Abstract
Background
Carboxylesterase 4A (CES4A) belongs to the member of the carboxylesterase family, yet there has been limited research into its malignant biological behavior in malignant tumors. Here, we aim to investigate the expression, cellular biological functions, and the potential underlying mechanism of CES4A in nasopharyngeal carcinoma (NPC).
Method
A standardized mean difference (SMD) analysis was used to analyze the dysregulation of CES4A based on the gene expression omnibus (GEO) database. qRT-PCR and immunohistochemical staining (IHC) were used to identify the mRNA and protein levels of CES4A in NPC cell lines and tissues, respectively. CCK-8, colony formation, wound healing and transwell assays were utilized to estimate cellular growth and metastasis, respectively. Western blot was conducted to evaluate the activity of PI3K/AKT signaling pathway.
Result
Both mRNA and protein expression of CES4A was significantly diminished both in NPC cell lines and primary tumor tissues. Ectopic expression of CES4A restrains the proliferation, colony formation, migration and invasion of NPC. Additionally, KEGG analysis based on GEO data and high-throughput transcriptome sequencing of cell lines all strongly suggested that CES4A was involved in regulating phosphoinositide 3-kinase (PI3K)/protein kinase B (AKT) signaling pathway. It was observed that AKT and phosphorylated AKT were remarkably reduced in CES4A overexpressing NPC cells, indicating that PI3K/AKT signaling pathway is hindered by CES4A.
Conclusion
CES4A expression is silenced in NPC, functioning as a tumor suppressor by negatively modulating the PI3K/AKT signaling pathway.
Introduction
Nasopharyngeal carcinoma (NPC) is a malignancy originating from the epithelium of the nasopharynx. 1 NPC exhibits strong regional characteristics and racial distribution patterns, being most prevalent in Southeast Asia, particularly in Guangdong Province, China. 2 The pathogenesis of NPC is complex and not yet been fully elucidated, involving Epstein-Barr virus (EBV) infection, hereditary susceptibility, and interactions with environmental factors.3456789–10 Due to its concealed anatomical location and sensitivity to radiation therapy, radiation therapy currently stands as the primary treatment method that effectively inhibits clinical progression of NPC.1112–13 However, some patients may still experience distant metastasis, 14 and radiation therapy can also cause concerning side effects such as fatigue and toxic effects on surrounding tissues.14,15 Therefore, conducting in-depth research on the pathogenesis of NPC, is urgently needed, as well as the search for more reliable diagnostic molecular markers and precise clinical treatment targets.
Carboxylesterase (CES) family consists of five members, namely CES1, CES2, CES3, CES4A, and CES5A. 16 Among them, CES1 and CES2 have been extensively researched in the context of tumor development. It has been observed that the expression of CES1 is downregulated due to promoter methylation, and its reactivation mediates tumor repression in prostate cancer. 17 Likewise, upregulation of CES1 in hepatocellular carcinoma (HCC) activates PKD1/PKCμ signaling pathway, exerting an anti-proliferative effect. 18 Conversely, overexpression of CES1 is detrimental, and targeting its activity enhances the sensitivity of HCC cells to cisplatin. 19 Besides, CES1 accelerates the malignant progression of invasive colorectal cancer by enhancing triacylglycerol catabolism. 20 The expression of CES2 are notably diminished in both cholangiocarcinoma and oral squamous cell carcinoma (OSCC) and correlate with a favorable prognosis of patients.21,22 In cases of colorectal cancer and ovarian cancer, a decrease in the expression of CES2 is noted as disease progression occurs.2324–25 The modulation of CES2 expression within tumors appears to be mediated by P53,26,27 although some studies dispute this. 28 Multiple studies have consistently demonstrated that CES2 enhances cancer cells’ sensitivity to irinotecan while leading to decreased viability.27,2930–31 However, an increase in CES2 expression does not always correlate with better outcomes. For instance, it has been shown to expedite pancreatic adenocarcinoma progression through upregulation of hepatocyte nuclear factor 4α (HNF4α). 32 CES3 is downregulated in colon cancer and affects the immunotherapeutic efficacy of colon adenocarcinoma. 33 CES5A is associated with chemotherapy toxicity of platinum-based drugs in the treating non-small cell lung cancer. 34 Collectively, the aforementioned studies indicate that members from the CES family play significant roles concerning tumor occurrence and progression. However, CES4A has not been studied in any tumor to date. In this study, we focus primarily on the expression of the CES4A in NPC, its impact on the malignant biological behavior of NPC, and the exploration of potential mechanisms. We aim to reveal the possible role of CES4A in tumorigenesis and development of NPC, which will provide a reference for further studies in other tumors.
Materials and Methods
A Standardized Mean Difference Analysis and KEGG Pathway Enrichment Analysis Based on GEO Database
To investigate potential changes in the transcription level of CES4A in NPC, a standardized mean difference (SMD) analysis was conducted using cDNA microarrays and RNA-sequencing data from the gene expression omnibus (GEO) database. The search terminology used were: (nasopharynx OR nasopharyngeal) AND (carcinoma OR cancer OR tumor OR adenocarcinoma OR malignance OR tumor OR neoplasm* OR oncology). Inclusion criteria for statistical analysis were as follows: (1) samples obtained from humans, (2) samples collected from nasopharyngeal tissue with a minimum of three cases of cancer or non-cancer, and (3) mRNA expression data containing CES4A. The analysis was performed using STATA15 software to implement a SMD analysis. Furthermore, a Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis based on GSE102349 was carried out to identify signaling pathways associated with CES4A in NPC.
Human Samples
A total of 16 newly diagnosed primary NPC samples were collected, and nine samples diagnosed with chronic mucositis were used as non-cancerous specimens(NNE). All donors were informed of the purpose of collecting pathological samples and obtained their verbal consent before collecting samples, and the samples were diagnosed by clinical as well as experienced pathologists based on the World Health Organization classification. In addition, a tissue array containing 110 cases of NPC was obtained from Outdo Biotech (HNasN110Su01, Shanghai, China). However, one case was fell off during staining, resulting in 109 samples being included in the analysis.
Cell Lines and Transfection
NP69 35 as the nonmalignant nasopharyngeal epithelial cell line was cultured in KSFM medium supplemented with epidermal growth factor. NPC cell lines CNE1, 36 HONE1, 37 5–8F, 38 and TW03 39 were cultured in DMEM media, while CNE2-EBV was cultured in RPMI 1640 media, containing with 10% Fetal Bovine Serum and 1% penicillin-streptomycin mixed solution, at 37°C and 5% CO2 atmosphere. The PCDH-entry plasmid was served as the vector for CES4A expression, while the empty vector regarded as the control group (PV822400, Paivi Biosciences Inc, China). Plasmids were instantaneously transfected into NPC cell lines (CNE1 and HONE1) using lipofectamine 3000 (L3000015, Invitrogen, USA), in accordance with the recommended guidelines of manufacturer. CNE1/HONE1-PCDH was considered as the control group; CNE1/HONE1-CES4A was the experimental group. CES4A expression was validated by qRT-PCR, as well as western blotting.
RNA Extraction and Quantitative Real-Time PCR (qRT-PCR)
Total RNA was isolated with EasyPure® Fast Cell RNA Kit (TransGen Biotech, China) and cDNA was synthesized by employing the TransScript One-Step gDNA Removal and cDNA Synthesis SuperMix reagent kit (TransGen Biotech, China). Analysis of transcriptional gene expression was implemented using PowerUpTM SYBRTM Green Master Mix (Applied Biosystems, USA) on the qRT-PCR System (QuantStudio5, Applied Biosystems, USA). The primer sequences were as follows:
CES4A-Forward, 5’-GATGTCACCCCTAGCCTCG- 3’; CES4A-Reverse, 5’-TGGCCACTTTCAGTGGGTTA-3’; GAPDH-Forward, 5’-GCTCAGACACCATGGGGAAG-3’; GAPDH-Reverse, 5’-CCCCTTCATTGACCTCAACTACA-3’.
The reaction conditions included pre-denaturation at 98°C for 30 s; denaturation at 98°C for 15 s, annealing at 65°C for 60 s, and extension at 72°C for 90 s, with a total of 40 cycles. The relative transcriptional level of CES4A was standardized to GAPDH and measured using the 2−ΔΔCt method.
40
Immunohistochemical Staining (IHC)
Tissues wax were cut into 4-μm-thick sections and subjected to deparaffinizate and rehydrate. Following high-pressure antigen retrieval using EDTA and blocking with peroxidase, the primary antibody (#43923, 1:100, Signalway Antibody, USA) was added dropwise and incubated at 4°C overnight. Subsequently, the secondary antibody labeled with horseradish peroxidase (ZSGB-BIO, China) was incubated at 37 °C for 20 min, followed by DAB (ZSGB-BIO, China) staining and hematoxylin for background staining.
Two experienced tissue pathology experts who were blinded to the clinical pathological information were scored independently. Immunohistochemical scoring was according to the percentage of positive cells and the intensity of staining. Scoring for the percentage of positive cells was as follows: 0% (0 points), > 0% to 25% (1 point), > 25% to 50% (2 points), > 50% to 75% (3 points), and >75% (4 points). Staining intensity scoring criteria were as follows: no staining (0 points), light yellow (1 point), brown-yellow (2 points), and brown (3 points). The total score was calculated by multiplying the percentage of positive cells by the staining intensity score. 41
Cell Proliferation Assay
Cell Counting Kit-8 (CCK-8) assay was employed to assess the impact of CES4A on cell growth. Briefly, the transiently transfected NPC cells were seeded into 96-well plates at a density of 1 × 103 cells per well. After the cells adhered to the well, the original medium was aspirated out, and then 10 μl of CCK-8 solution (Dojindo, Japan) was mixed with 100 μl of culture medium and added to each holes, followed by incubation for specific periods of time (1, 2, 3, 4 days). The absorbance (OD450 nm) was determined using a microporous plate reader.
Colony Formation Assay
3 × 102 processed cells were inoculated into the well of six-well plates. The cells were continuously cultured in an incubator for 10–14 days, after the clone clusters were washed with phosphate buffered saline, fixed with a 4% stationary solution and stained with crystal violet. The formed colonies were then photographed, and the area of the colonies was calculated using ImageJ software. These experiments were duplicated three times.
Wound Healing Assay
NPC cells (7 × 104 cells/wells) were seeded into 12-well plates with complete medium overnight. The ibidi Culture-Insert (ibidi, Germany) was utilized to generate scratch in the monolayer of cells. When the cells monolayer was completely overspread, the insert was taken out and the cells were switched to the serum-free media for further cultivation. Wound closure was observed and imaged under an inverted optical microscope (Nikon, Japan) at two-time points (0 and 6 h). The migratory area was analyzed using ImageJ software. These experiments were conducted in triplicate.
Transwell Assay
1 × 105 cells dispersed in 200μl of basal medium were implanted into the upper chambers of invasion chambers covered with Matrigel (Corning, USA). The lower layer of the invasion chamber was equipped DMEM added with 10% Fetal Bovine Serum. After 24 h of incubation, the non-invasive cells were wiped off with cotton swab, and the invasive cells on the surface of the inferior membrane were fixed with 4% tissue fixative, then stained with 0.1% crystal violet, photographed and counted with ImageJ software.
Western Blotting
Cytoplasmic proteins were extracted using RIPA lysis buffer (Beyotime, China) comprising phenylmethylsulfonyl fluoride and a protease inhibitor cocktail (Beyotime, China). The protein concentration was determined using the Enhanced BCA Protein Assay Kit (Beyotime, China). The equivalent amounts of protein were separated by electrophoresis on 4–12% Sodium Dodecyl Sulfate PolyAcrylamide Gel at 200 V, and then transferred to a polyvinylidene fluoride (PVDF) membrane (Sigma-Aldrich, USA). The membranes were blocked using a protein-free rapid sealing solution (PS108P, Epizyme, China) to prevent non-specific protein binding and then incubated with primary antibodies overnight at 4°C. Flag (AE092, 1:10000, ABclonal, China), AKT (9272S, 1:1000, Signalway Antibody, USA), P-AKT (4060S, 1:1000, Signalway Antibody, USA), PI3K (4292S, 1:1000, Signalway Antibody, USA), P-PI3K (4228S, 1:1000, Signalway Antibody, USA) and GAPDH (60004-1-Ig, 1:2000, Proteintech, China) served as an internal control. Subsequently, secondary antibodies that were opposite to the species of primary antibodies (anti-rabbit/mouse, dilution 1:10000) were used for 1.5 h at room temperature. Fluorescent signals were collected using the Odyssey DLX system (LI-COR, USA), and the expression of target proteins was standardized to GAPDH levels and quantified using ImageJ software.
Statistical Analysis
All data were analyzed using SPSS 17.0, and GraphPad Prism 8.0.1 was utilized for plotting. The independent Student's t-test was employed for comparing different groups. Statistical differences for IHC score were evaluated using the Mann-Whitney U-test. The SMD analysis was performed using STATA 15 software. P-value < 0.05 was considered to be statistically significant.
Results
CES4A is Downregulated in NPC Cell Lines and Primary Tumor Tissues
To investigate potential alterations in CES4A mRNA expression in NPC, we conducted a SMD analysis using data from five sets of microarrays in the GEO database, encompassing 42 non-cancerous samples and 86 NPC tissues. Results produced by the fixed-effects model suggested no heterogeneity among the individual chip datasets (I2 = 0%, p = 0.586). The SMD analysis evaluation revealed that the transcriptional expression of CES4A in NPC tissue was lower than in normal control tissue (SMD = -2.164; 95% confidence interval CI: -2.640 to -1.689; P < 0.001). Funnel plots demonstrated a symmetrical funnel plot (P > 0.05) and there was no significant publication bias according to the Egger test (P > 0.05). Sensitive analysis showed no obvious deviation upon excluding any GEO microarray dataset separately (P > 0.05) (Figure 1A-D). Details of the 5 sets of NPC microarrays employed in this research are provided in Table 1.
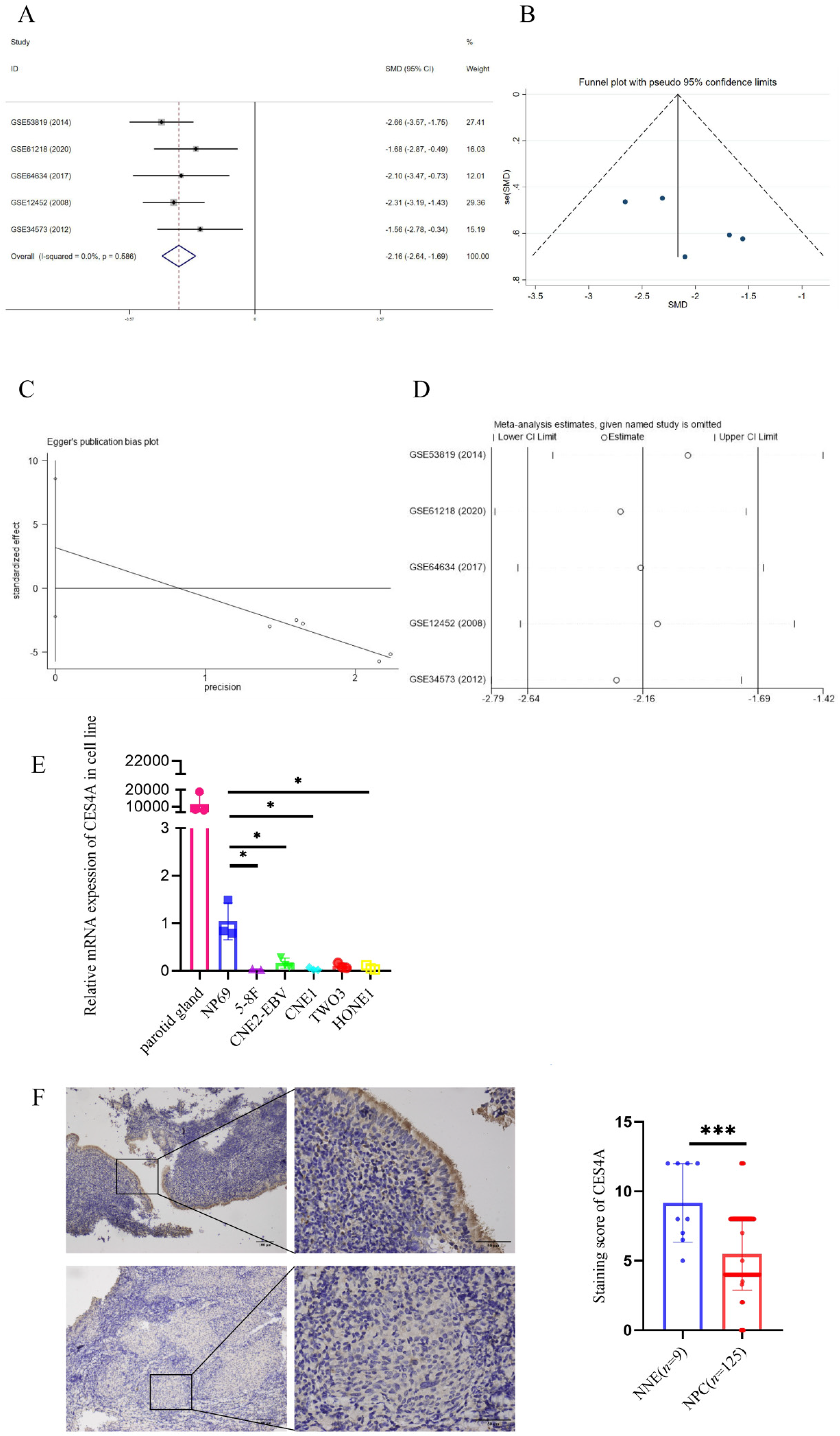
Downregulation of CES4A mRNA and protein expression in NPC. (A) SMD analysis of CES4A mRNA expression in NPC. Forest plot of five sets of microarrays, displaying significant differences between NPC and normal groups (SMD = -2.164; 95% confidence interval CI: -2.640 to -1.689; P < 0.001). (B) Funnel blot represented no publication bias (P > 0.05). (C) Egger test demonstrated no significant publication bias (P > 0.05). (D) Sensitive analysis showed no obvious deviation upon excluding any GEO datasets separately (P > 0.05). (E) mRNA level of CES4A was measured by qRT-PCR in five NPC cell lines and a normal nasopharyngeal epithelium cell line NP69. The independent Student's t-test was used for analysis, and data are displayed as the means ± SD. (F) IHC of CES4A in primary NPC tissues and NNE. Magnification: × 400. The Mann-Whitney U test was utilized for analysis.
Detailed Information on the GEO Profiling Dataset Used for SMD Analysis.

We further performed qRT-PCR to assess the consistency of abnormal CES4A transcription in NPC cell lines, with parotid gland tissue serving as a positive control. The results showed a significant reduction in CES4A mRNA levels in NPC cells (CNE1, HONE1, 5-8F, TWO3, CNE2-EBV) compared with NP69 (Figure 1E). Notably, CES4A protein was primarily localized in the cytoplasm and exhibited high expression in normal nasopharyngeal epithelium, characterized by deep yellow or brown staining, especially in ciliated columnar epithelial cells. In contrast, staining in NPC tissue appeared noticeably lighter, often presenting as light yellow or even colorless (Figure 1F). Our results were in accordance with the SMD analysis, hinting the inactivation of CES4A in NPC.
CES4A Inhibits the Proliferation and Colony Formation of Cells in Vitro
In order to investigate the potential impact of CES4A on the malignant biological behavior of NPC cells, we expressed CES4A in NPC cell lines (CNE1, HONE1) instantaneously. Ectopic overexpression of CES4A was assessed separately using qRT-PCR and western blotting (Figure 2A, B). Our results demonstrated that the proliferative rate of NPC cells with CES4A overexpression was notably slower compared to the control group (Figure 2C). Furthermore, CES4A overexpression significantly suppressed the colony-forming ability of both NPC cell lines (Figure 2D).
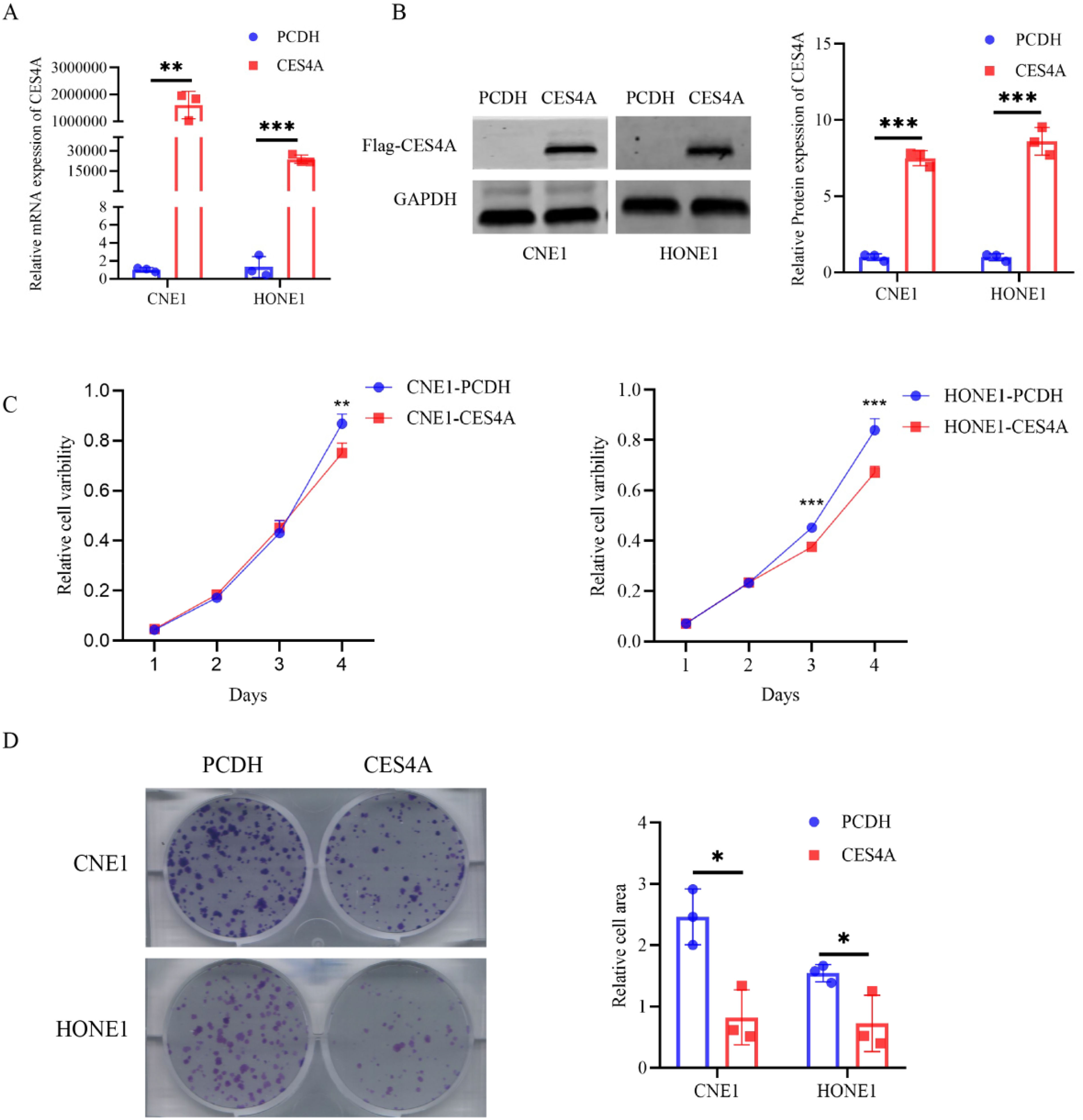
Overexpression of CES4A inhibits NPC cell proliferation. (A, B) The mRNA and protein levels of CES4A in transiently transfected NPC cells was validated by qRT-PCR and western blotting, respectively. (C) Cell proliferation was determined by CCK-8 assay (OD = 450 nm). (D) The colony formation assay was recruited to assess the colony-forming ability of NPC cells. The independent Student's t-test was utilized for analysis, and data are represented as means ± SD.
CES4A Restrains NPC Cell Migration and Invasion
Metastatic capability is significant in cancer, particularly in NPC. To further assess the impact of CES4A on the migratory and invasive abilities of NPC cells in vitro, we transiently expressed CES4A in NPC cells. Cells overexpressing CES4A displayed a reduced area of gap closure compared with the control cell line (Figure 3A). Additionally, the number of cells passing through the invasion chamber in the group overexpressing CES4A was fewer than the matched group (Figure 3B). Our findings demonstrated that restoring the expression of CES4A attenuates the motility of NPC cells. In addition, we found that the group overexpressing CES4A had fewer cells passing through the compartments compared to the control group.
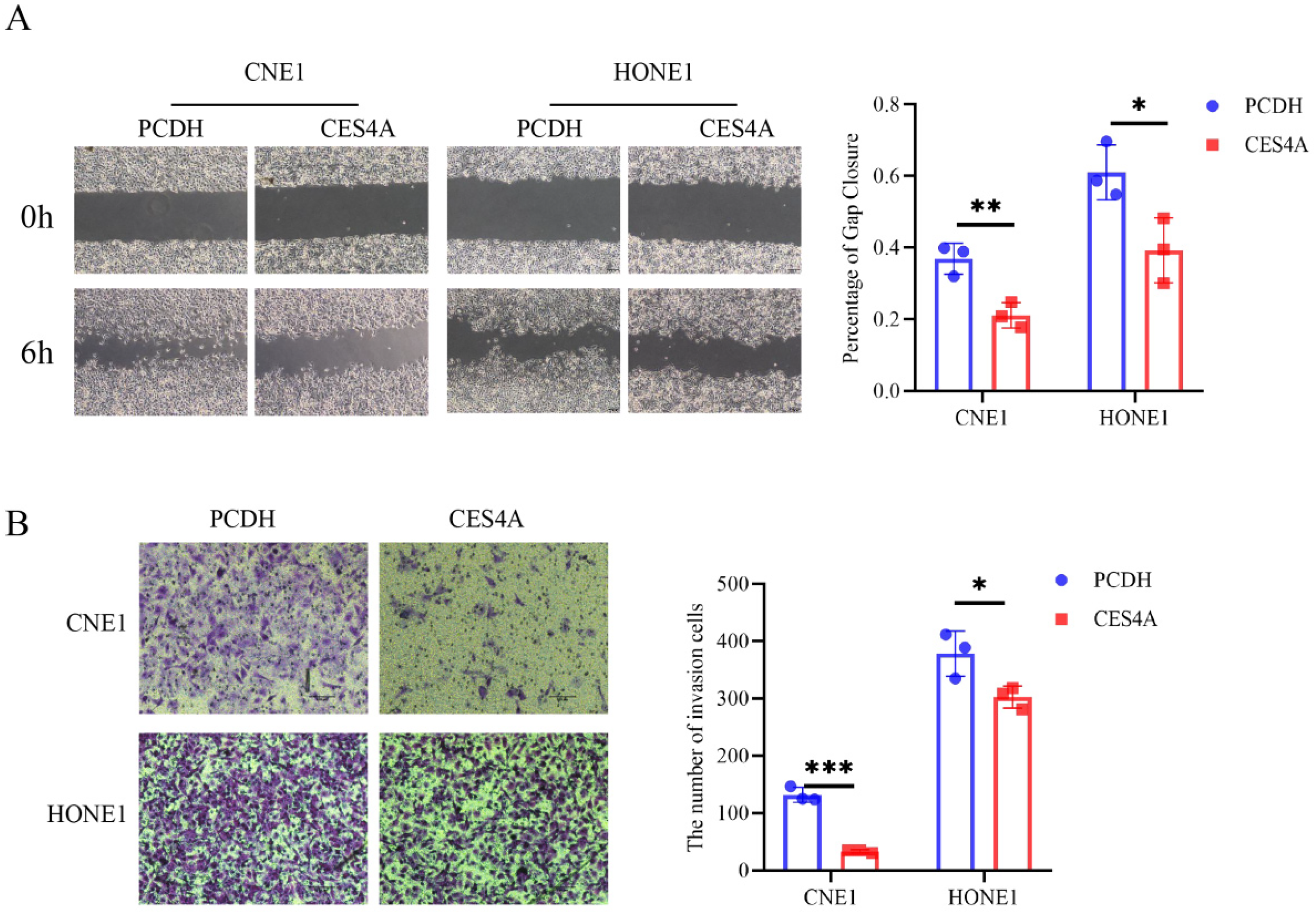
Overexpression of CES4A represses the motility of NPC cells lines in vitro. (A) Wound-healing assay was used to evaluate the migration characteristics of NPC cells transiently transfected with CES4A or empty vectors. The width of the wound was captured and measured at 0 h and 6 h, and the percentage of gap closure was calculated by ImageJ software. Magnification: ×200. (B) Evaluation of the differential invasion abilities of NPC cells by transwell assay. The blue dots represent invading cells, and their numbers are quantified and displayed in the histogram. Data are presented as means ± SD in the bar graph. The independent Student's t-test was employed for analysis.
CES4A Suppresses Malignant Characterization of NPC via the PI3K/AKT Signaling Pathway
To elucidate the specific molecular mechanisms through which CES4A impacts the malignant biological behavior of NPC, we conducted KEGG pathway enrichment analysis based on the GSE102349 dataset and high-throughput transcriptome sequencing of cell lines overexpressing CES4A. We identified a total of 18 overlapping pathways by integrating the results of cell sequencing and KEGG. Furthermore, the results indicated a close association between CES4A and the phosphoinositide 3-kinase (PI3K)/protein kinase B (AKT) signaling pathway (Table 2). Subsequently, we examined changes in the PI3K/AKT signaling pathway following CES4A overexpression using western blotting. Our findings indicated that restoring CES4A expression led to a reduction in AKT and phospho-AKT (P-AKT) expression in CNE1; AKT also exhibited a decreasing trend in HONE1, while P-AKT was downregulated. Additionally, PI3K and phospho-PI3K (P-PI3K) displayed a declining trend in CNE1 but an increasing trend in HONE1 (Figure 4). In conclusion, our results indicate that the PI3K/AKT pathway is inhibited following CES4A overexpression.

Ectopic overexpression of CES4A suppresses the PI3K/AKT signaling pathway. The expression of AKT, P-AKT, PI3K, P-PI3K were measured by western blot and quantified by grayscale values. GAPDH was employed as an internal reference. Data are presented as mean ± SD in a bar graph. The independent Student's t-test was utilized for analysis.
The Pathway of RNA Sequencing and KEGG Enrichment Analysis Pathways Based on Tissue Microarrays.

Discussion
The CES family is involved in the biotransformation of drugs and the hydrolysis of esterified molecules, as well as endogenous substrates containing thioester, amide, or carbamate bonds.42,43 They also play a critical role in the metabolism of fatty acid acyl and cholesterol esters and are responsible for the hydrolytic elimination of ingested esters.43,44 CES4A, also known as CES6, encodes a secreted enzyme that may contribute to the detoxification of drugs and xenobiotics in neural tissues, other peripheral tissues, and cerebrospinal fluid. 45
Our findings reveal that CES4A is downregulated in both cell lines and primary tissues of NPC. Additionally, overexpression of CES4A was found to inhibit the proliferation, migration, and invasion of NPC cells, suggesting that CES4A serves as a tumor suppressor gene in NPC.
To explore the mechanism by which CES4A is involved in the tumor signaling pathway of NPC cells, we conducted transcriptome sequencing of CES4A-overexpressing NPC cell. Our results, along with KEGG enrichment analysis based on the GEO dataset, suggest that CES4A is associated with the PI3K/AKT pathway, which is abnormally activated in various cancers.464748–49 Additionally, several studies have shown that many genes abnormally expressed in NPC have an impact on the malignant biological behavior of NPC cells, such as proliferation, cycle, migration and invasion, by regulating the PI3K/AKT signaling pathway.5051525354555657–58 Other studies also have reported that CES2 inhibits the expression of PI3 K and enhance the sensitivity of colorectal cancer cells to oxaliplatin. 59 Similarly, CES2 has demonstrated its inhibitory effect on PI3K/AKT pathway activity in OSCC. 22 The enzymes encoded by the members of the CES family share an amino acid sequence homology ranging from 39% to 46%. 16 Therefore, we hypothesize that CES4A inhibits the malignant progression of NPC by mediating the PI3K/AKT pathway. Our findings confirm that the protein levels of AKT and P-AKT, as the important markers of the PI3K/AKT pathway, are reduced in NPC cells overexpressing CES4A compared to control cells. AKT, a key regulator of the PI3K pathway, can be widely activated by growth signals and regulates many downstream proteins functions including angiogenesis, cell survival, migration, metabolism and proliferation. 60 These results indicate that the overexpression of CES4A may inhibit the malignant progression of NPC through the PI3K/AKT signaling pathway.
Some research on lipid metabolism in NPC has demonstrated that the regulation of adipose triglyceride lipase (ATGL) by UBE2L6, along with the modulation of ATGL by EBV latent membrane protein 2A (EBV-LMP2A), consistently indicates that NPC is characterized by dysregulation of lipid metabolism.61,62 Furthermore, multiple studies have shown that members of the CES family play a role in regulating various aspects of lipid metabolism, thereby influencing tumor progression.19,20,22,32 In addition, KEGG pathway analysis revealed a noteworthy association between CES4A and the lipid metabolism and atherosclerosis pathways, as well as the PI3K/AKT signaling pathway in NPC. Notably, lipid metabolism and the PI3K/AKT signaling pathway are intricately linked, playing pivotal roles in both physiological and pathological contexts.63,64 Activation of AKT promotes lipogenesis through upregulation of sterol regulatory element-binding protein 1c (SREBP-1c), which enhances the expression of key lipogenic enzymes, such as fatty acid synthase (FASN) and acetyl-CoA carboxylase (ACC). 63 In cancer, aberrant activation of the PI3K/AKT pathway is a hallmark feature, frequently accompanied by lipid metabolic reprogramming. Tumor cells exploit this pathway to support rapid proliferation by enhancing fatty acid synthesis and cholesterol accumulation, which are critical for membrane biosynthesis and energy production. 65 Moreover, CES2 has been shown to inhibit the PI3K/AKT/MYC signaling pathway through its regulation of signaling lipids in OSCC. 22 These findings underscore the close relationship between the PI3K/AKT pathway and lipid metabolism while highlighting their potential as therapeutic targets.
Although significant advances have been made in the understanding of CES4A, it remains unclear whether it regulates the PI3K/AKT signaling pathway by modulating lipid metabolism in NPC. Therefore, further exploration through metabolomics is warranted.
Conclusion
In summary, we discovered that CES4A expression is inactivated in both the cell lines and primary cancer tissue of NPC. Increasing CES4A expression inhibits the proliferation, colony formation, migration, and other malignant biological behaviors of NPC. In addition, CES4A exerts anti-cancer effects by inhibiting the activation of the PI3K/AKT signaling pathway in NPC. Our findings suggest that CES4A might serve as a novel tumor suppressor gene in NPC, potentially emerging as a potential therapeutic target and biomarker for NPC.
Footnotes
Abbreviations
Table: The full names and corresponding abbreviations used in this article
Acknowledgements
Not applicable.
Author Contribution
Conceptualization, WLZ, YH, PL, XYZ, ZZ; methodology, QLC, QXH, LML, DPL; investigation (data acquisition), QLC, QXH, YPY, YSL; resources, XX, YSL,YH; analysis, QLC, QXH, RZ, WQW, LXD, JMS, ZYL, SYT, YML; writing, QLC, XYZ,WLZ, PL; project administration and supervision, PL, WLZ,XYZ,XX; funding acquisition, PL, XYZ, ZZ,XX. All authors contributed to the article and approved the submitted version.
Conflict of Interest
The authors confirm that they do not have any conflicting financial interests or personal relationships that could affect the work of this paper.
Data Availability Statement
The data supporting the research results can be obtained from the corresponding authors on a reasonable grounds.
Funding
Our work was obtained by the Innovation Project of Guangxi Graduate Education (YCSW2023231), Natural Science Foundation of China (82202512, U22A20322, 82020511, 81960490) and the High-level Talent Introduction Plan of the First Affiliated Hospital of Guangxi Medical University (the fifth level).
Ethics Approval and Consent to Participate
The research involving human participants were reviewed and ratified by The Ethics Committee of the First Affiliated Hospital of Guangxi Medical University (2022-KT-243). Besides, informed consent of the participants was obtained.
