression of RUNX3 in vitro.
Conclusion
In summary, the current case-control and genotype–phenotype study provides convincing evidence that functional RUNX3 polymorphism (rs7528484) is related to colorectal cancer risk and is a plausible marker for the prediction of colorectal cancer.
Introduction
As one of the leading digestive cancers in the world, colorectal cancer (CRC) seriously threatens public health worldwide. 1 Despite the significant advance in CRC therapy, numerous CRC cases still faced a cancer-free survival challenge.2,3 Although the pathogenesis of CRC is not yet fully elucidated, recent molecular epidemiological studies have demonstrated that genetic polymorphisms in genes that relate to carcinogenesis play a vital role in susceptibility to CRC.4,5
As a member of the runt-domain transcription factor family, runt-related transcription factor 3 (RUNX3) is localized to chromosome 1p36, a region that exhibits frequent loss-of-heterozygosity events in multiple cancers including CRC; it is considered a tumor-suppressor gene involved in the transforming growth factor beta (TGF-β) signaling pathway. 6 RUNX3 participates in cell signaling pathways involved in cell growth, apoptosis, proliferation, cell cycle regulation, and angiogenesis of malignant tumors. 7 Its anticancer mechanism is mainly related to the TGF-β pathway that induces growth inhibition and apoptosis. 8 Inactivation of RUNX3 is caused by promoter hypermethylation, loss of heterozygosity, or incorrect localization. An important reason for the down-regulation of RUNX3 expression is the methylation of the CpG islands in the RUNX3 promoter region. 9 The RUNX3 can be used as a basis for judging the degree of malignancy of CRC and is an indicator for the assisted diagnosis of this neoplasm. 10 Furthermore, RUNX3 expression is frequently reduced in human intestinal metaplasia; polymorphisms within RUNX3 have been reported in digestive cancers such as gastric cancer.11–13 This fact raised the possibility that genetic polymorphisms within the functional region of RUNX3 would lead to aberrant expression and further confer a risk for CRC.
RUNX3 rs7528484 C > T polymorphism is a C to T transition leading to the DNA sequence change in the promoter. Since rs7528484 was firstly reported to involve in gastric cancer risk, 13 only one study reported a positive genetic association between rs7528484 and severe preeclampsia. 14 However, the association between RUNX3 rs7528484 and CRC susceptibility is still unclear.
To further confirm whether RUNX3 rs7528484 influences CRC risk, we performed a case-control study to investigate the plausible association between the RUNX3 rs7528484 and CRC risk in a Chinese Han population, and additional promoter activity analysis in vitro to study the potential function of RUNX3 rs7528484.
Materials and methods
Ethics approval and informed consent
Informed consent was obtained from all subjects according to the Internal Review and Ethics Boards of the Affiliated Hospital of Xuzhou Medical University, and the project was in accordance with the Helsinki Declaration of 1964.
Study population
All recruited subjects were local residents of Han Chinese population. A total of 427 patients with histopathologically diagnosed primary CRC and 503 cancer-free subjects were from the Affiliated Hospital of Xuzhou Medical University between January 2014 and February 2016. The diagnosis for primary CRC was conducted under guidelines of the American Joint Committee on Cancer and International Union Against Cancer standard (8th version). There was no sex, stage, and histology restriction for CRC cases. CRC patients aged > 18 years were enrolled in the present study. The selection criteria for controls included no family history of any other cancer and the distribution frequency of age and sex of matched to that of cases.
Tumor tissues from 77 patients with a diagnosis of CRC were collected according to the availability of frozen stored tissues of CRC resections from 2012 to 2014. These cases were confirmed by pathologic diagnosis, and none of these patients had ever received preoperative chemotherapy or radiotherapy. (These 77 patients were part of the 427 cases recruited.) Tumor stages were determined according to a modified American Joint Committee on Cancer and International Union Against Cancer standard. After surgical resection, fresh tissues were immediately stored at −80°C until DNA/RNA isolation and protein extraction.
DNA extraction and genotyping
Genomic DNA was extracted from peripheral blood leukocytes according to instruction of genomic DNA purification kit (Qiagen). The RUNX3 rs7528484 was detected using an allele-specific polymerase chain reaction (PCR) method. The PCR primers were designed based on the GenBank reference sequence and Primer 5.0. The sequences for primers are listed in Supplementary Table 1. A DNA fragment with the C allele could be amplified by the F1 and R1 primer pair; similarly, a DNA fragment with the T allele could be amplified by F2 and R2 primer pair PCR was performed in a 20 μL reaction mixture. Each primer pair was separately amplified: PCR conditions were 94°C for 5 min, followed by 35 cycles of 30 s at 94°C, 30 s at 57°C, and 30 s at 72°C, with a final elongation at 72°C for 5 min. The PCR products were 481 bp for primer pair 1, 525 bp for primer pair 2. The PCR products for two independent tubes were then mixed and separated on a 2% agarose gel stained with ethidium bromide for visualization under ultraviolet light. After electrophoresis, homozygous C alleles were represented by DNA bands with sizes at 481 bp; homozygous T alleles were represented by DNA bands with sizes at 525 bp; and heterozygous genotypes were displayed as a combination of 481 bp and 525 bp. To validate the genotyping result, approximately 20% of the case and control samples were randomly selected and tested in duplicate by direct sequencing, and the reproducibility was 100%.
qPCR
Total RNA was isolated from tumor tissue specimens and cell lines using RNA isolation kit of Qiagen and then converted to cDNA using random primers and Superscript II (Invitrogen). A SYBR® Green gene expression assay was performed using Roche LightCycler® 480 to quantify relative RUNX3 expression in these samples. Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was chosen as the internal control. The real-time primers are listed in Supplementary Table 1. The 25μL total volume final reaction mixture consisted of 1 μM of each primer, 12.5 μL of Master Mix (Applied Biosystems, Foster City, CA, USA), and 2.5μL of cDNA. Negative controls consisted of distilled H2O. The expression levels of target genes were normalized with GAPDH with an arbitrary unit. A melting curve analysis was performed for the PCR products to evaluate primer specificity. To further validate the specificity of primers for RUNX3, we analyzed 10 randomly selected RUNX3 PCR products by direct sequencing.
Western blotting
Approximately 50 µg of proteins from CRC tumor tissue lysates were electrophoresed on a 12% polyacrylamide gel. Proteins were transferred to a PVDF membrane (GE Healthcare) and reacted with antibodies against RUNX3 (ab49117) or GAPDH (Abcam Company) at a dilution of 1:1000 and 1:2000, respectively. The primary antibodies were detected by horseradish peroxidase conjugated secondary antibodies (Santa Cruz Biotechnology) at a 1:2500 dilution. Visualization of the immunoreactive proteins was accomplished with an enhanced chemiluminescence system (ECL, Cell Signaling Technologies).
Constructs, cell culture, and luciferase assay
The 250 bp DNA fragments with rs7528484 and extra Bgl II and Hind III cutting sequences were directly synthetized by Sangon Company (Shanghai). Then the C or T allele fragment was cloned into pGL3-basic vector (Promega) at the restriction cutting sites and labelled C construct or T construct. The sequence orientation was validated by direct sequencing.
Cell lines including HCT116 and LoVo were cultured in Dulbecco's Modified Eagle Medium with added 10% fetal bovine serum and 1% penicillin-streptomycin at 37°C with 5% CO2 in cell culture box (Thermofisher).
Cells were firstly seeded at a density of 3 × 106 cells per well in 6-well plates (Corning), then cells were transfected by Lipofectamine 2000 (Invitrogen) After 16 h. 500 ng pGL3-WT or pGL3-MT and 50 ng pRL-TK vector (Promega) were co-transfected per well. pGL3-basic empty was taken as negative control. 24 h after transient transfection, cells were harvested and the luciferase assay was applied conforming to the instructions of the Dual-Luciferase® Reporter Assay System kit manufacturer (Promega).
Statistical analysis
The genotype distribution was analyzed for the Hardy-Weinberg equilibrium using χ2 test. Logistic regression was used to analyze the association between rs7528484 and CRC risk, and adjusted for sex, age, smoking status, and family history of CRC. Unconditional logistic regression analysis was fitted to estimate odds ratios (ORs) and their 95% confidence intervals (CIs), with adjustment for age, gender, smoking status, and family history of cancer. The normalized expression levels of RUNX3 in different genotype tumor tissues were compared using the non-parametric Mann–Whitney U test. The luciferase activity comparison was tested with the Wilcoxon signed rank test. These statistical analyses were implemented in the Statistic Analysis System software (version 8.0, SAS Institute). P < 0.05 was used as the criterion of statistical significance, and all statistical tests were two sided.
Results
The association of rs7528484 with CRC susceptibility
The main characteristics of the 427 CRC patients and 503 controls included in the analysis are summarized in Supplementary Table 2). Genomic DNA was extracted from peripheral venous blood and tissue samples; after PCR amplification the target DNA fragment was identified as shown in Figure 1. There were no statistically significant differences between cases and controls in terms of the distribution of age, sex, body mass index (BMI), smoking status, and family history of cancer.
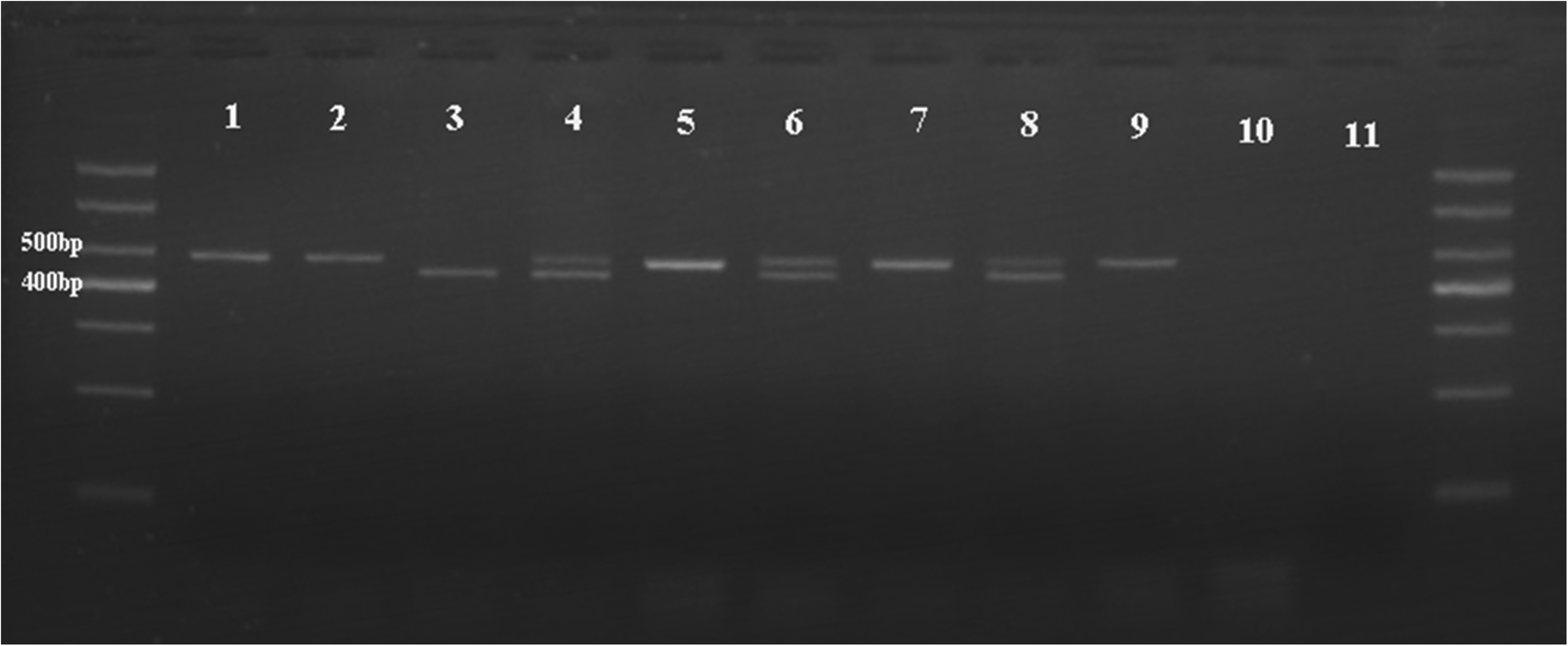
Rs7528484 genotype in 2% agarose gel. Lane 1, 2, 5, 7, 9 TT; Lane 4, 6, 8 CT; Lane 3 CC; Lane 10, 11 as the blank control using water. Nine randomly selected samples from the cases.
Genotype distributions had no deviation from HWE in control groups (P = 0.75), the detailed value of χ2 test is not shown in Supplementary Table 3. Based on all genetic model analyses, the results of the dominant model showed that rs7528484 was more pronounced in association with CRC susceptibility. Compared with the reference (CC), the heterozygote (CT), or homozygote (TT) of rs7528484 was associated with a significantly increased risk of CRC after controlling for age and sex (CT vs. CC: OR = 1.34, 95% CI = 1.04–1.73; TT vs. CC: OR = 1.74, 95% CI = 1.11–2.69). Meanwhile, frequencies of C or T allele were significantly different between CRC and control groups. In the additive model, each additional copy of the T allele was associated with a 33% increased risk of developing CRC (T vs. C: OR = 1.33, 95% CI = 1.09–1.65).
Further stratified analysis was performed for rs7528484 by clinical characteristics to explore the relationship of rs7528484 polymorphism, gene-environment interaction, and the risk of CRC. As shown in Supplementary Table 4, the risk of CRC in the T allele carriers was significantly increased in smokers (OR = 1.47; 95% CI 1.15–1.84; P = 0.001) compared with the C allele carriers. Regarding tumor stage (Supplementary Table 5), we divided cases into two subgroups: I + II stage and III + IV stage; significant correlation between the increased risk of CRC and the T allele were found in III + IV stage CRC cases (OR = 1.51; 95% CI 1.21–1.88; P = 0.0002). In addition, when considering the tumor site (Supplementary Table 5), CRC risk was increased in the T allele carriers (OR = 1.54; 95% CI 1.22–1.93; P = 0.0002). No positive correlation was found when stratified analysis was carried out with potential confounders such as age, gender, BMI, and family history of CRC (data not shown).
The phenotype–genotype correlation between rs7528484 and RUNX3 expression in tumor tissues
With such strong evidence of association, we then evaluated the effects of rs7528484 on its adjacent gene expression. Since rs7528484 was located in distal promoter of RUNX3, we examined the expression of RUNX3 in CRC tissue samples with different genotypes. As shown in Figure 2, results of qPCR demonstrated that subjects with variant genotypes (CT and TT) had significantly lower RUNX3 mRNA levels (mean ± SE) than those with the most common CC wild genotype.
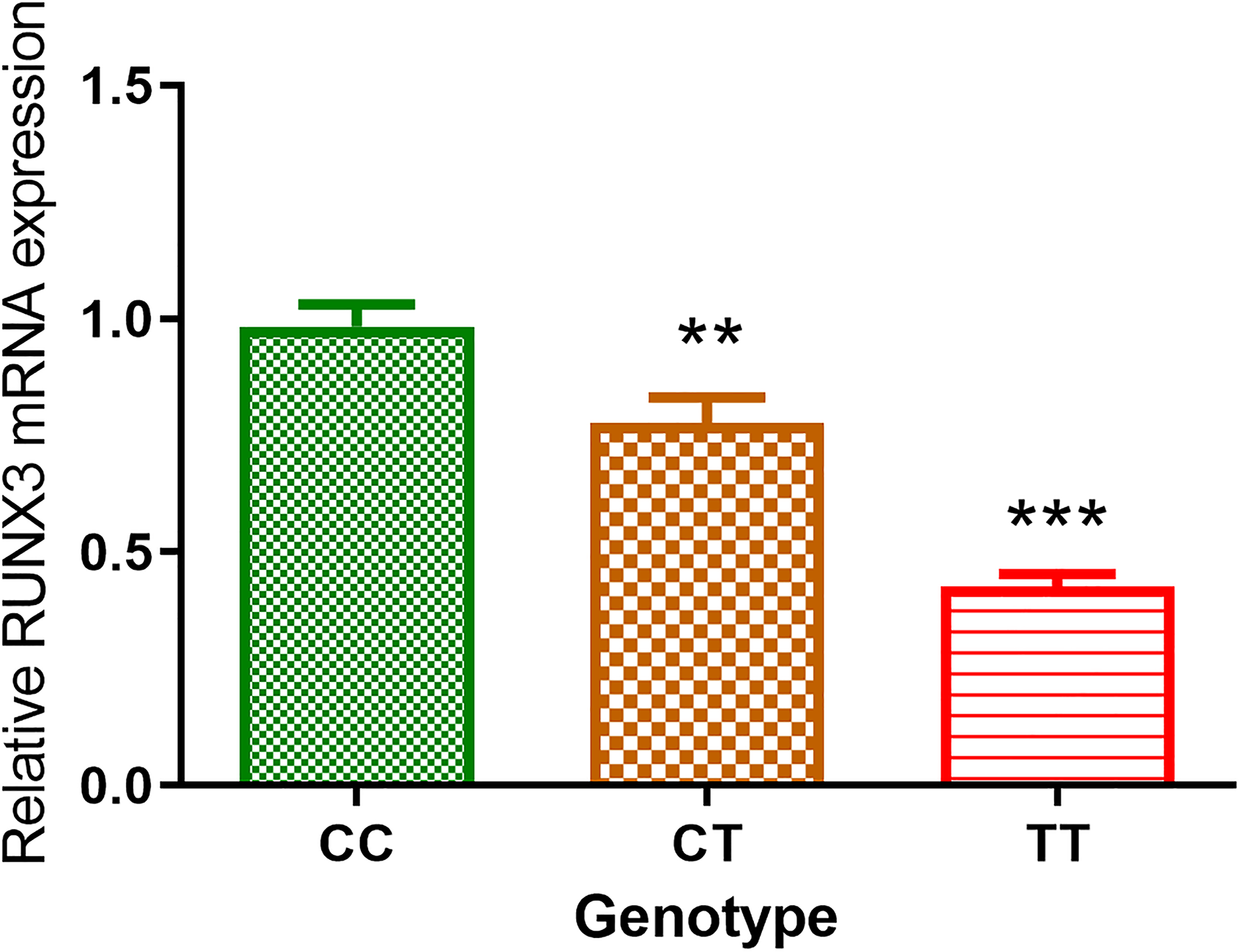
Relative RUNX3 mRNA expressions of CRC tissues in three genotypes. The CC group was the reference. *P < 0.05 vs. CC; **P < 0.01 vs. CC. All 77 CRC tissue samples were genotyped as 31 CC, 36 CT and 10 TT.
In order to further investigate this correlation at the protein level, we randomly selected five CRC tumor tissue samples with different genotypes and analyzed the RUNX3 expression level using western blotting. We found that the levels of RUNX3 protein of three cases carrying variant genotypes were significantly lower than that of wild genotype carriers (Figure 3). Further RUNX3 protein expression analysis of 77 CRC cases was carried out to validate the trend found in five CRC tissue samples. Quantification of western blot showed that RUNX3 expression was attenuated with the presence of the T allele. The RUNX3 expression of CT genotype carriers was approximately 69% of CC carriers in CRC tissues, while in TT genotype carriers, the ratio was about 57% (Figure 4). The data demonstrated that rs7528484 was significantly correlated with RUNX3 expression at both the mRNA and protein levels.
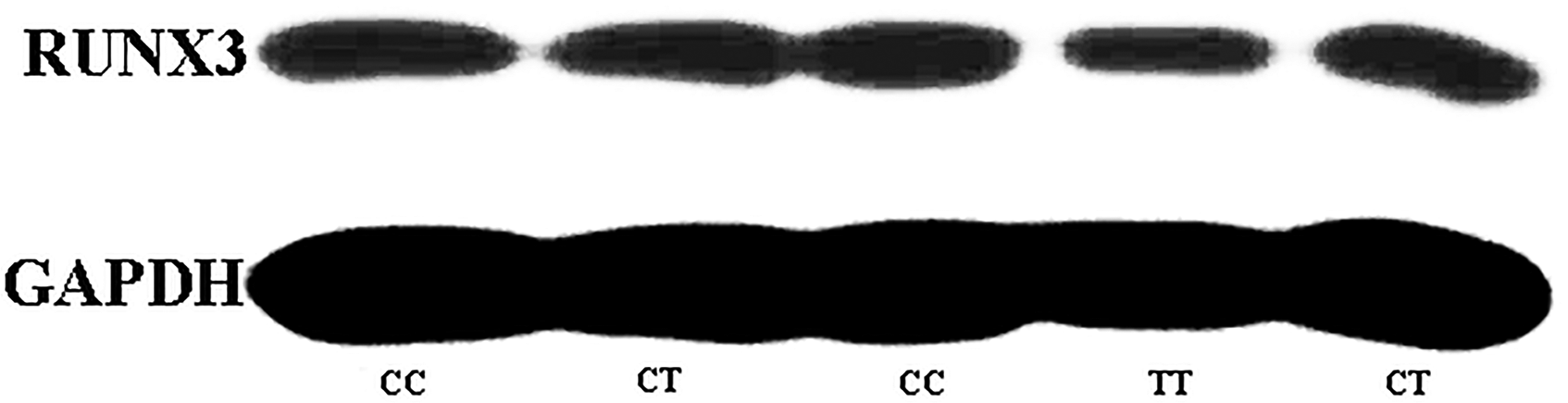

Western blot analysis results of rs7528484 genotyped 5 CRC tissues. Selected CRC tissues were analyzed. The CC group was the reference.
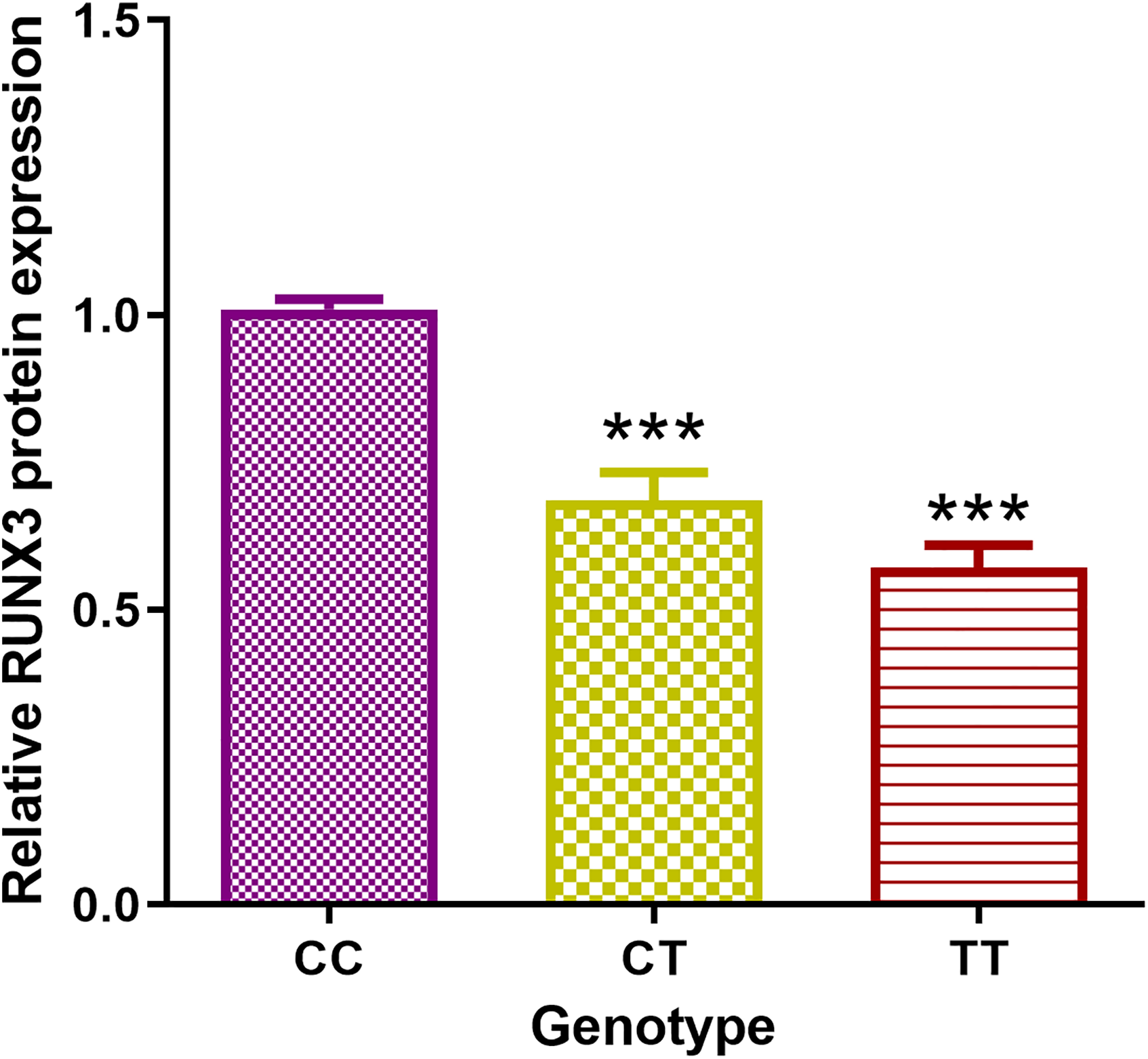
Western blot analysis results of rs7528484 genotyped CRC tissues. There were 77 CRC tissues genotyped as 31 CC, 36 CT, and 10 TT and analyzed. The CC group was the reference for the relative quantification for both CT and TT groups. *P < 0.05 vs. CC; **P < 0.01 vs. CC.
Rs7528484 modulate promoter activity of RUNX3
Given that rs7528484 could modulate the protein level of RUNX3 and its location in the distal promoter region of RUNX3, we speculated that rs7528484 could modulate the promoter activity; therefore, we did the luciferase assay to assess its possible mechanism. As shown in the data in Figure 5, the T allele vector had a marked decrease in promoter activity when compared to the C allele vector (P < 0.01).
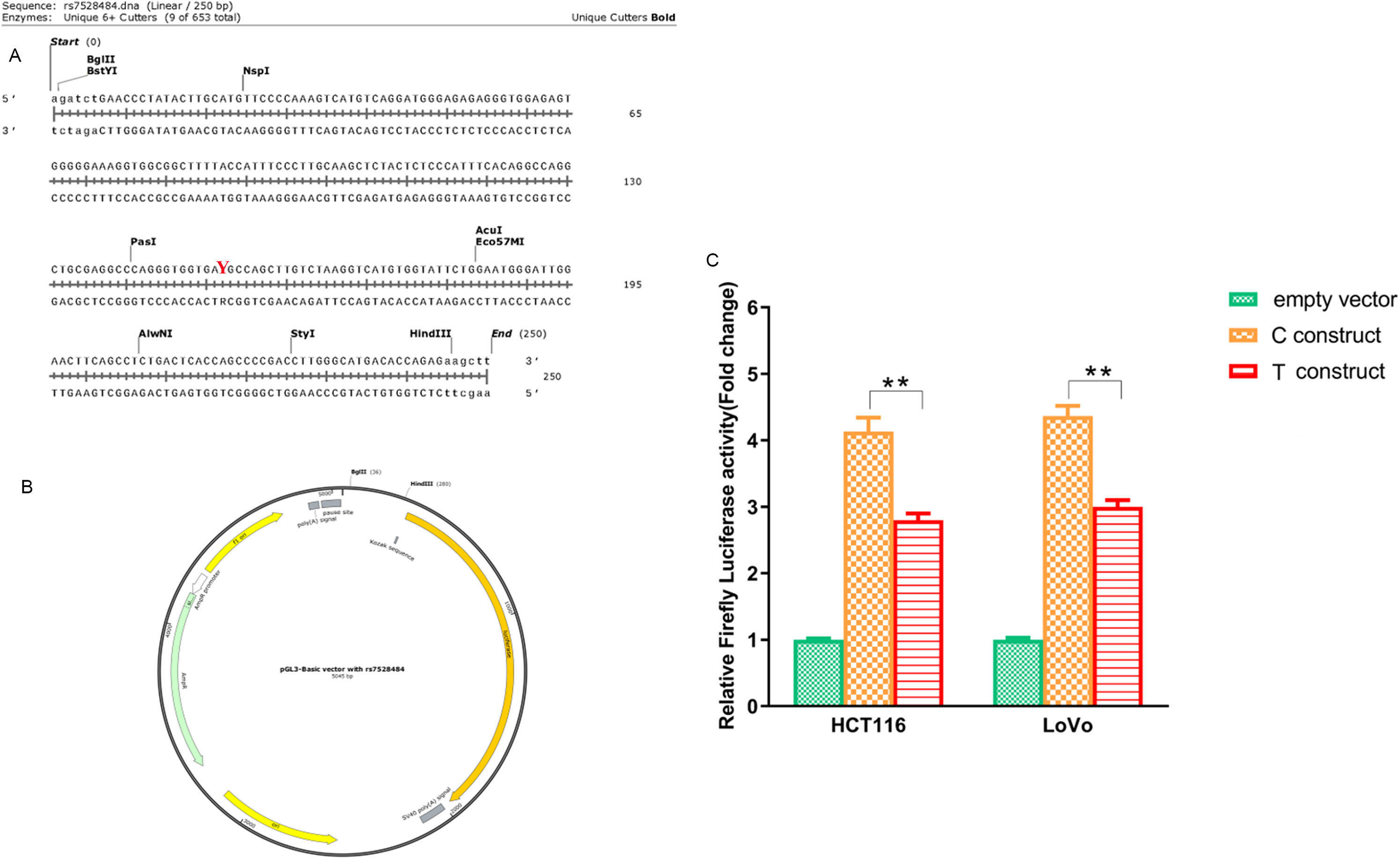
Luciferase reporter assay to investigate the possible function of rs7528484. *P < 0.05 vs. C construct; **P < 0.01 vs. C construct. (A) The direct synthesis sequence with rs7528484 (Y indicates the location). (B) The construct with DNA sequence shown in (A). (C) Influence of rs7528484 on transcription activity by luciferase reporter assay. *P < 0.05 vs. C construct; **P < 0.01 vs. C construct.
Discussion
This is the first case-control study evaluating the association between rs7528484 and CRC risk in a Chinese Han population, and the first report to investigate a functional SNP in RUNX3 distal promoter in CRC. On the basis of our current results, we found that the functional SNP rs7528484 influenced RUNX3 expression in CRC tissues.
RUNX3, although widely expressed, is thought to be especially correlated with gastrointestinal tract development. 15 Considering the role of RUNX3 in signaling cascades mediated by TGF-β, aberrant expression of RUNX3 would probably lead to a TGF-β signaling pathway disorder. The TGF-β signaling pathway was one of the canonical pathways investigated in CRC development and progression16,17; thus RUNX3 expression disorders have the potential for CRC etiology. 18 Even though previous reports mainly focused on the aberrant methylation of RUNX3,19,20 polymorphisms related CRC case-control studies remain in a smaller number. 15 Reports of functional polymorphisms on CRC did not include rs7528484 perhaps because our target polymorphism was not a tag SNP. 15 The functional SNP rs7528484 influenced RUNX3 expression in intestinal-type gastric cancer tissues was reported by Lim et al., 13 but the researchers only investigated the association in gastric cancer tissues and cell lines. It was inspiring that the rs7528484 could increase activity of the RUNX3 distal promoter in cancer cells. 13 With such strong evidence, we did the case-control study in a Chinese Han population to confirm our hypothesis that rs7528484 could increase the risk of CRC and therefore influence RUNX3 expression in human CRC tissues. Our data supported the concept that a similar trend was present in alimentary system cancers.
Interestingly, further stratification analysis found that the genotype T allele versus the C allele showed statistically significant differences in the smoker group, stage III + IV group, and the rectum location group; however, no statistically significant difference was found among non-smokers, stage I + II, and the colon location group. The difference in smoking status may indicate that the polymorphism is highly associated with smoking; tobacco may contribute to RUNX3 aberrant expression and thus may elevate the risk of CRC. 21 However, the exact mechanism is still unclear. RUNX3 expression was downregulated in high-grade tumor stage groups and the oncogene-repressor role was attenuated; thus, the tumor progression was enhanced. This may be the most feasible reason for the T allele increase with high-grade tumor stage. The high T allele frequency in the rectum sites was likely due to abuse of the rectum (e.g. drugs having been carried there, or rectal fingering having taken place), which induced chronic inflammation of epithelial cells of rectum and canceration.
The functional assay indicated that the rs7528484 could affect the expression of RUNX3 in CRC; the possible mechanism may involve with transcription factor binding disruption. Lim et al. reported that CREB1 was predicted to bind the sequence carrying risk-associated base G allele of rs7528484 but not with non-risk-associated base A in gastric cancer model.13 Similarly, we hypothesized that the CREB1 binding disruption induced the RUNX3 expression in the CRC model. Thus, future in vitro assays are warranted for a full understanding of the underlying mechanism.
Some limitations in the study are noted. First, the sample sizes for our case control association studies are relatively small. Therefore, further studies with larger sample sizes are needed to validate our findings. Second, without enough individual data, we could not determine the exact interaction between rs7528484 and environmental factors. Third, lack of functional analysis in vitro limited full elucidation of the potential role of rs7528484 in the predisposition for CRC. A low expression of RUNX3 was frequently detected in human cancer tissues; thus, the tumor suppressor role was weakened. Similarly, in our study, the T allele decreased the RUNX3 level, probably for its tumor repressor role in carcinogenesis; that is, the T allele could increase CRC risk and promote CRC progression. In addition to smoking status and family history, we did not adjust other known potential confounding factors associated with CRC risk (including other lifestyle factors) due to insufficient data collected; this would likely influence the reliability of the point estimates (ORs) of rs7528484. Obviously, our present finding needs more complex experiment support from other research centers in the future.
Collectively, we provided the experimental evidence of the positive correlation between rs7528484 and CRC susceptibility in the Chinese Han population. Therefore, rs7528484 may be a promising marker for predicting CRC.
Supplemental Material
sj-docx-1-jbm-10.1177_17246008211073342 - Supplemental material for A functional polymorphism within the distal promoter of RUNX3 confers risk of colorectal cancer
Supplemental material, sj-docx-1-jbm-10.1177_17246008211073342 for A functional polymorphism within the distal promoter of RUNX3 confers risk of colorectal cancer by Huiping Wang, Jin Wang, Danhua Li, Zhansheng Zhu and Dongsheng Pei in The International Journal of Biological Markers
Supplemental Material
sj-docx-2-jbm-10.1177_17246008211073342 - Supplemental material for A functional polymorphism within the distal promoter of RUNX3 confers risk of colorectal cancer
Supplemental material, sj-docx-2-jbm-10.1177_17246008211073342 for A functional polymorphism within the distal promoter of RUNX3 confers risk of colorectal cancer by Huiping Wang, Jin Wang, Danhua Li, Zhansheng Zhu and Dongsheng Pei in The International Journal of Biological Markers
Supplemental Material
sj-docx-3-jbm-10.1177_17246008211073342 - Supplemental material for A functional polymorphism within the distal promoter of RUNX3 confers risk of colorectal cancer
Supplemental material, sj-docx-3-jbm-10.1177_17246008211073342 for A functional polymorphism within the distal promoter of RUNX3 confers risk of colorectal cancer by Huiping Wang, Jin Wang, Danhua Li, Zhansheng Zhu and Dongsheng Pei in The International Journal of Biological Markers
Supplemental Material
sj-docx-4-jbm-10.1177_17246008211073342 - Supplemental material for A functional polymorphism within the distal promoter of RUNX3 confers risk of colorectal cancer
Supplemental material, sj-docx-4-jbm-10.1177_17246008211073342 for A functional polymorphism within the distal promoter of RUNX3 confers risk of colorectal cancer by Huiping Wang, Jin Wang, Danhua Li, Zhansheng Zhu and Dongsheng Pei in The International Journal of Biological Markers
Supplemental Material
sj-docx-5-jbm-10.1177_17246008211073342 - Supplemental material for A functional polymorphism within the distal promoter of RUNX3 confers risk of colorectal cancer
Supplemental material, sj-docx-5-jbm-10.1177_17246008211073342 for A functional polymorphism within the distal promoter of RUNX3 confers risk of colorectal cancer by Huiping Wang, Jin Wang, Danhua Li, Zhansheng Zhu and Dongsheng Pei in The International Journal of Biological Markers
Footnotes
Acknowledgments
We are grateful to Professor Ronghao Wang from Xi’nan Medical University for essential language editing and valuable advice for the current study.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author contributions
Huiping Wang and Jin Wang contributed equally for the current manuscript.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the NSFC (No. 81502428) and the Key Research and Development Program of Xuzhou City (No. KC21250).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
