Abstract
Background:
Previous studies have reported that polymorphisms in the interleukin-1 gene may be involved in tumorigenesis and tumor progression.
Aim:
The purpose of the present study was to evaluate whether an insertion/deletion polymorphism, rs3783553, located in the miR-122 target gene interleukin-1α, was associated with the risk of colorectal cancer.
Methods:
Genomic DNA was extracted from peripheral venous blood of 382 patients with colorectal cancer and 433 controls, and the polymorphism was genotyped using a polymerase chain reaction assay.
Results:
Significantly decreased colorectal cancer risk was observed to be associated with the interleukin-1α rs3783553 insertion/insertion genotype (P=0.0001; OR=0.41; 95% CI 0.26, 0.65) and the insertion allele (P<0.001; OR=0.68; 95% CI 0.55, 0.83). Stratification analysis based on clinical and pathological features also revealed that the “TTCA” insertion allele of rs3783553 contributes to slow the progression of colorectal cancer.
Conclusion:
These results suggest that the rs3783553 polymorphism could be a useful genetic marker to predict the size/extent of colorectal cancer.
Introduction
Colorectal cancer (CRC) is the third most commonly diagnosed cancer, and is responsible for a high proportion of cancer-related mortality worldwide. 1 Colon carcinogenesis is a multistep process in which normal colon epithelial cells are transformed into malignant cells. It is well known that gene mutations and inflammatory responses play key roles in the development of CRC. Previous studies have shown that KRAS rs712, interleukin (IL)-16 rs11556218, and CD86 polymorphisms are associated with the risk of CRC.2-4 In addition, the survival time of CRC patients is highly dependent upon the clinical stage of the tumor. Therefore, there is an urgent demand for determining the gene mutations that may act as predictors of clinical outcome in CRC patients, in order to improve their prognosis.
The IL-1 cytokine family consists of 11 agonist and antagonist molecules that are involved in regulating inflammatory responses. IL-1α, a member of the IL-1 family, is an abundant cytokine that activates a cytokine network and affects local, or even systemic, inflammation, and anti-tumor immune responses. 5 It is generally accepted that IL-1α is involved in all phases of the malignant process in CRC, including colon homeostasis, inflammation, carcinogenesis, and invasion. 5
MicroRNAs (miRNAs) are small non-coding single-stranded RNA molecules ~22 nucleotides long that regulate human gene expression by combining with the 3′ untranslated region (UTR) of target messenger RNAs (mRNAs) 6 to interfere with the translation of mRNA and protein synthesis. 7 It has been established that genetic variations at crucial loci can influence miRNA–mRNA binding, which may affect the target gene expression and influence an individual’s risk of developing cancer. Evidence has indicated that rs3783553, an insertion/deletion polymorphism located in the 3′-UTR of the IL-1α gene, can alter the strength of miRNA-122 and miRNA-378 binding to the IL-1α 3′-UTR. 8 Subsequently, further studies have demonstrated that the rs3783553 polymorphism is associated with various types of cancer, including breast cancer, 9 cervical carcinoma, 10 and ovarian cancer.8,11
In the current study, we aimed to clarify the relationship between the IL-1α rs3783553 polymorphism, and CRC risk in a Chinese population. Our data indicated that patients harboring the “TTCA” insertion genotype had a significantly reduced risk of CRC. Furthermore, rs3783553 was associated with not only susceptibility to CRC, but also the clinical features of CRC, including the pathological type and clinical stage.
Materials and methods
Samples
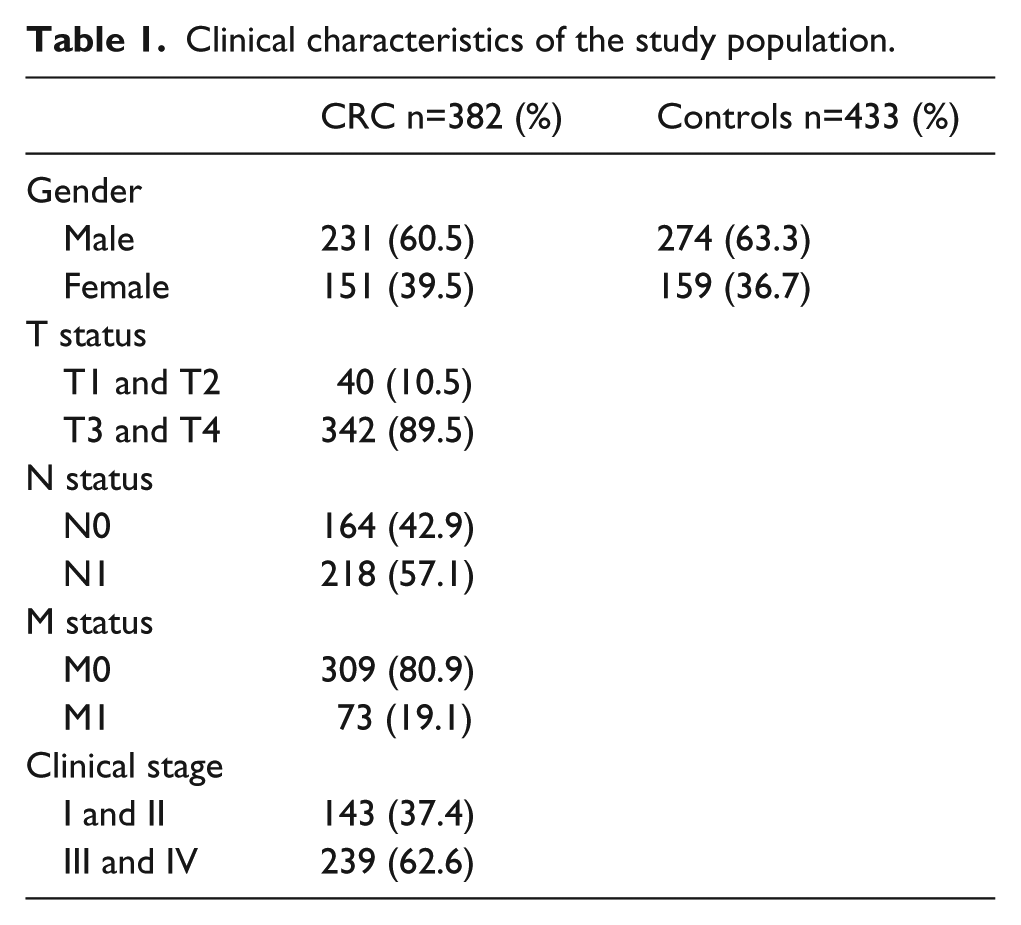
The study population consisted of 382 patients with CRC and 433 unrelated controls. The study was performed with the approval of the ethics committee of the West China Hospital of Sichuan University, and all subjects provided written informed consent. The diagnosis of CRC was confirmed by histopathological analysis. Clinical information, including gender, plus tumor, node, metastasis (TNM) status, and clinical stage, was obtained from surgical and pathological records. The basic characteristics of the study population are summarized in Table 1. The case group consisted of 231 males and 151 females. The controls were healthy individuals randomly selected from the same hospital; subjects with any personal or family history of CRC or other serious disease were excluded. The control group comprised 274 males and 159 females.
Clinical characteristics of the study population.
DNA extraction and genotyping
Peripheral venous blood samples were collected from each individual and stored at −20°C. Genomic DNA was extracted from 200 μL EDTA-anticoagulated peripheral blood samples with a commercial DNA kit (TIANGEN Biotech, Beijing, China) according to the manufacturer’s protocol. The sequences of the primers used for the genotyping of rs3783553 were 5′-ATTGGTCCGATCTTTGACTC-3′ and 5′-TGATAACAGTGGTCTCATGG-3′. 8 The PCR products were separated by 8% non-denaturing polyacrylamide gel electrophoresis and were analyzed by silver staining. Genotypes were determined by the numbers and sizes of the bands present in the gels. To verify the quality of the results, 20% of DNA samples were randomly selected to undergo a repeated assay, and 10% of the genotypes were confirmed by DNA sequencing, which showed that the results were 100% identical.
Statistical analysis
All data were analyzed using SNPstats (Free Software Foundation, Inc., Boston, MA, USA), assuming codominant, dominant, recessive, overdominant or log-additive genetic models. Allele and genotype frequencies were obtained by directed counting, and Hardy–Weinberg equilibrium (HWE) was evaluated by χ2 test. Odds ratios (ORs) and 95% confidence intervals (CI) were calculated to evaluate the effects of IL-1α rs3783553 genotype on CRC risk. Differences were considered significant at P<0.05.
Results
The rs3783553 polymorphism in the IL-1α gene was successfully genotyped in 382 patients with CRC and 433 controls. The distribution of genotypes fulfilled HWE in both cases and controls. The genotype and allele frequencies of the rs3783553 polymorphism are shown in Table 2. Significantly reduced CRC risk was observed to be associated with the insertion (I) allele (P<0.001, OR=0.68, 95% CI 0.55, 0.83). A significantly reduced risk of CRC was found to be associated with the insertion/insertion (I/I) genotype of rs3783553 in a codominant model compared with the deletion/deletion (D/D) and deletion/insertion (D/I) genotype (P=0.0002; OR=2.73; 95% CI 1.68, 4.44). In a dominant model, compared with the D/D genotype, I/I and D/I genotype individuals also had a reduced CRC risk (P=0.0097; OR=0.69; 95% CI 0.51, 0.92). Furthermore, in a recessive model, the I/I genotype was significantly associated with reduced CRC susceptibility (P=0.0001; OR=0.41; 95% CI 0.26, 0.65).
Genotype and allele frequencies of IL-1A gene rs3783553 polymorphism in CRC patients and control subjects.
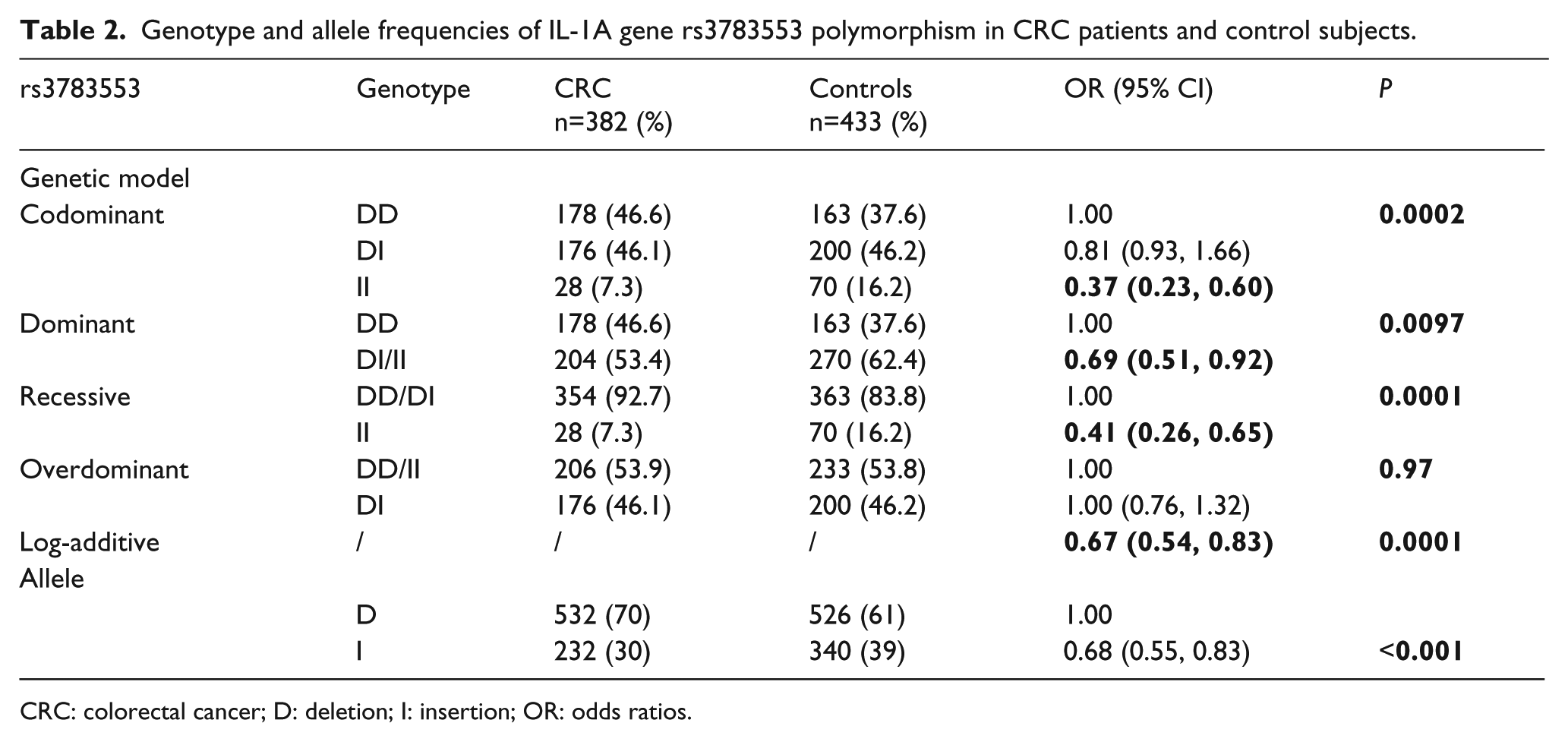
CRC: colorectal cancer; D: deletion; I: insertion; OR: odds ratios.
To further assess whether the rs3783553 polymorphism was associated with certain clinical features of CRC patients, we performed stratified analyses of genotype distribution in CRC patients with different ages, TNM stage, and clinical stage. As shown in Table 3, when considering the N status, a significant association between N1 status and the D/I and I/I genotypes was observed, compared with D/D genotype in a dominant model (P=0.001; OR=0.51; 95% CI 0.34, 0.77). Meanwhile, in a recessive model, the I/I genotype was associated with a reduced risk of lymph node metastasis compared with the D/D or D/I genotypes (P=0.018; OR=0.39; 95% CI 0.18, 0.87). The D/I and I/I genotypes were also significantly associated with clinical stage; D/I and I/I were increased in patients with clinical stage I/II (P<0.0001; OR=0.44; 95% CI 0.29, 0.68) compared with clinical stage III/IV in a dominant model. Furthermore, the D/I and I/I genotypes were higher in patients with M0 versus M1 status, thus indicating a significantly decreased risk of distant metastasis based on a dominant genetic model (P=0.037; OR=0.58; 95% CI 0.35, 0.97). Additionally, in a recessive model, the I/I genotype was associated with a reduced risk of developing stage III/IV CRC (P=0.008; OR=0.36; 95% CI 0.16, 0.79). An association between clinical stage and the heterozygous D/I genotype was identified in an overdominant model (P=0.018; OR=0.61; 95% CI 0.40, 0.92). Nevertheless, no association was identified between the genotypes of rs3783553 and other factors, including patient age or T status.
Association between the IL-1A rs3783553 polymorphism and clinical features of CRC patients.
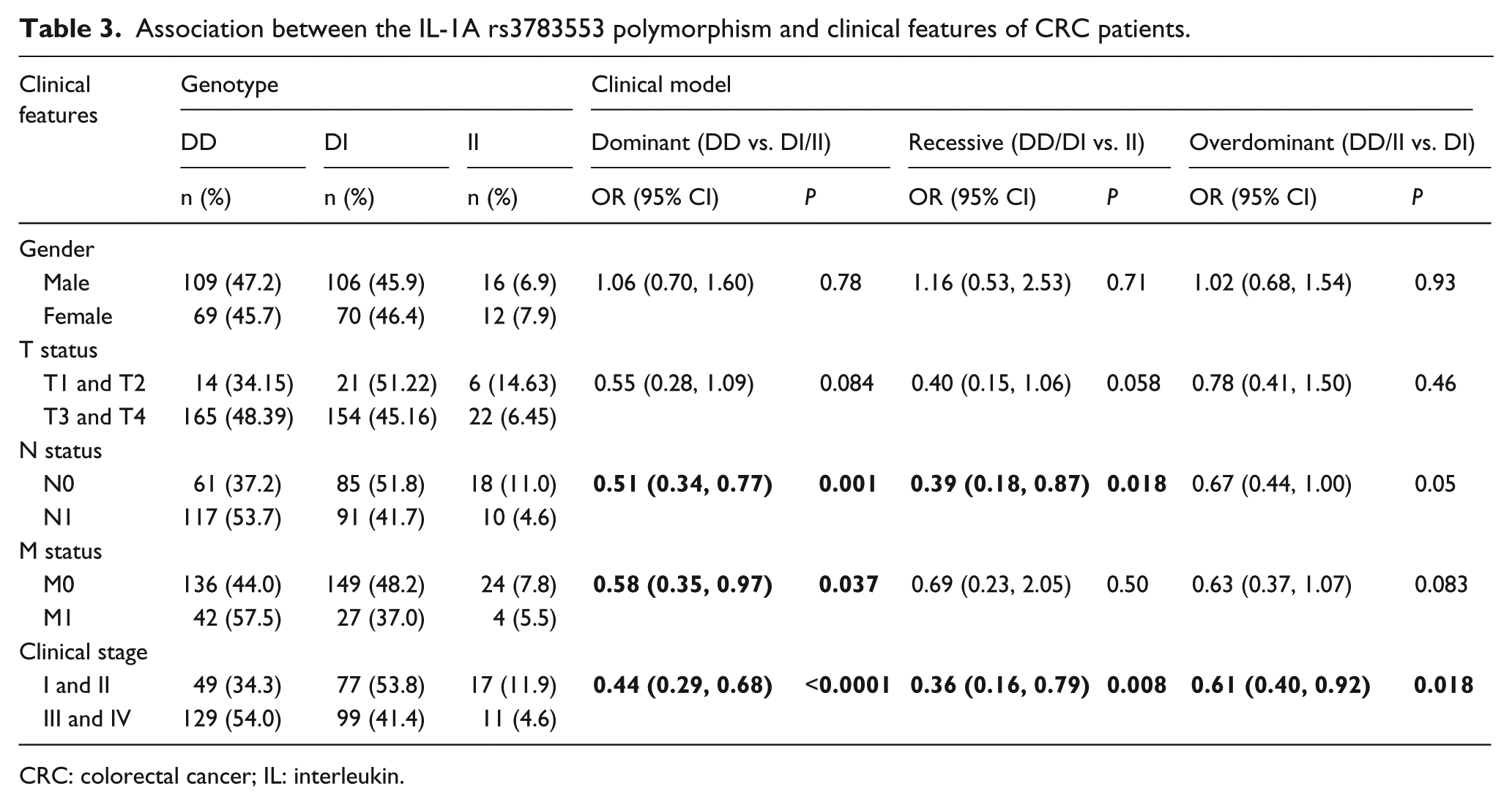
CRC: colorectal cancer; IL: interleukin.
Discussion
In the present study, we identified an association between the 4-base (TTCA/-) insertion/deletion polymorphism of rs3783553 in the IL-1α gene and CRC risk in a Chinese population. Our results suggested that the I allele of the rs3783553 polymorphism may be a crucial protective factor against CRC development. Stratification analyses also revealed associations between rs3783553 genotype and certain clinical features, including TNM and clinical stage. The example output from sequencing and genotyping assays of rs3783553 is shown in Figure 1.
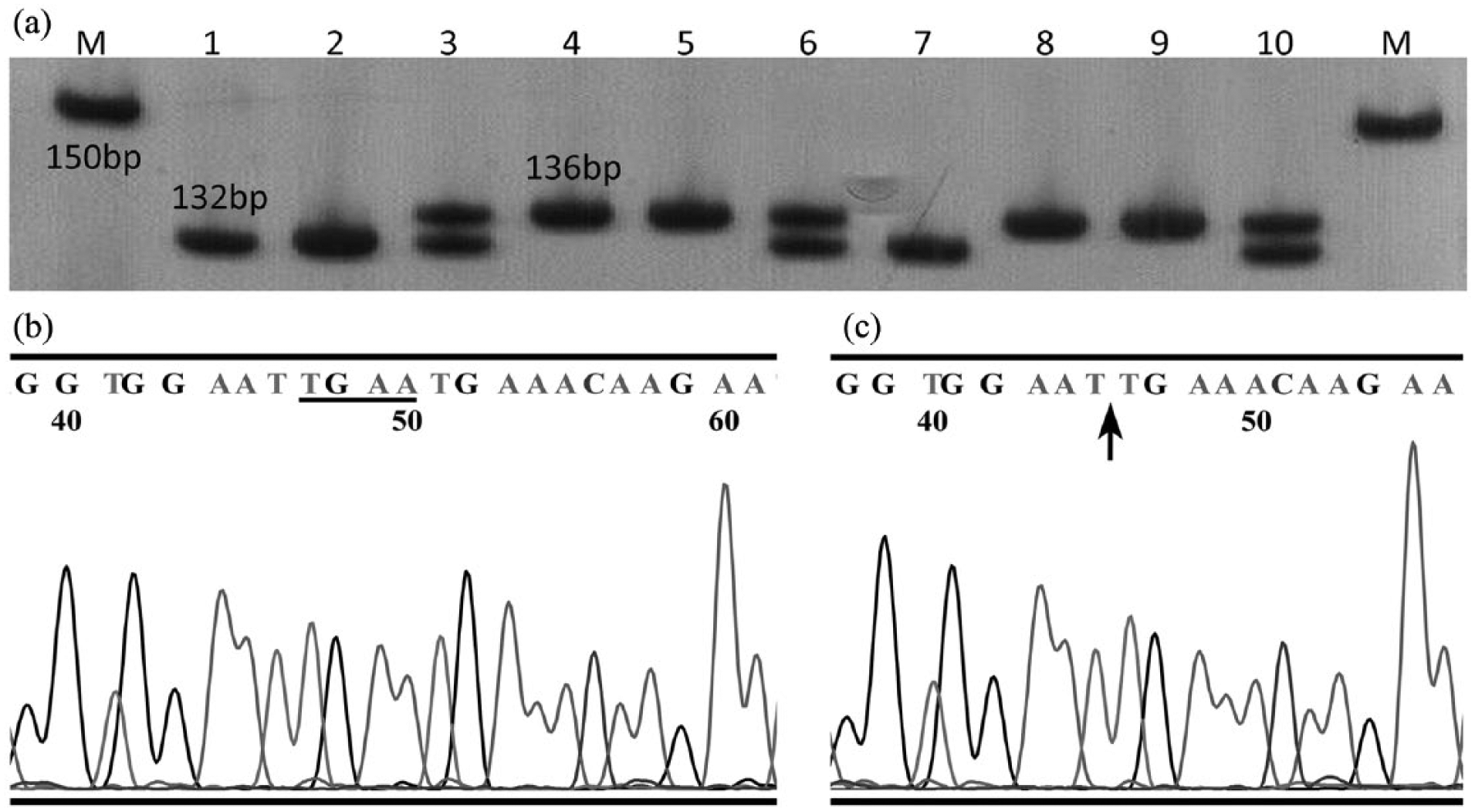
PAGE gel electrophoresis and sequencing results show the amplification pattern of rs3783553 (I/D) gene polymorphism. (a) Representative PAGE gel electrophoresis data. M represents the 150bp ladder. Lanes 4, 5, 8, and 9 show I/I genotype (136bp product); lanes 1, 2, and 7 show D/D genotype (132bp product); lanes 3, 6, and 10 show D/I genotype (132bp product and 136bp product). (b) and (c) The sequencing results of deletion and insertion allele, respectively. The underlined base pairs in (b) indicate the inserted reverse sequence (TGAA), and the arrow in (c) indicates the deletion allele.
Inflammation has pros and cons; although it is critical for host defense, the development of uncontrolled inflammation can lead to chronic immune-mediated inflammatory diseases or cancers. 12 IL-1 is a major upstream “alarm” pro-inflammatory cytokine, and is produced and secreted by malignant cells and infiltrating myeloid cells, as well as by normal colon epithelial cells and other stromal cells. 11 IL-1α, a member of the IL-1 family, has been reported to be involved in colon homeostasis, inflammation, and carcinogenesis, and in the invasion of CRC.5,13 Recognized as an immune mediator, IL-1α serves multiple roles in the development of cancer. A full understanding of the roles of IL-1α in these processes may contribute to the development of novel IL-1-modulating approaches.
Over recent years, miRNAs and long non-coding RNAs (lncRNAs) have been widely studied in various types of cancer, including CRC.14-19 Insertion/deletion polymorphisms may affect target genes expression through regulating the interactions between miRNAs or lncRNAs and target genes.7,19 Previous studies have shown that high IL-1 concentrations within the tumor microenvironment are associated with a more virulent tumor phenotype, and IL-1 upregulation has been observed in breast, colon, lung, and head and neck cancers, and melanomas. 20 Recently, miRNAs have emerged as important regulators responsible for altering IL-1α expression. For example, it was shown that an insertion/deletion polymorphism, rs3783553, disrupted a binding site for miRNA-122 and miRNA-378, thereby increasing the transcription of IL-1α. Higher IL-1α expression was observed in the liver tissue of hepatocellular carcinoma patients harboring a homozygous TTCA insertion genotype. 8 In our study, we analyzed the association between the rs3783553 insertion/deletion polymorphism in the predicted miRNA target site of IL-α gene, and the risk of CRC and clinical outcomes. We identified that significantly decreased CRC risk was associated with the IL-1α rs3783553 I/I genotype and the I allele. The results suggest that this polymorphism has a potential role in the progression and prognosis of CRC. Our results are in agreement with certain previous studies, which reported that the risks of gastric cancer, 21 epithelial ovarian cancer, 22 and nasopharyngeal carcinoma 23 were reduced by this polymorphism. In light of the crucial role of IL-1α in CRC tumorigenesis, we assumed that genetic variations at rs3783553 may affect the function of IL-1α and eventually influence the tumor growth, lymph node metastasis, and clinical stage.
In summary, we have demonstrated that the rs3783553 TTCA/- polymorphism influences not only the genetic susceptibility to CRC, but also the TNM and clinical stage of patients with CRC in a Chinese population. This risk may be due to its regulation of the expression of IL-1α, which may contribute to the pathogenesis of CRC. Collectively, these findings suggest that IL-1α rs3783553 has potential applications in patient screening, targeted drug administration, and guiding individualized clinical treatment.
Footnotes
Author contributions
Qizhao Ma and Zhigang Mao contributed equally to this work.
Declaration of conflicting interest
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by research grants from the Natural Science Foundation of China (No. 81471551, 81630054 and 81571861).
