Abstract
Background:
miR-142-3p has proved to be involved in tumorigenesis and the development of renal cell carcinoma. The present study aimed to explore the prognostic value of miR-142-3p.
Methods:
Total RNA was extracted from renal cell carcinoma specimens and the expression level of miR-142-3p was measured. Pearson Chi-square test, Kaplan–Meier analysis, as well as univariate and multivariate regression analysis were performed to determine the correlation between miR-142-3p and the prognosis of renal cell carcinoma patients. Receiver operating characteristic curves were constructed to evaluate the predictive efficiency of miR-142-3p for the prognosis of renal cell carcinoma patients. Data from The Cancer Genome Atlas (TCGA) were utilized to validate our findings.
Results:
Our results demonstrated that upregulation of miR-142-3p was correlated with shorter overall survival (P=0.002) and was, in the meantime, an independent prognostic factor for renal cell carcinoma patients (P=0.002). The receiver operating characteristic curve combining miR-142-3p expression with tumor stage showed an area under the curve of 0.633 (95% confidence interval 0.563, 0.702). The result of TCGA data was consistent with our findings.
Conclusions:
Our results suggest miR-142-3p expression is correlated with poor prognosis of renal cell carcinoma patients and may serve as a prognostic biomarker in the future.
Introduction
Renal cell carcinoma (RCC)—the most common type of adult kidney cancer—accounts for 90% of renal malignancies and 2%–3% of all human tumors.1–3 The mortality rate of RCC is the highest among the genitourinary cancers and the incidence of RCC has increased worldwide over recent decades.4, 5 For early-stage RCC, a surgical approach may provide the possibility of cure. 6 However, many RCC patients are still misdiagnosed at an early stage. Approximately 20%–30% of RCC patients had already suffered from metastases at the time of diagnosis, while 20%–40% of postoperative patients still develop metastasis later. 7 There is an urgent need of available molecular biomarkers in RCC that are capable of detecting it early, supervising tumor progression, and indicating prognosis, for the purpose of guiding our clinical management and individual therapy.
microRNAs (miRNAs)—a group of endogenous non-coding RNA molecules—act as gene expression regulators by translation inhibition and degradation of RNAs through a bind with 3′ untranslated regions (UTRs). 8 Studies have shown that miRNAs are involved in the occurrence and progression of various cancers including RCC, probably as a result of their participation in biological processes, such as proliferation, differentiation, migration, invasion, and apoptosis. 9 Furthermore, previous research had proved the potential of miRNAs to be the biomarkers for detection and prognosis and even be novel therapeutic targets.10–12
miR-142-3p is a promising miRNA that has been proved to be involved with several malignancies.13–17 Moreover, a previous study has verified the correlation between miR-142-3p and prognosis of cervical cancer, revealing its potential to be a prognosis biomarker. 18 In order to explore the clinical significance of miR-142-3p, more investigation is necessary. In the present study, we concentrated on the potential value of miR-142-3p for predicting the prognosis of RCC patients after surgery. The expression level of miR-142-3p in 284 RCC specimens was assessed to evaluate its correlation with the prognosis of RCC patients. A Pearson Chi-square test was used to evaluate the relationship between miR-142-3p expression and clinical–pathological variables. A Kaplan–Meier method was conducted for survival analysis. Next, we utilized regression analyses to judge if the expression level of miR-142-3p was an independent predictor for the prognosis of postoperative RCC patients. To evaluate the predictive efficiency of miR-142-3p for RCC patients, receiver operating characteristic (ROC) curves were constructed. Furthermore, we downloaded data from the TCGA data portal to validate our findings.
Materials and methods
Patients and samples
Patients who underwent partial or radical nephrectomy and pathological diagnosed as RCC at Peking University Shenzhen Hospital (Guangdong, China) from April 2003 to May 2013 were included in this study (n=284). None of them had accepted chemotherapy and radiotherapy before surgery. All of the patients were followed up until May 2018 or death. All of their formalin-fixed paraffin-embedded (FFPE) tumor tissue were obtained and classified according to the 2010 American Joint Committee on Cancer (AJCC) staging system. We collected the clinical and pathological features of them, including age, gender, tumor size, and tumor stage (summarized in Table 1). All patients included in the study signed informed consent forms and the study was approved by the Ethics Committees of Peking University Shenzhen Hospital.
Association between miR-142-3p expression level a and clinical–pathological information in FFPE renal cancer samples.
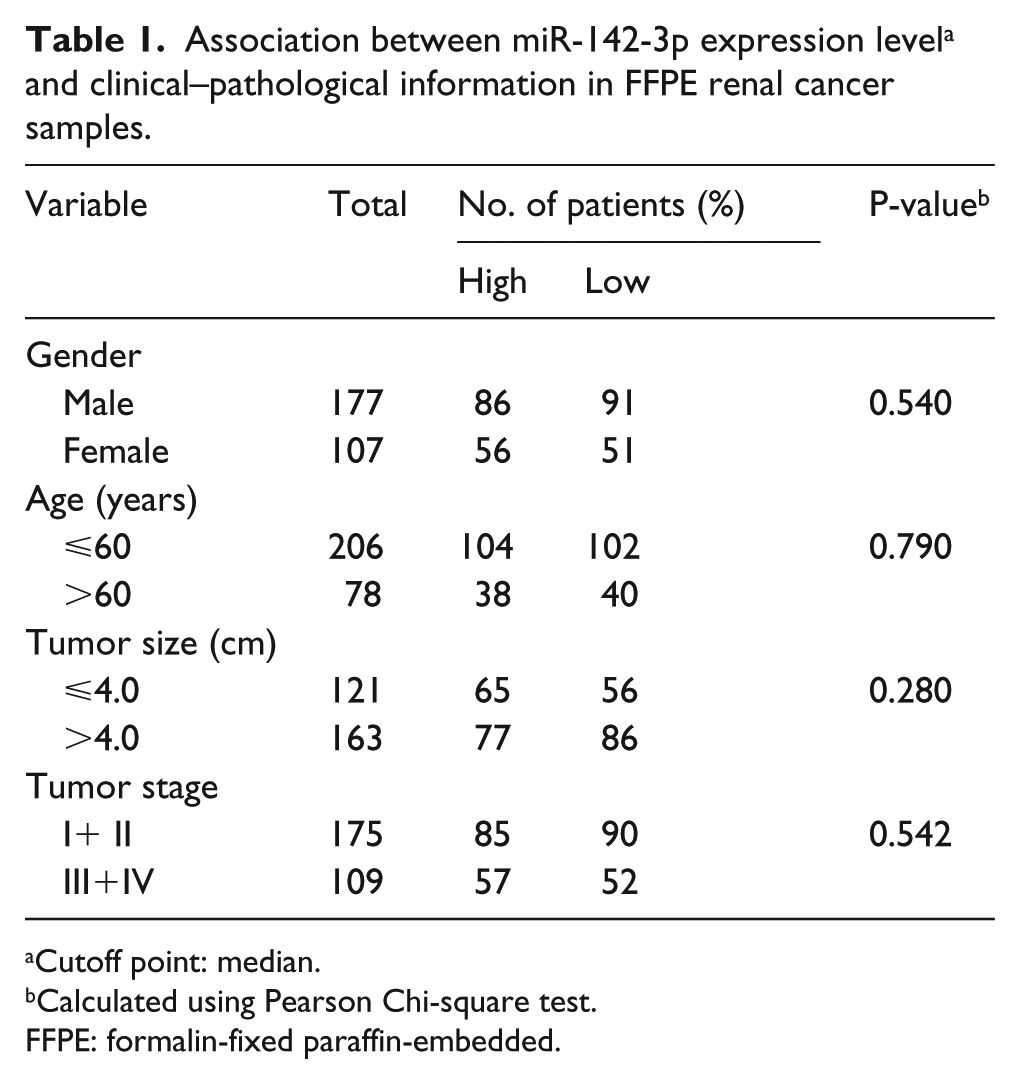
Cutoff point: median.
Calculated using Pearson Chi-square test.
FFPE: formalin-fixed paraffin-embedded.
RNA extraction and reverse transcription quantitative polymerase chain reaction
Total RNA was extracted using TRIzol (Invitrogen; Thermo Fisher Scientific, Inc., Waltham, MA, USA) from FFPE specimens and purified with the RNeasy Maxi kit (Qiagen), according to the manufacturer’s protocol. RNA concentration was determined on a NanoDrop 2000c (Thermo Fisher Scientific, Inc.). RNA samples with an optical density (OD) ratio at 260/280 of 1.8–2.1 were used for further investigation. Reverse transcription was performed with 1µg of total RNA using the miScript Reverse Transcription kit (Qiagen GmbH), as recommended by the manufacturer. The expression levels of miR-142-3p were detected with miScript SYBR-Green PCR kit (Qiagen) and quantitative polymerase chain reaction (qPCR) using the Roche LightCycler 480 Real-Time PCR system (Roche Diagnostics, Basel, Switzerland). The primers are listed as below: miRNA-142-3p forward: 5′-TGTAGTGTTTCCTACTTTATGGA-3′; reverse: universal primers (miScript SYBR Green PCR kit); U6 forward: 5′-CTCGCTTCGGCAGCACA-3′; reverse: 5′-ACGCTTCACGAATTTGCGT-3′. Thermocycling conditions were 95°C for 15 min, 40 cycles of 94°C for 15 sec, 55°C for 30 sec, and 72°C for 30 sec. (PCR was performed in triplicate.) The expression levels of miR-183-5p were determined using the 2(-Delta C(T)) method. 19
Statistical analysis
Pearson Chi-square tests were conducted to analyze the relation between miR-142-3p expression and clinical–pathological variables including gender, age, tumor size, and tumor stage. The Kaplan–Meier method was performed for survival analysis and the Log-rank test was used to evaluate the differences between the survival curves. We also classified the patients by gender, age, tumor size, and tumor stage, and conducted Kaplan–Meier analysis for each group. Univariate and multivariate Cox regression analyses were run to assess the prognostic significance of miR-142-3p expression as well as several clinical–pathological variables in RCC patients. ROC curves were constructed to examine the predictive efficiency of miR-142-3p. Statistical tests and P-values were two-sided. All the statistical analyses were performed using SPSS 19.0 (IBM SPSS, Armonk, NY, USA). All data were presented as the mean ± SD. A P-value < 0.05 was considered statistically significant.
Validation from The Cancer Genome Atlas
We compiled an in dependent data set of 506 cases of RCC patients from The Cancer Genome Atlas (TCGA; www.cancergenome.nih.gov). These data included miR-142-3p expression and survival time. Similarly, we conducted the Kaplan–Meier method, as mentioned, with our own data in order to validate our findings.
Results
Association between miR-142-3p expression level and clinical–pathological information
Our study utilized 284 RCC patients’ specimens (details on patients’ clinical–pathological information are listed in Table 1). We conducted a Pearson Chi-square test to assess the association between miR-142-3p expression level and clinical–pathological information. Clinical–pathological characteristics, including gender, age, tumor size, and stage, were irrelevant to the expression of miR-142-3p according to our results (Table 1).
Association between miR-142-3p expression level and survival
We performed the Kaplan–Meier method for survival analysis, finding that patients with higher miR-142-3p levels had a distinctly shorter overall survival (OS) compared with patients with low levels (P = 0.002) (Figure 1(b)). Later, we stratified our cases according to gender, age, tumor size, and tumor stage, conducting the Kaplan–Meier analysis correspondingly to evaluate if there was any difference between the survival of the high-level miR-142-3p group and the low-level group. The results revealed that the level of miRNA-142-3p expression had a better predictive value for prognosis in tumor <4m group (P=0.001) (Figure 2(e)) and stage Ⅲ/Ⅳ group (P=0.006) (Figure 2(h)). High-level patients have shorter OS in all ages (age<60y: P=0.032, age>60y: P=0.007), which may signify that age has no predictive value for RCC (Figure 2 (c) and (d)). Interestingly, we found that the miR-142-3p expression level was more preferable for the prediction of male (P=0.008) (Figure 2(b)).
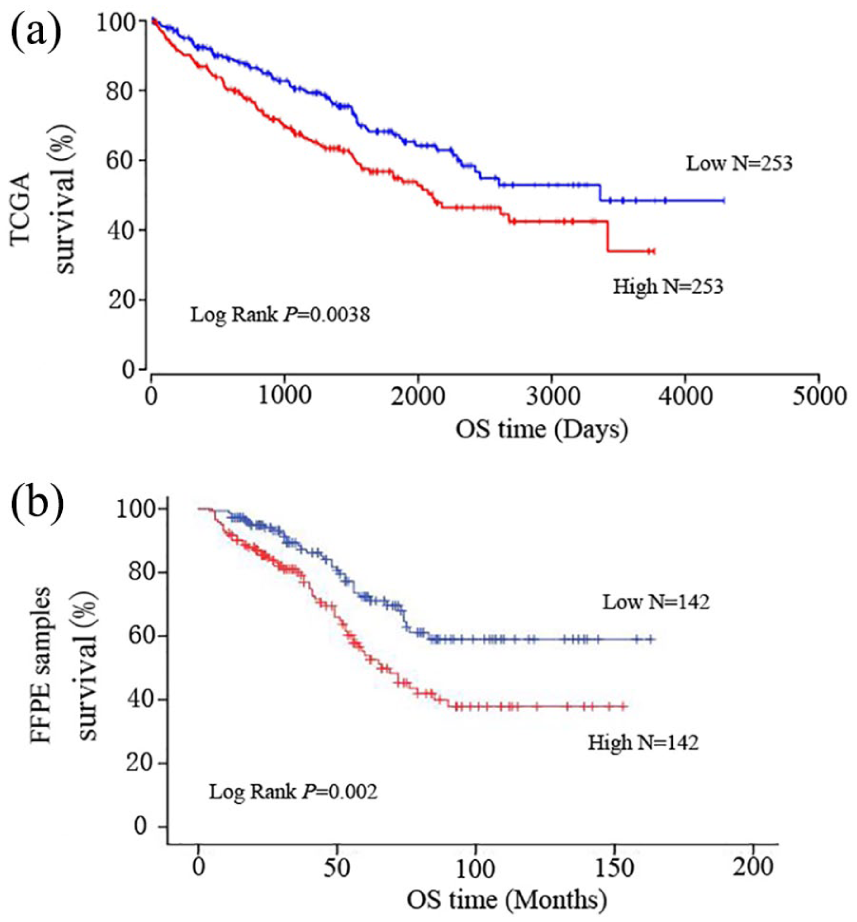
Kaplan–Meier analysis for renal cell carcinoma (RCC) patients based on the expression level of miR-142-3p, (a) with data from TCGA; (b) with data of our FFPE samples. P-value was calculated by log rank test.
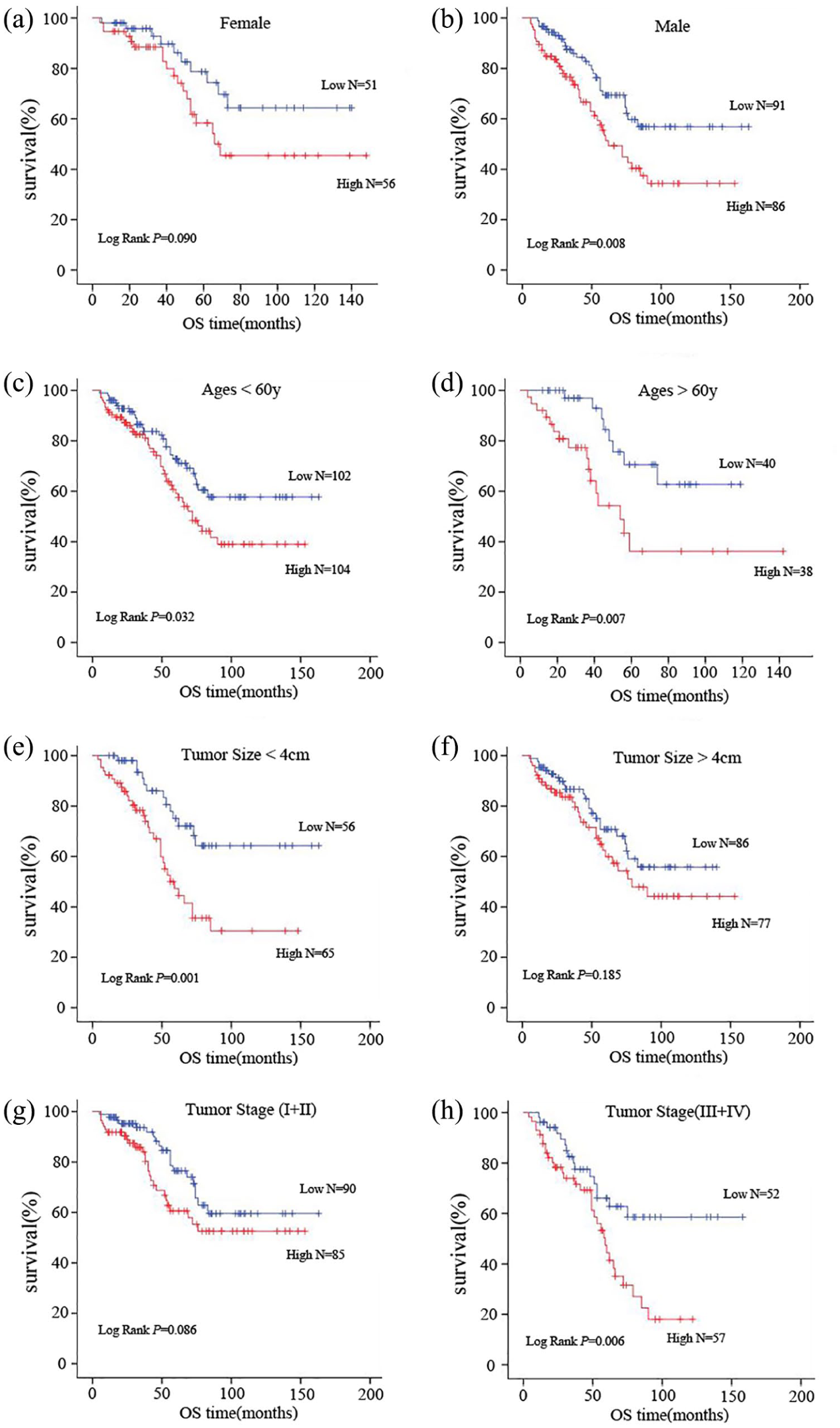
Kaplan–Meier analysis for renal cell carcinoma (RCC) patients based on the expression level of miR-142-3p in subgroups. Female (a) and male (b). Age<60y (c) and age>60y (d). Tumor size<4 cm (e) and tumor size>4 cm (f). Tumor stage (I+ II) (g) and tumor stage (III+IV) (H). P-value was calculated by log rank test.
Our results of univariate regression analysis negated the association between RCC patients’ survival and several variables, including age, gender, and tumor size. Statistical significance was seen in tumor stage (P=0.009) and miR-142-3p expression (P=0.002). On the basis of this result, we performed a multivariate regression analysis, verifying tumor stage (P=0.010) and miR-142-3p expression (P=0.002); both reached statistical significance (Table 2).
miR-142-3p expression and patients’ survival.
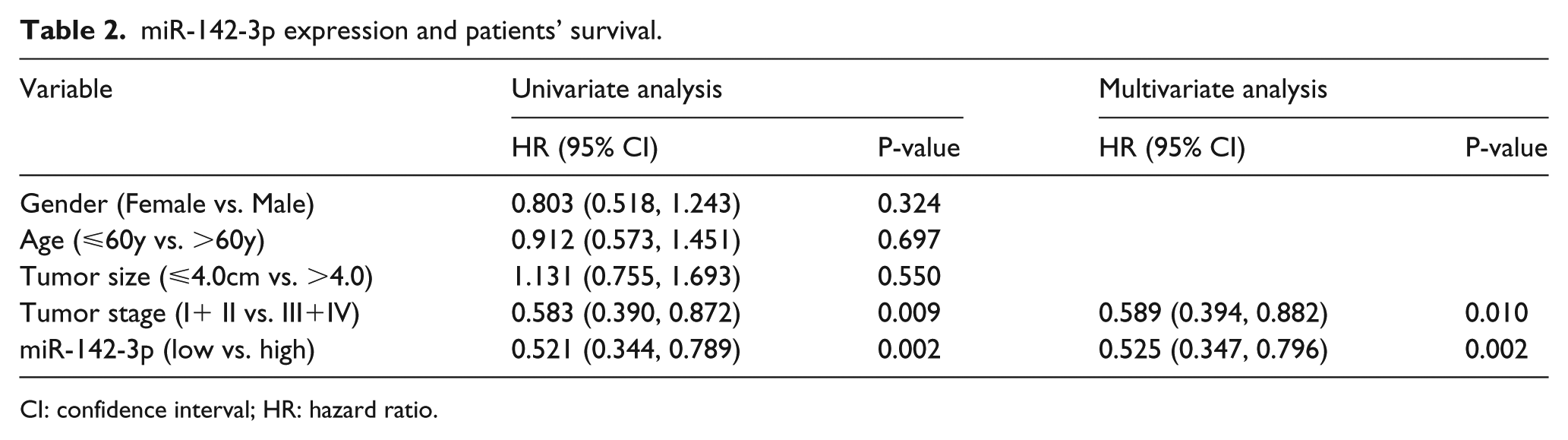
CI: confidence interval; HR: hazard ratio.
To evaluate the predictive efficiency of miR-142-3p for the prognosis of RCC patients, we constructed ROC curves with miR-142-3p expression, tumor stage, and a combination of both. The area under the curve (AUC) of miR-142-3p expression alone was 0.591 (95% CI 0.521, 0.661) with a sensitivity and specificity of 62% and 56%. The AUC of tumor stage alone was 0.591 (95% CI 0.520, 0.662) with a sensitivity and specificity of 51% and 68%. The combination of miR-142-3p expression and tumor stage was capable of improving predictive efficiency, showing an AUC of 0.633 (95% CI 0.563, 0.702) with a sensitivity and specificity of 34% and 87%, respectively (Figure 3).
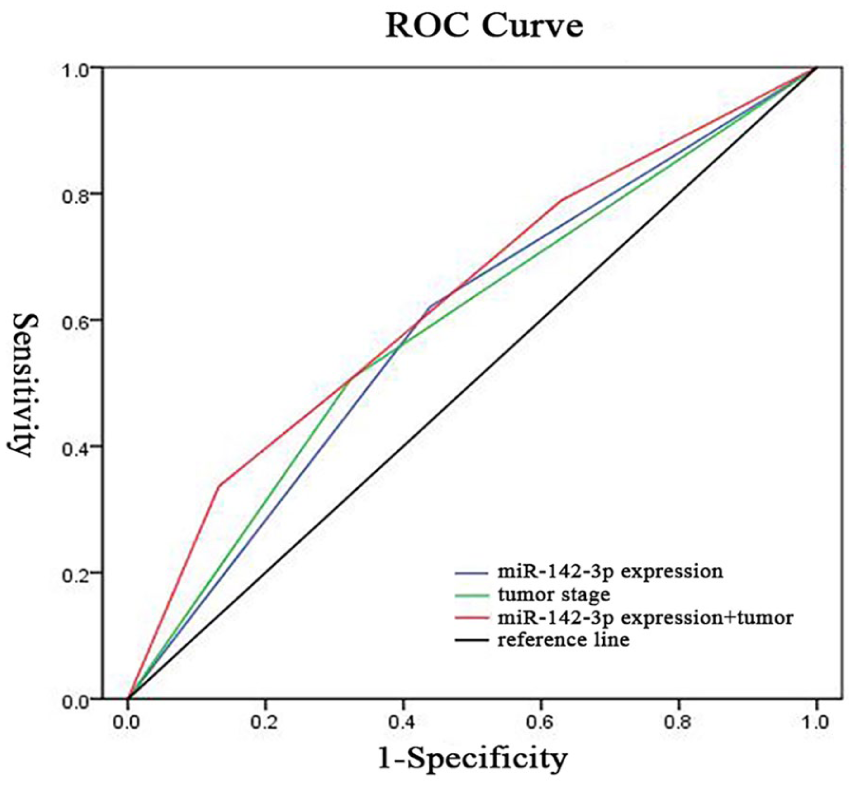
ROC curves for predictive efficiency of miR-142-3p expression and tumor stage in renal cell carcinoma patients. AUC of miR-142-3p expression and tumor stage combined was 0.633 (95% confidence interval 0.563, 0.702) with a sensitivity and specificity of 34% and 87%, respectively. AUC: area under the curve; ROC: receiver operating characteristic.
Combining the above results, it is reasonable to believe that miR-142-3p has the potential to be an independent prognostic marker.
Validation of miR-142-3p prognostic significance on TCGA
To validate our findings, we obtained data set of 506 RCC patients from TCGA. Kaplan–Meier analysis revealed that RCC patients with higher miR-142-3p expression level underwent a shorter OS, which is consistent with our findings (Figure 1(a)).
Discussion
The morbidity of RCC has gradually increased over the past 20 years. 20 Often, RCC are either asymptomatic or nonspecific in the early stage and are easily misdiagnosed. 21 Although the advances of detection and surgery have improved the survival and life quality of RCC patients to some extent, approximately 30% of patients undergoing curative nephrectomy still developed recurrence and metastasis. 22 For those patients with poor prognosis, timely use of comprehensive treatment may lead to a better outcome. For this purpose, discovering accurate prognostic biomarkers is necessary. However, up to now there have not been any biomarkers in clinical application to predict the prognosis of RCC patients.
In recent years miRNAs have gained focus worldwide. Some of them had already been confirmed to be potential prognostic biomarkers in tumors.11, 23–25 In the present study, we concentrated on miR-142-3p, which had already been proved to be involved in several malignancies. Previous studies have shown miR-142-3p was downregulated in various cancers, such as gastric cancer, cervical cancer, colorectal carcinoma, hepatocellular carcinoma, leukemia, acting as a tumor suppressor.26–30 Li et al. 18 confirmed the relation between the low expression of miR-142-3p and the poor prognosis of cervical patients, claiming that it may be a potential biomarker. Meanwhile, miR-142-3p was found to be a potential biomarker for esophageal squamous cell carcinoma by Lin et al. 31
miR-142-3p was also proved to be upregulated in RCC by Lokeshwar et al. 32 and was confirmed to be involved in the migration, proliferation, and apoptosis procedures of RCC by Li et al. 33 Our study aimed to explore the relationship between miR-142-3p expression and the prognosis of RCC patients. We assessed the expression level of miR-142-3p in 284 RCC specimens, and later utilized Pearson Chi-square test, regression analyses, and the Kaplan–Meier analysis to estimate its prognostic value. Our results revealed that the upregulation of miR-142-3p corresponded with shorter OS (P=0.02) (Figure 1(b)). In addition, the upregulation of miR-142-3p was an independent factor to predict a poor prognosis of RCC (P=0.02) (Table 2). Similar results were obtained with data from TCGA (Figure 1(a)). ROC curves analysis indicated that miR-142-3p was useful for differentiating patients with good prognosis from those with poor prognosis (AUC 0.633, 95% CI 0.563, 0.702, sensitivity and specificity 34% and 87%, respectively) (Figure 3). These results revealed that miR-142-3p was a novel prognostic biomarker for RCC, with a high specificity but a lack of sensitivity.
In fact, some researchers have tried to combine several miRNAs into a signature to improve the detectability of miRNA markers. For instance, Lokeshwar et al. 32 identified a three-miRNA signature that is potentially able to predict RCC patients’ outcome, showing higher accuracy. Our present study not only proves miR-142-3p as a novel RCC prognostic marker, but also lays a foundation for future research on a miRNA-combined signature.
It is worth mentioning that our present research findings are based on FFPE specimens, which are mainly worthy for the treatment and management of postoperative patients. In fact, serum and urine miRNAs as novel biomarkers have unique superiority for their non-invasion and wider application, 34 which reveal a new direction for our future research.
Conclusion
In summary, this is the first study to explore the prognostic value of miR-142-3p for RCC patients. Our research findings revealed that high miR-142-3p expression is correlated with shorter OS and is an independent factor to predict the poor prognosis of RCC patients. This reminds us of its potential to be a novel biomarker for predicting the prognosis of RCC patients. More investigation in the future may push it into clinical application.
Footnotes
Author contributions
Xiqi Peng and Xiang Pan contributed equally to this study.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by Basic Research Project of Peking University Shenzhen Hospital (JCYJ2017001, JCYJ2017004, JCYJ2017005, JCYJ2017006, JCYJ2017007, JCYJ2017012), Clinical Research Project of Peking University Shenzhen Hospital (LCYJ2017001), Science and Technology Development Fund Project of Shenzhen (no. JCYJ20180507183102747), and Clinical Research Project of Shenzhen Health Commission (no. SZLY2018023).
