Abstract
Background
Death-associated protein kinase (DAPK) has a strong function of tumor suppression involving apoptosis regulation, autophagy, and metastasis inhibition. Hypermethylation of CpG islands in DAPK gene promoter region is one of the important ways to inactivate this tumor suppressor gene, which might promote lung carcinogenesis. However, the clinicopathological significance of the DAPK promoter hypermethylation in lung cancer remains unclear. In this study, we performed a meta-analysis trying to estimate the clinicopathological significance of DAPK promoter hypermethylation in non-small cell lung cancer (NSCLC).
Methods
A detailed literature search for publications relevant to DAPK gene promoter methylation and NSCLC was made in PubMed, Embase, Cochrane Library, Web of Science, China National Knowledge Infrastructure, CSTJ, Wanfang databases, and SinoMed (CBM). The random-effects model and fixed-effects model were utilized to pool the relative ratio based on the heterogeneity test in the meta-analysis.
Results
A total of 41 studies with 3348 patients were included. The frequency of DAPK methylation was significantly higher in NSCLC than in non-malignant control (odds ratio (OR) = 6.88, 95% confidence interval (CI): 4.17–11.35, P < 0.00001). The pooled results also showed that DAPK gene promoter hypermethylation was significantly associated with poor prognosis for overall survival in patients with NSCLC (hazard ratio: 1.23, 95% CI:1.01–1.52, P = 0.04). Moreover, DAPK gene promoter hypermethylation was significantly associated with squamous cell carcinoma (OR: 1.25, 95% CI: 1.01–1.54, P = 0.04) and smoking behavior (OR: 1.42, 95% CI: 1.04–1.93, P = 0.03) but not with TNM stage, tumor differentiation, age, or gender.
Conclusion
DAPK promoter hypermethylation might be a candidate diagnostic and prognostic tumor marker for NSCLC.
Introduction
Globally, lung cancer is the leading cause of cancer incidence and mortality in all kinds of tumors. 1 It has been estimated that nearly 1.6 million people died of lung cancer in 2012, accounting for approximately 20% of all cancer deaths that year.1,2 Despite recent improvements in treatment, the prognosis of lung cancer remains poor, with a 5-year overall survival (OS) rate of 19%. 3 Of all primary pulmonary malignancies, approximately 85% of cases are non-small cell lung cancer (NSCLC) including adenocarcinoma (ADC), squamous cell carcinoma (SCC), adenosquamous carcinoma, and large cell carcinoma. It is now well established that genetic mutations and epigenetic modifications play an important role in lung cancer development.4,5 DNA methylation is one of the important molecular mechanisms in epigenetics. In contrast to the feature of genome-wide hypomethylation in malignancies, hypermethylation of CpG islands in tumor suppressor genes (TSGs) promoter regions, leading to inactivation of TSGs, were common in various cancers.6–8 Death-associated protein kinase (DAPK) belongs to the superfamily of calcium/calmodulin (Ca+2/CaM) regulated serine/threonine kinases (STKs). It is encoded by DAPK gene located on chromosome 9q34.1 and known for its role as a strong tumor suppressor, a regulator of apoptosis and autophagy and an inhibitor of metastasis.9,10 Several studies indicated that DAPK genes’ promoter methylation occurred frequently in NSCLC, and has a tight relationship with occurrence and development of NSCLC.11–15 However, the results were varied from different studies, and the clinicopathological significance of DAPK promoter hypermethylation remains unclear. In this study, 41 eligible studies were pooled, and we performed a meta-analysis trying to provide a more accurate estimate of the clinicopathological significance of DAPK promoter hypermethylation in NSCLC, including age, gender, smoking status, histological type, differentiation, tumor node metastasis (TNM) stage, and prognostic outcomes.
Materials and methods
Search strategy
PubMed, Embase, Cochrane Library, Web of Science, China National Knowledge Infrastructure, CSTJ, Wanfang databases and SinoMed (CBM) were reviewed by two reviewers comprehensively till April 10, 2021. Studies relevant to DAPK gene promoter methylation and NSCLC were identified using the Medical Subject Headings (MeSH) combined with Free-Word index term of: “Lung Neoplasms”, OR “Pulmonary Neoplasms”, OR “Neoplasms, Lung”, “Lung Neoplasm”, OR “Neoplasm, Lung”, OR “Neoplasms, Pulmonary”, OR “Neoplasm, Pulmonary”, OR “Pulmonary Neoplasm”, OR “Lung Cancer”, OR “Cancer, Lung”, OR “Cancers, Lung”, OR “Lung Cancers”, OR “Pulmonary Cancer”, OR “Cancer, Pulmonary”, OR “Cancers, Pulmonary”, OR “Pulmonary Cancers”, OR “Cancer of the Lung”, OR “Cancer of Lung”, OR “NSCLC”, OR “LC”; AND “death associated protein kinase methylation”, OR “death-associated protein kinase methylation”, OR “DAPK methylation”, OR “death-associated protein kinase”, OR “DAPK”, OR “death-associated protein kinase”, OR “methylation”.
Inclusion and exclusion criteria
The included criteria were as follows: (a) case-control or cohort studies related to DAPK gene promoter methylation and NSCLC; (b) patients in the case group included in each study were confirmed to be NSCLC by cytology or pathology; and (c) research published in English or Chinese. The following exclusion criteria were used: (a) studies published in languages other than Chinese or English; (b) articles with insufficient data; (c) the same population or repeat publication; (d) degree theses, reviews, conference abstracts, editorials, letters, case reports, dissertation and expert opinion; and (e) the studies utilized cell lines or basic research on animal experiments. (For the selection process, see Figure 1) After evaluation, 41 articles fulfilled the entry criteria of this meta-analysis, and the detailed information of these final included articles are listed in Table 1.
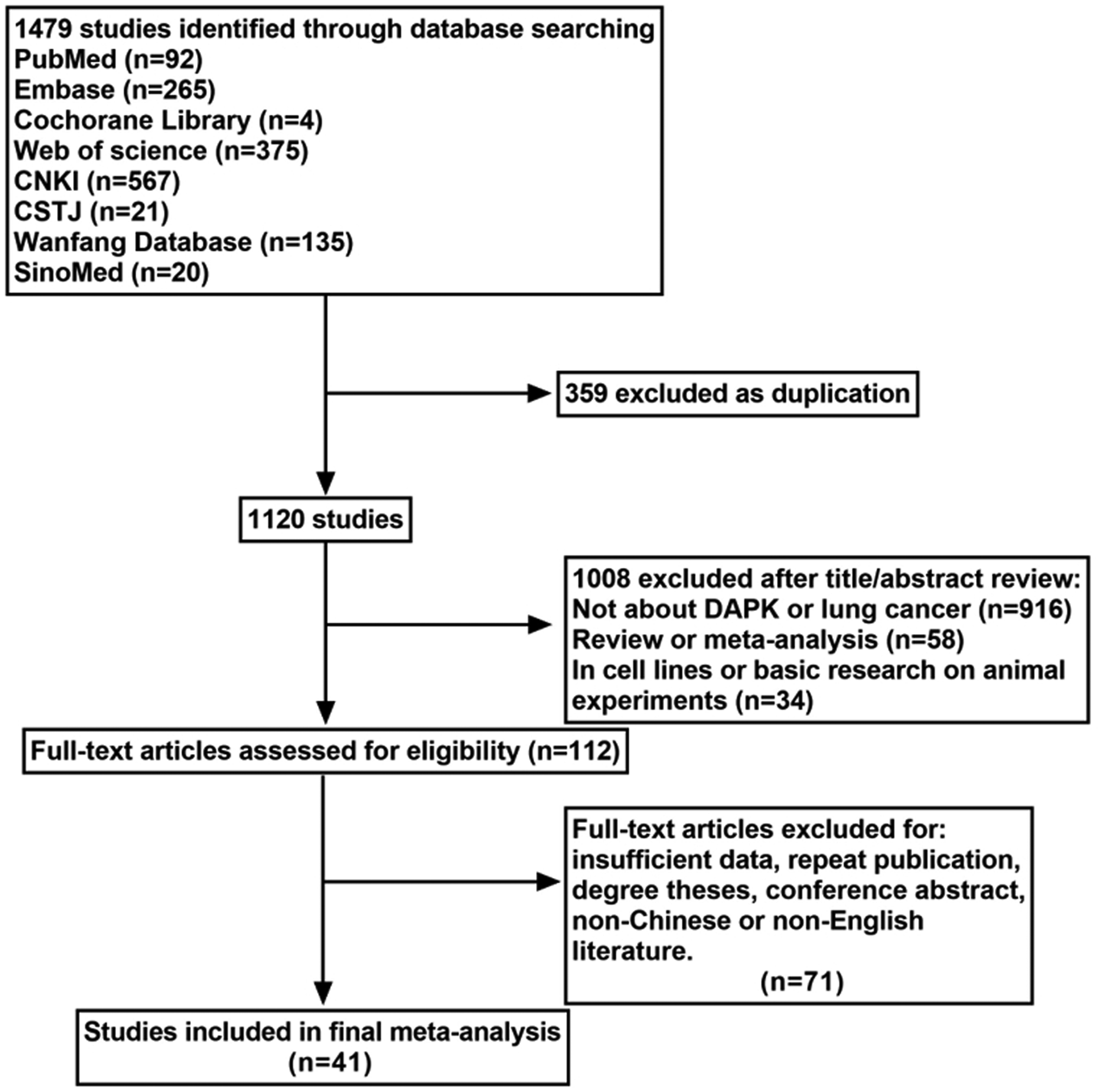
Flow chart of literature search and study selection.
The main characteristics of included studies.
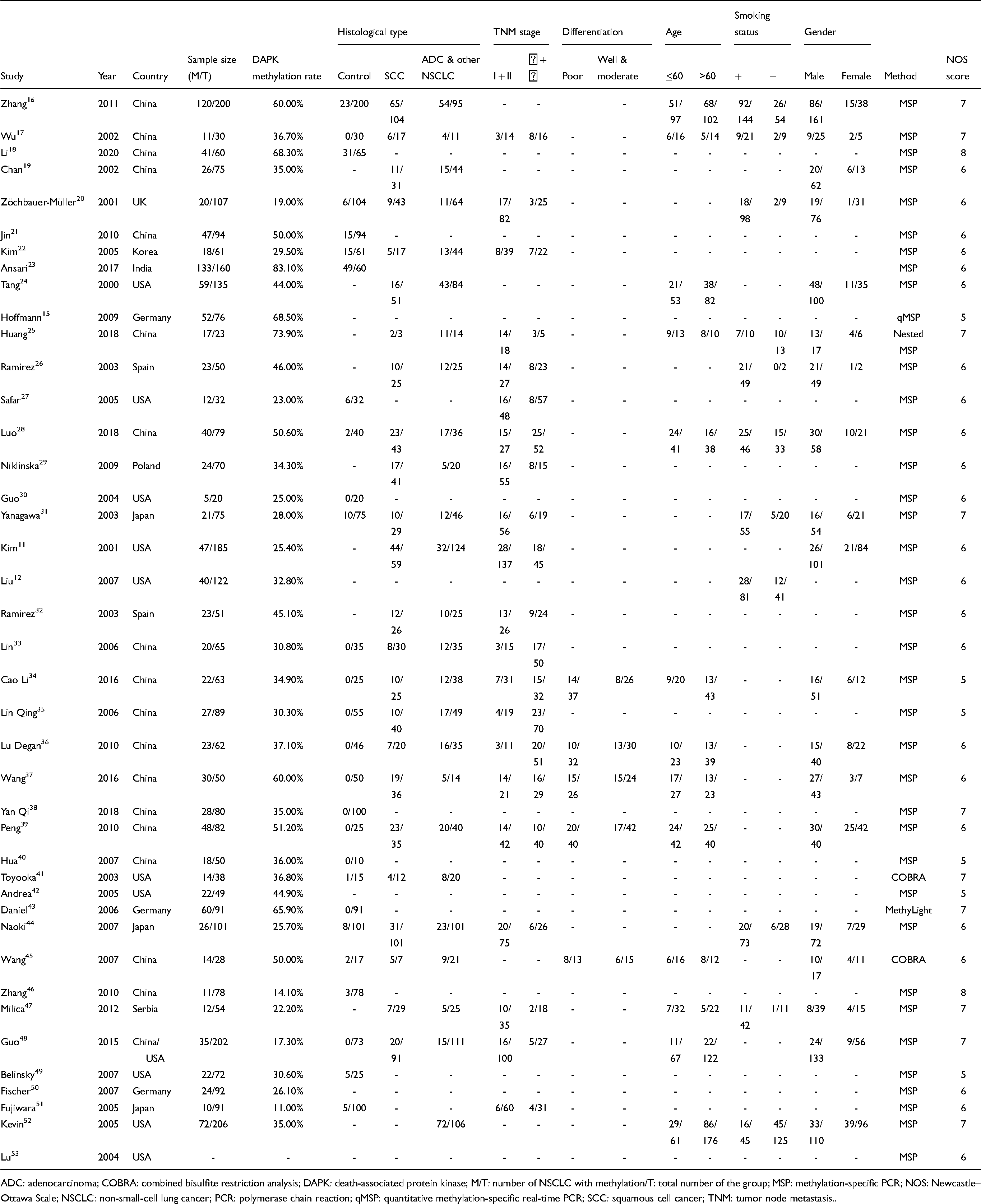
ADC: adenocarcinoma; COBRA: combined bisulfite restriction analysis; DAPK: death-associated protein kinase; M/T: number of NSCLC with methylation/T: total number of the group; MSP: methylation-specific PCR; NOS: Newcastle–Ottawa Scale; NSCLC: non-small-cell lung cancer; PCR: polymerase chain reaction; qMSP: quantitative methylation-specific real-time PCR; SCC: squamous cell cancer; TNM: tumor node metastasis..
Data extraction
The data extraction followed the procedure described by the published literature. Two investigators independently extracted data from eligible studies. Disagreements were resolved by discussion and consensus. Two investigators reviewed all of the articles that met the inclusion and exclusion criteria. The following information was recorded for each study: the first author’s name, country, year of publication, sample source, methylation detection method, number of cases, DAPK methylation status, and clinicopathological parameters including TNM stage, differentiation, age, gender, smoking status, and data of overall survival (OS). The study characteristics and clinicopathological parameters are summarized in Table 1. Heterogeneity of investigation was evaluated to determine whether the data from the various studies could be analyzed in a meta-analysis.
Quality assessment
We used the Newcastle–Ottawa Scale (NOS) to assess the quality of each study by two authors. The scores included three parts: the selectivity of patients (0–4), comparability of groups (0–2), and assessment of outcome (0–3). Studies with scores > 5 were included in final meta-analysis.
Statistical analysis
Data were analyzed using the RevMan 5.3 software and STATA14.0. When the hazard ratio (HR) values of OS were not directly reported, we obtained additional data from the original authors. When the request was not answered, the HR values were extracted from Kaplan–Meier curves by the Engauge Digitizer 4.1 software. Heterogeneity was calculated by the chi-squared test and I -squared statistics. If I2 ≥ 50% or P ≤ 0.01 was established, the results were pooled through a random-effects model; otherwise, a fixed-effects model was selected. In addition, sensitivity analysis was used to minimize the influence of heterogeneity. Publication bias was estimated qualitatively using Begg's and Egger's tests with funnel plots. If Begg's and Egger's results indicated that publication bias exists, the trim and fill method was used to examine the sensitivity of the result. A difference was considered statistically significant if two-sided P < 0.05.
Results
Study characteristics
The initial search information retrieved 1479 articles. According to the selection criteria, 41 studies with 3348 patients were eventually included in the meta-analysis (Figure 1). The basic characteristics of the eligible studies are shown in Table 1.
DAPK methylation and clinicopathological features of NSCLC
The total number of NSCLC from 27 studies is 2160. And 876 of them were with DAPK methylation, the methylation rate was 40.6%. The frequency of DAPK methylation was significantly higher in NSCLC than in non-malignant controls; and the pooled odds ratio (OR) was 6.88 with 95% confidence interval (CI) 4.17–11.35, Z = 7.55, P < 0.00001, I2 = 74% (Figure 2(a)). We also observed the similar effect when divided the included populations into two subgroups according to the geographic origin. Figure 2(a) showed that the frequency of DAPK methylation was significantly higher in NSCLC than in non-malignant controls both in European and American countries (OR: 8.71; 95% CI: 1.76–43.09; Z = 2.65, P < 0.001, I2 = 78%) and in Asian countries (OR: 6.77; 95% CI: 3.94–11.66; Z = 6.91, P < 0.00001, I2 = 74%). When stratified by SCC, ADC, and other NSCLC, the frequency of DAPK hypermethylation was significantly higher in SCC; and the pooled OR was 1.25 with 95% CI: 1.01–1.54, Z = 2.07, P = 0.04, I2 = 0% (Figure 2(b)). As shown in Figure 2(c), DAPK hypermethylation was associated with smoking behavior in NSCLC (OR: 1.42; 95% CI: 1.04–1.93; Z = 2.20, P = 0.03, I2 = 0%). In addition, the relevant results showed that DAPK hypermethylation was not significantly associated with TNM stage (Stage I + II vs. Stage III + IV, OR: 0.94; 95% CI: 0.72–1.23; Z = 0.43, P = 0.67, I2 = 13%), differentiation (poor vs. well and moderate, OR: 1.13; 95% CI: 0.70–1.81; Z = 0.50, P = 0.62, I2 = 0%), age (≤60 vs. >60, OR: 0.90; 95% CI: 0.71–1.15; Z = 0.83, P = 0.41, I2 = 0%), and gender (male vs. female, OR: 1.18; 95% CI: 0.94–1.49; Z = 1.44, P = 0.15, I2 = 0%)(Figure 2(d)–(g)). Analysis of the relationship between DAPK hypermethylation and OS showed that DAPK hypermethylation was significantly associated with poor prognosis in patients with NSCLC (HR: 1.23; 95% CI: 1.01–1.52; Z = 2.01, P = 0.04, I2 = 23% (Figure 2(h)).
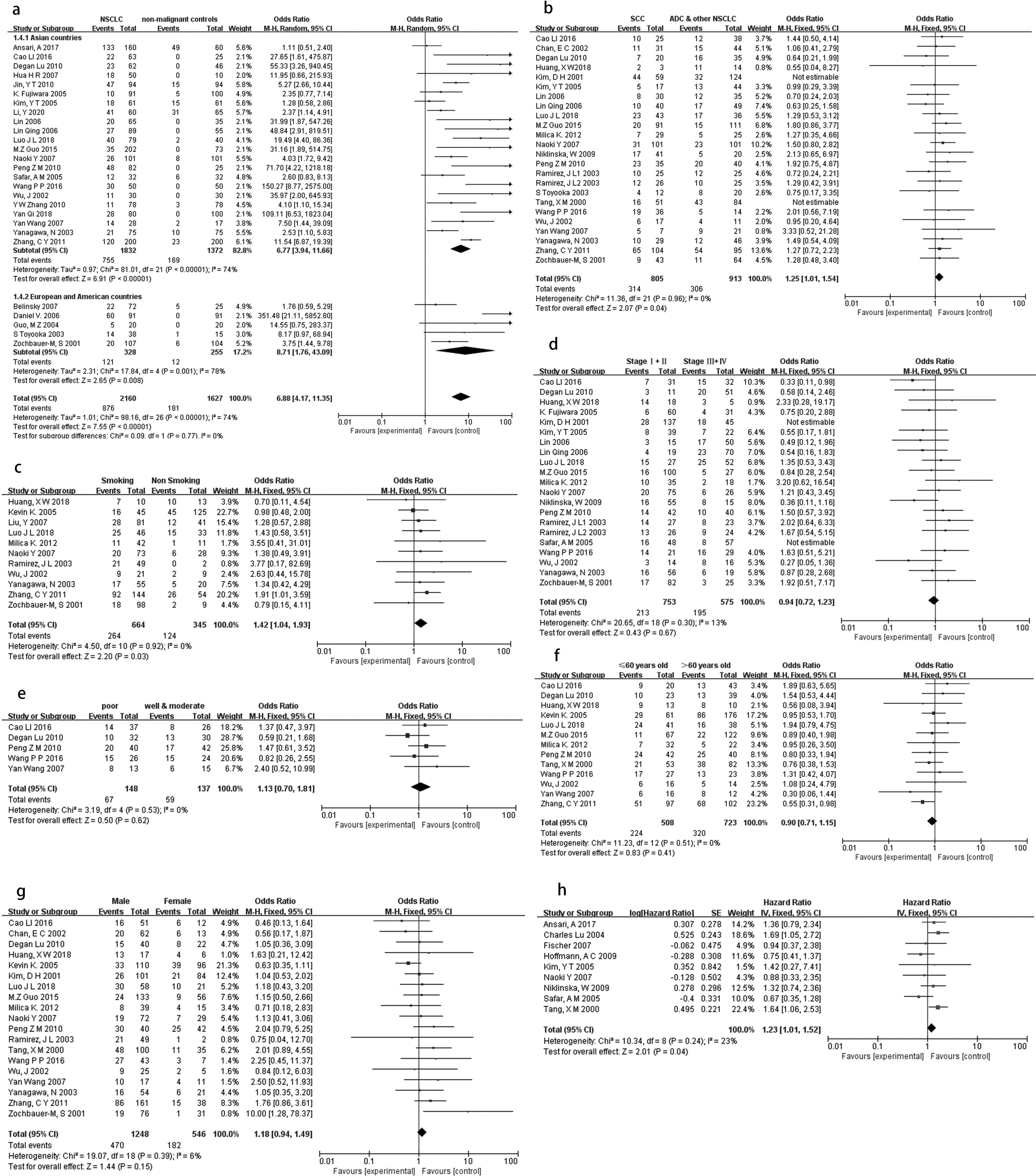
Forest plots for the association between DAPK methylation and in cancer patients: (a) NSCLC vs. non-malignant controls; (b) SCC vs. ADC and other NSCLC; (c) smoking vs. non-smoking; (d) TNM stage (Stage I + II vs. Stage III + Iv;); (e) differentiation (poor vs. well and moderate); (f) age (≤60 vs. >60); (g) gender (male vs. female); (h) forest plots for the association between DAPK methylation and OS.
Publication bias
This meta-analysis adopted Begg's test and Egger's test to evaluate publication bias. All the results except DAPK methylation in NSCLC versus non-malignant controls showed no significant publication bias as Begg's test showed pr > |z| = 0.05 or Egger's test showed P > |t| = 0.05 (Figure 3).
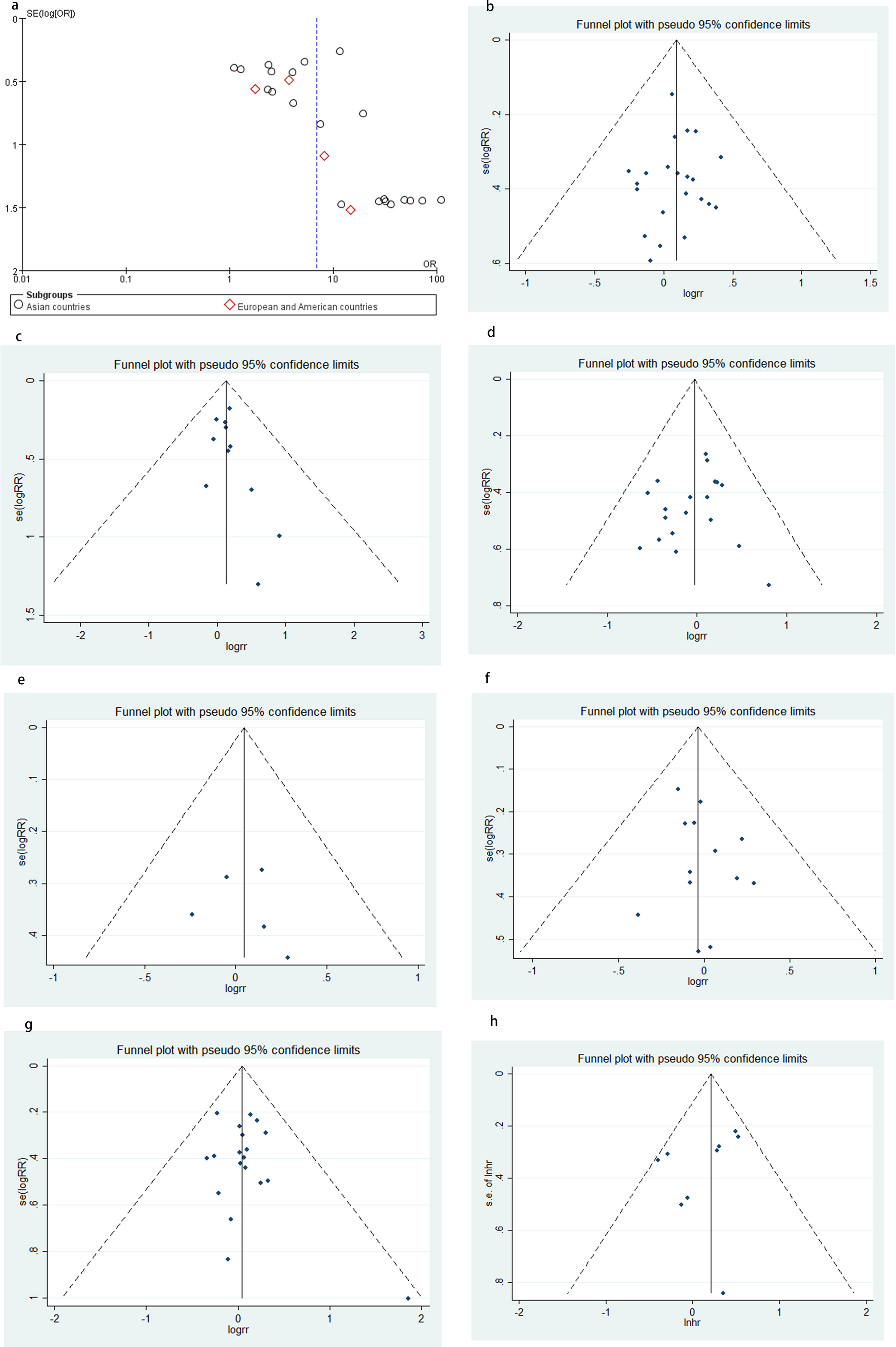
Funnel plot for publication bias. (a) DAPK methylation in NSCLC and non-malignant controls; (b) SCC vs. ADC and other NSCLC; (c) smoking vs. non-smoking; (d) TNM stage (Stage I + II vs. Stage III + IV); (e) differentiation (poor vs. well & moderate); (f) age (≤60 vs. >60); (g) gender (male vs. female); (h) funnel plots for the association between DAPK methylation and OS.
Discussion
Epigenetic and genetic changes have become established in recent years as being one of the most important molecular mechanisms of human tumors. Plus, DNA methylation constitutes an important molecular mechanism of epigenetics. CpG islands are 200–4000 bp stretches of DNA that have a significantly higher concentration of CpG dinucleotides than the bulk of the genome. Whereas 70–80% of all CpG dinucleotides in the human genome are methylated, CpG islands located in promoter region and exon region large remain unmethylated.54,55 Hypermethylation of the CpG islands affects certain tumor suppressor genes involved in cell cycle (p16INK4a, p15INK4b, Rb, p14ARF), DNA repair (BRCA1, hMLH1, MGMT), carcinogen-metabolism (GSTP1), cell-adherence (CDH1, CDH13), apoptosis (DAPK, TMS1), etc., which has been proved to be closely related to the occurrence and development of a variety of cancers. 56 With a strong function of tumor suppression involving apoptosis regulation, autophagy and metastasis inhibition, DAPK has received wide appreciation by researchers. It is established that hypermethylation of CpG islands in the DAPK encoding gene promoter region has a tight relationship with the occurrence and development of NSCLC. The methylation rate of the included studies varied from 11.0% to 83.1%. The present study pooled 27 studies including 2160 NSCLC and 1627 non-malignant controls, and 40.6% (876/2160) were with DAPK gene promoter hypermethylation. Therefore, DAPK promoter hypermethylation indicated high risk of NSCLC compared to non-malignant control (the pooled total OR: 6.88, 95% CI: 4.17–11.35, P < 0.00001). A similar effect was observed in the populations from European, American, and Asian countries (Figure 2(a)). However, due to the publication bias of these 27 included studies, the difference of DAPK promoter hypermethylation between NSCLC and non-malignant controls might be exaggerated to some extent. Based on this evidence, we recommend DAPK hypermethylation to be a candidate tumor marker for lung cancer diagnosis, especially for NSCLC. Moreover, we found that DAPK promoter hypermethylation correlated significantly with SCC (P < 0.05, Figure 2(b)) and smoking behavior in patients with NSCLC (P < 0.05, Figure 2(c)). However, we did not find a significant association of DAPK promoter hypermethylation with differentiation, age, and gender.
Since the first integration of the descriptors of tumor size, nodal status, and metastases by Denoix, the TNM staging system was widely used as an important guideline for treatment and prognostic prediction in various cancers. 57 With our advanced understanding of molecular pathways on tumor biology, an increasing number of research focused on identifying molecular prognostic factors. A 14-gene expression assay has proven to improve prognostic accuracy beyond the National Comprehensive Cancer Network criteria for stage I high-risk tumors, and differentiated low-risk, intermediate-risk, and high-risk patients within all disease stages. 58 Subsequently, a novel staging system, the TNMB (with the B denoting biology) system was the develop and validate to significantly improve prediction of OS greater than adoption of the seventh or eighth edition staging systems. This might enable us to better differentiate high risk patients from low-risk patients, which might help to select appropriate early intervention. 59 Buckingham et al. suggested that hypermethylation of a specific promoter cytosine in the death-associated kinase gene, DAPK was associated with shorter time to recurrence in surgically treated stage I and II NSCLC patients. 13 Lu et al. showed that DAPK promoter methylation is a significant negative prognostic factor for disease-specific survival in resected stage I NSCLC. 53 In this meta-analysis, we confirmed that DAPK gene promoter hypermethylation was significantly associated with poor prognosis for OS in patients with NSCLC (HR: 1.23, 95% CI: 1.01–1.52, P = 0.04), which makes it possible for DAPK gene promoter hypermethylation to be a prognostic predictive factor.
All the included literature used the assay based on a standard sodium bisulfite DNA treatment to assess DAPK gene methylation status, including methylation-specific real-time PCR (MSP), 60 quantitative (q)MSP, nested MSP, combined bisulfite restriction analysis (COBRA), 61 and MethyLight. 62 These technologies entail initial modification of DNA by sodium bisulfite, converting all unmethylated, but not methylated, cytosines to uracil, and subsequent amplification with primers specific for methylated versus unmethylated DNA. Thus, primers specificity is the most critical factor to avoid a false positive result and to ensure accuracy. Also, incomplete bisulfite conversion could have a negative impact on the experimental results. Most of the included literature used MSP as the detection assay, and the high enough methylation frequency of CpG sites in CpG islands renders this technique extremely sensitive for such regions. However, MSP theoretically only reflects the average methylation level of the entire regions, for the reason that not all CpG sites in CpG islands were completely methylated. Therefore, this method is mainly used for qualitative or semi-quantitative research. 60 Although quantitative techniques, such as qMSP, COBRA, and MethyLight, have been used in individual studies, the impact of the above limitations of these sodium bisulfite treatment-based technologies on the pooled results cannot be ignored.
There are still many limitations in our study: First, Begg's test and Egger's test showed that the comparison of DAPK methylation in NSCLC versus non-malignant controls existed significant publication bias, which might lead to exaggerated interpretation of the frequency of DAPK hypermethylation in NSCLC. In the future, we need to adopt the counterpart publications indicated in the published bias graph adjusted by the trim and fill method. Second, the majority of samples included in the analysis were serum and lung tissue. One of the original studies used pleural fluid samples to detect DAPK gene promoter methylation, while two of the original studies used sputum samples, which may have increased heterogeneity and had a subsequent influence on the pooled results. Third, although studies included in this meta-analysis were all ≥ 5 stars, and 35 out of 41 publications were of high quality (NOS ≥ 6 stars), most of included studies were retrospective, which means selection bias is inevitable. Therefore, further prospective research is warranted to investigate the clinical significance of DAPK promoter hypermethylation.
Conclusion
DAPK gene promoter methylation occurred frequently in NSCLC, and has a tight relationship with occurrence and development of it. DAPK promoter hypermethylation might be a candidate diagnostic and prognostic tumor marker for NSCLC.
Footnotes
Acknowledgments
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship and/or publication of this article.
Author contributions
Zhimao Chen and Yu Fan contributed equally to this work.
