Abstract
Background
Circular RNAs (circRNAs) are crucial in the regulation of gene expression and biological processes. However, in colorectal cancer, the expression characteristics and biological function of circRNA_0006174 (circ_0006174) is not fully understood. This work is aimed to investigate the biological function of circ_0006174 in colorectal cancer and its molecular mechanism.
Methods
Circ_0006174, microRNA-142-3p and X-linked inhibitor of apoptosis expression levels were detected in colorectal cancer tissues and cells using quantitative real-time polymerase chain reaction analysis or Western blot. The effects of circ_0006174 on colorectal cancer cell proliferation, apoptosis, migration and invasion were detected using the cell counting kit-8 method, bromodeoxyuridine experiments, flow cytometry analysis and Transwell experiments. The targeting relationship among circ_0006174, microRNA-142-3p and X-linked inhibitor of apoptosis was analysed by bioinformatics prediction, dual-luciferase reporter experiment and RNA immunoprecipitation experiment.
Results
Circ_0006174 was up-regulated in colorectal cancer tissues as well as in cell lines, and its high expression was remarkably associated with enlarged tumour volume and advanced tumour, node, metastasis stage of the patients. Circ_0006174 overexpression enhanced colorectal cancer cell proliferation, migration and invasion, and inhibited colorectal cancer cell apoptosis; while knocking down circ_0006174 caused the opposite effects. Circ_0006174 directly targeted and negatively regulated microRNA-142-3p expression, and X-linked inhibitor of apoptosis, a target gene of microRNA-142-3p, could be indirectly and positively modulated by circ_0006174.
Conclusion
Circ_0006174 facilitates colorectal cancer cell proliferation, migration and invasion, and represses colorectal cancer cell apoptosis by regulating microRNA-142-3p/X-linked inhibitor of apoptosis axis.
Introduction
Colorectal cancer (CRC) is a highly prevalent malignancy, and is the fourth most deadly cancer in the world after lung, gastric and liver cancers. 1 Currently, the main therapies for CRC include surgery, radiotherapy, targeted therapy and immunotherapy. 2 Despite recent advances in CRC therapy, the prognosis of patients remains unfavourable due to the high rate of CRC metastasis, with a 5-year survival rate of less than 65%. 3 Therefore, the search for early diagnostic biomarkers and potential therapeutic targets for CRC is of great importance.
Circular RNAs (circRNAs) are noncoding RNAs widely found in eukaryotic cells, and unlike linear RNAs, circRNAs form a covalently closed loop by reverse splicing, a structure that makes them highly stable and conservative. 4 Reportedly, circRNAs are vital regulators in the pathogenesis of multiple human diseases, including cancers. 4 For instance, has_circ_0000567 is down-regulated in both CRC tissues and cell lines, and knocking down has_circ_0000567 promotes CRC cell proliferation and migration in vitro. 5 CircBase database (http://www.circbase.org/) indicates that circRNA_0006174 (circ_0006174) is generated from the transcript of RAD23B (Supplemental Figure 1A). Reportedly, overexpression of RAD23B promoted the growth and metastasis of CRC cells both in vitro and in vivo by targeting the talin1/2/integrin/FAK/RhoA/Rac1/CORO1C signalling pathway. 6 Moreover, it is reported that circ_0006174 promotes CRC progression by regulating the miR-138-5p/MACC1 axis. 7 However, the molecular mechanism by which circ_0006174 exerts its effects has not been fully elucidated.
Reportedly, microRNA-142-3p (miR-142-3p) plays a tumour-suppressive role in multiple cancers. In CRC, miR-142-3p has been reported to induce apoptosis and inhibit proliferation, migration and invasion of CRC cells. 8 Nonetheless, the upstream regulatory mechanism of miR-142-3p in CRC remains unclear. X-linked inhibitor of apoptosis (XIAP) expression was up-regulated in CRC, and XIAP functioned as a tumour-promoting factor 9 ; meanwhile, bioinformatics analysis indicates that miR-142-3p is a potential target of circ_0006174, and miR-142-3p might target XIAP. This relationship suggests that circ_0006174/miR-142-3p/XIAP axis may exert a tumour-promoting effect in CRC. This study is performed to validate this scientific hypothesis mentioned above.
Materials and methods
Tissue specimens
CRC tissue samples and adjacent non-cancerous tissue samples were collected from 46 subjects who were diagnosed with CRC at the first time and received surgery at our hospital from May 2018 to August 2019. All tissue specimens were preserved in liquid nitrogen at −196°C immediately after surgical excision. All subjects had not received radiotherapy or chemotherapy before the surgery and had not taken targeted therapeutic drugs. All subjects provided written informed consent and the research was approved by the Guizhou Provincial People's Hospital Ethics Committee and strictly adhered to the Declaration of Helsinki.
Cell culture
Human normal colorectal mucosal cells (FHC) and CRC cell lines (DLD-1, CACO2, SW480, and HT-29 cells) were obtained from the American Type Culture Collection (ATCC, Rockville, MD, USA). The above cells were cultured in Dulbecco's Modified Eagle Medium (DMEM) (Thermo-Fisher Scientific, Waltham, MA, USA) containing 10% foetal bovine serum (FBS, Thermo-Fisher Scientific, Waltham, MA, USA), 100 U/mL penicillin, and 100 µg/mL streptomycins (Thermo-Fisher Scientific, Waltham, MA, USA) and incubated at 37°C with 5% CO2. Cell growth was observed routinely, and the medium was changed every 2–3 days, and the cells in the logarithmic growth phase were used for subsequent experiments.
Cell transfection
Circ_0006174 overexpression plasmid (circ_0006174), empty vector (NC), small interfering RNAs targeting circ_0006174 (si-circ_0006174#1: 5ʹ-AACAGCAACTGACAGGCAAAA-3ʹ, si-circ_0006174#2: 5ʹ-AACTGACAGGCAAAATCCTCA-3ʹ and si-circ_0006174#3: 5ʹ-GACAGGCAAAATCCTCAATGA-3ʹ), negative control (si-NC: 5ʹ-GGACUCUCGGAUUGUAAGAUU-3ʹ), miR-142-3p mimics (3′-AGGUAUUUCAUCCUUUGUGAUGU-5′), miR-142-3p inhibitor (5′-ACAUCACAAAGGAUGAAAUACCU-3′) and control (mimics NC: 5′-UUCUCCGAACGUGUCACGUTT-3′) were constructed by GenePharma (Shanghai, China). DLD-1 and HT-29 cells were transfected with the above vectors/oligonucleotides using Lipofectamine®3000 (Thermo-Fisher Scientific, Waltham, MA, USA) according to the manufacturer's instruction, respectively.
Quantitative real-time polymerase chain reaction analysis
Total RNA from tissues and cells was extracted using TRIzol reagent (Invitrogen, Carlsbad, CA, USA), tested for its purity, and reverse transcribed into cDNA. Subsequently, quantitative real-time polymerase chain reaction (qRT-PCR) analysis was performed using SYBR®Premix-Ex-Taq™ (Takara, Tokyo, Japan) on an ABI7300 PCR detection system (Thermo-Fisher Scientific, Waltham, MA, USA). The relative expressions of circ_0006174, miR-142-3p, and XIAP mRNAs were calculated using the 2−ΔΔCt method with glyceraldehyde-3-phosphate dehydrogenase (GAPDH) and U6 as the internal controls. The primer sequences are as follows: circ_0006174: Forward: 5′ TTCTGAACCTGCACCTGCTA 3′ and Reverse: 5′ GTGGACACTGCTTTGGGTTTG 3′; miR-142-3p: Forward: 5′ UGUAGUGUUUCCUACUUUAUGGA 3′ and Reverse: 5′ CAUAAAGUAGGAAACACUACAU 3′; GAPDH: Forward: 5′ AGAAGGCTGG GGCTCATTTG 3′ and Reverse: 5′ AGGGGCCATC CACAGTCT TC 3′; U6: Forward: 5′ CTCGCTTCGGCAGCACA 3′ and Reverse: 5′ AACGCTTCACGAATTTGCGT 3′.
Nuclear–cytoplasmic fractionation
To obtain nuclear and cytoplasmic fractions of DLD-1 (human colorectal adenocarcinoma) and HT-29 (human colorectal adenocarcinoma) cells, a NE-PER® Nuclear and Cytoplasmic Extraction kit (Thermo Fisher Scientific, Waltham, MA, USA) were used. Circ_0006174 expression in the nucleus and cytoplasm was detected by qRT-PCR, with GAPDH and U6 used as internal references for cytoplasmic and nuclear RNA, respectively.
Cell counting kit-8 experiment
After trypsinization, single-cell suspension was prepared and the cell concentration was modulated to 3 × 104 cells/mL. Then the cells were transferred into 96-well plate (3 × 103 cells/well). After 24 h of culture, 10 μL of cell counting kit-8 (CCK-8) solution (Dojindo Molecular Technologies, Kumamoto, Japan) was added to each well. After the cells were incubated for 2 h, the absorbance values were measured at 450 nm using a microplate reader. With this method, the changes in cell proliferation of DLD-1 and HT-29 cells after transfection and co-transfection at 24, 48, 72 and 96 h were determined, respectively.
Bromodeoxyuridine experiment
DLD-1 and HT-29 cells in logarithmic growth phase were employed to prepare single-cell suspensions, and the cells were inoculated into 24-well plate (1 × 105 cells/well). After the cells were cultured for 24 h, bromodeoxyuridine (BrdU) reagent (Beyotime, Shanghai, China) was supplemented, and the cells were incubated at 37°C with 5% CO2. After 48 h of continuous cell culture, the cells were fixed, and then stained with anti-BrdU antibody (Beyotime, Shanghai, China) and 4′,6-diamidino-2-phenylindole (DAPI) staining solution (Beyotime, Shanghai, China) in the dark. After the cells were washed with phosphate buffered saline (PBS), three fields of view were randomly selected and the cells were counted under a fluorescence microscope. Cell proliferation rate = the number of BrdU-positive cells/number of DAPI-positive cells, and the average of the proliferation rate of cells from the three visual fields were taken as the cell proliferation rate.
Flow cytometry
Apoptosis was detected using an AnnexinV-FITC and propidium iodide (PI) double staining assay kit (Southern Biotechnology, Birmingham, AL, USA). Transfected cells were plated in a 6-well plate, and the cell density was modulated to 2 × 106 cells/well. After the cells were cultured for 24 h, the supernatant was discarded, and the cells were rinsed with pre-chilled PBS 2 times, and the cells were resuspended with 1 × binding buffer. The cell suspensions were then mixed with 5 μL of AnnexinV-FITC staining solution and 5 μL of PI staining solution, incubated for 15 min in the dark, and subsequently, the apoptosis rate was examined by flow cytometry within 1 h.
Transwell experiment
Transwell experiments were employed to detect cell migration and invasion. The transfected DLD-1 and HT-29 cells were collected, trypsinised with trypsin, centrifuged, and resuspended in serum-free DMEM to a density of 1 × 105 cells/mL. Next, 200 μL of cell suspension was supplemented to the upper compartment of the Transwell chamber and 600 μL of medium containing 20% FBS was supplemented to the lower compartment, and then the cells were cultured in 5% CO2 at 37°C for 24 h. Subsequently, the compartment was removed, the cells on the upper surface of the membrane were gently wiped off with a cotton swab. Next, the migrated cells were fixed with 4% paraformaldehyde, stained with 0.1% crystalline violet, washed by tap water, dried, photographed and counted. Cell invasion experiment: the membrane should be coated with a layer of Matrigel (Millipore, Bedford, MA, USA) before the cells were inoculated, and the other experimental procedures were the same as migration experiments.
Luciferase reporter experiment
Wild type circ_0006174 (circ_0006174 WT), mutant circ_0006174 (circ_0006174 MUT), wild type XIAP (XIAP WT) and mutant XIAP (XIAP MUT) luciferase reporter vectors were constructed by Promega (Madison, WI, USA). DLD-1 and HT-29 cells were inoculated in 48-well plates and cultured to 70% confluence. DLD-1 or HT-29 cells were then co-transfected with the above reporter vector and miR-142-3p mimics or negative control, respectively, using Lipofectamine®3000 (Thermo Fisher Scientific, Inc; Carlsbad, CA, USA). After 48 h of culture, the luciferase activity of the cells in each group was measured with a Dual-Luciferase Reporter Assay System (Promega, Madison, WI, USA).
RNA immunoprecipitation experiment
Briefly, DLD-1 and HT-29 cells were lysed in RNA immunoprecipitation (RIP) lysis buffer, then the cell lysates were incubated with magnetic beads conjugated with anti-Ago2 antibody or negative control IgG in RIP buffer. Subsequently, the specimens were incubated with Proteinase K with shaking to remove the proteins, and then RNA precipitation was obtained. Furthermore, the purified RNA was used for qRT-PCR to validate the binding relationship between circ_0006174 and miR-142-3p.
Western blot
The medium was discarded and RIPA lysis buffer (Beyotime, Shanghai, China) was supplemented to lyse the cells, and the supernatant was collected by high-speed centrifugation (4°C, 10,000 rpm, 5 min). The 10 μg of total protein in each group was dissolved by sodium dodecyl sulphate polyacrylamide gel electrophoresis. It was then electrically transferred to the polyvinylidene fluoride membrane (Millipore, Bedford, MA, USA). Next, the membrane was incubated with the primary antibody (rabbit anti-XIAP antibody, ab21278, 1:1000; mouse anti-β-actin antibody, ab6276, 1:3000) overnight at 4 °C. After rinsing with tris buffered saline with Tween 20 (TBST), the membranes were incubated with the corresponding secondary antibodies (goat anti-rabbit IgG coupled with HRP, ab205718, 1:2000; goat anti-mouse IgG, ab205718, 1:2000) for 1 h at room temperature. After the membranes were washed by TBST again, the protein bands were detected with an ECL Western blot Substrate kit (Thermo-Fisher Scientific, Waltham, MA, USA). The grey values of each protein were analysed by Image J (NIH, Bethesda, MD, USA).
Statistical analysis
All data in this research were analysed using SPSS 20.0 statistical analysis software (SPSS Inc., Chicago, IL, USA), and the measurement data were expressed as ‘mean ± standard deviation’. Comparisons between the two groups were made using t-tests, the mean among multiple groups was compared using one-way ANOVA with Turkey's post-hoc test Counting data were expressed in a fourfold table, and differences between the two groups were analysed using χ2 test P < 0.05 signified statistical significance.
Results
Circ_0006174 expression is up-modulated in CRC tissues
The GSE138589 dataset was downloaded from the Gene Expression Omnibus database and analysed for screening out differentially expressed circRNAs using R software. The data revealed that circ_0006174 was overexpressed in CRC tissues (Figure 1(a) and (b)). Circ_0006174 expression in 46 cases of CRC tissues and their corresponding paracancerous tissues was subsequently examined by qRT-PCR. Circ_0006174 expression was found to be up-regulated in most CRC tissues compared with paracancerous tissues (Figure 1(c)). After nucleocytoplasmic separation, it was confirmed that circ_0006174 was mainly expressed in the cytoplasm (Supplemental Figure 1B). Additionally, the relationship between circ_0006174 expression and clinicopathological parameters of the CRC patients was analysed by the χ2 test, and the data showed that circ_0006174 overexpression was remarkably associated with enlarged tumour volume and advanced tumour, node, metastasis (TNM) stage (Table 1).
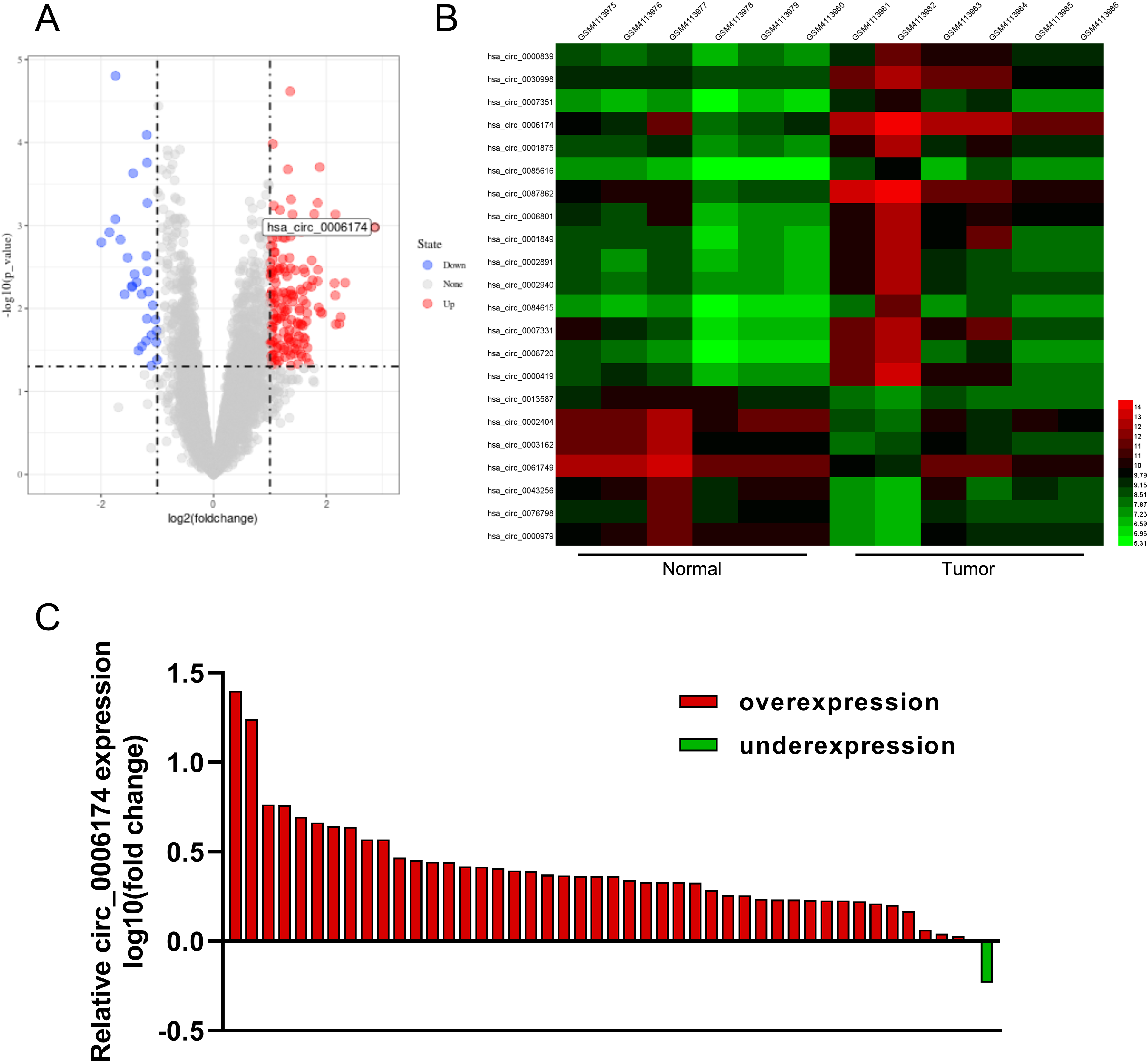
Circ_0006174 expression characteristics in CRC. (a) A volcano plot was used to show the changes of circRNA expression profile of CRC tissues compared with normal lung tissues in GSE138589. The red dots indicated the up-regulated circRNAs, and the blue dots indicated the down-regulated circRNAs. (b) Some differentially expressed circRNAs in GSE138589 were presented in the heatmap. (c) Circ_0006174 expression in tumour tissues and paracancerous tissues of 46 CRC patients was detected by qRT-PCR analysis.
Correlation between circ_0006174 expression and multiple clinicopathological characteristics in CRC patients (n = 46).
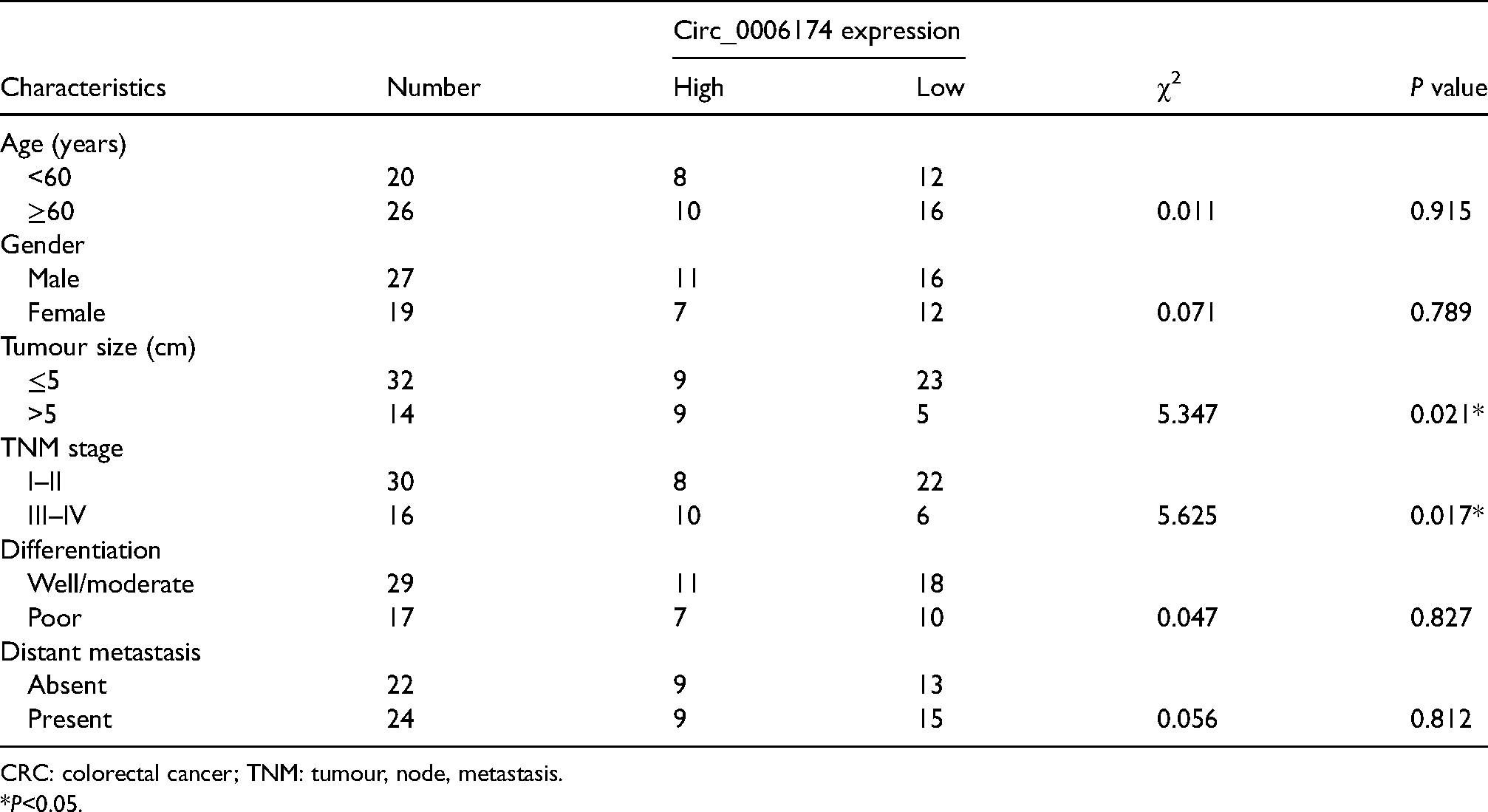

CRC: colorectal cancer; TNM: tumour, node, metastasis.
*P<0.05.
Circ_0006174 enhances the proliferation, migration and invasion of CRC cells and inhibited apoptosis
Circ_0006174 expression in FHC cells and human CRC cell lines (DLD-1, CACO2, SW480 and HT-29 cells) was examined by qRT-PCR. The data showed that circ_0006174 expression was markedly up-regulated in all four CRC cells relative to FHC cells (Figure 2(a)). The circ_0006174 overexpression plasmid was transfected into DLD-1 cells for constructing the circ_0006174 overexpression model. si-circ_0006174#1, si-circ_0006174#2 and si-circ_0006174#3 were transfected into HT-29 cells for constructing the circ_0006174 knockdown model. qRT-PCR showed that the transfection was successful (Figure 2(b)). Furthermore, cell proliferation was monitored by the CCK-8 experiment and BrdU experiment, which showed that circ_0006174 overexpression markedly enhanced the proliferation of DLD-1 cells and knockdown of circ_0006174 inhibited the proliferation of HT-29 cells relative to the control group (Figure 2(c) and (d)). Furthermore, flow cytometry showed that circ_0006174 overexpression significantly suppressed the apoptosis of DLD-1 cells, and knocking down circ_0006174 promoted the apoptosis of HT-29 cells (Figure 2(e)). Additionally, in Transwell experiments, circ_0006174 overexpression was found to promote the migration and invasion of DLD-1 cells, while knocking down circ_0006174 impeded the migration and invasion of HT-29 cells (Figure 2(f)).
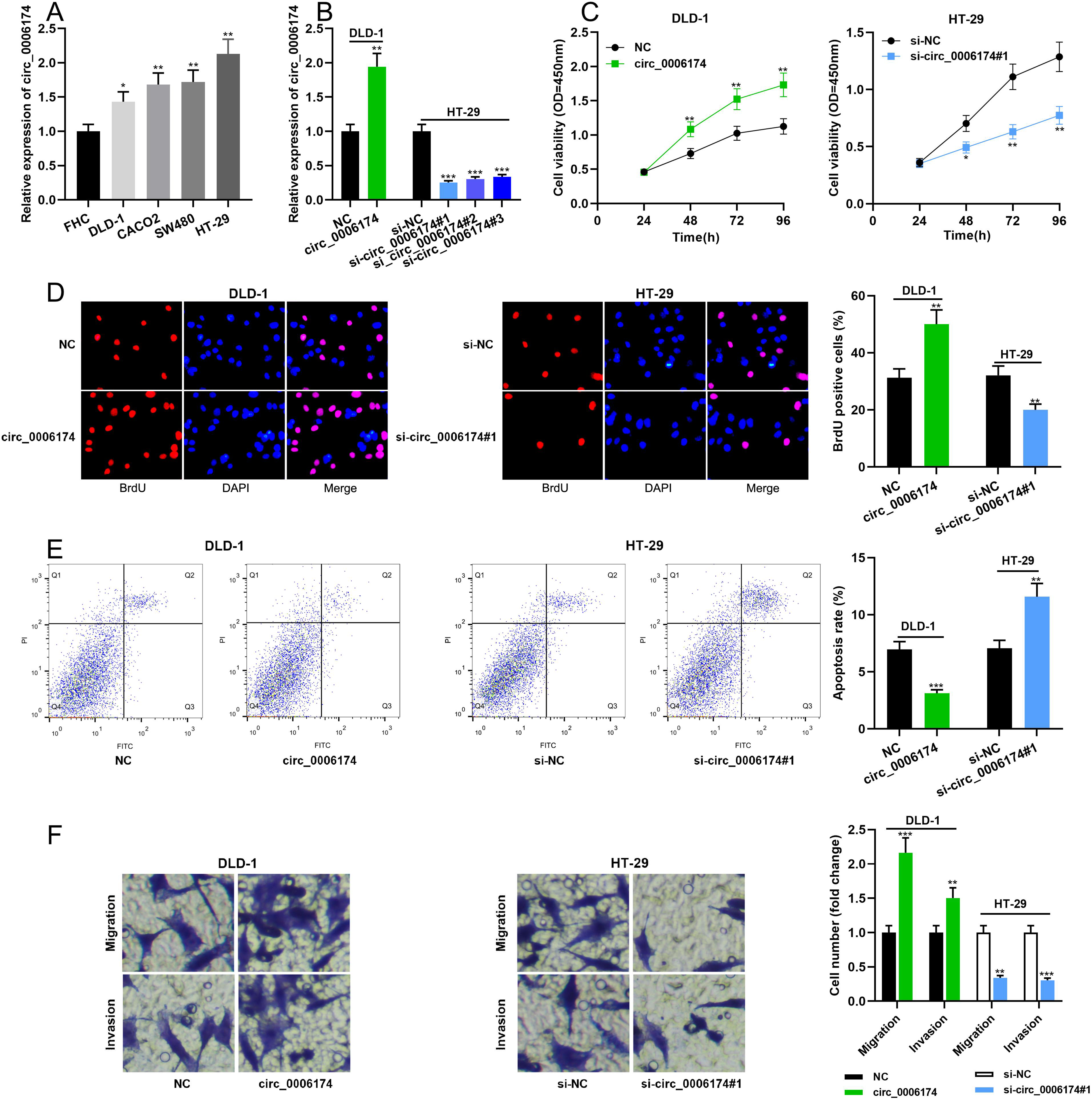
Effects of circ_0006174 on CRC cell proliferation, migration, invasion and apoptosis. (a) Circ_0006174 expression in FHC and four human CRC cell lines (DLD-1, CACO2, SW480 and HT-29 cells) was determined by qRT-PCR. (b) The transfection efficiency of circ_0006174 overexpression plasmid and circ_0006174 siRNAs was determined by qRT-PCR. (c and d) The effects of overexpression or knockdown of circ_0006174 on the proliferation of DLD-1 and HT-29 cells were detected using the CCK-8 method and BrdU experiment. (e) Apoptosis of DLD-1 and HT-29 cells was analysed by flow cytometry. (f) Transwell assay was used to detect migration and invasion of DLD-1 and HT-29 cells. *P < 0.05, **P < 0.01, ***P < 0.001.
Circ_0006174 directly targets miR-142-3p
Circular RNA Interactome database (https://circinteractome.nia.nih.gov/index.html) showed that there was a potential binding site between circ_0006174 and miR-142-3p (Figure 3(a)). Dual-luciferase reporter experiments unveiled that overexpression of miR-142-3p substantially inhibited the luciferase activity of circ_0006174 WT, while overexpression of miR-142-3p exerted no remarkable effect on the luciferase activity of circ_0006174 MUT (Figure 3(b)). The data of RIP experiments showed that relative to control IgG, circ_0006174 and miR-142-3p were remarkably enriched in Ago2-containing microribonucleoproteins, suggesting that the two could bind with each other directly (Figure 3(c)). Furthermore, circ_0006174 overexpression significantly suppressed miR-142-3p expression, while knocking down circ_0006174 exerted the opposite effect in CRC cells (Figure 3(d)). qRT-PCR revealed that miR-142-3p was lowly expressed in CRC tissue specimens (Figure 3(e)). Besides, the expression of circ_0006174 and miR-142-3p was negatively correlated in CRC tissues (Figure 3(f)).
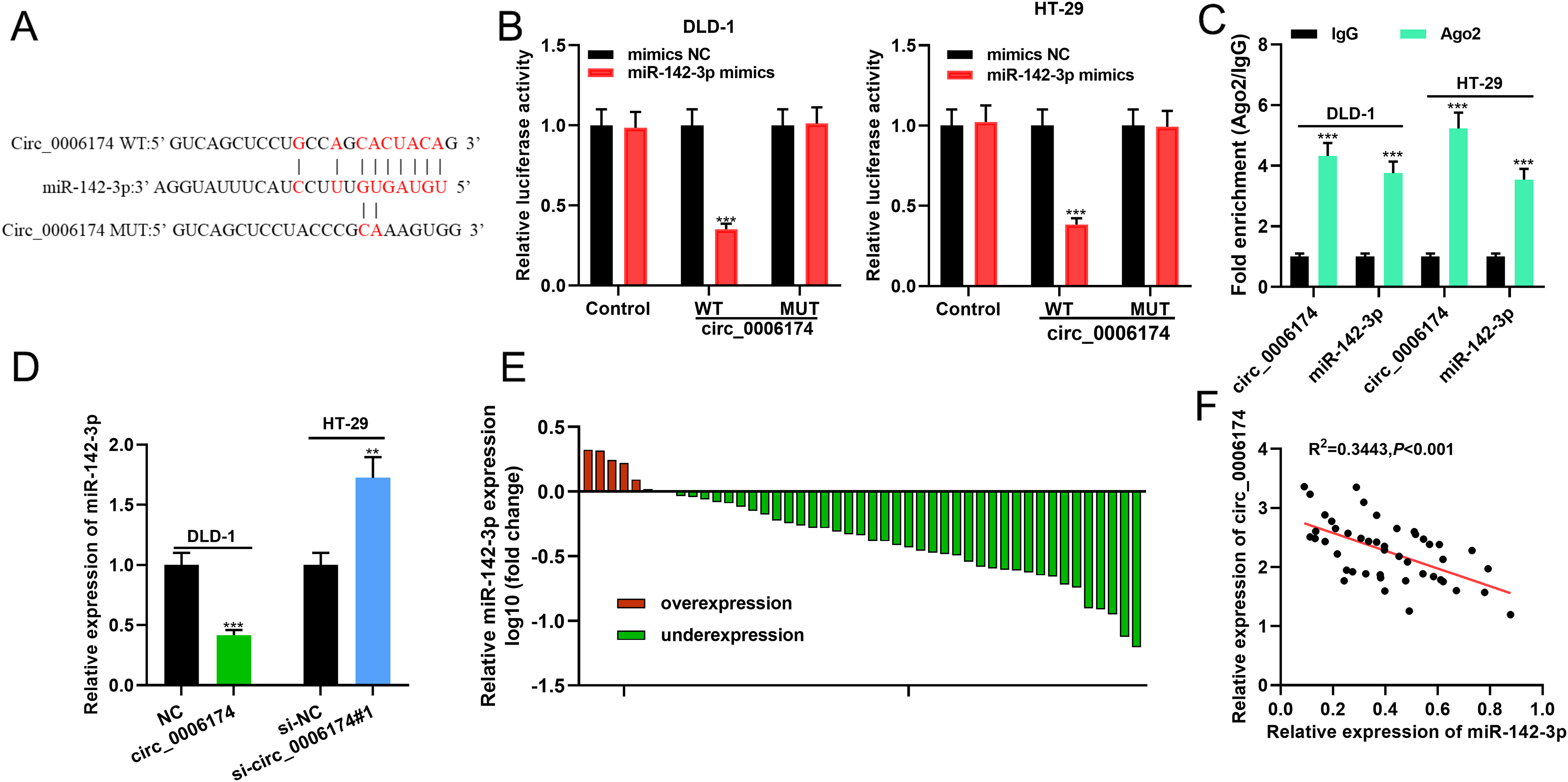
In CRC cells, miR-142-3p is the target of circ_0006174. (a) Bioinformatics was used to predict the binding sites between circ_0006174 and miR-142-3p. (b) The binding sequence was validated using the dual-luciferase reporter assay. (c) RIP experiments were used to further verify the binding relationship between circ_0006174 and miR-142-3p. (d) The effect of overexpression or knockdown of circ_0006174 on miR-142-3p expression in CRC cells was detected using qRT-PCR. (e) qRT-PCR was employed to detect miR-142-3p expression in 46 pairs of CRC tissues and paracancerous tissues. (f) Pearson's correlation analysis was utilised to analyse the correlation between circ_0006174 and miR-142-3p expression in CRC tissues. **P < 0.01, ***P < 0.001.
Circ_0006174 participates in modulating proliferation, apoptosis, migration and invasion of CRC cells through adsorbing miR-142-3p
To decipher whether circ_0006174 was involved in regulating the proliferation, apoptosis, migration and invasion of CRC cells through repressing miR-142-3p, circ_0006174 overexpression plasmid and miR-142-3p mimics were co-transfected into DLD-1 cells, and si-circ_0006174#1 and miR-142-3p inhibitors were co-transfected into HT-29 cells. qRT-PCR validated that the transfection was successful (Figure 4(a)). CCK-8 experiments, BrdU experiments, flow cytometry and Transwell experiments were performed and the data demonstrated that circ_0006174 overexpression enhanced the proliferation, migration, and invasion of DLD-1 cells and repressed apoptosis, while up-regulation of miR-142-3p attenuated this effect; conversely, knocking down circ_0006174 exerted the opposite effects, and down-regulation of miR-142-3p partially reversed the effects induced by circ_0006174 (Figure 4(b) to (f)).
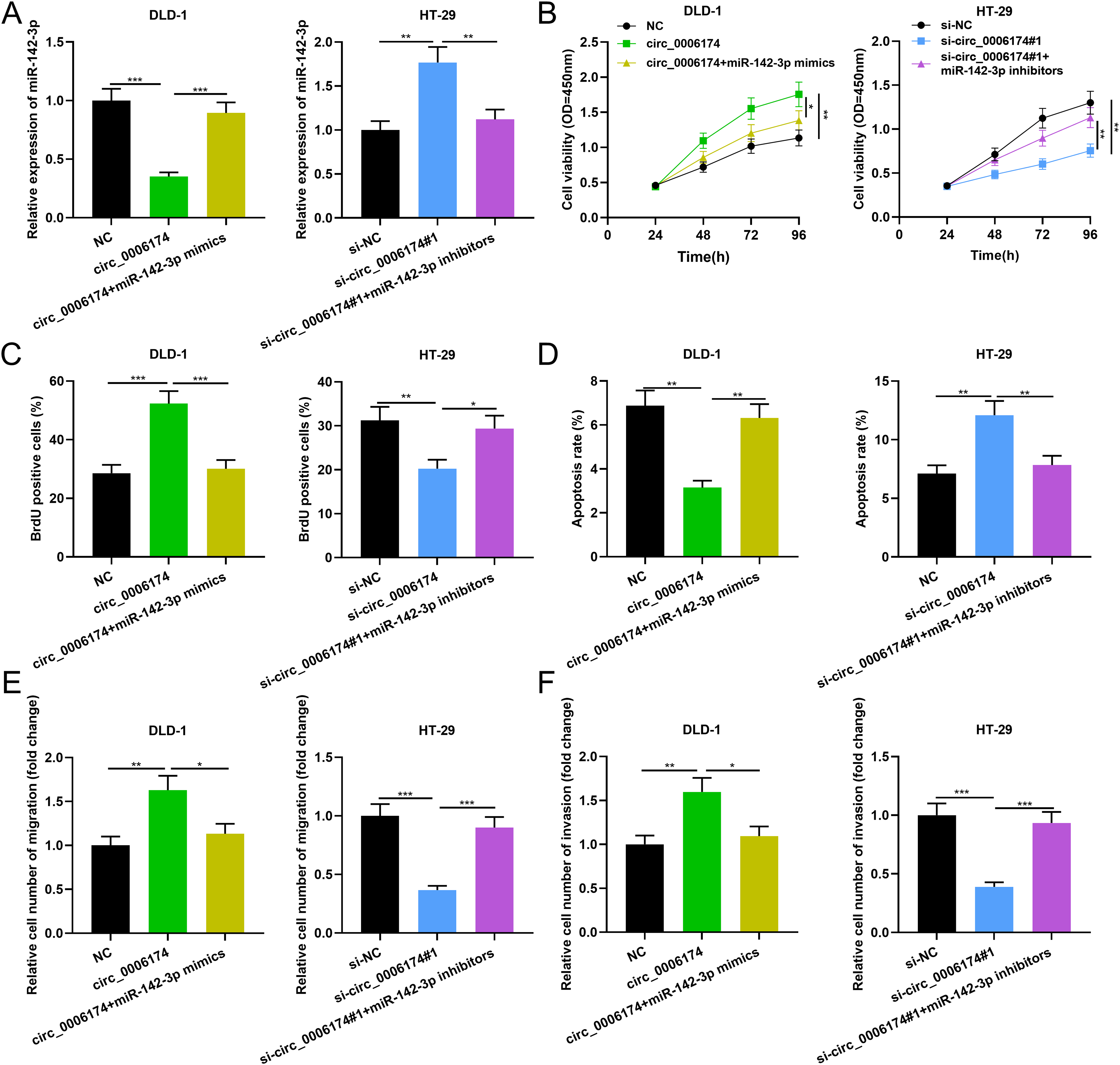
Effects of circ_0006174/miR-142-3p axis on proliferation, migration, invasion and apoptosis of CRC cells. (a) The transfection efficiency of co-transfected miR-142-3p mimics and circ_0006174 overexpression plasmids or co-transfected miR-142-3p inhibitors and si-circ_0006174#1 was determined using qRT-PCR. (b and c) The proliferation of DLD-1 and HT-29 cells was detected using the CCK-8 method and BrdU assay. (d) Apoptosis of DLD-1 and HT-29 cells was analysed by flow cytometry. (e and f) Transwell assay was used to detect migration and invasion of DLD-1 and HT-29 cells. *P < 0.05, **P < 0.01, ***P < 0.001.
Regulatory effects of circ_0006174/miR-142-3p axis on XIAP expression
Notably, a potential binding sequence between miR-142-3p and XIAP was identified by the StarBase database (http://starbase.sysu.edu.cn/) (Figure 5(a)). Dual-luciferase reporter gene experiments showed that up-regulation of miR-142-3p remarkably decreased the luciferase activity of XIAP WT, but exerted no remarkable effect on the luciferase activity of XIAP MUT (Figure 5(b) and (c)). Moreover, Western blot revealed that circ_0006174 overexpression or inhibiting miR-142-3p expression promoted the expression of XIAP in CRC cells while circ_0006174 knockdown or miR-142-3p overexpression worked oppositely (Figure 5(d)). Besides, the effects of circ_0006174 on XIAP expression could be reversed by the miR-142-3p (Figure 5(d)). Furthermore, XIAP mRNA was demonstrated to be overexpressed in CRC tissues (Figure 5(e)). In addition, XIAP mRNA expression was positively correlated with circ_0006174 expression and negatively correlated with miR-142-3p expression in CRC tissues (Figure 5(f)).
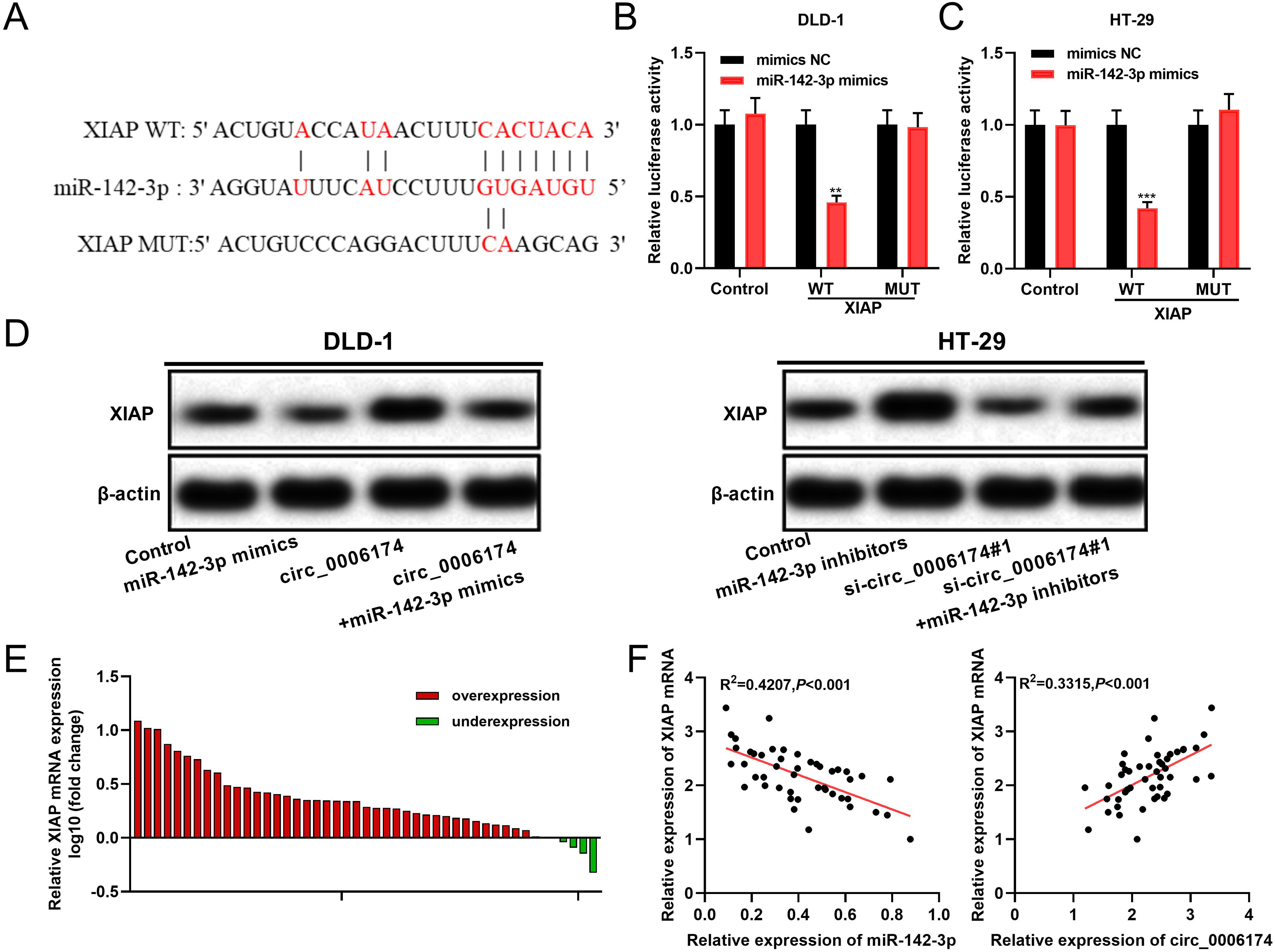
Circ_0006174 positively regulates XIAP expression via repressing miR-142-3p. (a) Bioinformatics was used to predict the binding site between miR-142-3p and XIAP 3′UTR. (b and c) The predicted binding site between miR-142-3p and XIAP 3′UTR was validated using the dual-luciferase reporter assay. (d) Western blot was performed to detect XIAP expression in DLD-1 and HT-29 cells, after circ_0006174 and miR-142-3p were selectively regulated. (e) XIAP mRNA expression in 46 pairs of CRC tissues and paracancerous tissues was detected by qRT-PCR. (f) Pearson's correlation analysis was employed to analyse the correlation between XIAP mRNA, miR-142-3p and circ_0006174 expression in CRC tissues. **P < 0.01, ***P < 0.001.
Discussion
With the development of second-generation sequencing technology and bioinformatics, a lot of studies have confirmed that circRNAs are essential in the progression of diverse diseases, including cardiovascular diseases, and viral infections and cancers.10-12 The high stability and tissue specificity of circRNAs endow them with the potential to be biomarkers, and circRNA has become a hotspot in cancer research in recent years. 13 Reportedly, circRNAs partake in tuomorigenesis through multiple mechanisms. 14 Specifically, circRNAs can form complexes with proteins to regulate the biological functions of the proteins, and circRNAs can also function as competitive endogenous RNAs to decoy miRNAs to modulate the regulatory function of miRNAs on their targets.15,16 CircRNAs is involved in regulating the malignant biological behaviours of multiple tumours, including CRC. 17 For instance, knockdown of circ_101951 blocks the epithelial–mesenchymal transition mediated by KIF3A, thereby suppressing the proliferation and metastasis of CRC cells. 18 Hsa_circ_0014717 overexpression restrains CRC cell proliferation and colony formation by up-regulating cell cycle inhibitory protein p16 expression. 19 Circ-ITGA7 exerts tumour-suppressive effects in CRC, impeding CRC cell proliferation and enhancing apoptosis of CRC cells in vitro. 20 In this work, circ_0006174 was revealed to be significantly overexpressed in CRC, and its high expression was significantly correlated with unfavourable clinicopathological characteristics of CRC patients. Functionally, circ_0006174 overexpression markedly enhanced the proliferation, migration and invasion of CRC cells and inhibited apoptosis. The results indicated that circ_0006174 was oncogenic in CRC. Notably, a recent study reports that circ_0006174 is up-regulated in CRC tissues, and overexpression of circ_0006174 promotes CRC cell proliferation, migration and invasion in vitro, and enhances xenograft tumour growth in BALB/c mice by up-regulating MACC1. 7 This report is similar to the findings of our study.
MiRNAs are crucial molecules implicated in the regulation of gene expression, by promoting the degradation of target mRNAs or impeding the translation process. 21 MiRNAs are involved in diverse biological processes such as cell differentiation, proliferation and apoptosis, and are vital in physiological and pathological processes.22,23 For example, up-regulation of miR-451a is validated to suppress CRC cell proliferation and induce apoptosis in CRC cells. 24 Moreover, miR-519b-3p is shown to impede the proliferation and invasion of CRC cells. 25 Importantly, miR-142-3p is down-regulated in CRC tissues, and up-regulation of miR-142-3p represses cell viability and colony formation by inducing cell cycle arrest. 26 Additionally, miR-142-3p expression is negatively regulated by circRNAs such as circ_0046701 and circ_VMA21.27,28 Herein, we verified that miR-142-3p substantially impeded the proliferation, migration and invasion of CRC cells and induced apoptosis, which is consistent with the previous report. 26 Furthermore, it was demonstrated that circ_0006174 directly targeted miR-142-3p to repress its expression, which partly explained the mechanism of miR-142-3p dysregulation in CRC.
XIAP is an important member of the inhibitor of apoptosis protein (IAP) family and the most potent apoptosis inhibitor in the IAP family. 29 XIAP is overexpressed in diverse tumour tissues, including oesophageal cancer, lung cancer and CRC, and its expression is closely correlated with tumour progression, recurrence, prognosis and chemoresistance.30,31 Caspases are the major executive proteins in apoptosis, and XIAP directly restrains caspases-3, caspases-7 and caspases-9 expression to impede apoptosis. 32 Reportedly, XIAP expression is markedly up-regulated in CRC, and its expression is correlated with tumour differentiation, vascular invasion, Duke stage and overall survival time of the patients.33,34 Additionally, XIAP expression is regulated by miRNA in CRC. For instance, miR-337-3p enhances the apoptosis and represses the proliferation and metastasis of CRC cells by targeting XIAP in CRC. 35 In this work, it was demonstrated that miR-142-3p directly targeted XIAP, and circ_0006174 overexpression up-regulated XIAP expression by adsorbing miR-142-3p. Collectively, we conclude that circ_0006174, miR-142-3p and XIAP forms a ceRNA network to regulate the progression of CRC.
In summary, this work reveals that circ_0006174 is overexpressed in CRC tissues and cells, and its overexpression is remarkably associated with unfavourable clinicopathological characteristics of CRC patients. Circ_0006174 enhances the proliferation, migration and invasion of CRC cells and impedes apoptosis via regulating the miR-142-3p/XIAP axis. The present study further deciphers the molecular mechanism of CRC development and provides a theoretical basis for exploring diagnosis biomarkers and therapeutic targets for CRC.
Supplemental Material
sj-docx-1-jbm-10.1177_17246008211034178 - Supplemental material for Circ_0006174 promotes colorectal cancer progression by sponging microRNA-142-3p and regulating X-linked inhibitor of apoptosis expression
Supplemental material, sj-docx-1-jbm-10.1177_17246008211034178 for Circ_0006174 promotes colorectal cancer progression by sponging microRNA-142-3p and regulating X-linked inhibitor of apoptosis expression by Bo Huang, Dejun Cui, Ying Ren, Xun Zhao, Fei Li and Wenqiang Yuan in The International Journal of Biological Markers
Footnotes
Authors contribution
Guarantor of integrity of the entire study: YR; Study concepts: YR; Study design: BH; Definition of intellectual content: DJC; Literature research: BH, DJC, YR, FL, and WQY; Conduction of experiments: BH, DJC, FL, and WQY; Clinical data acquisition: BH, DJC, FL, and WQY; Statistical analysis: BH, DJC, YR, FL, and WQY; Manuscript editing: BH, DJC, FL, and WQY. All authors read and approved the final manuscript.
Ethics statement
All experiments have been reviewed and approved by the Ethics Review Board of Guizhou Provincial People's Hospital. This study was performed following the Declaration of Helsinki.
Data availability statement
The data used to support the findings of this study are available from the corresponding author upon request
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Cultivation Fund of the National Natural Science Foundation ([2018]5764-11).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
