Abstract
X-linked inhibitor of apoptosis protein functions as an intrinsic regulator of apoptosis by inhibition of caspase activity and possesses a pivotal role in human cancer development and progression. A growing body of literature has demonstrated that microRNAs lead to the degradation or translational repression of messenger RNAs by binding to the non-coding region of messenger RNA at the 3′-untranslated region. Here, we revealed that the expression of HMGA2 is upregulated with X-linked inhibitor of apoptosis protein after transfection of X-linked inhibitor of apoptosis protein 3′-untranslated region in hepatocellular carcinoma cells, suggesting that X-linked inhibitor of apoptosis protein 3′-untranslated region serves as a competitor for microRNAs and prevent the co-targeted messenger RNA, HMGA2, from being suppressed. We further identified that let-7a-5p could bind to both the X-linked inhibitor of apoptosis protein 3′-untranslated region and HMGA2 3′-untranslated region. Moreover, we demonstrated that the forced expression of X-linked inhibitor of apoptosis protein 3′-untranslated region increases the oncogenicity of hepatocellular carcinoma cells in vitro. Cell functional analyses were performed to examine the association of HMGA2 status and X-linked inhibitor of apoptosis protein 3′-untranslated region. We have also measured the functional readout of let-7a-5p and HMGA2, an assay often employed to provide substantial evidence for the effects of X-linked inhibitor of apoptosis protein 3′-untranslated region on hepatocellular carcinoma cells. In general, our findings suggest that X-linked inhibitor of apoptosis protein 3′-untranslated region serves as a competitive endogenous RNA for HMGA2 to activate hepatocellular carcinoma progression by arresting endogenous let-7a-5p.
Keywords
Introduction
Hepatocellular carcinoma (HCC) has been reported to rank as the fifth most prevalent cancer globally, with a high overall rate of cancer-related deaths.1–3 A high mortality rate of HCC makes it one of the most lethal solid-organ cancers.4,5 Although some advances have been made in the diagnosis and treatment of HCC, the prognosis remains extremely poor due to late diagnosis and ineffective treatment. 6
X-linked inhibitor of apoptosis protein (XIAP) is a member of the inhibitor of apoptosis family of proteins (IAP). Among the IAPs, XIAP is the most extensively studied. 7 XIAP has been demonstrated to be powerfully anti-apoptotic by directly binding to caspases and inhibiting their activity. The loss or inhibition of XIAP results in cells that are sensitized to death receptor–induced cell death. 8 Increasing evidence has also indicated that aberrant expression of XIAP is relevant to many human cancers and is correlated with enhanced chemo- or radiation resistance.9–11
For many decades, cancer has been regarded as a genetic disease. 12 Only 1%–2% of the human genome is translated into proteins, while 70% of the genome is transcribed. Various experimental studies have demonstrated that most genomes are repressed by non-coding RNAs (ncRNAs), implicating involvement in the epigenetic regulation of gene expression. 13 MicroRNAs (miRNAs) include some ncRNAs which function in RNA interference. MiRNAs are 18–24 nucleotides in length and often function as negative regulators by partially binding to the non-coding region of the target protein coding gene.14,15 The non-coding region, especially the 3′-untranslated region (UTR), is the most important regulatory element of the messenger RNAs (mRNAs). The function of the 3′-UTR is generally achieved by its crucial function in the stability, nuclear transport, cellular localization, and translational efficiency of mRNAs.16,17 The stability of target mRNAs and their translation can be altered through miRNA–mRNA 3′-UTR interactions. 18 However, the role of XIAP 3′-UTR in the multistep process of human HCC remains to be clarified.
Protein-coding genes exert their functions not only by protein translation but also by regulating other mRNA through their ability to share multiple miRNA recognition elements (MREs). 19 The reduction of the targeting miRNAs is often accompanied by the de-repression of other mRNAs with common binding sites. 20 On the basis of this concept, we first confirmed that XIAP 3′-UTR regulates oncogenicity in SMMC-7721 and HepG2 cells. The regulatory roles of XIAP 3′-UTR in HCC cells were further confirmed as it upregulates the expression of HMGA2. We also found that the overexpression of let-7a-5p significantly abrogated XIAP 3′-UTR-induced HCC progression. Thus, our results suggest that XIAP 3′-UTR serves as a competitor for HMGA2 by arresting endogenous let-7a-5p in human HCC. Our study may provide novel perspectives into the development and progression of human HCC.
Materials and methods
Cell lines and culture
Two human HCC cell lines (SMMC-7721 and HepG2) were obtained from the American Type Culture Collection (ATCC, USA). Both cell lines were cultured in serum- or antibiotic-supplemented Dulbecco’s modified Eagle’s medium (DMEM) and maintained in a humidified incubator under 5% CO2 at 37°C.
Construct generation
To study the effects of XIAP 3′-UTR on HCC cell lines, we utilized complementary DNA (cDNA) that was obtained by reverse transcription polymerase chain reaction (RT-PCR) of RNA prepared from HepG2 cells. The primers for the cDNA were as follows: 5′-CTACTATAGAGTTAGAGGATCCTTAAGACATAAAAATTTTTGCTTG-3′ (forward) and 5′-GATATCTGCGGCCTAGCTAGCTGACGGACCGCGCCCGGTGTCTC-3′ (reverse). The PCR product was digested with BamH I and Hind III and inserted into pcDNA3.1 plasmid to create XIAP 3′-UTR, which was verified by DNA sequencing. Furthermore, we successfully established stable cell lines, including XIAP 3′-UTR or vector plasmid transfected SMMC-7721 and HepG2 cells. XIAP 3′-UTR and HMGA2 3′-UTR were cloned into psiCHECK2 luciferase reporter vector (Promega, USA) for luciferase reporter gene assays.
RNA oligonucleotides and transfection
MiRNA (let-7a-5p) mimic, small interfering RNAs (HMGA2), and negative control RNA were purchased from GenePharma (China). The negative control RNA was transfected as a contrast to eliminate the potential non-sequence-specific effects. Transfection was performed in the final concentration of 50 nM by LipofectamineTM 2000 Transfection Reagent (Invitrogen, USA). After 48–72 h, cells were collected and examined by quantitative RT-PCR (qRT-PCR) and western-blot analysis. For the stable transfection assay, Lipofectamine 2000 and XIAP 3′-UTR-pcDNA3.1 plasmid were co-transfected into the cell lines. Then, puromycin was used for screening and culturing surviving cells after 1 month. Establishment of stable cell lines provided convenient condition for follow-up experiments.
RNA extraction and qRT-PCR
Total RNA was isolated with RNA TRIzol reagent (Invitrogen), and cDNA was generated by TransScript Probe-One-Step qRT-PCR SuperMix (TransGen Biotech, China) according to the manufacturer’s protocol. cDNA was synthesized from 5 µL of total RNA, 4 µL all mix, 1 µL genomic DNA (gDNA) remover, and 10 µL RNase-free water. Gene-specific primers including XIAP 3′-UTR-A and XIAP 3′-UTR-B, HMGA2, and glyceraldehyde 3-phosphate dehydrogenase (GAPDH) were used (Table 1). qRT-PCR assays were performed on a Stratagene Mx3000P real-time thermocycler. Transfection efficiency was confirmed by qRT-PCR analysis. The threshold cycle (Ct) is defined as the fractional cycle number at which the fluorescence passes the fixed threshold. The fold change of mRNA expression was calculated using the 2−ΔΔCt method after normalization to GAPDH expression. 21
The sequence of the oligonucleotide primers used for qRT-PCR and transfection.
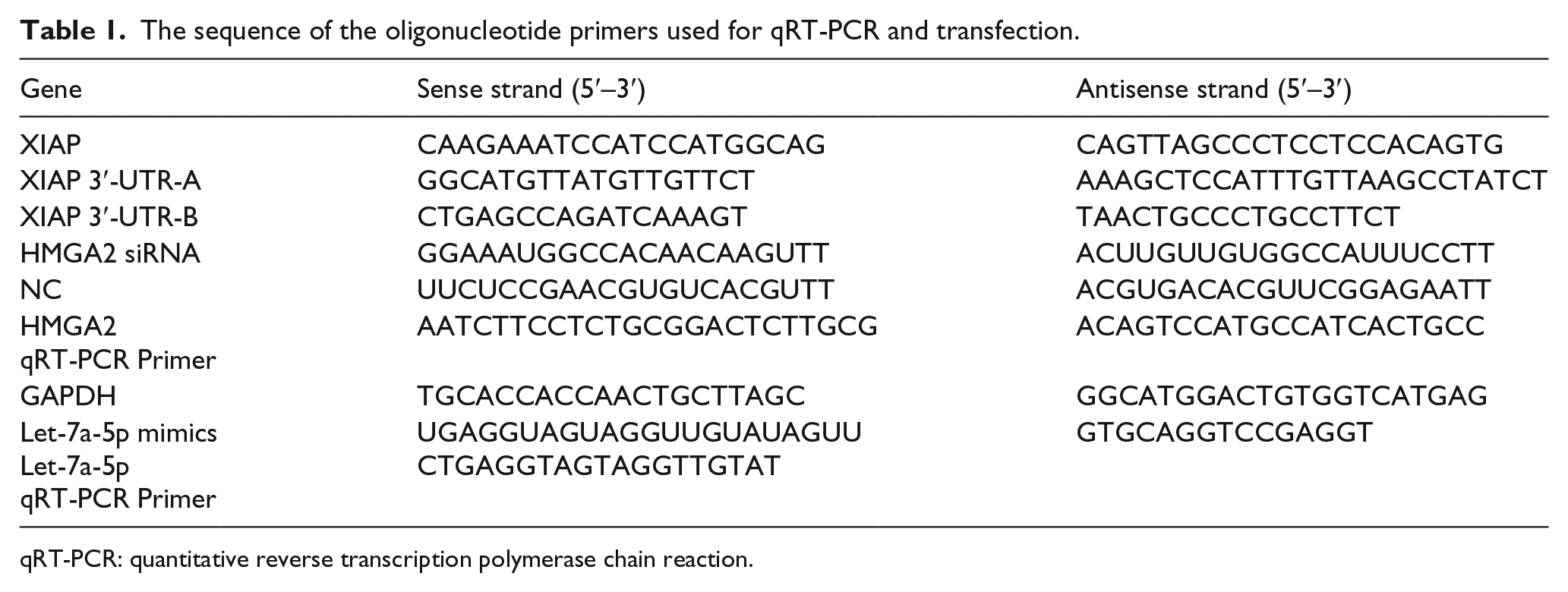
qRT-PCR: quantitative reverse transcription polymerase chain reaction.
Western-blot analysis
Proteins were extracted from cultured cells with radioimmunoprecipitation assay (RIPA) buffer (Beyotime, China) for 15 min at 4℃. Then, equal amounts (60 µg) of protein were used for 12% sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to nitrocellulose (NC) or polyvinylidene difluoride (PVDF) membranes.22,23 The antibodies used were as follows: β-actin (60008-1-Ig; Proteintech, USA), XIAP (#14334; Cell Signaling Technology, USA), and HMGA2 (20795-1-AP; Proteintech, USA).
Luciferase reporter gene assay
To analyze miRNA targets, XIAP 3′-UTR and HMGA2 3′-UTR were cloned into the downstream of Renilla luciferase gene in a psiCHECK2 vector (Promega). Then, 100 ng plasmids and 200 nmol/L let-7a-5p mimic or negative control were co-transfected in 24-well plates using Lipofectamine 2000. The cells broke open and luciferase activity was measured by Dual-Luciferase Assay (Promega) after 48 h. PsiCHECK2 vector (Promega) was co-transfected as an internal control.
Cell proliferation assay
A total of 1 × 105 cells per well were seeded into six-well plates in DMEM medium (2 mL) containing 10% fetal bovine serum (FBS), and three wells were included in each group. Cell suspension was mixed and diluted before taking aliquot cells to cell counting chamber. The cells were harvested by exposure to trypsin, the cell numbers were counted, and the cell density was estimated under microscope at same time points for up to 5 days. A minimum of three reliable counts was necessary.
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide tetrazolium reduction assay
A total of 2 × 103 cells per well in 96-well culture plates were incubated at 37℃ for a desired period of exposure. Then, 10 µL yellow 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) reagent was added into each well. The cultures were incubated for 4 h until purple precipitate was visible; 100 µL dimethyl sulfoxide (DMSO) was added to dissolve the formazan crystals and placed on a plate shaker for 15 min to ensure complete solubilization. Finally, the quantity of formazan (presumably directly proportional to the number of viable cells) was measured by recording changes in absorbance at 570 nm using an automatic microplate reader (Thermo Fisher Scientific, USA). The experiments were repeated in triplicate. The data were treated as mean ± standard deviation (SD).
Colony formation assay
For colony formation assays, logarithmic phase of monolayer cultured cells (empty vector- and XIAP 3′-UTR-transfected SMMC-7721 or HepG2 cells) were seeded at the density of 1 × 103 in six-well dishes in culture medium (2 mL) containing FBS. After 7–14 days, cells were fixed with 4% paraformaldehyde and stained with 1% crystal violet. Cells were photographed and cell numbers counted under a light microscope.
Migration and invasion assays
Migration and invasion assays were performed using uncoated (8 μm pore size; Corning Costar, USA) filters in 24-well plates. In both assays, XIAP 3′-UTR-transfected cells (1 × 105) were seeded or co-transfected transiently with HMGA2 siRNA or let-7a-5p mimics and control RNA in triplicate. For invasion assays, the inserts were coated with 100 µL of diluent Matrigel in advance. The upper chamber of each well was filled with 300 µL of culture medium while the lower chamber was provided with 800 µL of culture medium supplemented with 10% FBS. After incubation for 12–48 h at 37℃ with 5% CO2, cells that migrated to the lower surface of the membrane pores in the inserts were fixed with 4% paraformaldehyde and stained with 0.1% crystal violet. Migrated or invaded cells were photographed from representative areas for quantification under a microscope.
Statistical analyses
All the experiments were performed at least in triplicate, and statistical data were shown as mean ± SD. All statistical significance was assessed by the Student’s t-test; p < 0.05 was considered as statistically significant.
Results
The expression of XIAP and HMGA2 was upregulated by let-7a-5p in XIAP 3′-UTR-transfected cells
XIAP 3′-UTR (6798bp; Genebank access number: NM_001204401) was cloned and inserted into pcDNA3.1 plasmid, downstream of the cytomegalovirus (CMV) promoter, producing the XIAP 3′-UTR construct (Figure 1(a)). To investigate the effects of forced expression of XIAP 3′-UTR on HCC progression, we selected SMMC-7721 and HepG2 cells to further study. qRT-PCR assays were performed to confirm that the XIAP 3′-UTR was transcribed into RNA in HCC cell lines. To facilitate observation, we designed two sets of primers (named as XIAP 3′-UTR-A and XIAP 3′-UTR-B according to the sequence of XIAP 3′-UTR) using Primer Premier 5.0. We found that the mRNA levels of XIAP 3′-UTR in the stably transfected HCC cells were significantly upregulated compared to those of the control cells (Figure 1(b)).

Let-7a-5p interacted with XIAP 3′-UTR and regulated the expression levels of XIAP and HMGA2 in XIAP 3′-UTR-transfected cells. (a) The pcDNA3.1-CMV-XIAP 3′-UTR construct was made and was denoted as XIAP 3′-UTR. (b) Expression of XIAP 3′-UTR was analyzed by qRT-PCR after XIAP 3′-UTR stable transfection at mRNA level. XIAP 3′-UTR-transfected SMMC-7721 and HepG2 cells were successfully established. (c and d) qPCR assays and western-blot assays showed upregulation of XIAP and HMGA2 at RNA and protein expression levels. (e) Let-7a-5p could work as a common binding site of XIAP 3′-UTR and HMGA2 3′-UTR by three computational prediction algorithms. (f) Relative luciferase activity analysis for let-7a-5p overexpressing cells after stable transfection with XIAP 3′-UTR (*p < 0.05, **p < 0.01 (t-test)). (g) The expression of let-7a-5p was shown by qRT-PCR analysis of total RNA from XIAP 3′-UTR and negative control cells.
To determine how XIAP 3′-UTR exerted its biological functions, we analyzed perturbation of gene expression in SMMC-7721 and HepG2 cells after XIAP 3′-UTR transfection. The expression level of HMGA2 was significantly increased with XIAP 3′-UTR expression according to the microarray data in the XIAP 3′-UTR-transfected cells compared with that in control cells. 24 Since HMGA2 was reported to enhance cell proliferation and metastasis in HCC,25,26 we thus speculated HMGA2 is involved in the progression of HCC upon forced expression of XIAP 3′-UTR. In accordance with the results from the transcriptome analysis, qRT-PCR assays confirmed that the mRNA expression levels of both XIAP and HMGA2 in XIAP 3′-UTR-transfected cells increased over two-fold compared to negative control cells (Figure 1(c)). Similarly, the protein expression of XIAP and HMGA2 was also increased in XIAP 3′-UTR-transfected cells compared with control (Figure 1(d)). These results imply that HMGA2 is involved in the effect of XIAP 3′-UTR on HCC cells.
We hypothesized that the forced expression of XIAP 3′-UTR may antagonize specific miRNAs, thus relieving translational inhibition of the mRNA of HMGA2 from that particular miRNA. Three computational prediction algorithms (TargetScan, 27 miRanda, 28 and Pictar 29 ) were utilized to search for the potential downstream targets involved in this regulation. As shown in Figure 1(e), we observed that XIAP 3′-UTR provided 118 predicted target sites for miRNAs, while HMGA2 3′-UTR only possessed only 25. The shared binding sites of XIAP 3′-UTR and HMGA2 3′-UTR in let-7a-5p were of particular interest. To determine whether let-7a-5p directly targets XIAP 3′-UTR and HMGA2 3′-UTR, we then generated three luciferase constructs which contain the fragments of XIAP 3′-UTR and HMGA2 3′-UTR harboring the binding sites for let-7a-5p and a construct inserted with psiCHECK2 vector as a negative control. Cells were co-transfected with synthetic let-7a-5p mimics, and relative luciferase activity was analyzed. As shown in Figure 1(f), in HepG2 cells, luciferase activity was significantly reduced when constructs with XIAP 3′-UTR and HMGA2 3′-UTR were co-transfected with let-7a-5p mimics, as compared with negative control. To further detect whether let-7a-5p was relevant to XIAP 3′-UTR, we assessed the relative expression of let-7a-5p by qRT-PCR assays in XIAP 3′-UTR-transfected SMMC-7721 and HepG2 cell lines (Figure 1(g)). Compared with control, let-7a-5p was inhibited in XIAP 3′-UTR-transfected cell lines.
Forced expression of XIAP 3′-UTR promoted proliferation, viability, colony formation, migration, and invasion in SMMC-7721 and HepG2 cells
Functional assays were performed to examine the role of XIAP 3′-UTR in HCC cells. As shown in Figure 2(a), the expression of XIAP 3′-UTR increased cell proliferation in a time-dependent manner compared with the control. MTT assays were also used to determine the effects of XIAP 3′-UTR on cell viability. Cell viability was increased with forced expression of XIAP 3′-UTR in comparison to control (Figure 2(b)). In colony formation assays, cells transfected with XIAP 3′-UTR displayed larger and more numerous colonies than those in the control (Figure 2(c)). Furthermore, migration assay revealed that the forced expression of XIAP 3′-UTR increased cell migration through pores in the culture inserts than in the controls (Figure 2(d)). Similarly, as compared to the control group, the forced expression of XIAP 3′-UTR resulted in an increased invasion rate of HCC cells (Figure 2(e)). Taken together, our results strongly suggest that XIAP 3′-UTR could function as a promoter in progression of HCC.

The overexpression of XIAP 3′-UTR promoted proliferation, viability, colony formation, migration, and invasion in SMMC-7721 and HepG2 cells. (a) Cell proliferation assays were performed to assess the growth of XIAP 3′-UTR-transfected cells. (b) Cells were plated in 96-well plates; 48 h later, cell viability was examined by microplate reader. (c) Colony formation assay was performed to compare the size and number of colonies. (d) Migration rates were monitored using migration assays. The XIAP 3′-UTR cells traveled faster through the membrane. (e) Matrigel-coated Transwell chambers were used for cell invasion assay. XIAP 3′-UTR-transfected cells have a stronger capacity of invasion as compared to the controls. **p < 0.01.
XIAP 3′-UTR increased the progression of HCC by upregulating the expression of HMGA2
To determine whether the effect of XIAP 3′-UTR expression in HCC on the progression was directly mediated by HMGA2, we introduced HMGA2 siRNA in functional studies. HMGA2 siRNA was transiently transfected in cells with forced expression of XIAP 3′-UTR, and the efficacy of HMGA2 depletion was verified using western-blot assays (Figure 3(a)). We next performed a series of functional assays in cells with forced expression of XIAP 3′-UTR either with HMGA2 siRNA or negative control. As expected, the depletion of HMGA2 expression significantly abrogated the increased cell proliferation, viability, colony formation, migration, and invasion in either XIAP 3′-UTR -transfected SMMC-7721 or HepG2 cells (Figure 3(b)–(f)).

Knockdown of HMGA2 repressed malignant potential of XIAP 3′-UTR-transfected cells in vitro. (a) The efficiency of HMGA2 silencing was verified using western blotting in SMMC-7721 and HepG2-XIAP 3′-UTR-transfected cells. (b and c) Cell proliferating ability and growth density were detected by proliferation assays and MTT assays. It was found that XIAP 3′-UTR-transfected cells with knockdown of HMGA2 have a slower growth rate relative to the control cells. (d) Total colony numbers were shown in SMMC-7721 and HepG2 cells. Knockdown of HMGA2 repressed the capacity of colonies formation. (e and f) Transwell assays were performed to observe vicious behaviors of XIAP 3′-UTR-transfected cells after HMGA2 silencing. Fewer cells traveled through membrane pores that were detected in cells with HMGA2 siRNA transfection. Photographs of the cell migration and invasion through the polycarbonate membrane stained with crystal violet were shown.
Let-7a-5p reversed the effects of XIAP 3′-UTR in SMMC-7721 and HepG2 cells by targeting both XIAP and HMGA2
After HCC cells with forced expression of XIAP 3′-UTR were transfected with let-7a-5p mimics, the cell lysates were subjected to western-blot analysis. We observed a significant repression of XIAP and HMGA2 protein levels after transfection of let-7a-5p mimics, compared with negative control (Figure 4(a)). Notably, the effects of XIAP 3′-UTR on HMGA2 were mediated by let-7a-5p-induced de-repression.
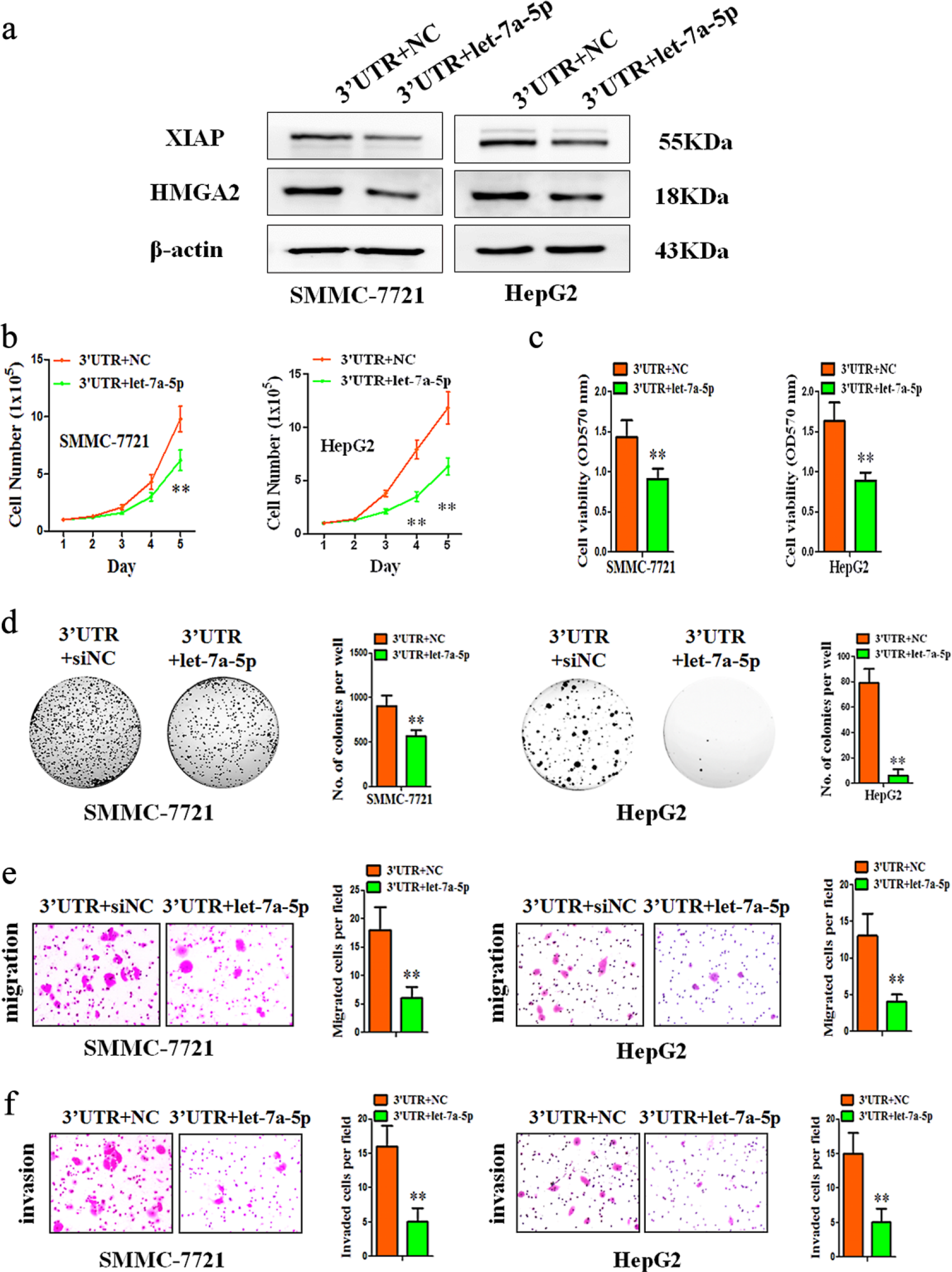
Overexpression of let-7a-5p significantly abrogated XIAP 3′-UTR-induced HCC progression. (a) Lysates of SMMC-7721-XIAP 3′-UTR cells or HepG2-XIAP 3′-UTR cells transfected with NC or let-7a-5p mimics were analyzed using western blot. Expression of let-7a-5p downregulated the expression of XIAP and HMGA2 protein. β-actin was used as input control. (b and d) The effects of let-7a-5p on cell proliferation, viability, and colony formation were tested. The regulation of XIAP 3′-UTR was mediated by let-7a-5p. (e and f) The number of migrated and invasive cells per field was counted after 12–48 h. The overexpression of let-7a-5p downregulated the migration and invasion rate.
To further determine whether the effects of the XIAP 3′-UTR expression on HCC progression were mediated by let-7a-5p, we performed functional studies in the presence of let-7a-5p on XIAP 3′-UTR-transfected cells. As expected, transfection of let-7a-5p mimics significantly abrogated increased cell proliferation, viability, colony formation, migration, and invasion in XIAP 3′-UTR-transfected HCC cells, compared with negative control (Figure 4(b)–(f)).
Discussion
Recent literature has reported that the majority of cellular transcripts undergo multipath and complex post-transcriptional regulation. 30 The sites of underpinning post-transcriptional regulations are located at both the mRNA 5′UTR and the coding region, but most target sites are present in the 3′-UTR. Binding sites for the same miRNA in 3′-UTR s exhibit a stronger capacity of regulation than those in coding sequences. 31 Therefore, we focused on the relationship between 3′-UTR and miRNAs. At the post-transcriptional level, these regulatory sequence elements within the 3′-UTRs of mRNAs provide multiple binding sites for regulatory factors.32,33 Ectopic expression of a 3′-UTR there often modulates miRNA functions and cell activities.34,35
In this study, we describe the ceRNA functions of XIAP 3′-UTR in SMMC-7721 and HepG2 cells. Notably, the forced expression of XIAP 3′-UTR, spanning 6798 bp, could sequester multiple miRNAs, functionally relieving translational inhibition of endogenous mRNAs, thereby increasing their protein expression levels. 3′-UTR s have been identified to sequester endogenous miRNAs and hence exert wide-spread influence on mRNA translation and stability. Therefore, modulation of gene expression by 3′-UTR may exert effects on tumor development and progression. 36 Publicly available data have indicated that TUSC2 3′-UTR sequesters miRNAs and leads to increased translation of TUSC2. 37 Hence, understanding the aberrant expression of transcripts as a result of modulation by XIAP 3′-UTR is of immense importance. Indeed, we observed that the expression of both XIAP and HMGA2 was increased in XIAP 3′-UTR-transfected cells. 24 These data are consistent with those obtained from the qRT-PCR and western-blot assays. This suggests that the regulatory effect of XIAP 3′-UTR on the expression of HMGA2 is at both the RNA and protein levels. However, it is increasingly obvious that the 3′-UTR s of the mature transcripts interacts with the expression of genes by competitively targeting the shared miRNAs. One example is that p21 3′-UTR may promote tumor development by interaction with miR-296, thus leading to the downregulation of p21WAF1. 38 Additionally, Wang et al. 36 reported that CYP4Z1 3′-UTR represses the expression of E-cadherin through binding to miR-9. Such information prompted us to explore the possible mechanism of miRNA–mRNA interaction of XIAP 3′-UTR. We hypothesized that the expression of XIAP 3′-UTR upregulated the expression of XIAP and HMGA2 by competing for binding to the same miRNA. Concordant with this idea, it is pertinent to note that the HMGA2 3′-UTR has seven conserved sites for let-7 miRNA. 39 Moreover, let-7-directed repression could be mediated by HMGA2 3′-UTR. 40 Although previous studies have proven that 3′-UTR can function both as a modulator of miRNA function and as a mediator of miRNA regulation, our analyses and results provide greater details. Thus, three target prediction algorithms were applied to screen common binding sites of XIAP and HMGA2 3′-UTR. To identify potential miRNA targets, we focused on let-7a-5p according to our bioinformatic analyses. Luciferase reporter gene assay and qRT-PCR analysis confirmed the existence of let-7a-5p binding sites in the 3′-UTR s of XIAP and HMGA2 mRNA. This is sufficient to further delineate the relationship of the respective miRNA and mRNA.
XIAP 3′-UTR might play a role in the progression of HCC. Nevertheless, our in vitro data revealed that XIAP 3′-UTR upregulated the proliferation and metastatic potential of HCC cells. Herein, we observed a high rate of cell proliferation in HCC cells with forced expression of XIAP 3′-UTR. Similar observations were also obtained from MTT and colony formation assays. Cell migration and invasion are generally associated with tumor progression and malignancy. 41 As such, we observed that forced expression of XIAP 3′-UTR in HCC cells increased cell migration and invasion. These results suggest that XIAP 3′-UTR would be functionally involved in tumor progression and that XIAP 3′-UTR should be further evaluated as a biomarker for predicting patient survival.
It is known that the IAPs affect cancer cell survival. 42 Increased expression of XIAP is observed in many cancer tissues and is associated with clinical outcome.43–47 HMGA2 serves as a transcription modulator through alteration of chromatin structure both locally and globally, modulating the transcriptional regulatory complex by recruitment of other proteins to transcriptional complexes. 48 HMGA2 protein is present at low levels in various normal adult human tissues, but is greatly increased in human cancers. 49 HMGA2 is involved in proliferation, differentiation, and invasion and metastasis of cancer cells. 50 It is therefore plausible that XIAP and HMGA2 may promote the metastatic capacity of HCC. Furthermore, we observed that the siRNA mediated depletion of HMGA2 downregulated malignant transformation of XIAP 3′-UTR -transfected cells. Our results implied that XIAP 3′-UTR indeed had a high affinity with tumorigenesis by upregulating HMGA2.
As described earlier, 3′-UTR s of XIAP and HMGA2 mRNA possess a common binding site for let-7a-5p. Thus, the XIAP 3′-UTR was analyzed to determine the sequence that interacts with let-7a-5p and affect the other transcript, HMGA2. Let-7a-5p inhibited XIAP expression by targeting its 3′-UTR. In contrast, let-7a-5p was downregulated with forced expression of XIAP 3′-UTR in HCC cell lines. Hence, the forced expression of the XIAP 3′-UTR may reduce the degradation of XIAP and HMGA2 by binding to let-7a-5p both on RNA and protein levels. As such, a novel concept involving ceRNAs has been observed. Typically, several miRNAs regulate UTRs, long non-coding RNA (lncRNA) or pseudogenes, indicative of cooperative translational control. Reciprocally, one miRNA may possess more than one target gene, reflecting target multiplicity. 28 On this basis, ceRNAs harbor MREs that can regulate other genes at a post-transcriptional level. 51 It is widely recognized that most binding sites are located in the 3′-UTR of mRNA. 20 XIAP 3′-UTR might act as a ceRNA for other genes by competing for limited pools of certain miRNA. Thus, our study indicates that XIAP 3′-UTR may function as a ceRNA to increase the expression of XIAP and HMGA2 in the development and progression of HCC, as increased expression of XIAP 3′-UTR provides more MREs for let-7a-5p.
In summary, we hypothesize that XIAP 3′-UTR could serve as a sink to activate let-7a-5p and inhibit degradation and translational repression of XIAP and HMGA2, thereby promoting oncogenicity of HCC cells. Hence, XIAP 3′-UTR functions as a ceRNA that promotes HMGA2 function in the progression of HCC by binding endogenous let-7a-5p. Although there are many reports of translational regulation by 3′-UTR s, the underlying mechanism remains to be clarified. The cellular functions of XIAP 3′-UTR can be exploited and potentially applied to gene therapy.
Footnotes
Acknowledgements
W.-Y.W. and S.-Q.T. have contributed equally to this work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported in part by grants from the National Nature Science Foundation of China (grant nos 81572305 and 81472493), the Program for Excellent Talents and the Scientific Research Program from Anhui Medical University (2013xkj006), Anhui provincial academic and technical leader reserve candidate (2016H074), Key Program of Outstanding Young Talents in Higher Education Institutions of Anhui (gxyqZD2016046 and 6), and Specialized Research Fund for the Doctoral Program of Higher Education (20133420120006).
