Abstract
Background:
Long non-coding RNA (lncRNA) PITPNA antisense RNA 1 (PITPNA-AS1) expression characteristics, function, and mechanism in papillary thyroid cancer are unclear.
Methods:
Quantitative real-time polymerase chain reaction (qRT-PCR) was applied for detecting PITPNA-AS1, UNC-5 netrin receptor B (UNC5B) mRNA, and miR-129-5p expressions in papillary thyroid cancer tissues and cell lines. EdU assay, cell counting kit-8 (CCK-8) assay, wound healing assay, and flow cytometry analysis were performed to investigate the biological functions of PITPNA-AS1 in papillary thyroid cancer. Dual-luciferase reporter assay was utilized for determining whether PITPNA-AS1 and miR-129-5p, as well as UNC5B and miR-129-5p could directly bind to each other. Western blot assay was employed for measuring UNC5B protein expression level in papillary thyroid cancer cell lines.
Results:
PITPNA-AS1 and UNC5B expressions were markedly increased in papillary thyroid cancer tissues and cell lines while miR-129-5p expression was down-regulated. Knockdown of PITPNA-AS1 could significantly inhibit papillary thyroid cancer cell growth and migration and promote cell apoptosis while UNC5B overexpression plasmids or miR-129-5p inhibitors counteracted the knockdown effect of PITPNA-AS1 on papillary thyroid cancer cells. PITPNA-AS1 targeted miR-129-5p to repress its expression and miR-129-5p targeted UNC5B to repress its expression. Silencing PITPNA-AS1 reduced the expression of UNC5B via regulating miR-129-5p expression.
Conclusions:
PITPNA-AS1 facilitated papillary thyroid cancer cell proliferation and migration, and suppressed apoptosis through miR-129-5p/UNC5B axis.
Keywords
Introduction
Thyroid cancer is recognized as the prevalent endocrine tumor in the world. Papillary thyroid cancer (PTC), the most common histological subtype, accounts for 85%–90% of all cases of thyroid cancer. 1 Over the last 40 years, PTC incidence has climbed steadily, though surgical resection and radioactive iodine therapy increase the 5-year survival rate of patients to 95%. 2 Nonetheless, the prognosis of PTC patients with distant metastasis and recurrence remains unfavorable.
Long non-coding RNAs (lncRNAs) are a type of RNA molecule with over 200 nucleotides and possess no protein-coding capability. It is revealed that lncRNAs participate in diverse biological processes and the pathogenesis of human diseases. PITPNA antisense RNA 1 (PITPNA-AS1) is a relatively newly discovered lncRNA, whose genes are located on chromosome 17p13.3. A study shows that PTTPNA-AS1 is abnormally expressed in hepatocellular carcinoma and associated with the worse prognosis of patients. 3 However, its expression pattern in PTC and its biological functions are not unclear.
Numerous studies manifest that lncRNAs may serve as molecular sponges (competitive endogenous RNAs (ceRNAs)), thus regulating microRNAs (miRNAs) expression and playing a role in cancer biology. In PTC, a large number of miRNAs are aberrantly expressed, such as miR-873-5p, 4 miR-320a, 5 miR-1178, 6 and so on. As a tumor suppressor, miR-129-5p takes part in the occurrence and development of multiple tumors, such as hepatocellular carcinoma 7 and breast cancer, 8 and, importantly, miR-129-5p plays a tumor-suppressive role in PTC. 9 However, the specific mechanism of its dysregulation in PTC still needs further investigation.
UNC-5 netrin receptor B (UNC5B), a netrin-1 receptor, is involved in axon migration and angiopoiesis. 10 Besides, UNC5B partakes in PTC progression as a cancer-promoting gene. 9 In the present study, it was unmasked that PITPNA-AS1 expression was up-regulated in PTC tissues, which was associated with unfavorable pathological parameters; its silencing repressed the malignant phenotypes of PTC cells and it functioned as a ceRNA to regulate miR-129-5p/UNC5B axis. These findings presented a potential molecular mechanism of PTC occurrence and development.
Methods and materials
Tissue samples
From March 2015 to March 2019, 66 PTC patients’ tumor tissues and adjacent tissues were collected (obtained from biopsy samples or during surgery) in the First Affiliated Hospital of Dalian Medical University. Before the samples were obtained, the subjects had not undergone any local treatment or systematic treatment (e.g. chemotherapy, radiotherapy and radioactive iodine therapy). After all tissues were confirmed by the pathologists, they were frozen at −80°C for follow-up experiments. All experimental schemes were approved by the Ethics Committee of the First Affiliated Hospital of Dalian Medical University.
Quantitative real-time polymerase chain reaction (qRT-PCR) assay and total RNA extraction
TRIzol reagent (Invitrogen, Carlsbad, CA, USA) was employed for extracting total RNA from cells and tissues. The Reverse Transcription Kit (Invitrogen) was utilized to reversely transcribe RNA samples to cDNA. qRT-PCR assay was conducted with SYBR Premix Ex Taq II (Invitrogen) on ABI7500 real-time PCR system (Applied Bio-systems, Carlsbad, CA, USA). Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) acted as the internal reference for PITPNA-AS1 and UNC5B, and U6 as that for miR-129-5p. Lastly, the 2−ΔΔCT method was utilized to calculate the relative gene expression level. The primer sequences were: PITPNA-AS1 (sense): 5′-GCAGGGTGGATAAAGAGGA-3′; PITPNA-AS1 (antisense): 5′-CCTACTGACAGGATGTCCT-3′; miR-129-5psense): 5′-GATCCGCAAGCCCAGACCGCAAAAAGTTTTTA-3′; miR-129-5p (antisense): 5′-AGCTTAAAAACTTTTTGCGGTCTGGGCTTGCG-3′; UNC5B (sense): 5′-AACCTGGCTGAGAAAAGCAGT-3′; UNC5B (antisense): 5′-TCACAATGTAGGCGTCCTGT-3′; U6 (sense): 5′-CTCGCTTCGGCAGCACA-3′; U6 (antisense): 5′-AACGCTTCACGAATTTGCGT-3′; GAPDH (sense): 5′-GAAGGTGAAGGTCGGAGTC-3′; GAPDH (antisense): 5′-GAAGATGGTGATGGGATTTC-3′.
Cell culture
Thyroid cancer cell lines (KAT5, Hth74, TPC-1 and IHH4) and human normal thyroid follicular epithelial cell line Nthy-ori3-1 were bought from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China). All cells were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM, Gibco, Grand Island, NY, USA) supplemented with 100 mg/mL streptomycin, 100 U/mL penicillin and 10% fetal bovine serum (FBS; Gibco) in a humidified incubator containing 5% CO2 at 37°C.
Cell transfection
PITPNA-AS1 small interfering RNA (siRNA), UNC5B overexpression plasmid (pcDNA3.1-UNC5B), miR-129-5p inhibitor as well as the corresponding controls were available from GenePharma (Shanghai, China). They were transfected into PTC cells by Lipofectamine 3000 (Invitrogen) according to the instructions provided by the manufacturer.
Cell growth assay
The cell proliferation was examined by cell counting kit-8 (CCK-8, DoJinDo, Kumamoto, Japan) assay. The transfected cells were inoculated into 96-well plates (2×103 cells/well). At 0, 24, 48, and 72 h of culture, each well was added with 10 μL of CCK-8 reagent. 1 h later, a microplate reader (Bio-Rad, Hercules, CA, USA) was employed for measuring the optical density (OD) at 450 nm. 72 h later, the growth curve of the cells in each group was plotted.
Ethynyldeoxyuridine (EdU) assay
The cell proliferation was examined by ethynyldeoxyuridine (EdU) assay kit (RiboBio, Guangzhou, China). To be specific, the transfected cells were inoculated into 96-well plates. 48 h later, each well was added with 50 μM of EdU labeling medium and the cells were incubated for 8 h. Subsequently, cells were fixed with 4% paraformaldehyde and then stained with Apollo Dye Solution. Hochest 33342 was used to mark the nucleus. Eventually, under the microscope, EdU-positive cells were observed and counted.
Wound healing assay
Transfected cells were inoculated into 6-well plates. When cells reached 90% confluence, the pipette tip of a 200 μL micropipette was utilized for scratching a cell-free gap across the cell layer. Then the width of the gap was observed and photographed under a microscope. The cells were cultured in serum-free medium for 24 h before the width of the gap was observed and photographed again. After that, the wound closure was evaluated.
Flow cytometry
After the transfected cells were washed with pre-cooled PBS, they were resuspended in loading buffer and then incubated with Annexin V-FITC staining solution and propidium iodide (PI) staining solution (Beyotime, Shanghai, China) in the dark at room temperature for 15 min, and flow cytometry (BD Biosciences) was utilized for apoptosis rate detection.
Western blotting assay
Protein extraction from tissues or cells was conducted with RIPA lysis buffer (Beyotime), and the protein was separated using sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and then transferred to polyvinylidene fluoride (PVDF) membranes (Invitrogen). Subsequently, 5% skim milk was employed to block the membranes for 2 h. Following this, they were incubated overnight at 4ºC with primary antibodies, including anti-GAPDH and anti-UNC5B (Abcam, Cambridge, UK), and then with secondary antibodies (Abcam) for 2 h at room temperature. Ultimately, protein bands visualization was conducted with enhanced chemiluminescence (ECL) detection system (Thermo Scientific, Rockford, IL, USA).
Luciferase assay
The sequence containing mutant-type (MUT) or wild-type (WT) PITPNA-AS1 and UNC5B 3′UTR were integrated into pGL3 luciferase reporter vector (Promega, Madison, WI, USA). Lipofectamine 3000 (Invitrogen) was employed to transfect miR-129-5p mimics or miR-NC and the above luciferase reporter plasmids into 293T cells. Forty-eight hours later, the Dual-luciferase reporter assay system (Promega) was utilized to detect the luciferase activity.
Statistical analysis
All of the experiments were conducted at least three times, and mean ± SD was the expression form of the results. Student’s t-test was applied to conduct the comparison between the two groups. Chi-square test was utilized for detecting the correlation of clinicopathological characteristics with PITPNA-AS1 expression. The correlation between the expressions of two genes was analyzed employing Pearson’s correlation analysis. P < 0.05 meant that differences were of statistical significance.
Results
PITPNA-AS1 expression was increased in PTC cell lines and tissues
In adjacent tissues and PTC tumor tissues, qRT-PCR assay was used for detecting PITPNA-AS1 expression, and it was demonstrated that PITPNA-AS1 expression in PTC tumor tissues was markedly higher compared to that in adjacent tissues (Figure 1(a)). Additionally, we detected PITPNA-AS1 expression in thyroid cancer cell lines and Nthy-ori3-1, the results of which unearthed that PITPNA-AS1 expression in thyroid cancer cells was higher compared with that in Nthy-ori3-1 (Figure 1(b)). Since PITPNA-AS1 was the most highly expressed in KAT5 and Hth74 cell lines, they were employed for follow-up functional experiments.
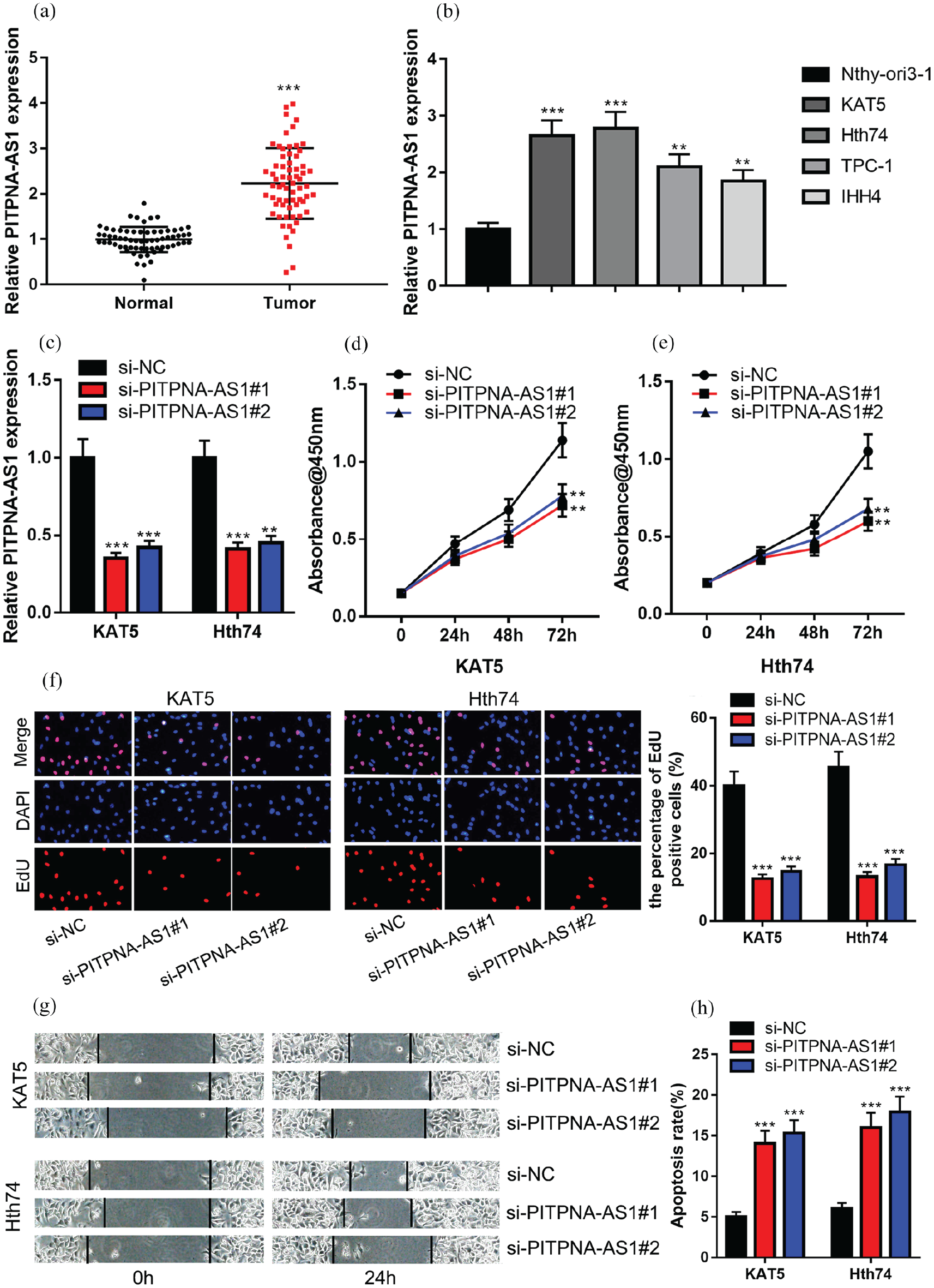
PITPNA-AS1 promoted PTC cell proliferation and migration, and inhibited the apoptosis. (a) qRT-PCR assay was utilized for comparing the difference in PITPNA-AS1 expression between adjacent tissues and PTC tissues. (b) qRT-PCR assay was employed to compare the difference in PITPNA-AS1 expression between Nthy-ori3-1 and PTC cell lines (KAT5, Hth74, TPC-1 and IHH4). (c) PITPNA-AS1 knockdown model was successfully constructed. (d) to (e) CCK-8 assay was utilized for detecting cell proliferation after the transfection. (f) EdU assay was utilized for detecting cell proliferation after the transfection. (g) Wound healing assay was utilized for detecting cell migration ability after the transfection. (h) Flow cytometry was utilized for detecting cell apoptosis after the transfection.
To make further analyses of PITPNA-AS1’s clinical significance in PTC, the relationship between PITPNA-AS1 expression level and clinicopathological features was analyzed. The samples were divided into PITPNA-AS1 high and low expression groups and the results displayed the positive correlation between high PITPNA-AS1 expression and the larger tumor size (Supplementary Table 1).
PITPNA-AS1 facilitated PTC cell proliferation and migration, and suppressed cell apoptosis
Next, PITPNA-AS1 siRNA#1 (si-PITPNA-AS1#1) and PITPNA-AS1 siRNA#2 (si-PITPNA-AS1#2) were used to silence PITPNA-AS1 expression in KAT5 and Hth74 cell lines (Figure 1(c)). CCK-8 assay results showed that knockdown of PITPNA-AS1 could significantly reduce cell proliferation in comparison to that in the control group (Figure 1(d) to (e)). Additionally, EdU assay indicated that EdU-positive cell number in the si-PITPNA-AS1 group was observably less compared to that in the si-NC group (Figure 1(f)); the wound healing assay suggested that the migration of PTC cells was markedly decreased after the knockdown of PITPNA-AS1 (Figure 1(g)); flow cytometry analysis showed that after PITPNA-AS1 were silenced in PTC cells, the apoptosis rate of PTC cells was dramatically higher compared to that in the control group (Figure 1(h)). The above findings indicated that PITPNA-AS1 might facilitate PTC progression by promoting proliferation and migration, and inhibiting cell apoptosis.
PITPNA-AS1 could absorb miR-129-5p in PTC as a sponge
StarBase V.3.0 database (http://starbase.sysu.edu.cn/) analysis manifested that PITPNA-AS1 contained a possible binding site for miR-129-5p (Figure 2(a)). Dual-luciferase reporter gene assay verified that in the miR-129-5p mimics+PITPNA-AS1-WT group, the luciferase activity was markedly lower compared to that in the miR-NC+PITPNA-AS1-WT group while no differences could be observed in the PITPNA-AS1-MUT groups (Figure 2(b)). qRT-PCR assay was utilized for detecting miR-129-5p expression in PTC tissues and cell lines and it was shown that compared to in adjacent tissues, miR-129-5p expression was observably down-regulated in PTC tissues; compared to in Nthy-ori3-1, miR-129-5p expression level in PTC cell lines was lower (Figure 2(c) to (d)). Pearson’s correlation analysis authenticated a negative correlation between PITPNA-AS1 expression and miR-129-5p expression in PTC samples (Figure 2(e)). Notably, in the si-PITPNA-AS1#1 group, miR-129-5p expression was markedly promoted (Figure 2(f)). Collectively, our results suggested that PITPNA-AS1 could adsorb miR-129-5p to repress it.
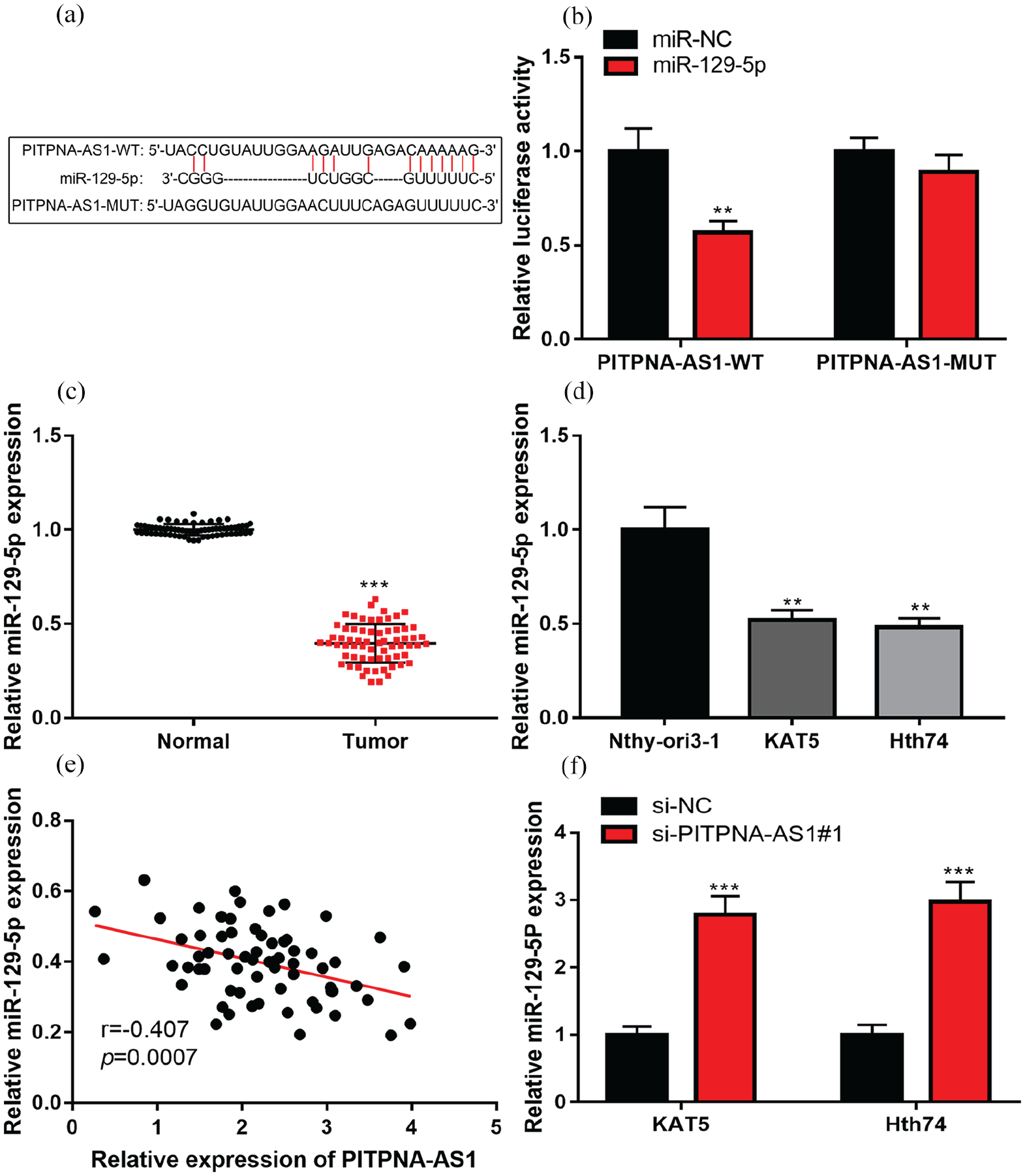
PITPNA-AS1 could absorb miR-129-5p in PTC. (a) It was predicted that PITPNA-AS1 contained a potential binding site to miR-129-5p.(b) Luciferase reporter assay validated that miR-129-5p could directly bind to PITPNA-AS1. (c) qRT-PCR assay was employed for comparing miR-129-5p expression difference between adjacent tissues and PTC tissues. (d) qRT-PCR assay was utilized for comparing miR-129-5p expression difference between PTC cell lines (KAT5 and Hth74) and Nthy-ori3-1. (e) Correlation analysis of miR-129-5p and PITPNA-AS1 expressions in PTC tissues. (f) qRT-PCR assay was utilized for detecting the effect of PITPNA-AS1 knockdown on miR-129-5p expression in PTC cells.
MiR-129-5p could partly counteract PITPNA-AS1’s effect on PTC cells
Subsequently, a rescue assay was conducted for validating whether miR-129-5p could partially reverse PITPNA-AS1’s cancer-promoting effect on PTC cells. MiR-129-5p inhibitors and PITPNA-AS1 siRNA#1 were co-transfected into KAT5 and Hth74 cell lines. Subsequently, EdU, CCK-8, and wound healing assays were performed, and the results suggested that miR-129-5p inhibitors could partly eliminate PITPNA-AS1 siRNA#1’s inhibitory effect on PTC cell proliferation, migration, and proliferation (Figure 3(a) to (d)). Moreover, flow cytometry displayed that miR-129-5p inhibitors could counteract the promotional effect of PITPNA-AS1 siRNA#1 on PTC cell apoptosis (Figure 3(e)). Therefore, it was supposed that PITPNA-AS1 could exert its cancer-promoting effect via repressing miR-129-5p.
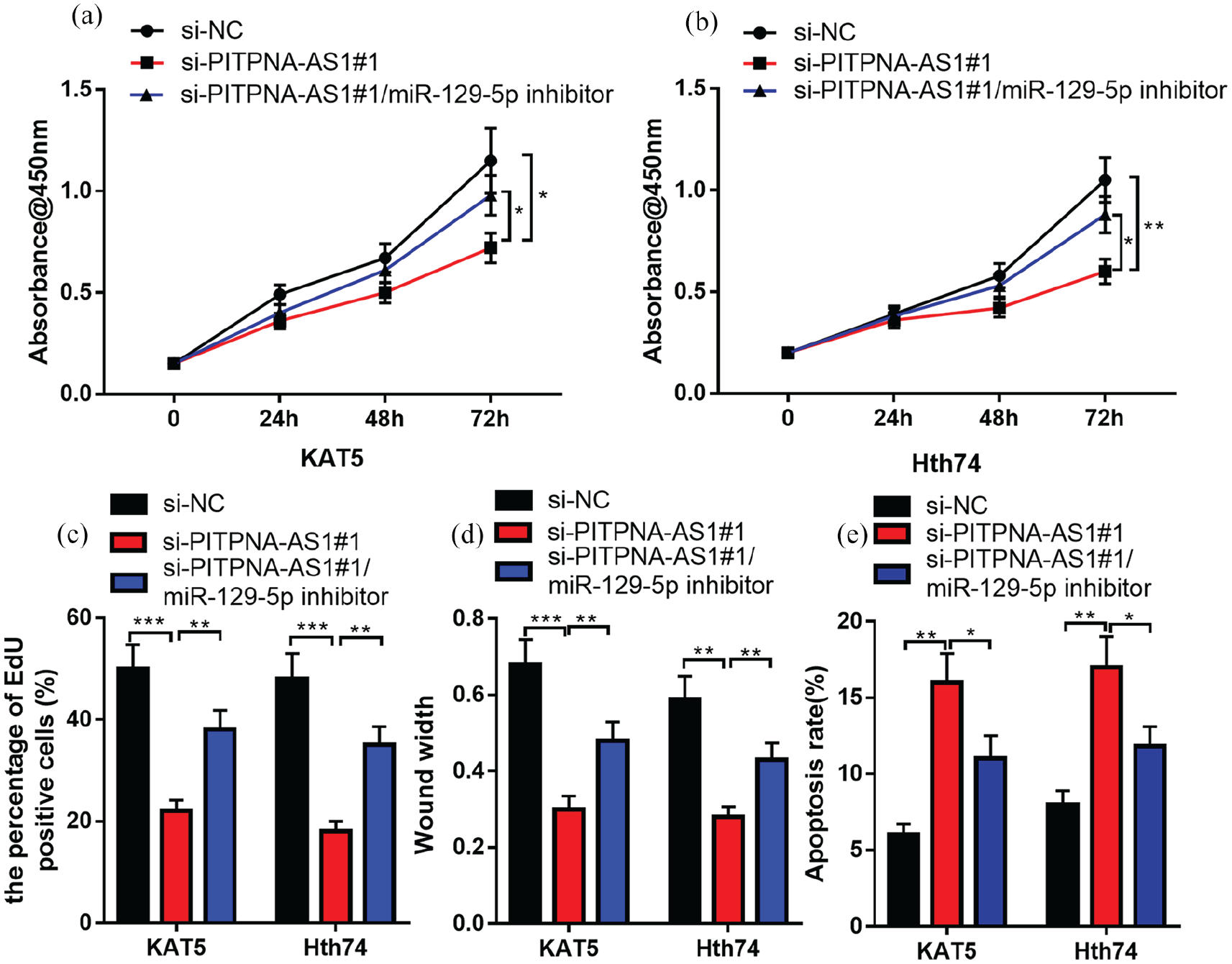
MiR-129-5p inhibitor could partly counteract the effect of PITPNA-AS1 knockdown on PTC cells. (a) to (b) CCK-8 assay was utilized for testing cell proliferation after the co-transfection of miR-129-5p inhibitors and PITPNA-AS1 siRNA. (c) EdU assay was utilized for detecting the cell proliferation. (d) Wound width experiment was conducted for detecting the cell migration ability. (e) Flow cytometry was employed to detect apoptosis.
UNC5B was a target of miR-129-5p
The miRNA.org database (http://www.microrna.org/microrna/getMirnaForm.do) showed that the 3’UTR of UNC5B contained a binding site for miR-129-5p (Figure 4(a)). To verify the binding relationship between UNC5B and miR-129-5p, dual-luciferase reporter gene assay was performed, and it was found that luciferase activity was significantly reduced in 293T cells co-transfected with WT 3′UTR UNC5B and miR-129-5p compared to that in miR-NC, but no change was seen in the group containing MUT 3′UTR UNC5B plasmids (Figure 4(b)). Furthermore, qRT-PCR assay elucidated that UNC5B expression was significantly up-regulated in PTC tumors (Figure 4(c) to (d)), and UNC5B mRNA was negatively correlated with miR-129-5p expression in PTC tissues (Figure 4(e)). Transfection of miR-129-5p inhibitor could remarkably promote UNC5B expression level (Figure 4(f)). Moreover, miR-129-5p inhibitor could partially abolish PITPNA-AS1 siRNA#1’s inhibitory effect on UNC5B expression in PTC cells (Figure 4 (g)). Therefore, it was confirmed that in PTC, UNC5B—a target gene for miR-129-5p—was probably one of the downstream genes of PITPNA-AS1.
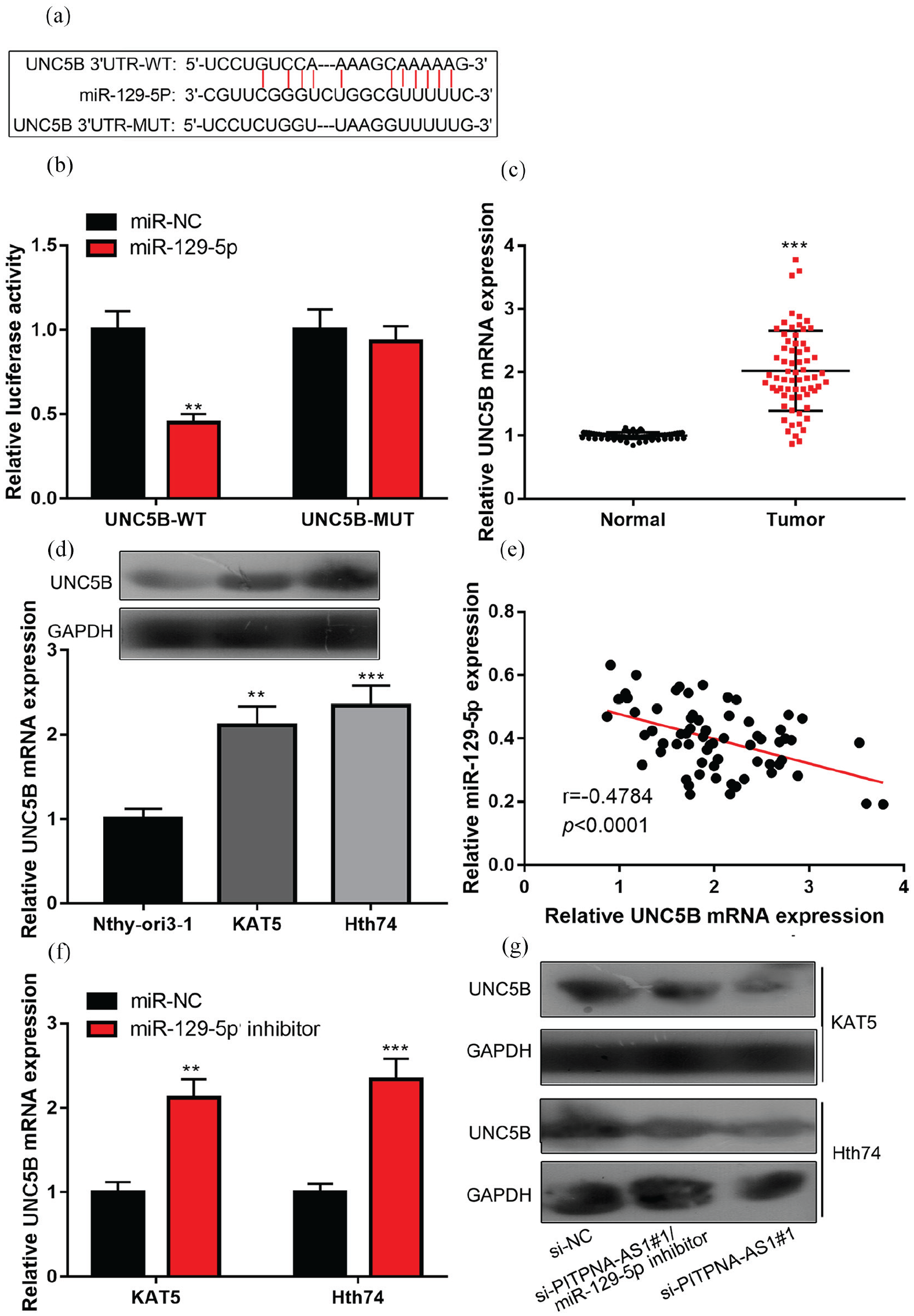
UNC5B was a target of miR-129-5p. (a) It was predicted that the 3’UTR of UNC5B mRNA contained a potential binding site to miR-129-5p. (b) Luciferase reporter assay confirmed that UNC5B mRNA could be targeted by miR-129-5p directly. (c) qRT-PCR assay was employed to compare UNC5B expression differences between adjacent tissues and PTC tissues. (d) Western blot and qRT-PCR assays were employed to compare UNC5B expression differences between PTC cell lines (KAT5 and Hth74) and Nthy-ori3-1. (e) Correlation analysis of UNC5B mRNA expression with miR-129-5p expression in PTC tissues. (f) qRT-PCR assay was utilized for detecting the effects of miR-129-5p inhibitor on UNC5B mRNA expression level. (g) Western blot detection displayed that miR-129-5p inhibitor could counteract the effect of the knockdown of PITPNA-AS1 on UNC5B expression level.
Overexpression of UNC5B could partially counteract the inhibitory effect of PITPNA-AS1 knockdown on PTC progression
To expound whether knockdown of PITPNA-AS1 affected PTC cells by regulating UNC5B expression, pcDNA3.1-UNC5B and PITPNA-AS1 siRNA#1 were co-transfected into KAT5 and Hth74 cell lines. Functional experiments confirmed that overexpression of UNC5B could partly abolish the regulatory function of PITPNA-AS1 knockdown on PTC cell proliferation (Figure 5(a) to (c)), migration (Figure 5(d)), and apoptosis (Figure 5(e)). Therefore, it was concluded that UNC5B overexpression could work as a downstream gene of PITPNA-AS1 and counteract the inhibitory effect of PITPNA-AS1 knockdown on PTC progression.
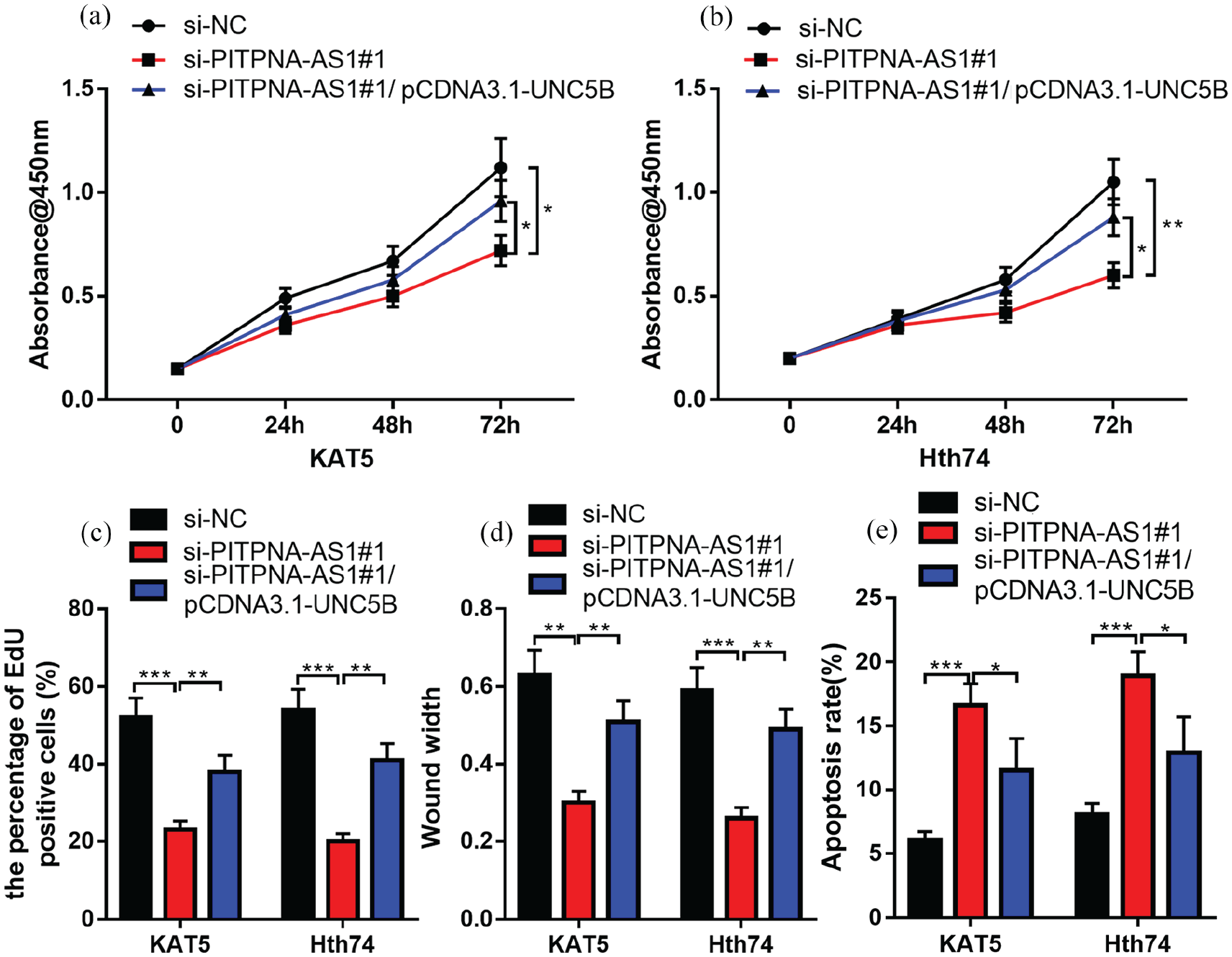
Overexpression of UNC5B could partially counteract the inhibitory effect of PITPNA-AS1 knockdown on PTC progression. (a) to (b) CCK-8 assay was utilized for testing cell proliferation after the co-transfection of UNC5B overexpression plasmid and PITPNA-AS1 siRNA. (c) EdU assay was utilized for detecting cell proliferation. (d) Wound healing assay was conducted for detecting the cell migration ability. (e) Flow cytometry was employed to detect the apoptosis.
Discussion
lncRNAs figure prominently in gene expression regulation and a lot of cellular life activities, such as cell differentiation, proliferation, migration, and apoptosis. 11 Besides, the aberrant expressions of lncRNAs probably contribute to tumorigenesis and cancer progression. At present, an enormous amount of research emphasizes the role of lncRNAs in PTC. For example, LINC00520 can function as a cancer-promoting molecule to participate in PTC tumorigenesis and development 12 ; SNHG3 can increase the malignancy of PTC cells and promote the progression of PTC 13 ; LINC00460 expression is dramatically enhanced in PTC cell lines and tissues; and LINC00460 can target miR-539 to up-regulate MMP expression, to facilitate PTC cell proliferation and migration, and to inhibit the apoptosis. 14 It is reported that PITPNA-AS1 takes part in hepatocellular carcinoma development as a cancer-promoting factor. 3 It facilitates the malignant biological behaviors of hepatocellular carcinoma cells via repressing miR-876-5p expression and up-regulating WNT5A expression. 3 In this study, we examined PITPNA-AS1 expression level in PTC tissues and its regulatory functions in the malignant phenotypes of PTC cells. It was found that PITPNA-AS1 expression was markedly up-regulated in PTC tissues and cell lines and high PITPNA-AS1 expression was linked to larger tumor size in patients with PTC. Functionally, both selected siRNAs targeting PITPNA-AS1 inhibited the malignancy of PTC cells. In summary, as a novel regulator, PITPNA-AS1 played a cancer-promoting role in PTC and might be a biomarker and therapy target.
lncRNAs can absorb miRNAs as ceRNA to reduce the availability of miRNAs, thus inducing the expressions of the target genes, and this network contributes to cancer progression.3,14 In this study, miRNAs that were probably targeted by PITPNA-AS1 were predicted through the bioinformatics database. Among these miRNAs, we focused on the tumor suppressor miR-129-5p, which participates in the development of many tumors. MiR-129-5p can suppress gastric cancer cell proliferation and invasion via repressing COL1A1 expression. 15 In oral squamous cell cancer, miR-129-5p represses cell growth and invasion via suppressing HMGB1 expression. 16 Reportedly, miR-129-5p expression is markedly down-regulated in PTC cell lines and tissues and lncRNA NEAT1 regulates miR-129-5p/KLK7 axis to modulate the biological behaviors of PTC cell. 17 Another study demonstrates that as a ceRNA of miR-129-5p, TNRC6C-AS1 represses PTC cell proliferation, invasion and migration through modulating UNC5B. 9 In this study, dual-luciferase reporter assay verified the direct binding of miR-129-5p to PITPNA-AS1. Additionally, Pearson’s correlation analysis indicated that miR-129-5p expression was negatively associated with PITPNA-AS1 in PTC tissues and miR-129-5p expression was up-regulated after PITPNA-AS1 was silenced in PTC cells. Moreover, miR-129-5p inhibitor could counteract the effect of down-regulating PITPNA-AS1 expression on PTC cell migration, proliferation, and apoptosis. It turned out that PITPNA-AS1 could serve as an endogenous miRNA sponge and repress miR-129-5p expression during PTC progression.
Netrin-1, a member of the netrin family, is a type of 60-80 kD amino-protein and can promote cell invasion and angiogenesis, and inhibit cell apoptosis. 18 UNC5H family proteins (UNC5A, UNC5B, UNC5C, and UNC5D) are netrin-1 receptors. 19 Besides, UNC5B activates PI3K/Akt signaling to enhance tumor proliferation, increase metastasis and worsen prognosis in breast cancer. 20 Research demonstrates that UNC5B expression is up-regulated in thyroid cancer and UNC5B can function as a target gene for miR-129-5p to facilitate PTC cell proliferation, migration and invasion. 9 In the present study, we verified that UNC5B expression was remarkably enhanced in PTC cell lines and tissues and it was a target gene for miR-129-5p in PTC. Additionally, it was found that PITPNA-AS1 indirectly up-regulated UNC5B expression in PTC and UNC5B could counteract the effect of PITPNA-AS1 knockdown on PTC cell proliferation, migration and apoptosis. However, we did not conduct animal and follow-up experiments to confirm the role of the PITPNA-AS1/miR-129-5p/UNC5B axis in PTC. More studies are required to validate our findings in the future.
To sum up, this study suggests that PITPNA-AS1 can facilitate PTC development by regulating the miR-129-5p/UNC5B axis. PITPNA-AS1 may be a prospective marker or therapeutic target of PTC.
Supplemental Material
sj-docx-1-jbm-10.1177_1724600820985528 – Supplemental material for Long non-coding RNA PITPNA-AS1 regulates UNC5B expression in papillary thyroid cancer via sponging miR-129-5p
Supplemental material, sj-docx-1-jbm-10.1177_1724600820985528 for Long non-coding RNA PITPNA-AS1 regulates UNC5B expression in papillary thyroid cancer via sponging miR-129-5p by Shijuan Mei, Huafeng Zong and Haicheng Zhou in The International Journal of Biological Markers
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
