Abstract
Background and aims
Oral cancer is the second most common cancer in men and accounts for 50%-70% of the total cancer mortality in India. PTEN is a tumor suppressor gene that plays a critical role in controlling cell growth and survival. Promoter hypermethylation of the PTEN gene has been reported in many tumors. However, little is known about the association between promoter methylation and oral squamous cell carcinoma (OSCC). Therefore, we aimed to detect the role of PTEN hypermethylation in OSCC patients in the Indian population.
Methods
Genomic DNA was isolated from 100 fresh oral tumor specimens and was subjected to bisulfite conversion. Methylation-specific PCR was employed on the converted DNA to investigate the methylation status.
Results
Of the total cases examined for PTEN promoter methylation we found that 35% were positive and 65% were negative. When evaluated in connection with tumor differentiation it was found that 82% of poorly differentiated, 24% of moderately differentiated and 32% of well differentiated OSCC samples were methylated. Methylation was further correlated with patient age, sex and tumor grade. Interestingly, we found that patient age and grade of differentiation were significantly associated with PTEN promoter methylation (p=0.05 and 0.0019, respectively) while sex was not (p=0.9).
Conclusions
The present study underlines the importance of PTEN hypermethylation among Indian OSCC patients.
Introduction
Oral cancer consistently ranks as 1 of the 10 most frequently diagnosed cancers in the world (1) with over 363,000 cases reported annually worldwide and a mortality rate of about 50% (2, 3). In India, head and neck squamous cell carcinoma accounts for 30%-40% of cancers at all sites, out of which 9.4% is oral cancer (4). As the incidence of oral cancer is continuously increasing, the disease has gradually become an important public health issue. Although oral cancer sites are readily amenable to clinical examination, the lack of a suitable molecular marker has proved to be a major handicap for early detection and risk evaluation of this disease. This is very distinctly reflected by the fact that more than 50% of all patients with oral squamous cell carcinoma (OSCC) have advanced disease at the time of diagnosis (5, 6).
PTEN is a tumor suppressor gene located on chromosome 10q23.3 (7). PTEN downregulates the signaling of Akt, which suppresses apoptosis and promotes cell survival (8). It induces G1 cell cycle arrest (9) and inhibits focal cell adhesion, spreading, and migration by dephosphorylating focal adhesion kinase (10). Gene silencing by genetic and epigenetic alterations of the PTEN gene has been reported in a subset of cancers. Involvement of PTEN promoter methylation was reported in breast cancer (10), cervical cancer (11), endometrial cancer (12), colorectal cancer (13) and glioblastoma (14); however, no such study has evaluated the role of the PTEN tumor suppressor gene in OSCC among the Indian population. The correlation of disease progression with promoter hypermethylation seems an interesting hypothesis and we therefore aimed to analyze PTEN promoter methylation in OSCC samples and normal tissues of the same patients.
Materials and Methods
Patients
A total of 100 primary OSCC tissues were obtained from biopsy or surgical specimens. The normal tissues taken from the same OSCC patients were blood samples. The patients were diagnosed and treated at the L.N.J.P. Hospital at Maulana Azad Medical College, New Delhi, in the period 2007 to 2011. The Institutional Review Committee's approval was obtained and all patients gave their consent to the collection of tumor tissue and blood. The histological grades of the tumors were classified according to the World Health Organization (WHO) criteria (15).
Specimen characteristics
Fresh tumor tissues as well as control (blood) samples were collected and preserved at −20°C for further processing.
Assay methods
DNA extraction
DNA was extracted from carcinoma tissues and blood as previously described (16). In brief, DNA was isolated by SDS/proteinase K treatment followed by phenol-chloroform extraction and ethanol precipitation and finally dissolved and stored in Tris EDTA buffer.
Bisulfite modification and methylation-specific PCR assay
Genomic DNA from the oral carcinoma and normal specimens was subjected to bisulfite treatment using the EZ DNA Methylation-Gold kit (Zymo Research, USA) according to the manufacturer's protocol; here methylated DNA is protected, and unmethylated cytosine is converted into uracil. The modified DNA was used as a template for methylation-specific PCR using primers specific for either the methylated or modified unmethylated sequences. Appropriate negative and positive controls were included in each PCR reaction. The primer sequences used to amplify the unmethylated product were 5′-TATTAGTTTGGGGATTTTTTTTTTGT-3′ (sense) and 5′-CCCAACCCTTCCTACACCACA-3′ (antisense), while for the methylated product the sequences were 5′-GTTTGGTTATTTTTTTTTCGC-3′ (sense) and 5′-AACCCTTCCTACGCCGCG-3′ (antisense), which amplify 173-bp and 155-bp amplicons, respectively. The MethPrimer online utility was used to define the CpG islands by importing the 5′ flanking region of the PTEN gene sequence from the Ensembl genome browser to MethPrimer. After defining the CpG islands, methylation-specific PCR primers were designed. PCR reactions were performed in a 25-μL reaction volume of 1× PCR buffer containing 1 mM MgCl2, 200 μM deoxynucleoside triphosphates (dNTPs), 0.4 μM of each PCR primer, 0.75 unit of Taq polymerase, and approximately 25 ng of bisulfite-modified DNA. Reactions were hot-started at 95°C for 5 minutes. This was followed by 33 cycles at 95°C for 45 seconds, 57°C for 30 seconds and 72°C for 30 seconds, followed in turn by 10-minute extension at 72°C in a Bio-Rad Thermocycler. Methylation-negative bacterial strain DNA (pUC19) artificially methylated by Sss/methyltransferase (Zymo Research) followed by bisulfite treatment was used as positive control for the methylated allele. pUC19 DNA without pretreatment was used as negative control. Water blank was used as negative control with every PCR amplification.
Bisulfite conversion and promoter region amplification
The promoter region was amplified by 2 rounds of hot-start PCR, which was carried out in 25 μL reaction mixture containing 1 μL bisulfite-modified DNA, 2.5 μL PCR buffer containing 1 mM MgCl2, 0.5 μL dNTPs (200 μM dNTPs each), and 1 μL (0.4 μM) forward and reverse primers; 0.5 μL Sigma Taq polymerase was added after we placed the tubes into the block. PCR was performed in a Bio-Rad Mastercycler gradient machine. Reactions were hot-started at 95°C for 4 minutes. This was followed by 2 rounds of PCR and a total of 30 cycles. The first round consisted of 5 cycles of denaturation for 30 seconds at 95°C, annealing for 90 seconds at 57°C, and extension for 120 seconds at 72°C. The second round (25 cycles) was performed with denaturation for 30 seconds at 95°C, annealing for 90 seconds at 57°C, and extension for 90 seconds at 72°C. The last cycle was followed by 4-minute extension at 72°C. Universal methylated human DNA standard (Zymo Research) was used as control sample. Water blank was used as negative control with every PCR amplification.
Statistical analysis
The chi-square (χ2) test was used to analyze the association of PTEN hypermethylation in OSCC with tumor grade and age and sex of the patients (17). P<0.05 was considered statistically significant.
Results
PTEN promoter methylation was analyzed in tumor as well as normal tissues from 100 OSCC patients. Figure 1 shows the results of amplified products using methylated and unmethylated specific primers. The 173-bp band represents the unmethylated PTEN promoter and the 155-bp band represents the methylated PTEN promoter. Aberrant promoter methylation of PTEN was detected in 35% of the tumor tissues and none of the normal samples. The data were confirmed by bisulfite sequencing on 2 randomly selected OSCC samples that appeared methylated for PTEN promoter (Fig. 2). These data clearly depicted heavy methylation of the PTEN gene promoter because the majority of the individual CpG sites were methylated in both cases. Of the 35 OSCC cases found positive for PTEN promoter methylation, 32% (18 of 56) were well differentiated, 24% (8 of 33) were moderately differentiated, and 82% (9 of 11) were poorly differentiated. PTEN promoter methylation was further correlated with age, sex and tumor grade of the patients. We found that patient age and tumor differentiation were significantly associated with PTEN promoter methylation (p=0.05 and 0.0019, respectively; Tab. I), while patient sex was not (p>0.05). The present study revealed that PTEN hypermethylation occurs frequently in OSCC among the Indian population.
Correlation Between Clinicopathological Parameters and PTEN Hypermethylation
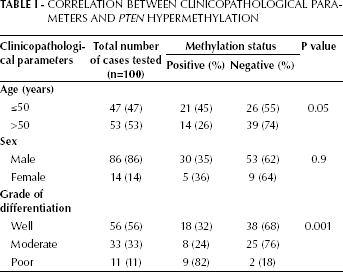
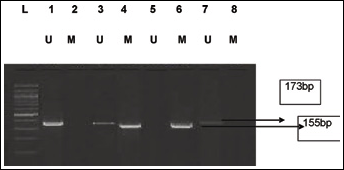
Methylation-specific PCR of the PTEN promoter region in OSCC. Bisulfite-treated DNA was amplified with methylated and unmethylated specific PTEN primers. The 173-bp product is indicative of an unmethylated PTEN allele, whereas the 155-bp product indicates a methylated PTEN allele.
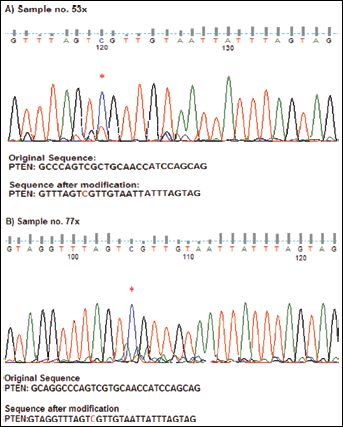
Bisulfite sequencing data of the PTEN promoter region of oral cancer tissue. Bisulfite sequencing of the PTEN promoter region was performed on the cases that were found to be methylated by methylation-specific PCR assay. The methylated CpG sites are marked by asterisks. (
Discussion
The role of the PTEN tumor suppressor gene has been studied in a variety of cancers. Promoter hypermethylation was observed in many human tumor types and was found to be associated with transcriptional inactivation and silencing of defined tumor suppressor genes (18, 19). For that reason, PTEN promoter hypermethylation represents an equivalent to coding region mutations in eliminating the function of tumor suppressor genes (19). We have recently reported that PTEN promoter hypermethylation is closely associated with clinicopathological parameters and disease progression in colorectal cancer (13). The methylation pattern increased from 51% to 82% from poorly differentiated cases to metastatic CRC. Likewise, hypermethylation was more prevalent among late-stage colorectal tumors (13). We used DNA sequencing with bisulfite modification to obtain an accurate picture of the hypermethylation in the PTEN promoter region.
Kurasawa et al demonstrated that methylation of the PTEN promoter region may be related to downregulation of PTEN expression in OSCC (20). This result is consistent with our findings and supports our hypothesis that methylation in the PTEN promoter region (and thereby reduced expression of the gene) contributes to the carcinogenesis of oral cancer. However, no hypermethylation was observed in 65 of our total of 100 cases and only 35 cases were positive (Tab. I). The present study indicated that PTEN promoter hypermethylation is more frequent in poorly differentiated OSCC among the Indian population, as was also reported for other types of cancer (11, 13). The cause of the carcinogenesis in the 65 methylation-negative cases could not be elucidated through this root and further assessment will be required to reach a conclusion about the genes associated with carcinogenesis in this group.
In conclusion, our data suggest that PTEN methylation might prove to be a biomarker for this type of cancer as it leads to the important event of gene silencing and thereby loss of gene functions associated with the development of cancer. PTEN promoter hypermethylation testing can be taken as a promising strategy in the diagnosis of OSCC; however, it needs further investigation on a larger sample size. No studies had been conducted so far to elucidate the role of the PTEN gene in OSCC in the Indian population and our study is the first to report on the level of PTEN methylation in Indian OSSC. We suggest that additional studies are required to elucidate the role of PTEN in this cancer type more precisely.
