Abstract
Background:
Long non-coding RNAs (lncRNAs) have been confirmed to participate in the regulation of nasopharyngeal carcinoma. Here, we endeavored to explore the character of lncRNA ILF3-AS1 in the nasopharyngeal carcinoma and its function.
Methods:
A total of 68 nasopharyngeal carcinoma tissues and adjacent normal nasopharyngeal tissues were collected. Expressions of lncRNA ILF3-AS1 in these tissues were detected using quantitative real-time polymerase chain reaction (qRT-PCR). The relationship between the expression level of lncRNA ILF3-AS1 and clinical pathological characteristics was analyzed. Inhibition of lncRNA ILF3-AS1 was done using small interference RNA.
Results:
lncRNA ILF3-AS1 expression was significantly up-regulated in the 68 nasopharyngeal carcinoma tissue samples compared to their adjacent normal tissue samples. Increased lncRNA ILF3-AS1 level was related to the advanced tumor node metastasis stage and the metastasis of nasopharyngeal carcinoma. Also, increased lncRNA ILF3-AS1 indicated poor prognosis of nasopharyngeal carcinoma patients. Inhibition of lncRNA ILF3-AS1 reduced proliferation, invasion and migration of nasopharyngeal carcinoma cells. MicroRNA-320a (miR-320a) was determined as a direct target for lncRNA ILF3-AS1 in nasopharyngeal carcinoma. Furthermore, lncRNA ILF3-AS1 could sponge miR-320a to promote BMI1 expression. The expression of BMI1 was significantly inhibited by the down-regulation of lncRNA ILF3-AS1.
Conclusions:
For the first time, we demonstrated that lncRNA ILF3-AS1 was markedly over-expressed in nasopharyngeal carcinoma tissues and cells. Elevated lncRNA ILF3-AS1 expression was correlated with severe cancer stage and poor prognosis. lncRNA ILF3-AS1 could promote proliferation, invasion, and migration of cells, which might indicate a novel target site for the future diagnosis and therapy of nasopharyngeal carcinoma.
Introduction
Nasopharyngeal carcinoma (NPC) is one of the malignant tumors in the head and neck. According to the study of Global Cancer Statistics 2018, NPC contributes to 0.7% of new cases and 0.8% of deaths among all cancers worldwide. 1 Due to high invasiveness and metastasis, 60% to 85% of patients with NPC had clinical metastasis at the time of diagnosis. Early metastasis is one of the main causes of failure in treatment of NPC. 2 Although comprehensive treatment based on radiotherapy, surgery, and chemotherapy has improved in recent years, the therapeutic effect on progressive metastatic NPC is still not satisfactory. 3 Therefore, early diagnosis and targeted therapy are the hotspots for NPC research.
Non-coding RNAs—including microRNAs (miRNAs) and long non-coding RNAs (lncRNAs)—have been verified to be involved in the genesis and development of tumors. 4 lncRNAs longer than 200 nt are recognized as “transcription noise” without any functions. However, in recent years, lncRNAs have been verified to regulate cell proliferation, angiogenesis, invasion, migration, apoptosis, and cell circle transition in several tumors.5,6 In lung squamous cell carcinoma, lncRNA HULC accelerates tumor progression by changing the expression of PTPRO through regulating nuclear factor-κB (NF-κB). 7 In breast cancer, lncRNA NONHSAT101069 regulates the miR-129-5p/Twist1, promoting epirubicin resistance, migration, and invasion. 8 In osteosarcoma, lncRNA AFAP1-AS1 acts as an oncogene to promote the development and epithelial-mesenchymal transition (EMT) via RhoC/ROCK1/p38MAPK/Twist1 signaling axis. 9 A novel lncRNA LINC00997 is related with the metastasis of kidney renal clear cell carcinoma by regulating S100A11. 10 Also, LINC00511 indicates the malignant status of cervical cancer and encourages cell proliferation and motility. 11
lncRNA ILF3-AS1, located at 19p13.2, has been identified to participate in the progression of several cancers including cervical cancer, melanoma, and colon cancer.12-14 However, the relative expression level of lncRNA ILF3-AS1 in the NPC has not been detected before, and its function in NPC remains unknown. Here, we measured the level of lncRNA ILF3-AS1 in 68 paired NPC and adjacent normal tissues for the first time. Also, we described the relationship between lncRNA ILF3-AS1 and clinical pathological features of the 68 NPC patients. Using NPC-derived cell line CNE-1, we elucidated the function and mechanism of lncRNA ILF3-AS1 in NPC. With these experiments, we might provide lncRNA ILF3-AS1 as a potential target for the diagnosis and therapy of NPC in the future.
Patients and methods
Clinical specimens
A total of 68 patients with NPC who were admitted to our hospital from June 2016 to June 2018 were selected. The inclusion criteria were: (a) patients underwent biopsy of nasopharyngeal tumor and its adjacent tissue in our hospital; (b) diagnosed as NPC by two pathologists, and had no radiotherapy or chemotherapy before biopsy; (c) underwent ultrasound, radiography, computed tomography scans and nuclear medicine imaging for tumor node metastasis (TNM) staging; (4) no other complications and serious cardiopulmonary diseases. TNM staging was conducted in accordance with the Union for International Cancer Control Version 7. NPC and its adjacent specimens were preserved in liquid nitrogen for later use.
This study has the approval of the Ethics Committee of Jiangsu Province Hospital of Chinese Medicine and all patients gave informed consent to participate in the study.
Cells lines and cell transfection
NPC-derived cell line CNE-1 and 5-8F and normal nasopharyngeal epithelial cell line NP69 were purchased from American Type Culture Collection (ATCC, Manassas, VA, USA). The CNE-1, 5-8F and NP69 cells were maintained in Dulbecco’s modified eagle medium (DMEM) (Gibco, New York, NY, USA) involving 10% fetal bovine serum (FBS; Gibco, New York, NY, USA) and 1% penicillin-streptomycin (Gibco) in a humid environment of 37°C, containing 5% CO2. 15 With using Lipofectamine 3000 (Invitrogen, Carlsbad, CA, USA), siRNA for lncRNA ILF3-AS1 (siRNA-lncRNA ILF3-AS1) or negative control (siRNA-NC) was added to each well at a concentration of 50 nmol/L.16,17 The siRNAs are synthesized by the GenScript Corporation (Nanjing, China). The siRNA-lncRNA ILF3-AS1 sequence is: 5′-TCGCCCTGAGTACGTTAGCGGAAAT-3′. Transfection efficiency was verified by quantitative real-time polymerase chain reaction (qRT-PCR).
RNA isolation and qRT-PCR
Total RNA from NPC tissues and experimental cells were extracted using TRIzol reagent (Invitrogen). The first strand of DNA was synthesized using PrimeScript RT Master Mix Kit (TaKaRa, Tokyo, Japan). Real-time PCR was then performed using SYBR Green I (TaKaRa) with ABI PRISM 7900 Real-Time system (Applied Biosystems, Foster City, CA, USA). As internal control, U6 was used for miR-320a and glyceraldehyde 3-phosphate dehydrogenase (GAPDH) for lncRNA ILF3-AS1 and BMI1. The primer sequences were synthesized by GeneWiz (Suchou, China) as follows: GAPDH: Forward: 5′-CTCAGTTGCTGAGGAGTCCC-3′, Reverse: 5′-ATTCGGAGAAGGGAGGGCT-3′; lncRNA ILF3-AS1: Forward: 5′-GTATCTCGGCCGGGGATTTT-3′, Reverse: 5′-TGTACATCGGGGGCCTCATA-3′; BMI1: Forward: 5′-GCTGCCAATGGCTCTAATGAA-3′, Reverse: 5′-TGCTGGGCATCGTAAGTATCTT-3′. The measurement was repeated three times.
Colony formation assay
The experimental CNE-1 cells in the logarithmic growth phase were adjusted to 1 × 103 cells/well, inoculated into a 6-well plate, and maintained in the incubator. After 15 days of culture, CNE-1 cells were washed with phosphate buffered saline (PBS) three times and fixed with 4% paraformaldehyde for 15 min. After crystal violet staining for 10 min, colonies with >50 cells were counted and compared.
5-Ethynyl-2′- deoxyuridine (EdU) assay
After transfecting the CNE-1 cells with siRNA for 24 h, cells were prepared as single cell suspension. Then, the suspension was inoculated into a 96-well plate and incubated for 12 h in a cell culture incubator. Reagents were then added to the well for testing sequentially according to the instructions. Then, the reaction solution was eluted with PBS and the well was photographed under a fluorescence microscope. Blue fluorescence represented the total number of cells detected, and red fluorescence represented proliferating cells.
Cell proliferation rate = number of red fluorescent cells / number of blue fluorescent cells × 100%.
Transwell assay and Matrigel assay
For Transwell assay, single established CNE-1 cell suspension was prepared in serum free DMEM medium. After counting, the cell density was adjusted to 1 × 105 per mL. A total of 200 μL of the cell suspension was placed to the upper layer of the chamber, and 600 μL of 10% FBS DMEM medium was added to the lower layer of the 8-μm chamber (Corning, Corning, NY, USA). After incubated for 48 h, chambers were placed in the paraformaldehyde for 10 min and stained in crystal violet for 30 min. CNE-1 cells at the bottom of the chamber were wiped out with a cotton swab; the cells on the outer side of the chamber were the migrated cells. Cells were observed under the microscope. For the Matrigel assay, a layer of Matrigel (BD Biosciences, San Jose, CA, USA) was placed on the bottom of the chamber. The rest of the steps were the same as the Transwell experiment.
Dual-luciferase assay
The Dual-Luciferase Reporter Assay Systems (Promega, Madison, WI, USA) was used to detect the luciferase activity. To construct a dual luciferase reporter vector, wild-type lncRNA ILF3-AS1and mutant lncRNA ILF3-AS1or wild-type BMI1 and mutant BMI1 (mutant miR-320a binding site) were cloned into the GP-miRGLO vector by Genepharma Co. (Shanghai, China). The WT, Mut, miR-320a mimics, or mimic negative controls were transfected into CNE-1 cells with lipofectamine 2000. Luciferase activity was measured 48 hours after cell transfection according to the manufacturer's instructions. The experiment was repeated three times.
Western blot analysis
Western blot analysis was employed to determine the protein expression level. RIPA reagent (Beyotime, Shanghai China) containing the PMSF protease inhibitor (Vazyme, Nanjing, China) was applied to isolate the total protein, and the BCA Kit (Beyotime) was applied to determine the protein concentration. A total of 20 mg protein was taken and added to 10% sodium dodecyl sulphate-polyacrylamide gel electrophoresis (SDS-PAGE) for electrophoretic separation, and then transferred to polyvinylidene fluoride (PVDF) membrane (Millipore, Billerica, MA, USA). The protein band was cultured overnight with specific primary antibody at 4°C. After washing, the membrane was immersed in the goat anti-rabbit IgG antibody labeled with horseradish peroxidase at 37°C for 1 h. After washing, the membrane was exposed using electrochemiluminescence (ECL) Kit (Millipore). The antibodies were: anti-BMI1 (1:1000, Cell Signaling Technology, Danvers, MA, USA), and anti-GAPDH (1:2000, Cell Signaling Technology).
Statistical analysis
Statistical Product and Service Solutions (SPSS) 20.0 Software (IBM, Armonk, NY, USA) was used to perform the statistical analysis. The t-test was used for analyzing measurement data. Differences between the two groups were analyzed by using the Student's t-test. Comparison between multiple groups was done using the one-way ANOVA test followed by a post hoc test (least significant difference). The difference was statistically significant at P<0.05.
Results
lncRNA ILF3-AS1 expressed significantly higher in NPC tissues and indicated poor prognosis of NPC
To expound the role of lncRNA ILF3-AS1 in NPC, we collected 68 paired NPC and adjacent tissue samples from NPC patients together with their clinical pathological characteristics. Using qRT-PCR, we detected the expression level of lncRNA ILF3-AS1 in these tissues and found an obvious increase of the lncRNA ILF3-AS1 level in the NPC tissue group compared with the adjacent normal tissue group (Figure 1(a)). At the same time, we measured the lncRNA ILF3-AS1 expression in NPC-derived CNE-1 and 5-8F cells compared to the normal nasopharyngeal epithelial cell NP69 cells. The CNE-1 and 5-8F cells showed a markedly up-regulated lncRNA ILF3-AS1 expression than the NP69 cells (Figure 1(b)). This indicated that lncRNA ILF3-AS1 might act as an oncogene in NPC.
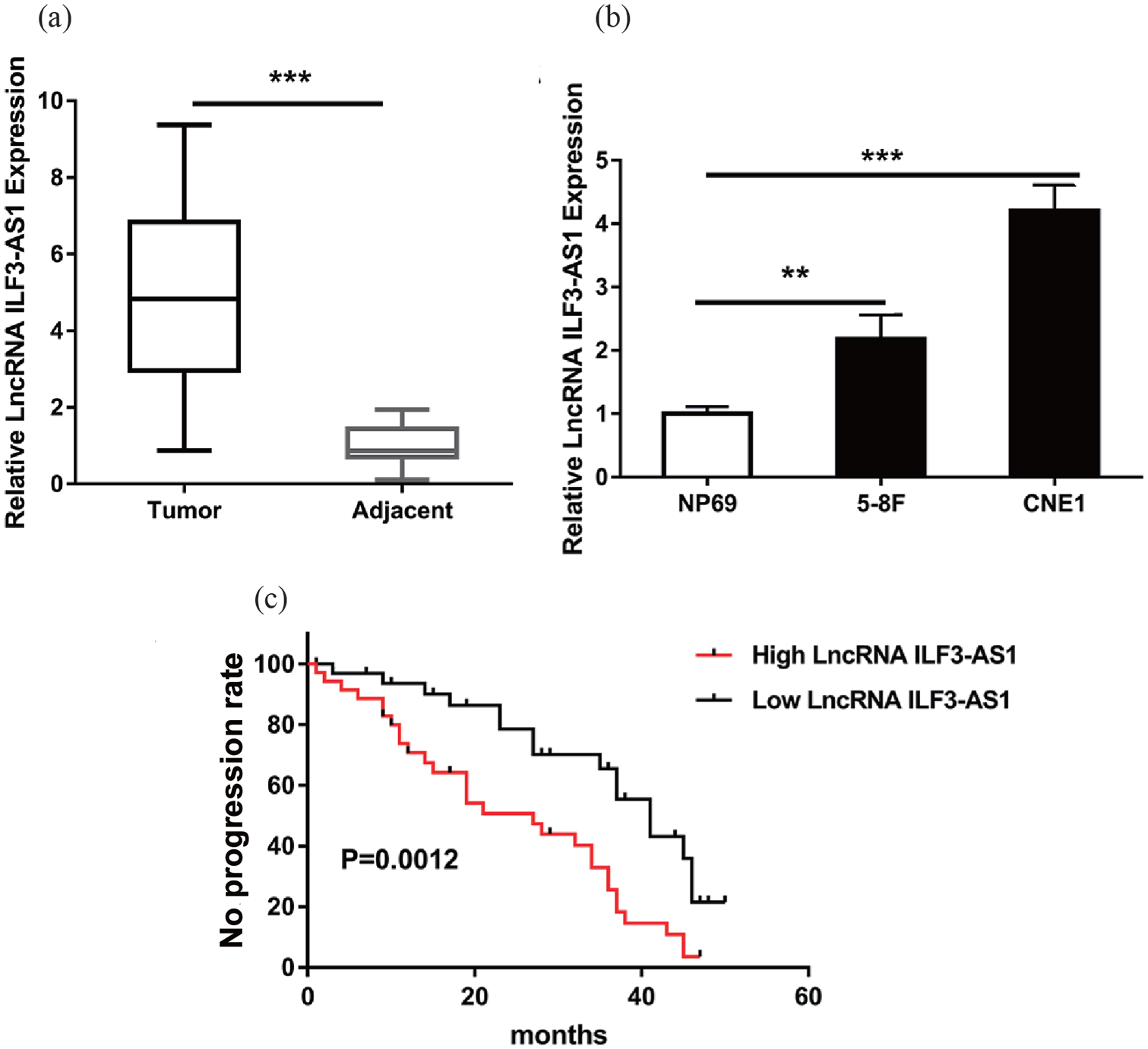
lncRNA ILF3-AS1 was increased in NPC tissues and cells. (a) qRT-PCR showed the lncRNA ILF3-AS1 expression level in total of 68 paired NPC tissues and adjacent para-tumor tissues. (b) lncRNA ILF3-AS1 expression level in NPC cell lines (CNE-1, 5-8F) and human normal nasopharyngeal epithelial cell line NP69. (c) Kaplan–Meier survival plot indicated that patients with high lncRNA ILF3-AS1 expression in tumor tissues showed significantly decreased survival compared with patients with low lncRNA ILF3-AS1 expression.
Next, we analyzed the relationship between the lncRNA ILF3-AS1 level and clinical pathological features including sex, TNM stage, metastasis and Epstein-Barr virus (EBV) infection. As shown in Table 1, higher lncRNA ILF3-AS1 level correlated with TNM stage and metastasis but had no concern with sex and EBV infection. Higher lncRNA ILF3-AS1 indicated advanced TNM stage and higher rate of metastasis in NPC patients. Furthermore, survival analysis of the 68 NPC patients suggested that the low-level lncRNA ILF3-AS1 group had a better prognosis than the high-level group (Figure 1(c)). All these results elucidated that lncRNA ILF3-AS1 was over-expressed in NPC tissues and cells and indicated poor prognosis of NPC patients.
Relationship between lncRNA ILF3-AS1 level and the clinical pathological characteristics of 68 patients with NPC.
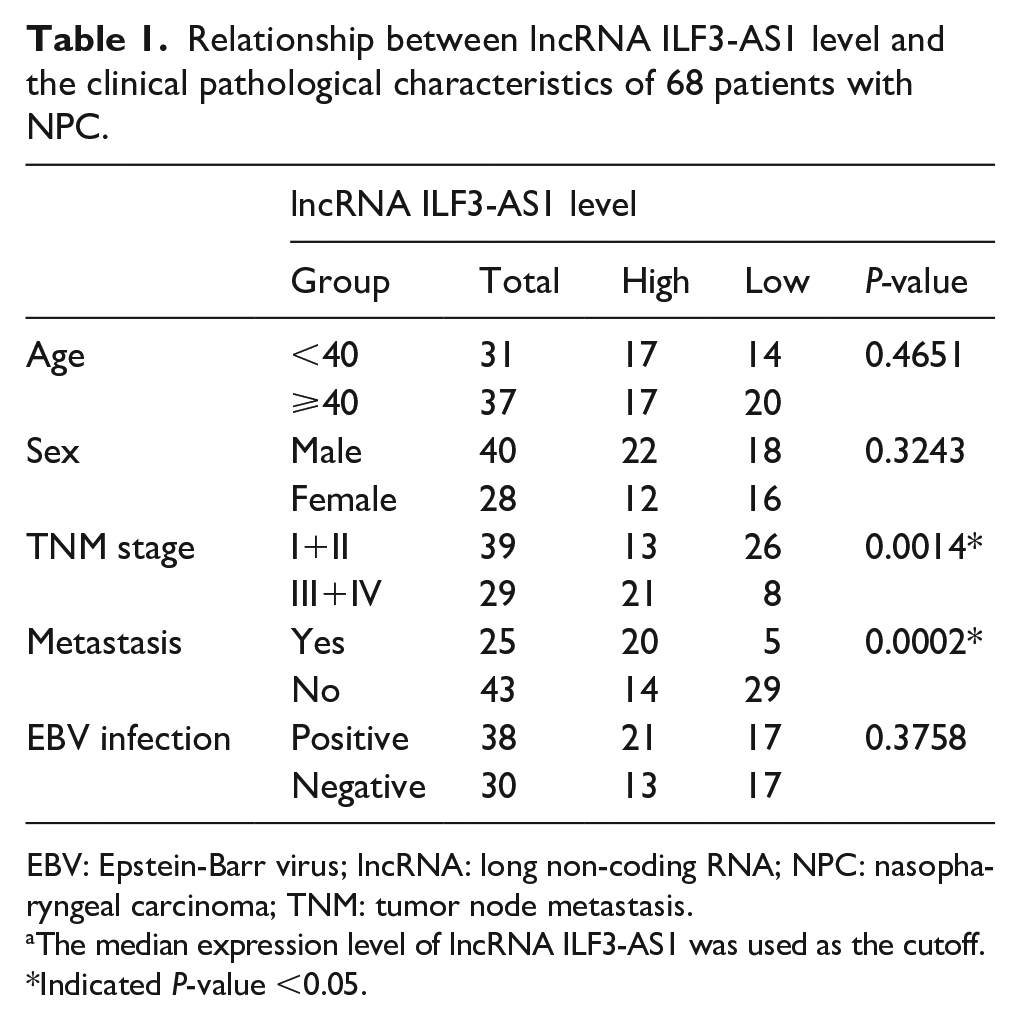
EBV: Epstein-Barr virus; lncRNA: long non-coding RNA; NPC: nasopharyngeal carcinoma; TNM: tumor node metastasis.
The median expression level of lncRNA ILF3-AS1 was used as the cutoff.
Indicated P-value <0.05.
Inhibition of lncRNA ILF3-AS1 reduced cell proliferation of CNE-1 cells
Next, to further explain the role of lncRNA ILF3-AS1 in NPC in vitro, we inhibited lncRNA ILF3-AS1 expression in CNE-1 cells using siRNA. Expression of lncRNA ILF3-AS1 was significantly decreased after siRNA-lncRNA ILF3-AS1 treatment compared to the siRNA-NC group (Figure 2(a)). Next, we employed colony formation assay and EdU assay to describe the CNE-1 cell proliferation. Cell ability of colony formation was impaired by lncRNA ILF3-AS1 inhibition in CNE-1 cells (Figure 2(b)). Similarly, EdU assay demonstrated that growth of CNE-1 cells was remarkably weakened by siRNA-lncRNA ILF3-AS1 compared to the siRNA-NC group (Figure 2(c)). These data suggested that inhibition of lncRNA ILF3-AS1 reduced cell proliferation of NPC cells in vitro.
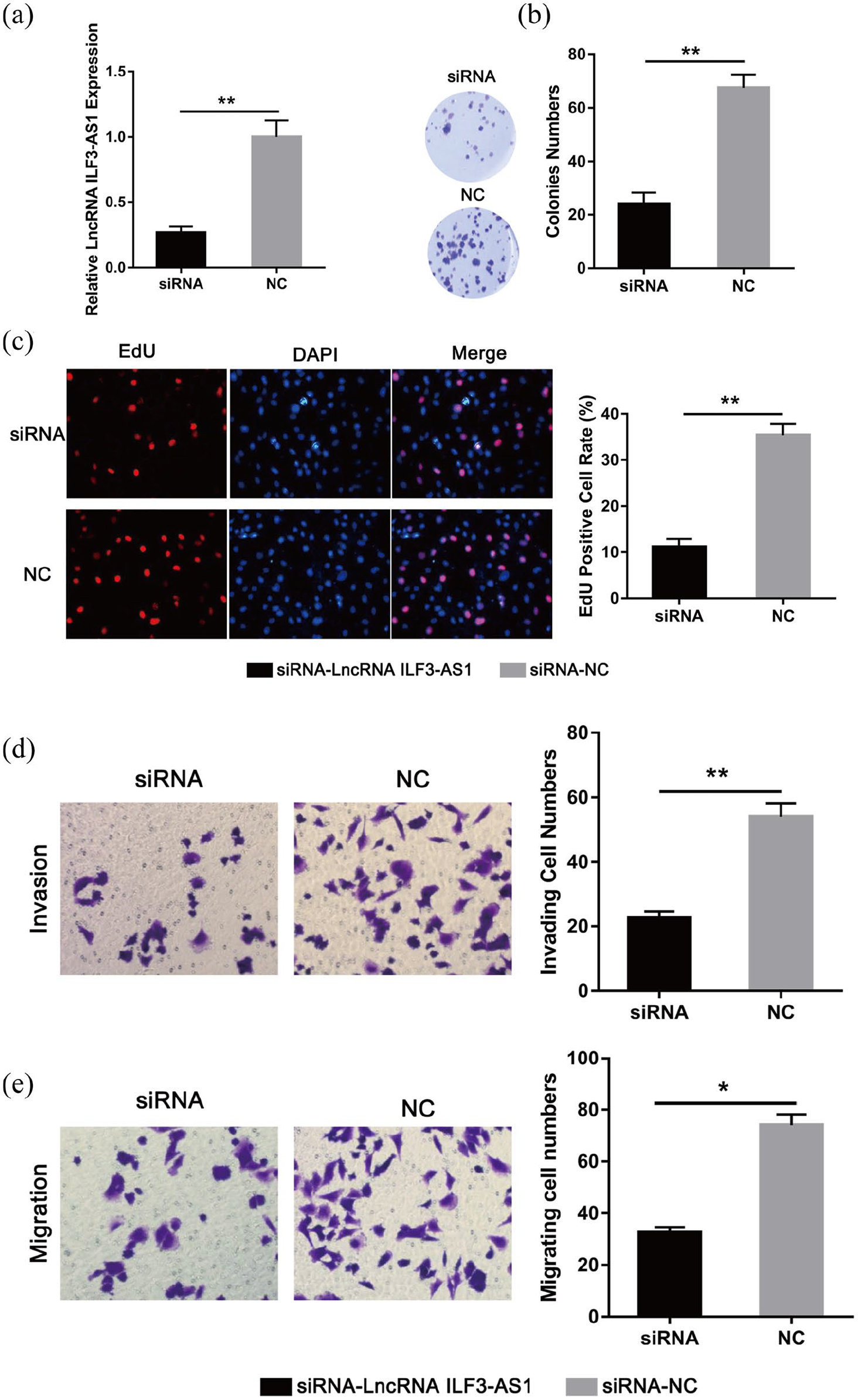
lncRNA ILF3-AS1 affected the proliferation, invasion and migration of NPC cells in vitro. (a) SiRNA targeting lncRNA ILF3-AS1 and negative controls (siRNA-NC) were transfected into CNE-1 cells. (b) Colony formation assay showed the proliferation ability of CNE-1 cells transfected with siRNA-lncRNA ILF3-AS1 and siRNA-NC. (c) EdU assay showed the growth of CNE-1 cells transfected with siRNA-lncRNA ILF3-AS1 and siRNA-NC. (d) Matrigel assay showed the invaded cell number in established CNE-1 cells. (e) Transwell assay showed the migrated cell number in established CNE-1 cells.
Ectopic lncRNA ILF3-AS1 level changed the cell invasion and migration of CNE-1 cells
Next, we employed Transwell and Matrigel assays to subscribe the cell invasion and migration abilities. As displayed in Figure 2(d), the inhibition of lncRNA ILF3-AS1 significantly reduced the number of CNE-1 cells invaded compared to the siRNA-NC group. Also, the cell migration ability was impaired by lncRNA ILF3-AS1 over-expression compared to the siRNA-NC group (Figure 2(e)). These data indicated that lncRNA ILF3-AS1 could promote cell invasion and ability of NPC cells.
MiR-320a was verified as a target for lncRNA ILF3-AS1 in NPC
Several studies have shown that lncRNAs could act as competitive endogenous RNA via direct sponging of their target gene including miRNAs. Herein, with searching several database including Starbase, Diana, and Deepbase, miR-320a was found to be a potential target for lncRNA ILF3-AS1 (Figure 3(a)). To confirm this hypothesis, we constructed plasmids containing the wild type or mutant lncRNA ILF3-AS13′-UTR and separately cloned them into the luciferase vector. After detecting the expression of luciferase fluorescence, we found that the fluorescence activity of the mutant group was obviously reduced comparing to the control, while the wild group did not change significantly (Figure 3(b)). In experimental CNE-1 cells, we found an elevated level of miR-320a in the lncRNA ILF3-AS1 inhibited group compared with the siRNA-NC group (Figure 3(c)). Furthermore, expression levels of miR-320a in the 68 paired NPC and normal tissues were detected. As expected, the expression of miR-320a was markedly decreased in the NPC tissues compared with the adjacent normal tissue samples (Figure 3(d)). These data suggested that miR-320a acted as a target for lncRNA ILF3-AS1 in NPC.
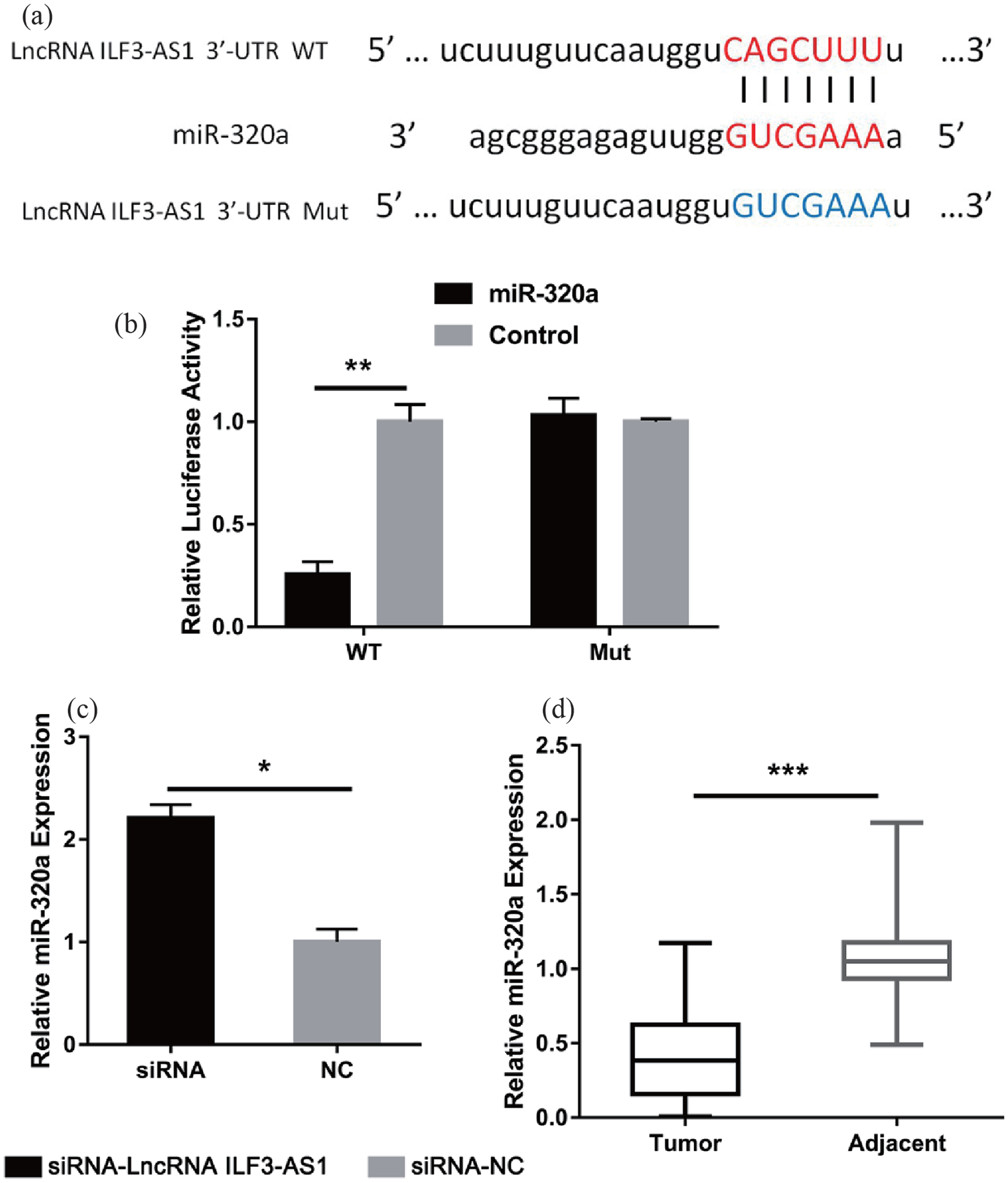
lncRNA ILF3-AS1 functioned as a sponge of miR-320a. (a) The sequences of lncRNA ILF3-AS1 with miR-320a, including wild type and mutant binding site. (b) Luciferase reporter assay verified the relationship within lncRNA ILF3-AS1 and miR-320a. (c) qRT-PCR showed the miR-320a expression level in CNE-1 cells transfected with siRNA-lncRNA ILF3-AS1 or siRNA-NC. (d) Expression of miR-320a level in 68 NPC tissues and compared normal tissues.
lncRNA ILF3-AS1 inhibited the function of BMI1 via miR-320a
In the above, we confirmed that lncRNA ILF3-AS1could work through miR-320a. Next, through bioinformatics analysis, we found that BMI1 might act as a target gene for miR-320a. The predicted binding sites are shown in Figure 4(a). Similarly, we mutated the 3′-UTR of the miR-320a-binding BMI1 and conducted dual-luciferase assay. The fluorescence intensity of the mutant group was not significantly different from that of the control group, while the wild type group showed a significant decrease (Figure 4(b)). This result indicated that miR-320a could directly combine with the 3′-UTR of BMI1. Next, we measured the mRNA expression levels of miR-320a and BMI1 in the experimental CNE-1 cells. Compared with the siRNA-NC group, the expression level of miR-320a in the siRNA-lncRNA ILF3-AS1 group was significantly increased while the expression of BMI1 was significantly reduced (Figure 4(c)). Next, we examined the protein expression of BMI1 in the experimental cells by the Western blot experiment. The BMI1 protein expression in the siRNA-lncRNA ILF3-AS1 group was significantly lower than that in the NC group (Figure 4(d) and (e)). These experiments demonstrated that BMI1 could act as a target molecule for miR-320a, whereas lncRNA ILF3-AS1 could promote the expression of downstream BMI1 by inhibiting the expression of miR-320a.
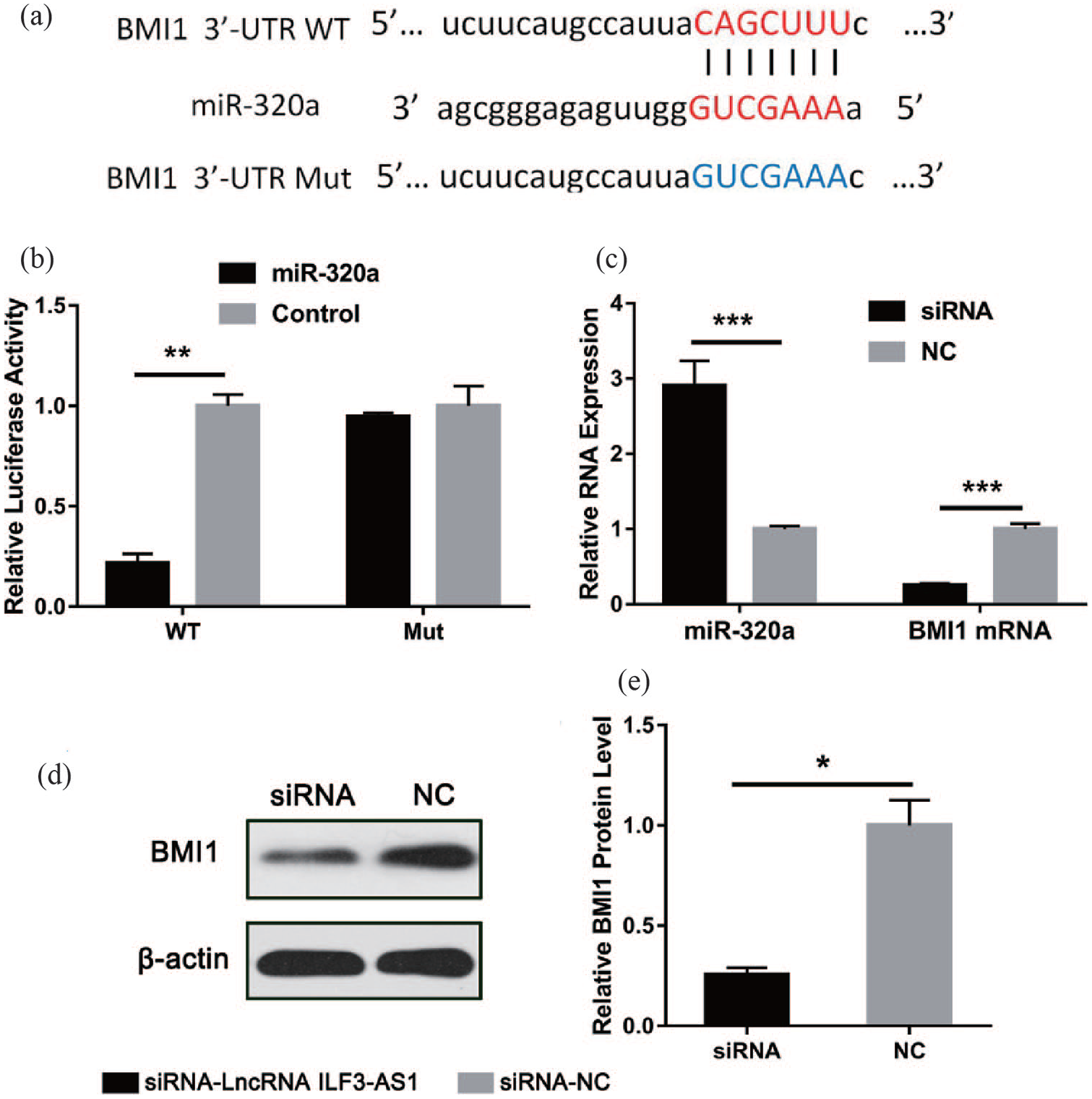
BMI1 was identified as the target protein of miR-320a. (a) The sequences of BMI1 mRNA 3′-UTR and miR-320a, including wild type and mutant. (b) Luciferase reporter assay indicated the relationship within BMI1 and miR-320a. (c) Expression of miR-320a and BMI1 mRNA in established CNE-1 cells. (d) and (e) Western blot assay indicated the BMI1 protein expression in established CNE-1 cells.
Discussion
NPC is a polygenic disease that occurs predominantly in southern China, Southeast Asia, and North Africa. 1 At present, through combined treatments—including surgery, radiotherapy, and chemotherapy—the survival rate of early stage patients has been improved. However, the overall survival rate of NPC has not been improved.18,19 Exploration of the underlying mechanism of NPC development and progression remains urgent. lncRNAs have been considered to play an important role in the genesis and development of tumors including NPC in recent years.5,20 lncRNA NKILA could suppress NPC carcinogenesis and metastasis through regulating the NF-κB pathway. 21 lncRNA FAM225A promotes NPC development and invasion by increasing the ITGB3 level via miR-590-3p/miR-1275. 22 lncRNA PVT1 regulates the growth of NPC cells by HIF-1α stability and KAT2A acetyltransferase activation, while lncRNA LOC284454 promotes NPC cell invasion and migration via the Rho/Rac signaling pathway.23,24 LINC01420 modulates NPC cell migration and invasion and indicates an unfavorable prognosis of NPC patients. 25
lncRNA ILF3-AS1 was found to act as a potential prognostic biomarker in cervical cancer and is associated with a danger of recurrence in colon cancer patients.12-14 It could promote proliferation and metastasis of melanoma via a positive feedback loop with ILF3, and regulates miR-200b/a/429 negatively.26,27 In the current study, we found that lncRNA ILF3-AS1 was significantly over-expressed in NPC tissues and cells. An elevated lncRNA ILF3-AS1 level was related to the advanced TNM stage and metastasis of NPC. These indicated that lncRNA ILF3-AS1 functions as an oncogene in NPC. To further explain the role of lncRNA ILF3-AS1 in NPC in vitro, we inhibited lncRNA ILF3-AS1 expression in CNE-1 cells using siRNA. The results of colony formation assay and EdU assay showed that the inhibition of lncRNA ILF3-AS1 reduced the cell proliferation of NPC. In addition, Transwell and Matrigel assays were employed to subscribe the cell invasion and migration abilities. The results indicated that lncRNA ILF3-AS1 could promote cell invasion and migration of NPC cells.
Endogenous competition accounts for a large part of the mechanism of action of lncRNAs. lncRNA can competitively bind to the targeted miRNA, thereby inhibiting its action and affecting the development of tumors. 28 Based on this mechanism, we found that miR-320a might function as a target miRNA of lncRNA ILF3-AS1. Through dual-luciferase experiments, we confirmed that lncRNA ILF3-AS1 could directly sponge miR-320a, affecting its expression. In vitro, an elevated level of miR-320a was found in the lncRNA ILF3-AS1 inhibited group. Furthermore, expression levels of miR-320a were markedly decreased in the NPC tissues compared with the adjacent normal tissue samples. These results suggested that miR-320a acted as a target for lncRNA ILF3-AS1 in NPC.
Next, we further analyzed (through bioinformatics) that BMI1 acted as a direct target gene for miR-320a. BMI1 was the first polycomb group (Pc G) family protein, which was discovered in 1991 to interact with C-myc to promote the formation of mouse B-cell lymphoma.29-31 The BMI1 gene is closely involved in the self-renewal of stem cells, and the occurrence, development, and prognosis of a series of malignant tumors. It has been found that the BMI1 gene is highly expressed in various malignant tumors such as leukemia, glioma, breast cancer, lung cancer, prostate cancer, etc., and is associated with tumor invasion, metastasis, and prognosis.31-35 In the present study, dual-Luciferase reporter assay system was used to verify the correlation of miR-320a and BMI1. We found that the fluorescence activity of the mutant group was obviously reduced comparing to the control, suggesting that BMI1 acted as a target gene for miR-320a. More intuitively, in CNE-1 cells that inhibited lncRNA ILF3-AS1 expression, miR-320a expression was significantly increased, while BMI1 mRNA and protein expression levels were evidently reduced. This series of experiments shows that lncRNA ILF3-AS1 can function through the miR-320a/BMI1 axis.
In summary, this study first found and verified the high-expression of lncRNA ILF3-AS1 in NPC tissues and cells. Moreover, increased lncRNA ILF3-AS1 expression suggested advanced TNM stage and metastasis of NPC. lncRNA ILF3-AS1 could promote the proliferation, invasion, and migration of NPC cells via sponging miR-320a to accelerate the expression of BMI1. These findings might provide a possible point for the future diagnosis, prognosis prediction, and bio-therapy for NPC.
Footnotes
Author contributions
XY designed the study and performed the experiments; FG and FL collected the data; XY and FG analyzed the data; and XY prepared the manuscript. All authors read and approved the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
