Abstract
Objective:
NTRK mutations and clinicopathological factors in patients with lung cancer in northeast China were analyzed by next-generation sequencing (NGS), and references were provided for patients with NTRK mutations undergoing targeted therapy in northeast China.
Methods:
A total of 224 specimens in 173 patients with lung cancer were collected. This included 51 patients with matched tissue and whole blood samples,133 tissue samples, 84 whole blood samples, and 7 pleural effusion samples. NGS (520 genes) was used to detected NTRK mutations and clinicopathologic factors.
Results:
NTRK mutation was detected in eight patients (8/173, 4.6%), including four NTRK missense mutations (4/173, 2.3%), two NTRK fusion gene mutations (2/173, 1.2%), and two NTRK copy number deletions (2/173, 1.2%). Among the eight patients with NTRK mutations, four were associated with lung cancer driver gene mutations (3/4 EGFR, 1/4ALK); NTRK in two patients was inconsistent in tissue and paired whole blood testing; NTRK missense mutation was detected in one patient, and NTRK copy number deletion was detected in the other; and NTRK wild type was detected in two patients. There was no correlation between NTRK mutation and clinicopathologic factors (including gender, age, pathological type, smoking status, metastasis site).
Conclusion:
NTRK mutation was only 4.6%, effective fusion gene mutation was 1.2%, and common driver gene mutation in lung cancer was evident in 50% of patients. The results of NTRK were inconsistent with matched tissues and whole blood. Therefore, patients with NTRK mutation should use a variety of specimen types and large target area sequencing (panel) analysis method to provide individualized treatment.
Introduction
A cancer-driving gene, tropomyosin receptor kinase (TRK)—a gene fusion protein of non-small cell lung cancer—was found at the University of Colorado in Denver and the Dana-Farber Cancer Institute in Boston. TRK protein was encoded by the neurotrophic tyrosine receptor kinase.1-2 The NTRK genes (NTRK1, NTRK2, NTRK3) encode TRKA, TRKB and TRKC, respectively. The TRK gene is normally only active in early embryonic development; it is mainly expressed in nerve cells and is mostly silent in the cells of mature individuals. NTRK may be reactivated when accidentally fused with some ligand genes, activating downstream signaling pathways and leading to cancer. 3
On November 27, 2018, the FDA approved Vitrakvi® (larotrectinib), a pan-cancer-targeted drug developed by Bayer and Loxo Oncology, to treat NTRK gene fusion in adults and children with locally advanced or metastatic solid tumors. In February 2018, the New England Journal of Medicine published three clinical studies on safety and efficacy. 4 For patients aged 4 months to 76 years, the overall response rate (ORR) for 17 different cancers was 75%. According to the latest data reported at the ESMO annual meeting, in 55 patients measured by RECIST criteria with TRK fusion, the ORR was 80%.
Drugs targeting NTRK had indicated good clinical response and may become new targeted drugs in the future. However, the mutation frequency, mutation type and dominant population of NTRK in Chinese patients with lung cancer have not been reported. Due to the different gene expression profiles between eastern and western populations, 5 the purpose of this study was to detect NTRK gene mutations in lung cancer patients in northeast China by next-generation sequencing (NGS). Combined with the analysis of clinicopathologic factors, this study provides data for larotrectinib in lung cancer patients with an NTRK fusion gene mutation in northeast China.
Materials and methods
Research subjects
A retrospective analysis of 173 lung cancer patients (224 specimens) were treated at the Jilin Cancer Hospital from June 2018 to June 2019, which included 51 patients with matched tissue and whole blood samples. The median age was 58 years (range 34–79), and the mean age was 53 years; the specimens were tissue (n=133), whole blood (n=84), and pleural effusion (n=7); the proportions of women and non-smokers were 53.75% and 52.02%, respectively. The majority of patients were diagnosed with adenocarcinoma (89.59%), stage IV (98.26%), with adenosquamous carcinoma (5.78%).
Experimental methods
Reagents and instruments
Nucleic acid extraction reagents and quantitative instruments
DNA was extracted from Paraffin nucleic acid DNA extraction kit (AmoyDx, China) and ctDNA extraction kit (QIAGEN, Germany), and the concentration of DNA was quantified by Nanodrop2000C and Qubit3.0 Fluorometer (Thermo, USA).
NGS analysis
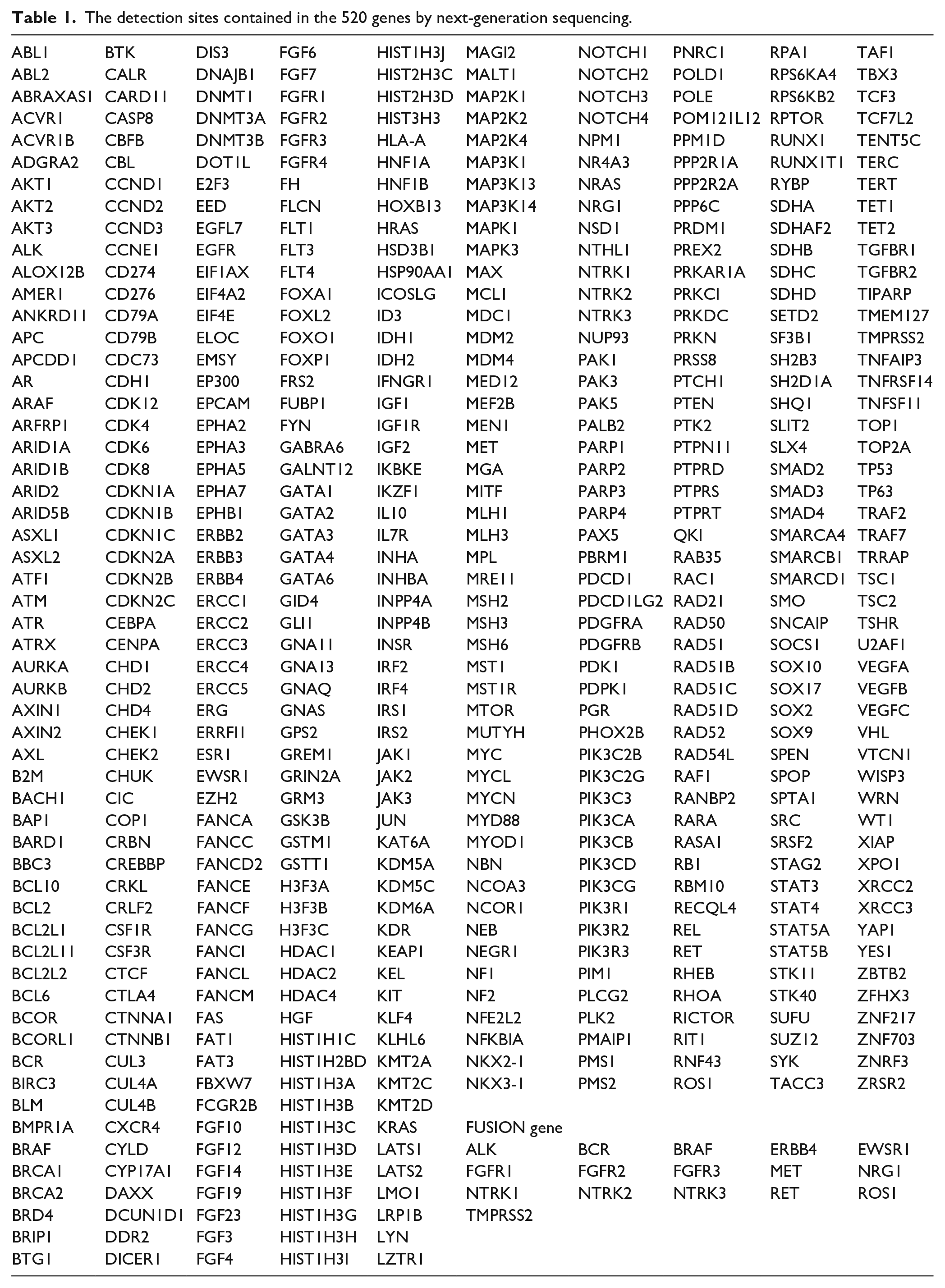
The NGS mutational analysis was performed by target region probe capture technology and the Illumina sequencing platform. This panel allowed for the simultaneous evaluation of the mutational status of 520 genes (Table 1). Instrument platform: Illumina Novaseq 6000/Nextseq 500 Sequencing Platform; Reagents: Burning Rock Dx Oncoscreen plus TM kit. The tissue sequencing depth was 2000×, and the blood sequencing depth was 10,000×. Copy number variation (CNV) was detected by in-house analysis scripts based on the depth of coverage data of the capture intervals. Coverage data was firstly corrected against sequencing bias resulting from Guanine and Cytosine content and probe design. The average coverage of all captured regions was utilized to normalize the coverage of different samples to comparable scales. The copy number was calculated based on the ratio between the depth of coverage in tumor samples and the average coverage of an adequate number (n>50) of samples without CNV as a reference at each capture interval. Gene CNV is called if the coverage data of the gene region was quantitatively and statistically significantly different from its reference control. Two criteria should be met for CNV: (a) the coverage of more than 60% capture intervals of the genes should be significantly different from the reference. The significance was evaluated by z-test comparing the coverage of each capture interval to the mean coverage of the interval in all control samples (P<5e-3 for hotspot genes and P<1e-3 for others); (b) the copy number should reach the minimum threshold of gain and loss (copy number gain: CN>2.25 for hotspot genes and CN>2.5 for others; copy number loss: CN<1.75 for hotspot genes and CN<1.5 for others). When the detection sequence does not match the reference sequence by one or more bases, it is considered that a single nucleotide mutation (SNV) occurs at this position. When the detection sequence and the reference genome are two or more and overlap the genes, it is considered a chromosome translocation (FUSION). For hot spot mutations, when the mutation abundance is ⩾2%; For non-hot spot mutations, the threshold of mutation abundance is ⩾5%, it is considered as SNV.
The detection sites contained in the 520 genes by next-generation sequencing.
Statistical methods
Statistical analysis was performed by SPSS 18.0. Fisher’s exact probability test was used to compare the two groups with theoretical frequency <5. P<0.05 was considered statistically significant.
Results
NTRK mutation in 173 patients
NTRK mutations were detected in 8 of the 173 patients (4.6%), including NTRK missense mutations (4/173, 2.3%), NTRK fusion gene mutations (2/173, 1.2%), and NTRK copy number deletions (2/173, 1.2%).
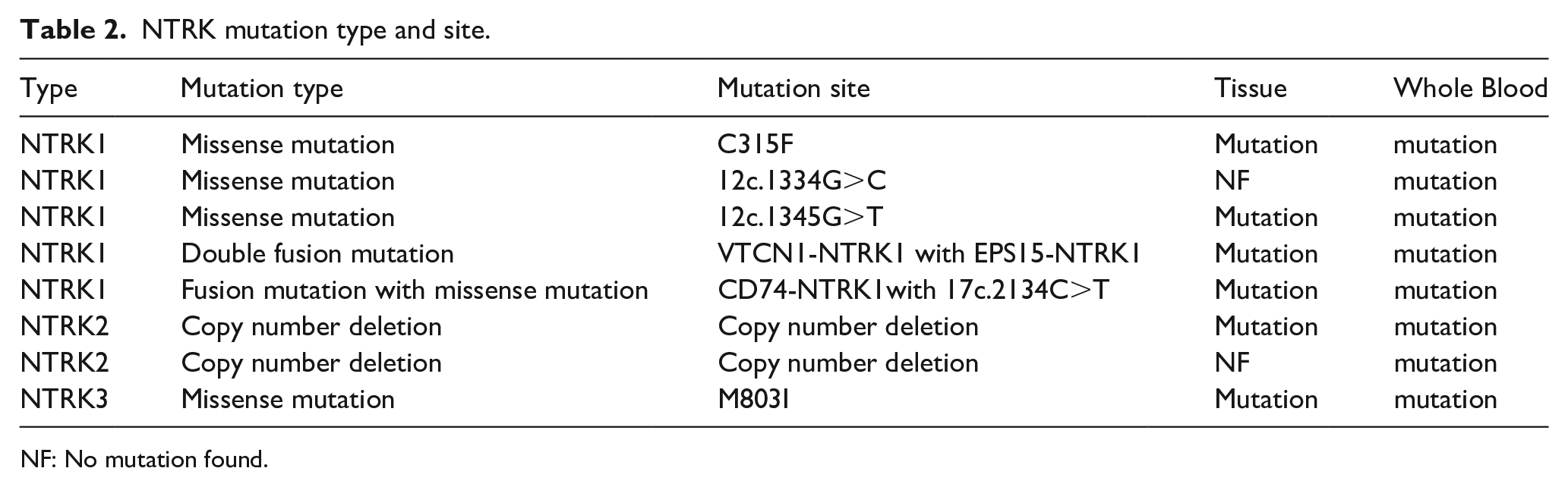
Four cases of NTRK missense mutations were: NTRK3 M803I, NTRK1 C315F, NTRK1 12 c.1334G>C, and NTRK1 12 c.1345G>T. Two cases of NTRK fusion genes were: CD74-NTRK1 (C6:N12) gene rearrangement accompanied by NTRK1 missense mutation 17 c.2134C>T p.Arg712Trp, and 1VTCN1-NTRK1 (Vintergenic:N10) gene rearrangement accompanied by EPS15-NTRK1 (E21:N10) gene rearrangement. NTRK copy number deletion: two cases were missing NTRK2 copy number (Table 2).
NTRK mutation type and site.
NF: No mutation found.
Four patients with NTRK mutations had the lung-cancer-driver gene
The 2019 National Comprehensive Cancer Network (NCCN) guidelines recommend advanced non-small cell lung cancer (NSCLC) patient should detect EGFR, ALK, ROS1, KRAS, BRAF, HER2, RET, MET. In this study, eight patients with NTRK mutations had one of lung-cancer-driver genes: EGFR(3/4), ALK(1/4), ROS1(0/4), KRAS(0/4), BRAF(0/4), HER2(0/4), RET(0/4), or MET(0/4).
Differences in NTRK mutation between tissue and paired whole blood
Tissue and paired whole blood NTRK mutations were inconsistent in two patients; one patient had an NTRK missense mutation, and one had NTRK copy number deletion. Two patients had NTRK wild type.
Tumor mutation burden in NTRK mutation patients
The median tumor mutation burden (TMB) in the tissue was 11.1 mutations/Mb (range 4.8–15.1) in NTRK mutations samples compared with median TMB of the blood was 6.3 mutations/Mb (range 1.6–15.9) in NTRK mutations samples.
Correlation analysis between NTRK mutation status and clinicopathologic factors
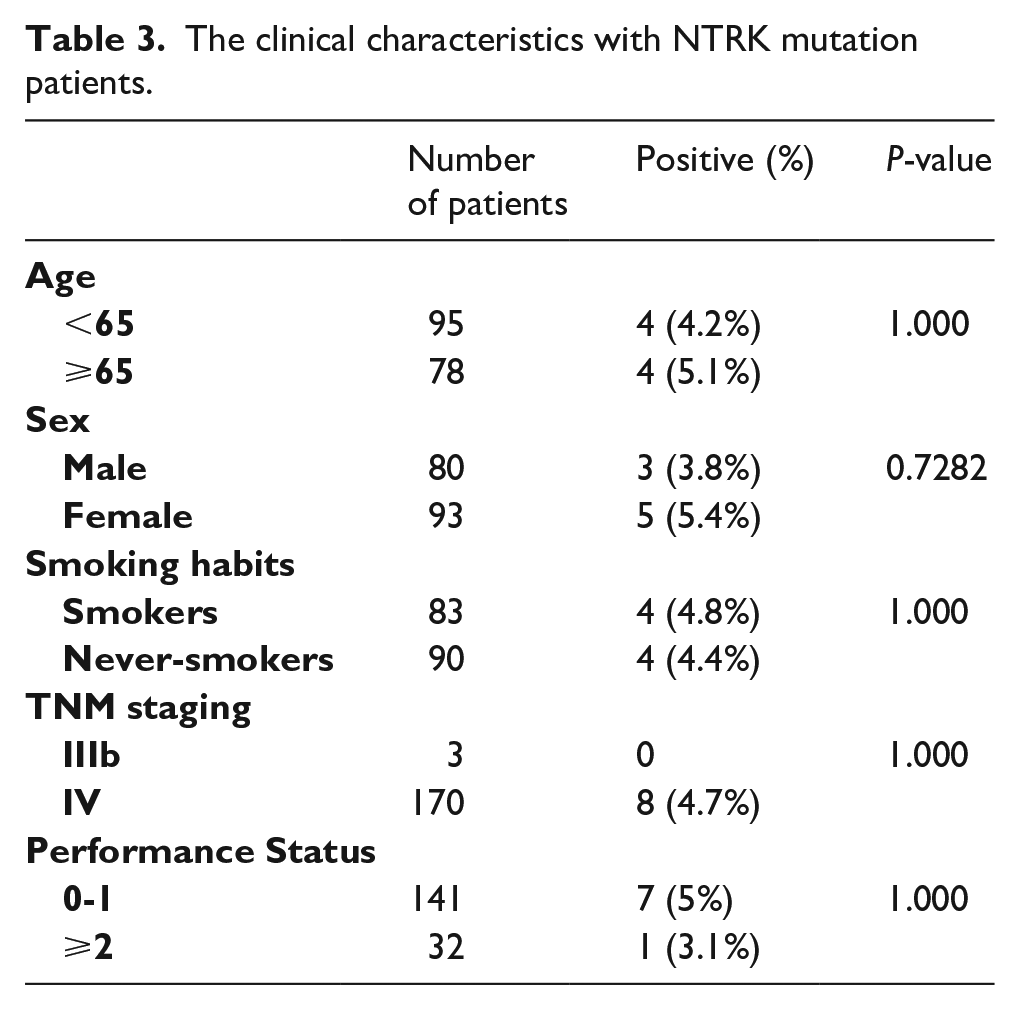
There was no significant correlation between NTRK mutation status and clinicopathologic factors, such as age and stage (P > 0.05) (Table 3).
The clinical characteristics with NTRK mutation patients.
Discussion
The development of nucleic acid (DNA and RNA) sequencing technology has promoted the understanding of the molecular characteristics of genetic diseases. With the characteristic of highly efficient, sensitivity and high-throughput, NGS has become an important process and provides a molecular basis for clinical molecular diagnostic and precision medicine.6-7 The NTRK gene can only be found in NGS large target region sequencing (panel) with the incidence of approximately 0.2%–1%.1,2 Currently, there are still some limitations in clinical testing: fluorescence in-situ hybridization and immunohistochemistry have not been certified as the kit products and the types tested must be formalin-fixed paraffin-embedded samples. Therefore, this study retrospectively analyzed the NTRK mutations in the lung cancer samples detected by the NGS 520 genes.
In the updated 2019 NCCN guidelines, the NTRK gene was used as the ninth driver gene for NSCLC, and larotrectinib is recommended as the first-line treatment for patients with NTRK gene fusion-positive metastatic NSCLC. The NTRK fusion gene has become the new “star” target for the treatment of NSCLC.
At present, the NTRK mutation in lung cancer has mainly been reported in European countries and Japan, while there are relatively few studies on NTRK in China. In October 2013, Vaishnavi et al. 8 investigated the fusion gene NTRK1 (as the first reported gene) in lung cancer patients in Nature Medicine. They used NGS to detect 36 patients with lung adenocarcinoma and found NTRK1 gene fusion (MPRIP-NTRK1 and CD74-NTRK1) in two patients. In November 2017, the European Society for Medical Oncology Asia Conference reported the fusion of NTRK in the Japanese NSCLC population. Only one (0.04%) ETC6-NTRK3 fusion mutation was found in 2668 patients; NTRK1 and NTRK2 were not found. 9 In May 2017, Nature Medicine published a large-scale, prospective, clinical sequencing study using the MSK-IMPACT method at the Memorial Sloan Kettering Cancer Research Center. They performed more than 300 detailed tumor types in more than 10,000 patients with advanced cancer by NGS, including 10 cases NTRK1 fusion mutation, lung adenocarcinoma accounted for 0.13% (2/1563), and co-mutation with P2RY8 and VANGL2, respectively. 10 In our study, the NTRK fusion gene mutation was only 1.2% with CD74, VTCN1, and EPS15; furthermore, one patient had a co-mutation with two fusion genes. In subsequent clinical studies, we will closely follow-up this patient, providing data support for individualized treatment.
Changes in the NTRK signaling pathway have been the cause of many tumors. NTRK mutations include protein overexpression and single nucleotide changes besides the most specific gene fusion. In addition to the statistical NTRK fusion mutations, two other types of mutations were analyzed, including four cases of missense mutations and two cases of copy number deletions. The current clinical significance of NTRK missense mutations and copy number deletions is not very clear; our study results analyze the mutations and clinicopathologic factors and provide data for future research.
A total of 51 tissue and whole blood paired samples were tested in this study; the NTRK results were found to be different. Although no effective NTRK fusion gene locus was detected in the differential samples, this suggests that blood will have its own unique genotype. In addition, this study counted TMB in eight patients with NTRK mutations; the TMB of all patients showed a higher trend compared with the NTRK wild type. In the next stage of research, we will closely monitor the treatment of these patients and provide data for individualized treatment.
This study has some limitations; for example, fewer specimens, fewer indicators for the analysis of mutation influencing factors, and shorter follow-up time. Discovering the most important factors of NTRK mutations will require more samples and more detailed analysis.
In conclusion, when using NTRK targeted drugs, patients should select a variety of sample types and large target area sequencing (panel) analysis methods for individual treatment.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by Scientific Research Project of Jilin Provincial Health and Family Planning Commission (grant numbers2017J023 and 2018Q007); and Science and Technology Agency of Jilin Provincial Project (grant numbers 20200201518JC).
