Abstract
Background:
Genetic variations in oxidative stress-related genes may alter the coded protein level and impact the pathogenesis of breast cancer.
Methods:
The current study investigated the associations of functional single nucleotide polymorphisms in the NFE2L2, HMOX1, P21, TXNRD2, and ATF3 genes with the early-stage breast cancer clinicopathological characteristics and disease-free survival, metastasis-free survival, and overall survival. A total of 202 Eastern European (Lithuanian) women with primary I–II stage breast cancer were involved. Genotyping of the single nucleotide polymorphisms was performed using TaqMan single nucleotide polymorphisms genotyping assays.
Results:
The CA+AA genotypes of P21 rs1801270 were significantly less frequent in patients with lymph node metastasis and larger tumor size (P=0.041 and P=0.022, respectively). The TT genotype in ATF3 rs3125289 had significantly lower risk of estrogen receptor (ER), progesterone receptor (PR) negative, and human epidermal growth factor receptor 2 (HER2) positive status (P=0.023, P=0.046, and P=0.040, respectively). In both, univariate and multivariate Cox analysis, TXNRD2 rs1139793 GG genotype vs. GA+AA was a negative prognostic factor for disease-free survival (multivariate hazard ratio (HR) 2.248; P=0.025) and overall survival (multivariate HR 2.248; P=0.029). The ATF3 rs11119982 CC genotype in the genotype model was a negative prognostic factor for disease-free survival (multivariate HR 5.878; P=0.006), metastasis-free survival (multivariate HR 4.759; P=0.018), and overall survival (multivariate HR 3.280; P=0.048).
Conclusion:
Our findings suggest that P21 rs1801270 is associated with lymph node metastasis and larger tumor size, and ATF3 rs3125289 is associated with ER, PR, and HER2 status. Two potential, novel, early-stage breast cancer survival biomarkers, TXNRD2 rs1139793 and ATF3 rs11119982, were detected. Further investigations are needed to confirm the results of the current study.
Introduction
Breast cancer (BC) is the most common cancer in women worldwide and is the second leading cause of cancer-related deaths among women. 1 It is a heterogeneous disease due to many phenotypes and different clinical and morphological presentations. Despite early detection, up to 10%–30% of early-stage BC patients still develop incurable metastatic disease. 2 Understanding the factors related to BC survival is essential in order to identify women at risk for fatal metastatic BC at earlier stages of the disease.
Oxidative stress has a key role in breast carcinogenesis as well as BC progression. 3 Overproduction of reactive oxygen species (ROS) stimulates the malignant phenotype through stimulus to sustained proliferation, death evasion, angiogenesis, invasiveness, and metastasis. 4 Antioxidants protect healthy cells from DNA damage induced by ROS, while malignant cells may be defended against chemotherapy or radiotherapy. 5
One of the oxidative stress-related proteins is Nuclear Factor, Erythroid 2 Like 2 (NFE2L2). NFE2L2 mediates protection against oxidants itself and controls the expression of other antioxidant response element–dependent genes. 6 In BC, NFE2L2 activation leads to cell-cycle progression, reduced cellular adhesion, increased migration, inhibition of the estrogen-related receptor α expression, and tumor chemo- and radioresistance.5,7,8 Heme oxygenase-1 (HO-1), encoded by the HMOX1 gene, is one of the NFE2L2-dependent antioxidants. Through degradation of pro-oxidant heme, HO-1 exhibits cytoprotective, anti-apoptotic, and anti-inflammatory properties. HO-1 is usually upregulated in tumors. Its expression is induced upon chemo- and radiotherapy, which decreases the effectiveness of the treatment. 9
Another gene in the NFE2L2 signaling pathway involved in antioxidant processes is P21. Despite its role in antioxidant protection, P21 also regulates many other cellular processes in a P53-dependent and -independent manner. P21 can promote cellular differentiation and senescence, and inhibit gene transcription and apoptosis. For BC patients, higher P21 expression is associated with more aggressive disease phenotype, lower survival, and worse response to systemic therapy.10,11
For the malignancies where oxidative stress that hinders tumor progression overwhelms NFE2L2-dependent anti-oxidant defense, upregulation of another gene thioredoxin reductase 2 (TXNRD2) could be beneficial. The TXNRD2 protein is critical for maintenance of intracellular red-ox status and ROS detoxification. TXNRD2 depletion generates ROS. It has been found that reduced expression of an antioxidant enzyme TXNRD2 may suppress BC progression.12,13 ROS superinduces expression of another oxidative stress-related activating transcription factor 3 (ATF3) gene. Under physiological conditions, ATF3 controls the expression of cell-cycle regulators and tumor suppressors, DNA repair, and apoptosis genes. However, under pathological conditions, such as BC, ATF3 upregulation aid in tumor development, survival, metastasis, and poor prognosis. 14
While several oxidative stress-related molecular markers for BC have been proposed, no biomarkers are well established to assess BC prognosis. Functional germline DNA polymorphisms are particularly attractive biomarkers since they may influence their coded protein production; they are present at the time of diagnosis and are not influenced by the state of the disease. Recently, several genome-wide association studies (GWASs) have investigated single nucleotide polymorphisms (SNPs; including those related to oxidative stress) that are associated with BC survival,15,16 no oxidative-stress-related variants were shown to be prognostic; however, these studies included large heterogeneously treated cohorts with all-stage BC patients. Knowing the important role that the abovementioned oxidative-stress-related genes play in the course of BC, we aimed to assess the contribution of functional polymorphism in the NFE2L2, HMOX1, P21, TXNRD2, and ATF3 genes to the clinicopathologic features and survival in a highly important early-stage BC group seeking to identify patients early who are likely to have a more aggressive disease.
Methods
Study subjects
The patients were investigated at the Hospital of Lithuanian University of Health Sciences Kaunas Clinics, Kaunas, Lithuania. Study was approved by Kaunas Regional Ethics Committee for Biomedical research; Protocol number BE-2-10. Informed consent for participation in the study and use of their tissue was obtained from all participants. A total of 202 clinically and histologically confirmed I–II stage BC patients were included in this cohort study. The exclusion criteria were other malignancies, significant comorbidities, and/or incomplete medical documentation. After informed consent, 5 mL venous blood was collected from each subject. Cancer treatment was provided as per institutional guidelines, in compliance with international guidelines at the time of diagnosis. The characteristics of clinical and pathological features and the course of the disease were obtained for all study subjects. The patients were prospectively followed until April 30, 2019 (censoring date).
SNP selection and genotyping
Nine known SNPs within the HMOX1, NFE2L2, TXNRD2, P21, and ATF3 genes were selected based on the following parameters: (a) functional single nucleotide polymorphisms, predicting alterations in the protein level; (b) SNP relevant to outcomes in other settings; (c) SNP with a minor allele frequency (MAF) greater than 10% in the European population. MAF of SNPs in the European population was identified using the 1000 Genomes Project database, 17 which contains the whole-genome sequencing data of a diverse set of individuals from multiple populations. We used the final phase of the project (phase 3) data, where common human genetic variation information of European ancestry individuals is provided (accessible online at http://www.ensembl.org/Homo_sapiens/Info/Index). For this study we selected nine SNPs: the HMOX1 gene rs2071746; the NFE2L2 gene rs10183914, rs35652124 and rs6721961; the TXNRD2 rs1139793; the P21 rs1801270; the ATF3 rs3125289, rs11119982 and rs1877474.
Peripheral blood samples from the study population were collected during 2009–2017. DNA was extracted from peripheral blood leukocytes, using a commercially available DNA extraction kit (i.e. Thermo Fisher Scientific Baltics, Lithuania). The DNA was stored at −20ºC prior to usage. Alleles were genotyped using TaqMan genotyping assays, in an Applied Biosystems 7900HT Real-Time Polymerase Chain Reaction System. Each sample genotyping was repeated twice for accuracy. Results were analyzed using the TaqMan Genotyper Software. Allelic discrimination was done by using the SDS 2.3 software provided by Applied Biosystems.
Statistical analysis
The statistical analysis was done to evaluate the association of screened polymorphisms with BC pathomorphological characteristics and survival. The allele frequency distributions of the investigated SNPs were compared with the European population data from the 1000 Genomes project phase 3 database 17 using the Chi-square (χ2) test. Hardy Weinberg Equilibrium (HWE) was tested by comparing the observed to expected genotype frequencies using the χ2 and Fisher’s exact tests. Univariate association of clinical, pathomorphological characteristics, and SNPs was performed using logistic regression, χ2 and Fisher's exact tests. The odds ratios (OR) with 95% confidence interval (CI) were estimated. Survival plots were performed using Kaplan–Meier methods. Univariate differences between survival rates were tested for significance using the log-rank test, while multivariate analysis for survival was tested using Cox proportional hazards models. Multivariate analysis was adjusted for patient age at diagnosis (⩽50 years/>50 years), tumor size (⩽2 cm/>2–5 cm), lymph node status (positive/negative), estrogen receptor (ER) status (positive/negative), progesterone receptor (PR) status (positive/negative), human epidermal growth factor receptor 2 (HER2) status (positive/negative) and tumor differentiation grade (G1 + G2/G3). The hazard ratios (HR) with 95% CI were estimated. A value of P<0.05 was considered statistically significant. All the statistical values were calculated using SPSS (Version 20, SPSS Inc., Chicago, IL). In the survival analysis, the date of histological BC verification was time zero. The endpoints of interest were disease-free survival (DFS), metastasis-free survival (MFS), and overall survival (OS).
Results
Study sample
Overall, 202 early-stage BC patients meeting eligibility criteria were available for SNPs evaluation. Patients' demographic and clinicopathological characteristics are presented in Table 1. Of 202 patients, 100% underwent surgery, 77% chemotherapy, 71% hormone therapy, 19% anti-HER2 therapy, and 97% radiation therapy as a primary radical treatment for early-stage BC. In the analyzed population, the observed genotype frequencies of eight polymorphisms were in HWE (P>0.05). Only in the NFE2L2 rs35652124 polymorphism, we observed a deviation from HWE, which could be attributed to selection bias (P<0.01). Comparing the determined genotype and allele frequencies in our cohort with 1000 Genomes project European population data, we observed that our cohort had slightly—but statistically significant—lower MAF of NFE2L2 rs10183914, rs35652124, and ATF3 rs1801270, rs1877474 SNPs. The basic information of candidate SNPs as well as genotype and allele frequency data are presented in Table 2.
Patient demographics and clinicopathological characteristics.

ER: estrogen receptor; HER2: human epidermal growth factor receptor 2; PR: progesterone receptor.
Basic information of candidate SNPs, genotype, and allele frequencies of the analyzed polymorphisms in the study population and the European population data from the 1000 Genomes Project Phase 3 database 17 .

Significant differences.
Analysis performed by 2 × 2 table for each SNP using major homozygotes versus others in cases and controls.
Analysis performed by a 2 × 2 table for the number of each allele in cases and controls.
1000 Genomes: 1000 genomes project phase 3 European population data. 17
MAF: minor allele frequency; N: number of valid subjects who were successfully genotyped for each of SNP; SNPs: single nucleotide polymorphisms.
Association of SNPs with clinical and pathological features
The CA+AA genotypes of P21 rs1801270 were significantly less frequent in patients with lymph node metastasis and larger tumor size (OR 0.426; 95% CI 0.149, 0.865; P=0.041 and OR 0.426; 95% CI 0.189, 0.958; P=0.022, respectively). In genotypic analysis model, patients with TT genotype in ATF3 rs3125289 had significantly lower risk of ER, PR negative, HER2 positive status (OR 0.226; 95% CI 0.063, 0.817; P=0.023; OR 0.336; 95% CI 0.115, 0.980; P=0.046; and OR 0.116; 95% CI 0.015, 0.911; P=0.040, respectively). ATF3 rs11119982 CC and TXNR2 rs1139793 GG genotypes were associated with higher rates of distant metastases, further investigated in MFS analysis.
No other significant associations were observed. Data are shown in Supplementary File 1 (Supplementary Tables 1–3).
Association of SNPs with survival
To assess a prognostic role of selected SNPs, we further tested their association with relevant disease outcomes, such as DFS, MFS, and OS (data presented in Table 3). Mean follow-up time for survival analysis was 67 months (28–202 months). The DFS event rate was 16% (33/202), of which 28 patients (13.9%) developed distant metastasis, and 22 patients (10.9%) died during the follow-up period. All deaths were cancer related.
Univariate and multivariate survival analysis data.

Only the combined results of the heterozygous and homozygous variant alleles are shown due to the small number of homozygous variant alleles. **Not included into multivariate model due to correlation between variables.
Multivariate – variables significant in univariate analysis were included into multivariate analysis and adjusted for age at diagnosis (⩽ 50 years/> 50 years), tumor size (⩽ 2 cm/>2–5 cm), lymph node status (positive/negative), ER status (positive/negative), PR status (positive/negative), HER2 status (positive/negative) and tumor differentiation grade (G1 + G2/G3).
CI: confidence interval; DFS: disease-free survival; ER: estrogen receptor; HER2: human epidermal growth factor receptor 2; HR: hazard ratio; MFS: metastasis, free survival; OS: overall survival; PR: progesterone receptor; ref.: reference.
In a univariate and multivariate Cox analysis, the risk of a DFS event was higher in the group of patients carrying ancestral TXNRD2 rs1139793 GG genotype, compared to A allele carriers (GA+AA) (HR 2.248; 95% CI 1.105, 4.572; P=0.025; and HR 2.211; 95% CI 1.084, 4.509; P=0.029, respectively). The same GG genotype was also associated with a higher risk of metastasis (univariate HR 2.257; 95% CI 1.041, 4.892; P=0.039; multivariate HR 2.214; 95% CI 1.017, 4.817; P=0.045). Kaplan–Meier curves for TXNRD2 rs1139793 associations with DFS and MFS in the allelic model are presented in Figure 1.
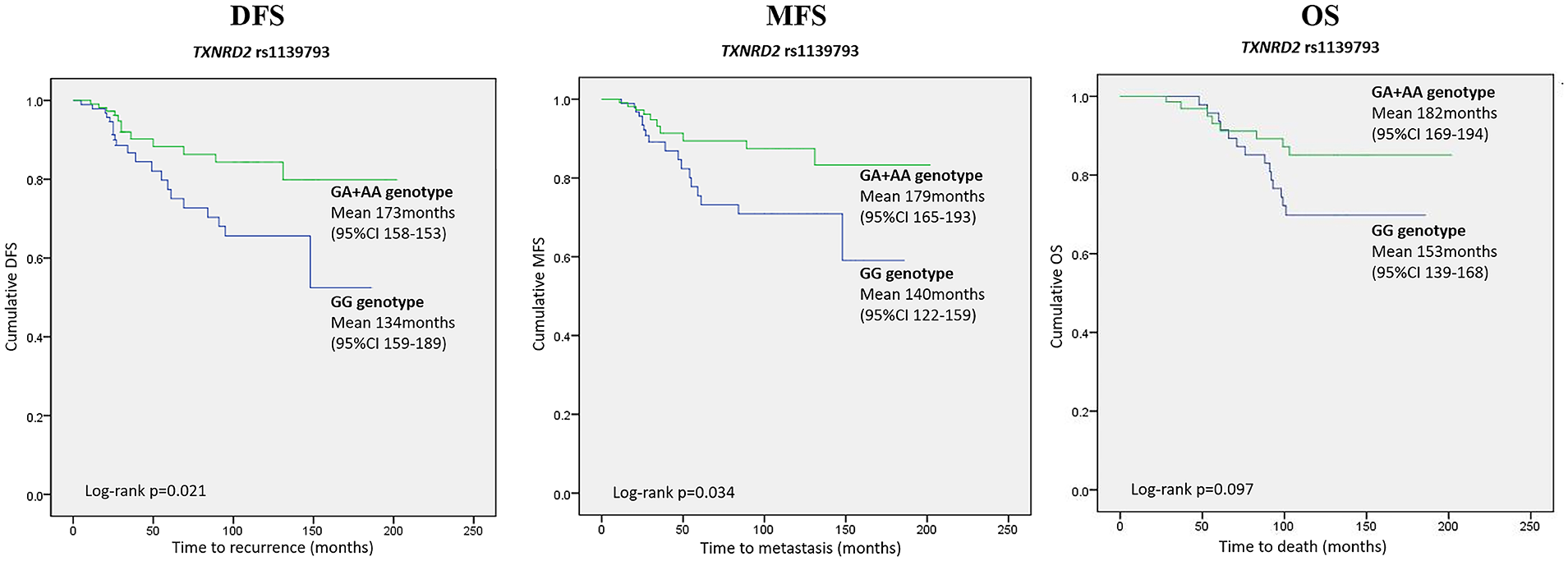
Kaplan–Meier survival curves for TXNRD2 rs1139793 polymorphism in allelic model (GA+AA vs. GG genotype) demonstrating DFS, MFS, and OS differences. The X-axis displays the number of months from the breast cancer confirming biopsy date to the event of interest (any recurrence for DFS, distant recurrence for MFS, and death for OS). The Y-axis displays the Kaplan–Meier survival probability. Means and 95% confidence intervals as well as Log rank P values are provided.
By analyzing the SNPs of the ATF3 gene, we found that the rs11119982 CC genotype (in the genotypic model: TT vs. TC vs. CC) was a negative prognostic factor for DFS (univariate HR 6.264; 95% CI 1.810, 21.680; P=0.004; multivariate HR 5.878; 95% CI 1.660, 20.806; P=0.006), MFS (univariate HR 5.036; 95% CI 1.434, 17.687; P=0.012; multivariate HR 4.759; 95% CI 1.310, 17.290; P=0.018) and OS (univariate 3.451; 95% CI 1.120, 12.748; P=0.042; multivariate HR 3.280; 95% CI 1.103, 12.809; P=0.048). Kaplan–Meier curves for ATF3 rs11119982 associations with DFS, MFS, and OS in genotype model are presented in Figure 2. No other associations between analyzed SPNs and survival endpoints were observed.
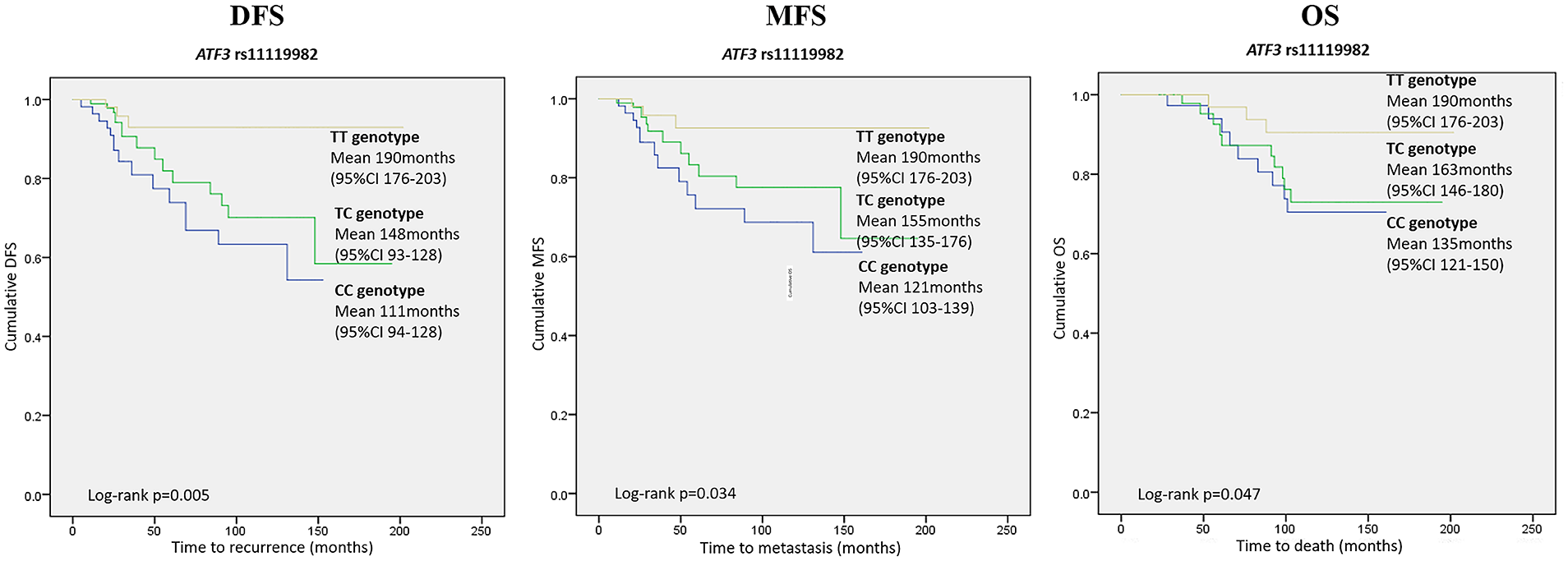
Kaplan–Meier survival curves for ATF3 rs11119982 polymorphism in genotype model (TT vs. TC vs. CC genotype) demonstrating DFS, MFS, and OS differences. The X-axis displays the number of months from the breast cancer confirming biopsy date to the event of interest (any recurrence for DFS, distant recurrence for MFS, and death for OS). The Y-axis displays the Kaplan–Meier survival probability. Means and 95% confidence intervals as well as Log rank P values are provided.
Discussion
In this cohort study of early-stage BC patients, we found that the functional P21 rs1801270 polymorphism is associated with tumor size and lymph node status, while ATF3 rs3125289 SNP was significantly linked with hormone receptor and HER2 status. In the survival analysis, the common homozygous TXNRD2 rs1139793 GG genotype was associated with shorter DFS and MFS, but not OS compared to patients carrying at least one rare A allele. Furthermore, the common homozygous ATF3 rs11119982 CC genotype in the genotypic model was a negative prognostic factor for DFS, MFS, and OS.
The P21 rs1801270 polymorphism, which occurs in codon 31, results in an amino acid substitution of arginine for serine. This polymorphism is located in a highly conserved region of P21 and affects molecular function of the P21 gene. The rare AA genotype of rs1801270 is associated with decreased P21 mRNA expression. 18 Lower P21 expression is known to be associated with smaller tumor size, lower tumor differentiation grade, less lymph node metastasis, longer disease-free survival and better anticancer drug response in BC.11,19 Several studies analyzed the role of P21 rs1801270 polymorphism on BC risk. A meta-analysis of such studies showed that the P21 rs1801270 CC genotype increases the risk of BC by 1.496 times for Caucasians. 20 In concordance with all these data, we found that patients carrying the P21 rs1801270 A allele, which is associated with lower P21 expression, had less aggressive tumor phenotype, a smaller primary tumor, and less lymph node involvement. We found no other studies evaluating the associations between P21 rs1801270 and clinicopathological BC parameters. Despite the association with more aggressive clinical features, P21 rs1801270 had no impact on BC prognosis in our study. These results are consistent with several GWAS studies of BC survival.15,16
ATF3 gene promotes survival, motility, and invasiveness of BC cells as well as radioresistance. It is shown, that the ATF3 expression is markedly increased in BC tissues compared to adjacent normal breast tissues, and ATF3 expression correlates with tumor node metastasis stage, invasion, lymph node metastasis, and the number of metastatic lymph nodes.21,22 Major ancestral alleles of the three investigated ATF3 SNPs alter the DNA landscape for potential transcriptional factors to attach, resulting in changes in transcriptional factor binding sites. 23 In our study, patients with a major CC genotype in ATF3 rs11119982 SNP had worse DFS, MFS, and OS. To the best of our knowledge, we are the first to investigate and report this association in the candidate gene approach. In the GWAS approach,15,16 this polymorphism was not associated with OS in BC; however, the cohorts of GWAS studies included patients with all cancer stages and evaluated only OS, while in our study only early-stage BC patients were involved. Another investigation of the ATF3 rs3125289 polymorphism was linked with hormone receptor and HER2 status. There is no known explanatory mechanism for this association; therefore, further investigations are needed.
The role of TXNRD2 in cancer progression is less investigated. In one study it was demonstrated that reduced expression of an antioxidant enzyme TXNRD2 may suppress BC progression. 13 However, in a hepatocellular carcinoma study, a lower level of TXNRD2 was associated with higher tumor stage. 24 In our study, we found that TXNRD2 rs1139793 common homozygote GG genotype, which is associated with low TXNRD2 protein level, 25 was a negative prognostic factor for DFS and MFS, while the rare A allele had a protective effect. Further studies are required to clarify the potential underlying biological mechanisms of this association. We are not aware of previous germline studies reporting on TXNRD2 rs1139793 polymorphisms and BC prognosis.
The results of this study are strengthened by several facts. First, we have evaluated a well-defined, early-stage BC cohort in the current study with patients from European ancestry, making this the largest study of its kind. Second, all patients were followed prospectively after treatment of localized BC. Third, survival analysis results remained significant after adjusting for clinical and pathological covariates. Of note, there is an observed deviation from HWE for rs35652124 in our study due to the observed higher minor homozygote rate than expected. The reasons that may contribute to this deviation include selection bias, since we only included patients with BC but without a control group. We acknowledge that the lack of the HWE is a limitation of this study, and the association results of rs35652124 from our study should be regarded with caution.
Other potential limitations of our study include a limited sample size, the risk of other confounders, possible gene, gene and gene, environment interactions, and non-random sampling. However, this study is the first to analyze the impact of selected polymorphisms on early-stage BC survival, and it supports the relevance of functional TXNRD2 rs1139793 and ATF3 rs11119982 germline polymorphisms to the BC prognosis.
In conclusion, our findings suggest that the P21 rs1801270 is associated with lymph node metastasis and larger tumor size, and the ATF3 rs3125289 is associated with ER, PR, and HER2 status. TXNRD2 rs1139793 and ATF3 rs11119982 may have an impact on early-stage BC survival. Further investigations, preferably on larger cohorts with different ethnic origins, are needed to confirm the results of the current study.
Supplemental Material
sj-pdf-1-jbm-10.1177_17246008211011177 – Supplemental material for The role of functional polymorphisms in oxidative stress-related genes on early-stage breast cancer survival
Supplemental material, sj-pdf-1-jbm-10.1177_17246008211011177 for The role of functional polymorphisms in oxidative stress-related genes on early-stage breast cancer survival by Erika Korobeinikova, Rasa Ugenskiene, Ruta Insodaite, Viktoras Rudzianskas, Jurgita Gudaitiene and Elona Juozaityte in The International Journal of Biological Markers
Footnotes
Acknowledgements
Not applicable.
List of abbreviations
ATF3 – Activating Transcription Factor 3 (gene)
ATF3 – Activating Transcription Factor 3 (protein)
BC – breast cancer
CI – confidence interval
DFS – disease-free survival
ER – estrogen receptor
GWAS – genome-wide association studies
HER2 – human epidermal growth factor receptor 2
HMOX1 – Heme Oxygenase 1 (gene)
HO-1 – Heme oxygenase-1 (protein)
HR – hazard ratio
HWE – Hardy Weinberg Equilibrium
MAF – minor allele frequency
MFS – metastasis-free survival
NFE2L2 – Factor, Erythroid 2 Like 2 (gene)
NFE2L2 – Factor, Erythroid 2 Like 2 (protein)
OR – odds ratio
OS – overall survival
P21 – Cyclin dependent kinase inhibitor 1A (gene)
P21 – Cyclin dependent kinase inhibitor 1A (protein)
PR – progesterone receptor
ROS – reactive oxygen species
SNP – single nucleotide polymorphism
TXNRD2 – Thioredoxin Reductase 2 (gene)
TXNRD2 – Thioredoxin Reductase 2 (protein)
χ2 – Chi-square
Authors’ contributions
EK, RU, and EJ designed the research study; EK, RU, RI, GJ, VR, and EJ performed the research; EK and RU analyzed the data; RU and RI contributed essential reagents or tools; EK wrote the manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The present study was partially supported by the Lithuanian University of Health Sciences science foundation.
Availability of data and materials
The datasets used and/or analyzed during the present study are available from the corresponding author on reasonable request.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
