Abstract
Objective
We aimed to determine whether single nucleotide polymorphisms in the
Methods
Nine single nucleotide polymorphisms of
Results
rs4143815C>G was associated with better pathological complete response in the addictive and over-dominant models and with poorer disease-free survival in the recessive model. Patients with different genotypes had different adverse events. Overexpression of miR34c resulted in the downregulation of PD-L1 mRNA expression.
Conclusion
The
Introduction
Breast cancer (BC) is one of the most common malignancies in women worldwide. Neoadjuvant therapy (NAT) has become a regular treatment procedure for BC in recent decades. Pathological complete response (pCR) refers to the status when no tumor cell or only ductal carcinoma in situ remains in the surgical specimens after preoperative treatment. A number of large-scale clinical trials have confirmed that patients who reach pCR in NAT have a much better prognosis than patients without pCR.1–3 Therefore, it is crucial to predict the patient’s response to NAT early to adjust the treatment strategy in time. In addition, the adverse events of NAT either related to chemotherapy or targeted therapy could often lead to the discontinuation of the treatment. This might weaken patient compliance or cause disease progression. Thus, it is just as important to minimize adverse events as to achieve pCR during NAT.
Programmed death-ligand 1 (PD-L1) is a membrane-spanning glycoprotein. The protein is mainly expressed on the surface of T cells, natural killer cells, dendritic cells, and the tumor cell membrane, functioning as an immunosuppressive molecule. When combined with its ligand programmed death 1 (PD-1), it triggers T-cell apoptosis and induces the differentiation from cytotoxic T cells to inhibitory T cells.4,5 The subsequent change in PD-L1 expression is often related to tumor immune escape. In clinical studies, PD-L1 expression is associated with poorer prognosis in a variety of solid tumors, such as melanoma, renal cancer, and lung cancer.6–9 In fact, antibodies against PD-1/PD-L1 have now been approved for the clinical management of melanoma and renal cancer. Laboratory experiments suggest that PD-L1 is regulated and influenced by many cellular pathways, such as the MAPK and PI3K/AKT pathways.10–12 Epigenetically, PD-L1 is also regulated by many microRNAs (miRNAs) and is involved in tumor progression, such as epithelial-mesenchymal transition.13–15 In addition, there are reports that in gastric cancer and lung cancer, single nucleotide polymorphism (SNP) of the
SNP refers to the polymorphism of a specific base in the genome. Different from gene mutations, a mutation base needs to reach a certain frequency (more than 1/1000) in the population to become a SNP site. SNP changes are often associated with disease development, drug metabolism, and resistance.19,20 According to the distribution sites on the genome, SNPs can be catalogued into coding SNPs and noncoding SNPs. Although it is not involved in protein synthesis, SNPs in noncoding regions can still affect cell function by affecting a variety of biological processes, such as gene cleavage, transcription factor binding, and mRNA degradation. Noncoding SNP mutations are often associated with tumor development.21–25 Currently, studies of SNPs have mainly focused on coding SNPs. The role of noncoding SNPs in disease development remains obscure.
Preliminary neoadjuvant clinical trials (SHPD001 and SHPD002) were conducted in our department with weekly paclitaxel and cisplatin treatment, demonstrating a high pCR rate.
26
We previously examined the expression of PD-L1 protein in patients from a clinical trial and found that PD-L1 protein expression is a predictive biomarker of pCR and a prognostic factor of disease-free survival (DFS) in BC patients and hormone receptor positive (HorR-positive) subgroups.
27
However, the underlying mechanism is yet to be clarified. In this study, we aimed to explore the potential PD-L1-regulatory SNPs through bioinformatics analysis and experimental verification. We hypothesize that the SNPs located at the 3′untranslated region (3′UTR) of
Materials and methods
Patient cohort
BC patients from two weekly paclitaxel- and cisplatin-based neoadjuvant clinical trials were included. The two trials were separately registered in ClinicalTrials.gov as SHPD001 (NCT02199418) and SHPD002 (NCT02221999). Detailed trial procedures and dosing schedule, detailed entry criteria, exclusion criteria, and the primary study endpoints and definitions for the trial were described in our previous publication.26,27
Blood samples were taken for the subsequent detection of SNPs before any treatment. All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. Both clinical trials were approved by the Ethics Committee of Renji Hospital, School of Medicine, Shanghai Jiaotong University. All patients provided informed consents.
SNP selection and genotyping
Potentially functional SNPs were selected according to the following criteria using a public database (NCBI/TargentScan)
28
: (a) SNPs with a minor allele frequency (MAF) >0.05; and (b) SNPs located in the 3′UTR of
DNA from the patient samples was extracted from peripheral blood using the TIANamp Genomic DNA Kit (Tiangen Biotech Co., Ltd., Beijing, China) according to the manufacturer’s protocols. Primers for quantitative polymerase chain reaction (qPCR) and single-base extension were designed by Assay Design Suite V2.0 Online (Agena Bioscience, San Diego, USA). qPCR was performed as previously described. 29 SNP genotyping was carried out on the MassARRAY system (Agena Bioscience, San Diego, CA, USA) using the matrix-assisted laser desorption ionization-time of flight (MALDI-TOF) mass spectrometry method. Genotype calling was performed in real time with MassARRAY RT software Version 3.0.0.4 and analyzed using MassARRAY Typer software version 3.4 (Sequenom). DNA sample extraction, qPCR and SNP genotyping were performed by Shanghai Benegene Biotechnology Co., Ltd. (Shanghai, People’s Republic of China).
Bioinformatics analysis
The gene locus of rs4143815 was found through the NCBI database (https://www.ncbi.nlm.nih.gov/snp), and the possible target miRNAs were found through the TargetScan database (http://www.targetscan.org/vert_71/).
The gene expression information of 500 patients with BC was downloaded from the TCGA database (https://cancergenome.nih.gov/) to analyze the relationship between the expression of
Statistical analysis
Hardy-Weinberg equilibrium (HWE) was determined using the chi-square test. Linkage disequilibrium among individual SNPs was measured by HaploView software (http://broad.mit.edu/mpg/haploview). Four models were used in SNP genotype analysis. Assuming “A” was the minor allele of one SNP locus, the addictive model compares TT vs TA vs AA; the dominant model compares TT vs TA+AA; the recessive model compares TT+TA vs AA; and the over-dominant model compares TT+AA vs TA. Haplotypes and their frequencies were estimated by HaploView for univariate analysis and Phase v2.1.1 30 for multivariate analysis adjusted for patient's age, tumor size, estrogen receptor (ER), progesterone receptor (PR), human epidermal growth factor receptor 2 (HER2), and Ki67. Considering linkage disequilibrium, SNP loci within a distance of 500 kb and both D and r2 values >0.8 were initially designed for further haplotype analysis. However, no two loci met the criteria of r2 >0.8; thus, r2 was adjusted to >0.49. With these criteria, rs822336 and rs822337, rs822336 and rs2282055, rs822337 and rs2282055, rs4143815 and rs4742098 were tested for any potential association between different haplotypes and the pCR rate.
Univariate and multivariate logistic regression tests were used to analyze the associations between different SNP genotypes, haplotypes and pCR outcomes. DFS was defined as the time from breast surgery to the first disease relapse, including one of the following events: distant disease metastasis, recurrence of ipsilateral locoregional invasive disease, or contralateral BC. Survival curves were derived from the Kaplan–Meier method, and the log-rank test was used to compare survival differences. A Cox proportional hazards model was used to calculate the adjusted hazard ratio (HR) with a 95% confidential interval (CI). Patient's age, tumor size, ER, PR, HER2, and Ki67 were adjusted.
After the qPCR data were normalized, a paired t-test was used to compare the mRNA and miRNA expression levels among different groups (cells transfected with miR34c mimics vs corresponding negative controls). Spearman’s test was used to observe the correlation between
Cell culture and construction
Human BC cell lines MCF-7, SK-BR-3, MDA-MB-468, MDA-MB-231, MDA-MB-453, Bcap-37, and T47D and the nontumorigenic epithelial cell line MCF-10A were obtained from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China) and were authenticated by routine detection of morphology, cell vitality, DNA fingerprinting, isoenzymology, and mycoplasma. Cell lines were maintained in McCoy’s 5A and Leibovitz’s L-15 medium (BasalMedia, Shanghai, China). All media were supplemented with 10% fetal bovine serum (Gibco, Waltham, MA, USA) and 1% penicillin-streptomycin (BasalMedia, Shanghai, China).
The miRNA mimics and a negative control (Biotend Company, Shanghai, China) were transfected into tumor cells by Lipofectamine 2000 (Invitrogen Life Sciences, Carlsbad, CA, USA), and gene changes were detected 24–72 hours after transfection.
Chemical reagents
MiRNA mimics and negative control were purchased from Biotend (Shanghai, China).
RNA extraction and qPCR
Total RNA from cultured cells and tissue samples was isolated using TRIzol reagent (Invitrogen Life Sciences, Carlsbad, CA, USA). The miRcute miRNA First-strand cDNA Synthesis Kit and the miRcute miRNA qPCR Kit (Tiangen Biotech Company, Beijing, China) were used to detect miRNA expression levels. The primers used for U6 and miR34c were also obtained from Tiangen Biotech. The mRNA expression level was detected by qPCR using the Takara Reverse Transcriptase Kit and SYBR Green PCR Kit (Takara, Dalian, China). Each sample was examined in triplicate. The primers used for
Predictive Value of SNPs for pCR outcome.
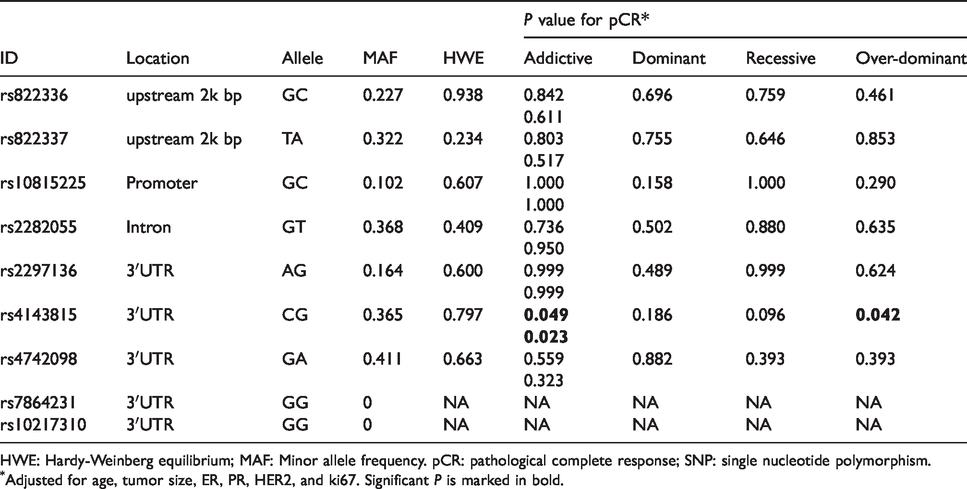
HWE: Hardy-Weinberg equilibrium; MAF: Minor allele frequency. pCR: pathological complete response; SNP: single nucleotide polymorphism.
Adjusted for age, tumor size, ER, PR, HER2, and ki67. Significant
Results
Basic clinical and pathological features of patients
A total of 145 patients provided whole blood specimens for SNP testing (Supplementary Table 2). Of the 145 patients, 62/145 (43%) had a tumor greater than 5 cm, 68/145 (47%) had HER2 gene overexpression, and 109/145 (75%) had a Ki67 expression level of more than 30%, which was consistent with the locally advanced nature of the included patients.
Predictive and prognostic value of
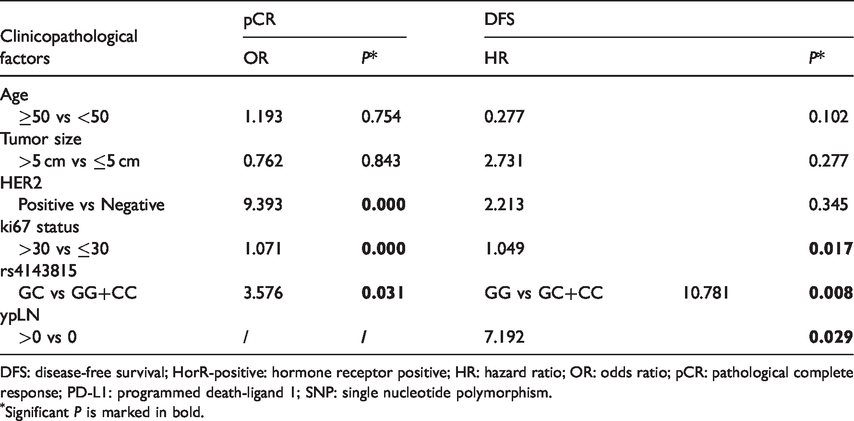
DFS: disease-free survival; HorR-positive: hormone receptor positive; HR: hazard ratio; OR: odds ratio; pCR: pathological complete response; PD-L1: programmed death-ligand 1; SNP: single nucleotide polymorphism.
Significant
Predictive value of the PD-L1 SNP genotype
We examined nine SNP loci of the
rs822336 and rs822337, rs822336 and rs2282055, rs822337 and rs2282055, rs4143815, and rs4742098 were loci with potential linkage disequilibrium (Figure 1(a) and (b)); however, no significant correlation was found between these haplotypes and pCR in either univariate or multivariate analysis (Supplementary Tables 3 and 4).
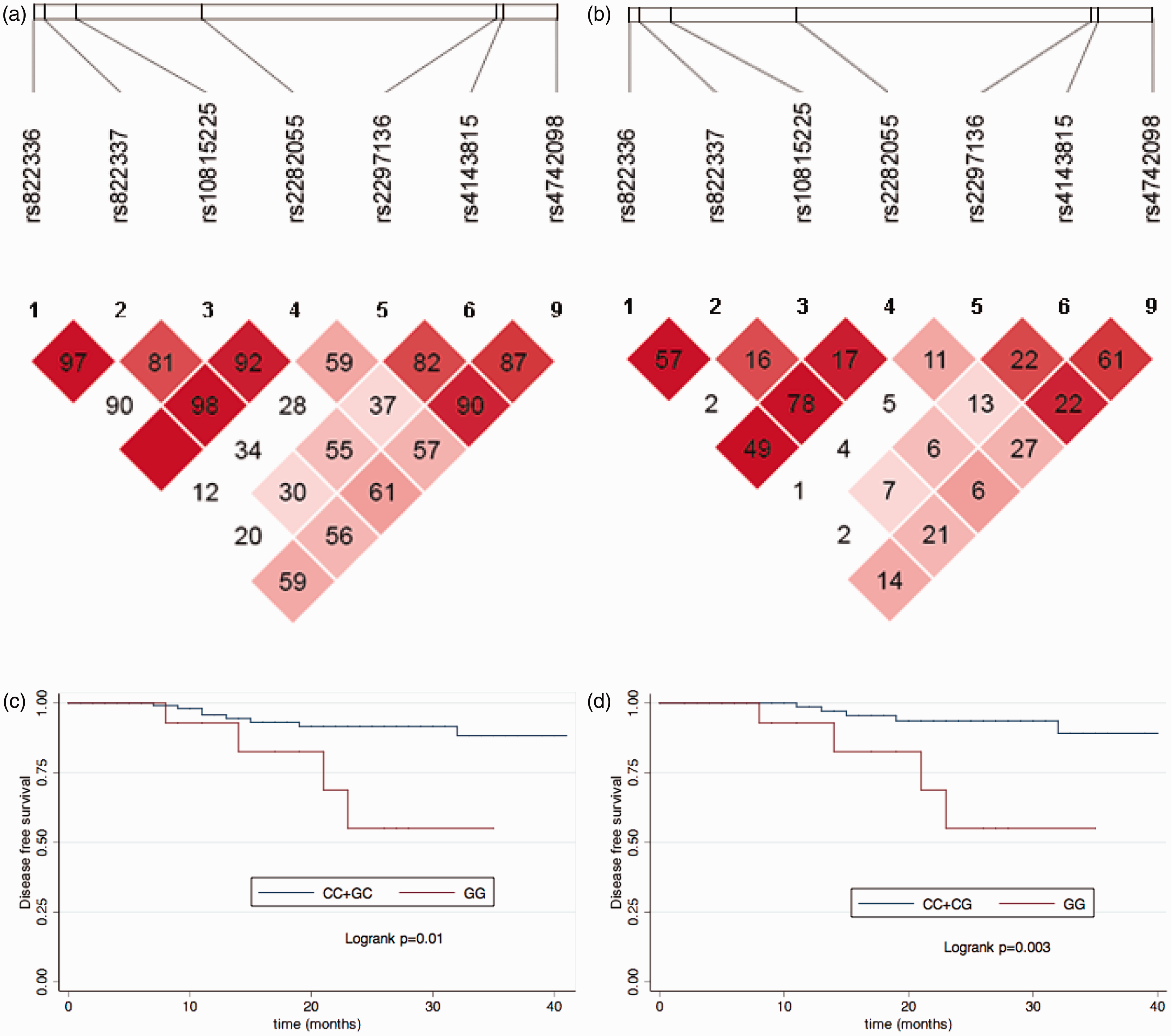
D and r2 value of paired SNP loci and the KM plot for different PD-L1 SNP genotypes in breast cancer patients. (a) D value of paired SNP loci. (b) r2 value of paired SNP loci. (c) KM plot for all patients with different rs4143815 genotype; blue: CC+GC, red: GG, Log-rank
Prognostic value of the PD-L1 SNP genotype
With a median follow-up time of 18 months, patients with a rs4143815C>G alteration exhibited a worse DFS. Kaplan–Meier plots showed that patients with the rs4143815 GG genotype had worse survival than those with the GC+CC genotype (log-rank test
Multivariable analyses of associations between clinicopathological factors and disease-free survival.
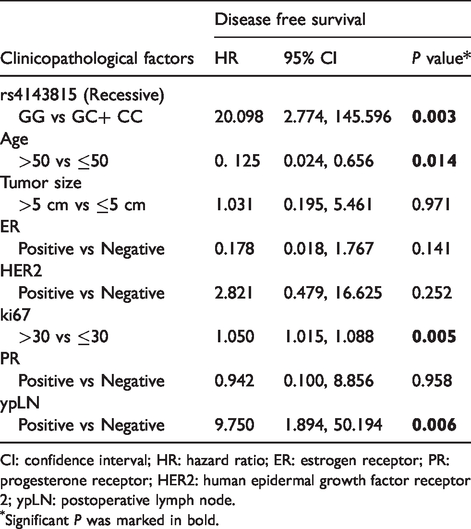
CI: confidence interval; HR: hazard ratio; ER: estrogen receptor; PR: progesterone receptor; HER2: human epidermal growth factor receptor 2; ypLN: postoperative lymph node.
Significant
In HorR-positive BC, Kaplan–Meier plots showed that the GG genotype had worse survival than the GC + CC genotype (log-rank
Adverse events
Analysis of adverse effects in patients with different rs4143815 genotypes revealed that neutropenia was more likely to occur in CC+CG than in GG patients (rho=−0.183,
Adverse events according to different genotype of rs4143815.
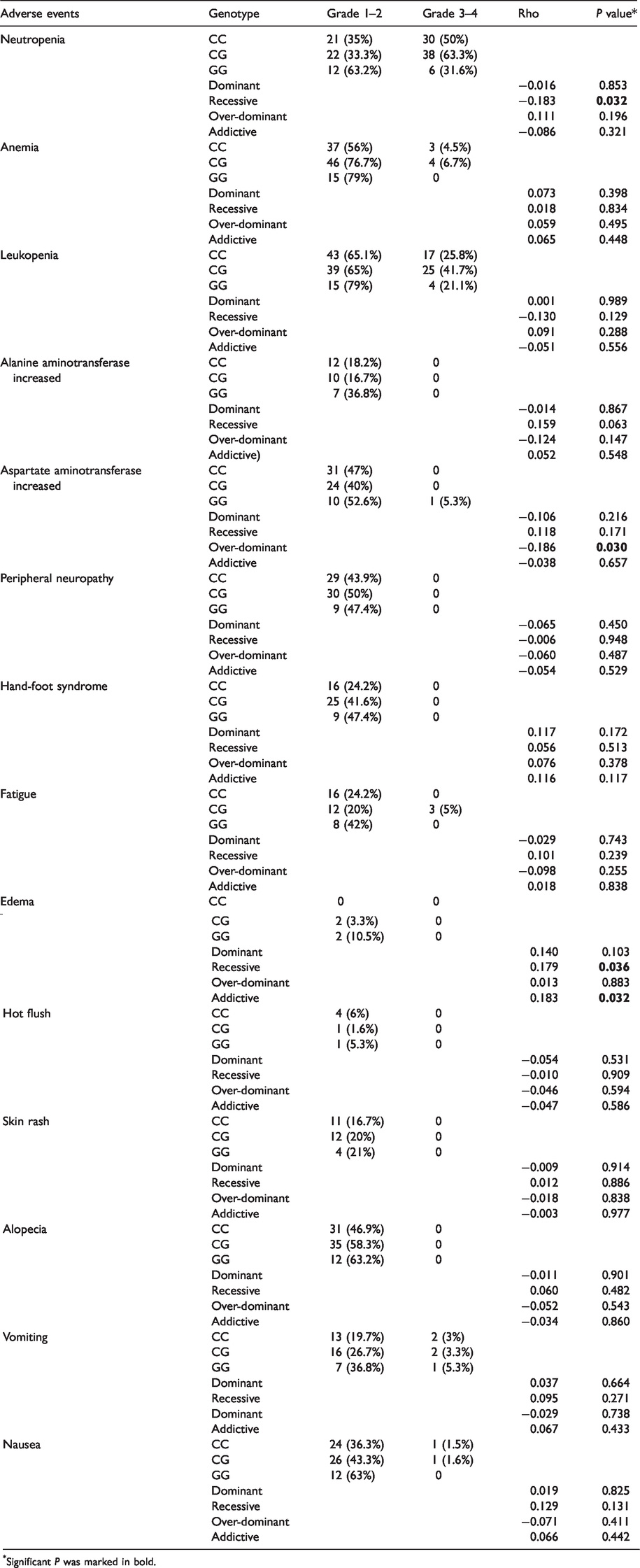
Significant
Bioinformatics prediction
Since rs4143815 is the only SNP site that has been demonstrated to be associated with pCR and survival in patients with NAT in our research, we further explored its possible molecular biological mechanisms.
Through the USCS database, we found the location of rs4143815 in the
We did find a negative correlation between
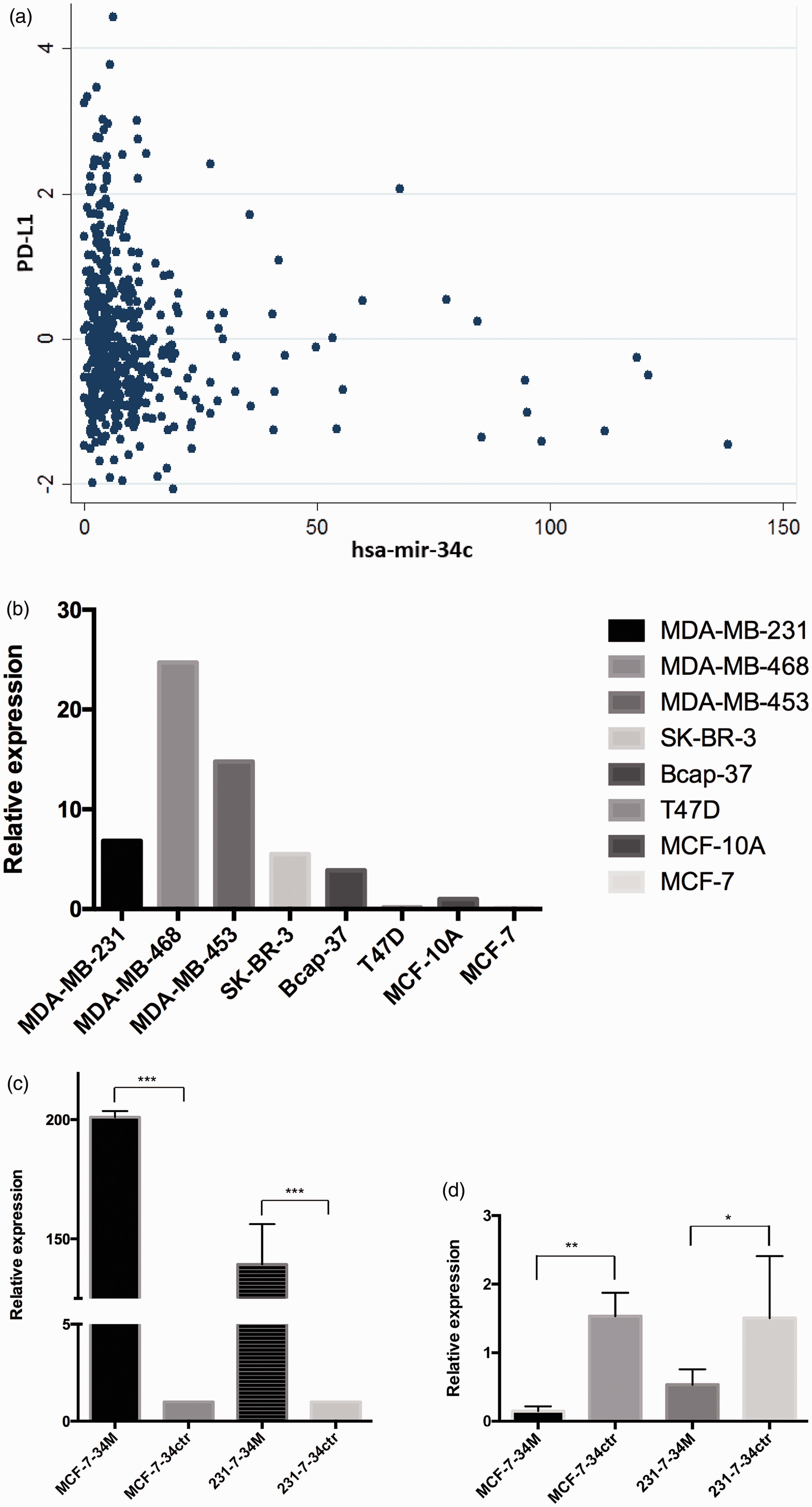
Correlation of PD-L1 mRNA expression with miR34c expression and miR34c overexpression. (a) Spearman’s rho=−0.1612,
Regulatory miRNA of PD-L1
We examined the level of
miR34c mimics, and corresponding negative controls were transfected into, MDA-MB-231 and MCF-7 cell lines to overexpress miR34c within 2–3 days. Compared with the control group, the expression of miR34c in the overexpression group increased significantly (Figure 2(c)). Overexpression of miR34c revealed a significant decrease in
Discussion
The
Our results suggest that patients with the rs4143815C>G were more likely to achieve pCR, whereas patients with the GG genotype were more likely to suffer from disease recurrence than those with the GC and CC genotypes. SNPs at this site have been previously reported in other tumors. Studies by Lee et al.16,17 showed that polymorphisms in rs4143815 were associated with pCR and long-term overall survival (OS) in neoadjuvant chemotherapy in patients with non-small-cell lung cancer (NSCLC). In their studies, patients carrying the GG genotype were better responders to neoadjuvant chemotherapy but had more OS events, which was consistent with our results. It was also reported that an SNP at this site was associated with the responses to the PD-1 antibody nivolumab in patients with NSCLC. 31 They discovered that the rs4143815 GG genotype of PD-L1 was significantly associated with a worse objective response rate and progression-free survival in NSCLC patients treated with nivolumab.
The rs4143815 polymorphism is also associated with neutropenia, elevated aspartate aminotransferase levels, and edema in our patient cohort. Currently, studies concerning adverse events have mainly focused on the polymorphism of metabolic enzyme-related genes such as CYP2D6 and CYP3A5.32,33 We are the first to find that the SNP of
rs4143815 is located in the noncoding 3′UTR region of the
Therefore, we continued to investigate the regulatory mechanism of
rs4143815 is located 400 bp upstream of the predicted binding site of miR34c and
There are still many imperfections in this chapter due to time limitations. The detailed regulatory mechanism of rs4143815 on
Conclusion
Our results showed that the
Supplemental Material
sj-pdf-1-jbm-10.1177_1724600820926172 - Supplemental material for Programmed death-ligand 1 single nucleotide polymorphism affects breast cancer chemosensitivity and adverse events in the neoadjuvant setting
Supplemental material, sj-pdf-1-jbm-10.1177_1724600820926172 for Programmed death-ligand 1 single nucleotide polymorphism affects breast cancer chemosensitivity and adverse events in the neoadjuvant setting by Jinglu Lu*, Ziping Wu*, Jing Peng, Shuguang Xu, Liheng Zhou, Yanping Lin, Yan Wang, Wenjin Yin and Jinsong Lu in The International Journal of Biological Markers
Footnotes
Abbreviations
3′UTR: 3′untranslated region; BC: breast cancer; CI: confidential interval; DFS: disease-free survival; ER: estrogen receptor; HorR-positive: hormone receptor-positive; HR: hazard ratio; HWE: Hardy-Weinberg equilibrium; mRNA: messenger RNA: miRNA: microRNA; NAT: neoadjuvant therapy; NSCLC: non-small-cell lung cancer; OR: odds ratio; OS: overall survival; pCR: pathological complete response; PD-1: programmed death 1; PD-L1: programmed death-ligand 1; PR: progesterone receptor; qPCR: quantitative polymerase chain reaction; SNP: single nucleotide polymorphism; MAF: minor allele frequency.
Acknowledgments
Not applicable.
Authors' contributions
Jinglu Lu, Ziping Wu contributed equally to the manuscript. Conceptualization: Ziping Wu, Wenjin Yin and Jinsong Lu; Data curation: Jinglu Lu, Ziping Wu, Shuguang Xu, Liheng Zhou, Yanping Lin and Yan Wang; Methodology: Jing Peng; Supervision: Wenjin Yin and Jinsong Lu; Writing—original draft: Ziping Wu; Writing—review and editing: Jinglu Lu and Jinsong Lu. All authors read and approved the final manuscript.
Declaration of conflicting interest
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Shanghai Collaborative Innovation Center for Translational Medicine (TM201908), the Shanghai Natural Science Foundation (19ZR1431100), the Shanghai “Rising Stars of Medical Talent” Youth Development Program for Outstanding Youth Medical Talents (2018-16), the Multidisciplinary Cross Research Foundation of Shanghai Jiao Tong University (YG2017QN49 and ZH2018QNA42), the Nurturing Fund of Renji Hospital (PYMDT-002), the Science and Technology Commission of Shanghai Municipality (15JC1402700), the Clinical Research Plan of SHDC (16CR3065B, 1201623). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Ethics approval and consent to participate
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. Both clinical trials (NCT02199418 and NCT02221999) have been approved by the Ethics Committee of Renji Hospital, School of Medicine, Shanghai Jiaotong University.
Patient consent for publication
All patients provided informed consents.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
