Abstract
Background:
High mobility group A proteins are involved in chromatin remodeling, thereby influencing multiple fundamental biological processes. HMGA2 has been linked to oncogenic traits among a variety of malignancies.
Objective:
To determine the prognostic implications of subcellular distribution patterns of HMGA2 in breast cancer.
Methods:
Nuclear and cytoplasmic HMGA2 was evaluated in 342 breast cancer specimens and matched with clinico-pathological parameters.
Results:
Overall and cytoplasmic, but not nuclear, levels of HMGA2 correlated with better survival prognoses in our collective (hazard ratio (HR) 0.34, P = 0.001 and HR 0.34, P < 0.001, respectively). The protective effect of cytoplasmic HMGA2 persisted in the Luminal A and triple negative breast cancer subgroups. Evaluating Luminal A and B subgroups jointly, only cytoplasmic, but not overall or nuclear HMGA2 levels were associated with better survival (HR 0.42, 95% confidence interval 0.21, 0.86, P = 0.017), irrespective of tumor size and node status. The addition of HMGA2 overall and cytoplasmic scores strengthened the prognostic selectivity in a model of conventional breast cancer risk factors. No predictive significance with regard to endocrine or chemoendocrine therapies was observed.
Conclusion:
Unexpectedly, we found a favorable survival probability upon overall levels of HMGA2 in our breast cancer collective, which was predominantly determined by the presence of HMGA2 in the cytoplasm.
Introduction
The high mobility group (HMG) proteins belong to a family of nuclear, non-histone architectural transcription factors: HMGA, HMGB, and HMGN. High mobility group A (HMGA) genes encode for HMGA1a-c and HMGA2, which share high sequence homologies. Structurally, HMGA2 proteins contain three domains, which bind to AT-rich DNA strands, and an acidic C-terminal tail. 1 Although HMGAs do not display intrinsic transcriptional activities, they are involved in chromatin remodeling and gene transcription by assembling or enhancing protein–protein and protein-DNA interactions of different transcription factors.2,3 Thus, HMGA activity influences multiple fundamental biological processes, such as cell differentiation, chromatin remodeling, DNA replication and repair, cell cycle progress and senescence, growth, and death. 4 HMGA proteins are highly expressed in human tissues during embryogenesis, but only sparsely in adult differentiated and healthy tissues.5,6 Excessive growth of benign tumors or increased adipogenesis in lipomas were observed upon rearrangements of HMGA2 genes,7-9 and genetic HMGA2 knockout studies in mice led to growth restriction and “pygmy” phenotypes, thereby indicating a critical role of HMGA2 genes in the regulation of growth. 10 An overexpression of HMGA2 was attributed to oncogenic traits and described in malignancies of different entities.11-20 Expression of HMGA2 in breast cancer has been discussed as a prognostic or even predictive biomarker upon expression in primary tumors or circulating tumor cells.20-23
Until very recently, physiologic and oncogenic properties of HMGA2 proteins were mainly linked to functions of HMGA2 in the nucleus. However, it has been recognized long since that HMGA2 is at least transiently present also in extranuclear cell compartments, 24 and functional analyses of extranuclear HMGAs have been reported progressively. Méndez et al. 25 could show that the HMGA1 is enriched in the cytoplasm of primary triple negative breast cancer (TNBC) at the invasive tumor front and in metastatic lesions, and raised the hypothesis that cytoplasmic HMGA1 could serve as a surrogate marker for oncogenic HMGA1 secretion into the tumor surrounding. Hawsawi et al. 26 observed a shift from predominantly cytoplasmic HMGA2 staining in low-grade prostate tumors to nuclear staining in high-grade tumors and metastatic disease. In placental tissues, translocation of HMGA1 from the nucleus to the cytoplasm was accompanied by a reduced migration capacity in trophoblastic cells of a preeclampsia model. 27 Translocation of HMGA2 from the cell surface to the nucleus caused oncogenic cell transformation on skin carcinomas. 28
To further explore the possibility of diverse prognostic implications of HMGA2 in dependence on specific subcellular localizations, we immunohistochemically evaluated the staining patterns of HMGA2 in a well-characterized cohort of 342 breast cancer patients. Surprisingly, we found a favorable survival probability associated with high levels of HMGA2 in our collective, which was predominantly determined by the presence of HMGA2 in the cytoplasm.
Materials and methods
Tissue microarray
An established tissue microarray (TMA) comprising two representative tumor areas of 400 samples from breast cancer patients has been previously described. 29 Patients were treated at the University Hospital of Kiel, Germany, between 1997 and 2008. Formalin-fixed and paraffin-embedded primary tumor specimens were retrieved from the local Department of Pathology, and patients’ reports were retrospectively reviewed for clinico-pathological parameters and follow-up data with approval of the local ethics committee (Christian-Albrechts-University of Kiel, Faculty of Medicine: #D450/10). Breast cancer specimens were classified in accordance to the World Health Organization and the TNM (tumor, node, metastases) criteria. Estrogen (ER) and progesterone receptor (PR) status was determined according to the Remmele Score (0–12, positive >2) by immunohistochemistry. HER2 (human epidermal growth factor receptor 2) was determined by immunochemistry and valued with a score of 0–3+ and in situ hybridization in case of 2+. Tumor samples with 3+ scores or positive HER2 amplification were regarded as positive. Specimens were allocated to intrinsic breast cancer subtypes on the basis of the receptor status, if available. HER2-negative, ER-positive carcinomas were further divided into Luminal A (Ki67 <14%/G1) and Luminal B (Ki67 >14%/G3) subtypes. Breast cancer treatment included primary tumor resection, adjuvant radiation, endocrine (tamoxifen or aromatase inhibitors) or chemotherapeutic (CMF or anthracycline-taxane-based) therapies. Of note, 48 patients from the HER2-positive subgroups received either chemotherapy alone (n=12) or chemoendocrine therapy (n=19), but no HER2-targeted therapies were routinely administered.
Immunohistochemistry
Immunohistochemistry was performed utilizing a monoclonal rabbit antibody directed against HMGA2 (dilution 1:200, D1A7, Cell Signaling, Leiden, Netherlands, [#8179]) using the autostainer Bond™ Max System (Leica Microsystems GmbH, Wetzlar, Germany). Pretreatment with ER2 (Leica Biosystems) for 20 minutes. The Bond™ Polymer Refine Detection Kit was used for the visualization of immunoreactions (DS 9800; brown labelling; Novocastra; Leica Microsystems GmbH, Wetzlar, Germany [#DS9800]).
HMGA2 scoring system
Without knowledge of the corresponding clinical data, staining intensities of HMGA2 in the nucleus and cytoplasm were examined by experienced pathologists (FV, SK, and CR). As uniform staining patterns for HMGA2 were observed among the majority of breast cancer specimens, a semi quantitative scoring system from absent to strong staining was chosen to discriminate different HMGA2 levels. 30 In cases of divergent staining patterns, the cell population with the highest staining intensity was chosen for calculation. Cells with no HMGA2 staining were scored “0.” A weak staining (<10%) was scored “1,” an intermediate staining (11%–50%) was scored “2,” and a strong staining (>50%) was scored “3.” Based on this scoring system, a score of 0–1 was regarded as low HMGA2 expression in the nucleus and cytoplasm, and a score of 2–3 was regarded as high HMGA2 expression. To calculate an overall cellular staining intensity of HMGA2, scores of nuclear and cytoplasmic staining were summed up to an overall score of 0–6. Based on median HMGA2 levels in our collective, overall scores of 0–3 were regarded as low and scores of 4–6 as high overall HMGA2 levels.
Statistics
The statistical software package IBM SPSS Statistics 2013 31 was used to calculate descriptive statistics and to perform analyses, which were all based upon dichotomous HMGA2 values. For correlations between HMGA2 scores (overall, nuclear, and cytoplasmic), intrinsic subtypes, and clinical parameters, we applied Spearman’s rank correlation coefficient. The Kaplan–Meier curves were calculated for survival data and the log-rank test was applied to unravel possible differences in survival times between low and high HMGA2 levels. Hazard ratios (HRs) were determined using single Cox regression analyses for each HMGA2 score separately. To investigate whether the prognostic value of HMGA2 for breast-cancer-associated deaths exceeds the prognostic value of other established prognostic factors we used a two-step hierarchical Cox regression analysis for each HMGA2 value separately. That is, after entering established prognostic factors in the prediction model, we added HMGA2 values as an additional predictor variable. Changes in model fit were determined with χ2 tests. In all analyses, P values < 0.05 were considered as statistically significant.
Results
Study population
The mean age of 342 primary breast cancer patients in our collective was 58.3 years. Most patients presented with early tumor stages and node-negative disease. Of the 342 patients, 118 specimens classified as Luminal A carcinomas (34.5%), 138 as Luminal B (40.4%), 31 as Luminal HER2-positive (9.1%), 17 as HER2-positive (5.0%), and 38 as TNBC (11.1%). Based on the patients’ medical reports, 144 of the 256 patients (56.3%) of the Luminal A and B subgroups received an adjuvant state of the art endocrine therapy alone, and 75 patients (29.3%) received endocrine therapy and chemotherapy. The mean follow-up was 48 months. A total of 57 breast-cancer-associated deaths were documented (16.7% of all patients). Detailed characteristics of the cohort are shown in Table 1.
Cohort table of primary tumor specimens from a total of 342 breast cancer patients.
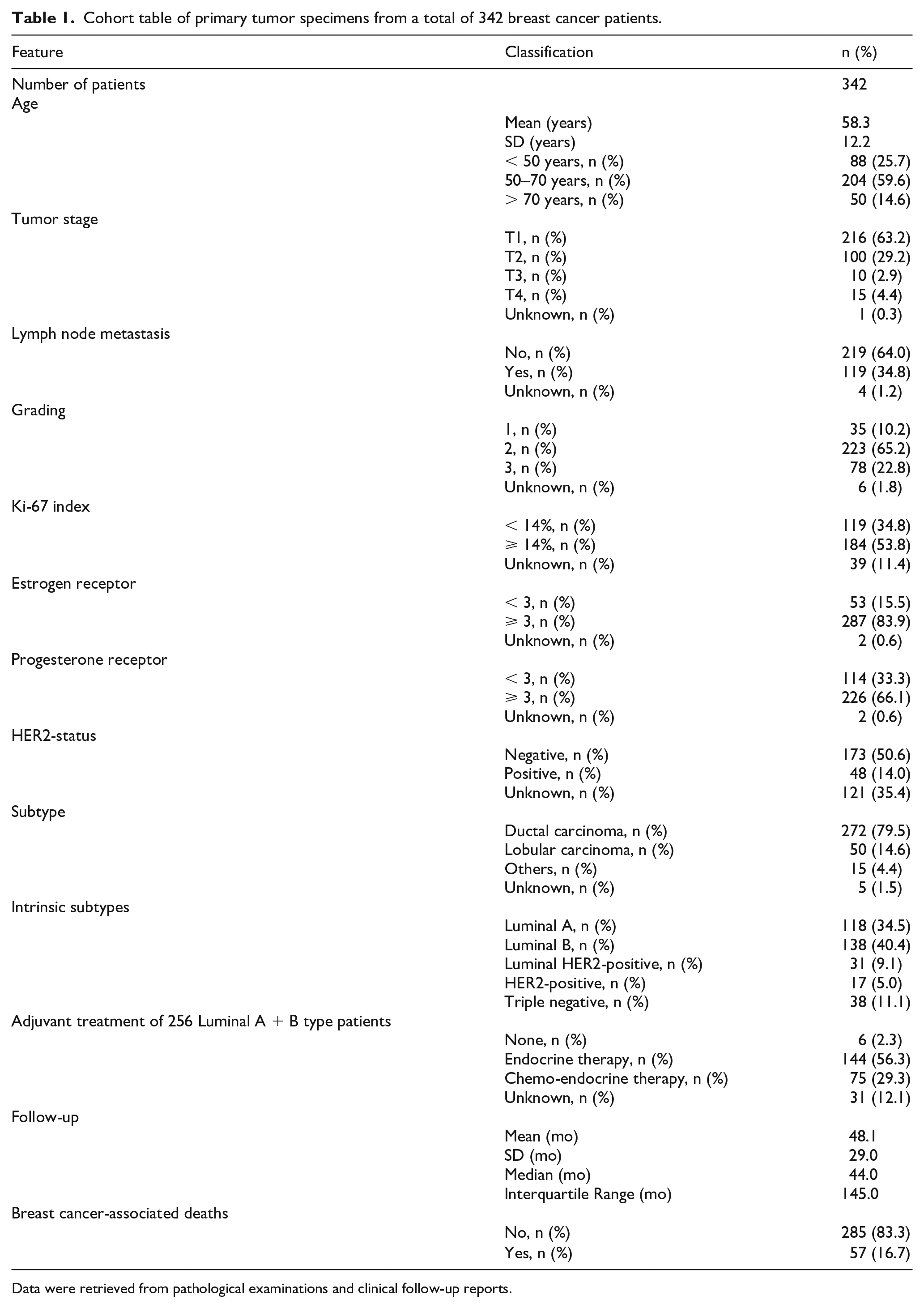

Data were retrieved from pathological examinations and clinical follow-up reports.
Subcellular expression of HMGA2
A total of 174 breast cancer specimens showed low overall HMGA2 levels (50.9%), whereas 168 showed high levels (49.1%). In the nucleus, low HMGA2 levels were found in 143 cases (41.8%), and high levels in 199 cases (58.2%). In the cytoplasm, low HMGA2 levels were found in 136 cases (39.8%), and high levels in 206 cases (60.2%). A weak positive correlation was observed between levels of HMGA2 in the nucleus and cytoplasm (r = 0.32, P < 0.001). Representative HMGA2 staining is shown in Figure 1.
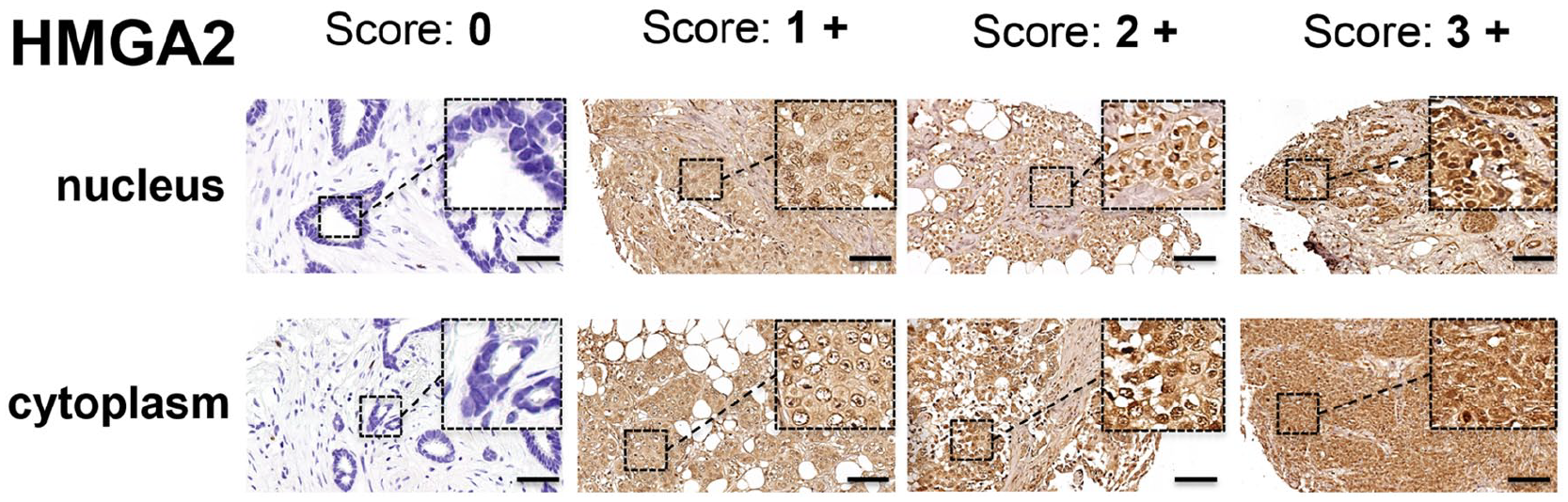
Representative staining for nuclear and cytoplasmic HMGA2 levels (score 0–3) in primary tumor specimens from the breast cancer tissue micro array. The addition of nuclear and cytoplasmic HMGA2 scores resulted in the overall score for each tumor specimen (score 0–6). The black scale bars indicate 100 µm. The dashed tumor area in each picture is presented in 7.5-fold magnification (400x).
Correlation of HMGA2 levels with breast cancer risk factors and intrinsic subtypes
The overall score of HMGA2 inversely correlated weakly with the HER2-positive subtype (r = −0.12, P < 0.05), and positively correlated with the progesterone receptor status (r = 0.19, P < 0.001) and the Ki-67 index (r = 0.11, P < 0.05) (Figure 2). No statistically significant associations with either subtypes or risk factors were observed for nuclear HMGA2 levels, whereas the cytoplasmic HMGA2 levels positively correlated with the progesterone receptor status (r = 0.12, P < 0.05) and Ki-67 index (r = 0.16, P < 0.01) (Figure 2).
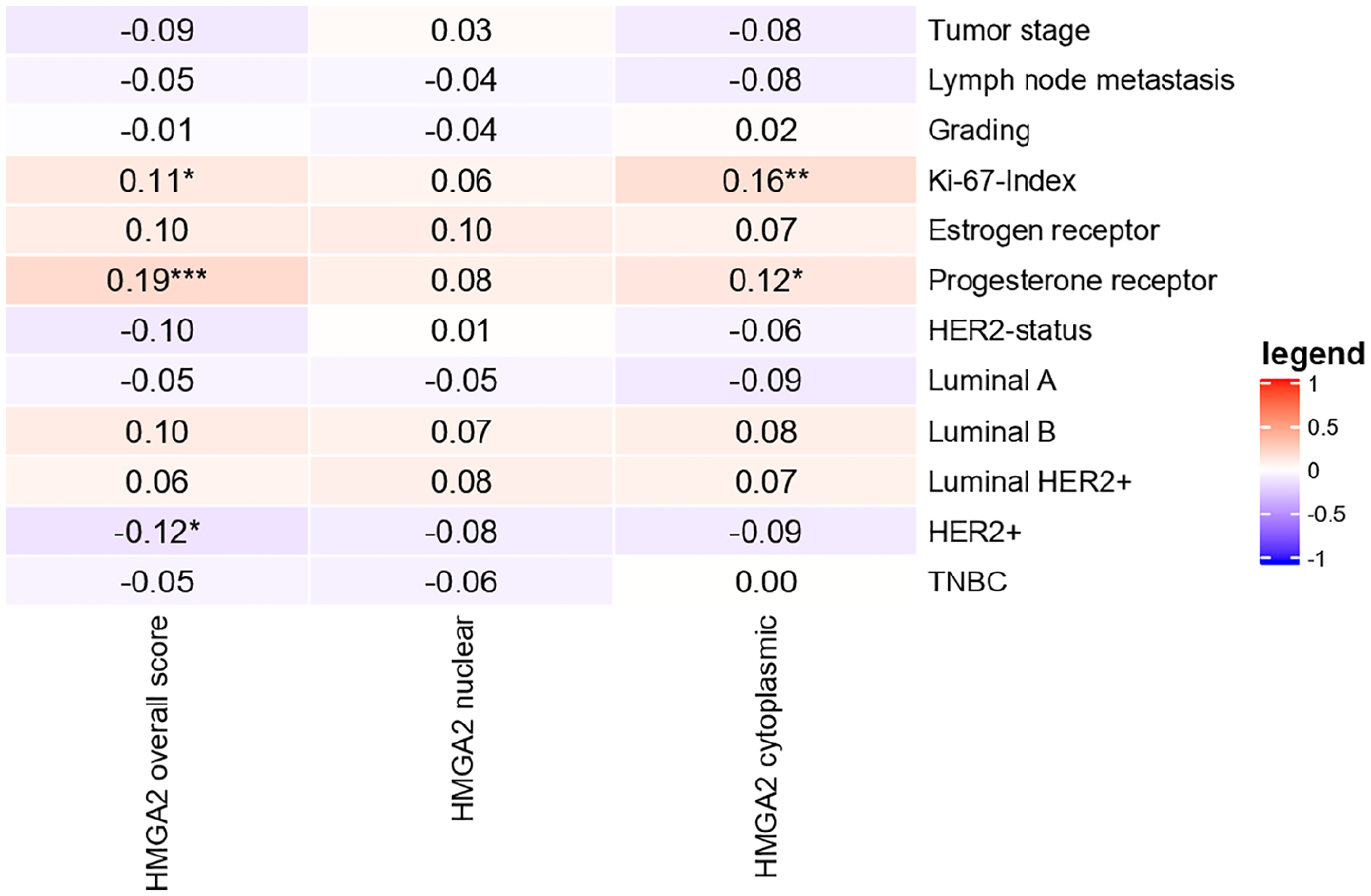
Expression patterns of HMGA2 divided by subcellular localization and correlation with breast cancer pathological features and intrinsic subtypes. Correlation coefficients r for positive and negative correlations (0–0.16).
3.4 HMGA2 levels and breast cancer overall survival
The impact of subcellular HMGA2 distribution on the survival probability in our patients’ collective without and with additional stratification by different intrinsic breast cancer subtypes was determined. When our collective was dichotomized depending on overall HMGA2 levels, we observed a highly significant survival benefit of high HMGA2 (HR 0.34; 95% confidence interval (CI) 0.18, 0.64; P = 0.001) (Figure 3(b)). The nuclear presence of HMGA2 did not impact the survival probability, whereas the cytoplasmic HMGA2 presence led to a higher survival probability (HR 0.34; 95% CI 0.19, 0.60; P < 0.001) (Figure 3(b)). Aligning HMGA2 staining intensities with different intrinsic breast cancer subtypes (Figure 3(d)) it became evident that overall and nuclear HMGA2 levels were not significantly associated with survival probability within any of the different intrinsic subtype groups, whereas cytoplasmic HMGA2 was associated with a better survival in the Luminal A and the TNBC type subgroups (HR 0.21; 95% CI 0.05, 0.97; and HR 0.27; 95% CI 0.08, 0.88—both P < 0.05, respectively).
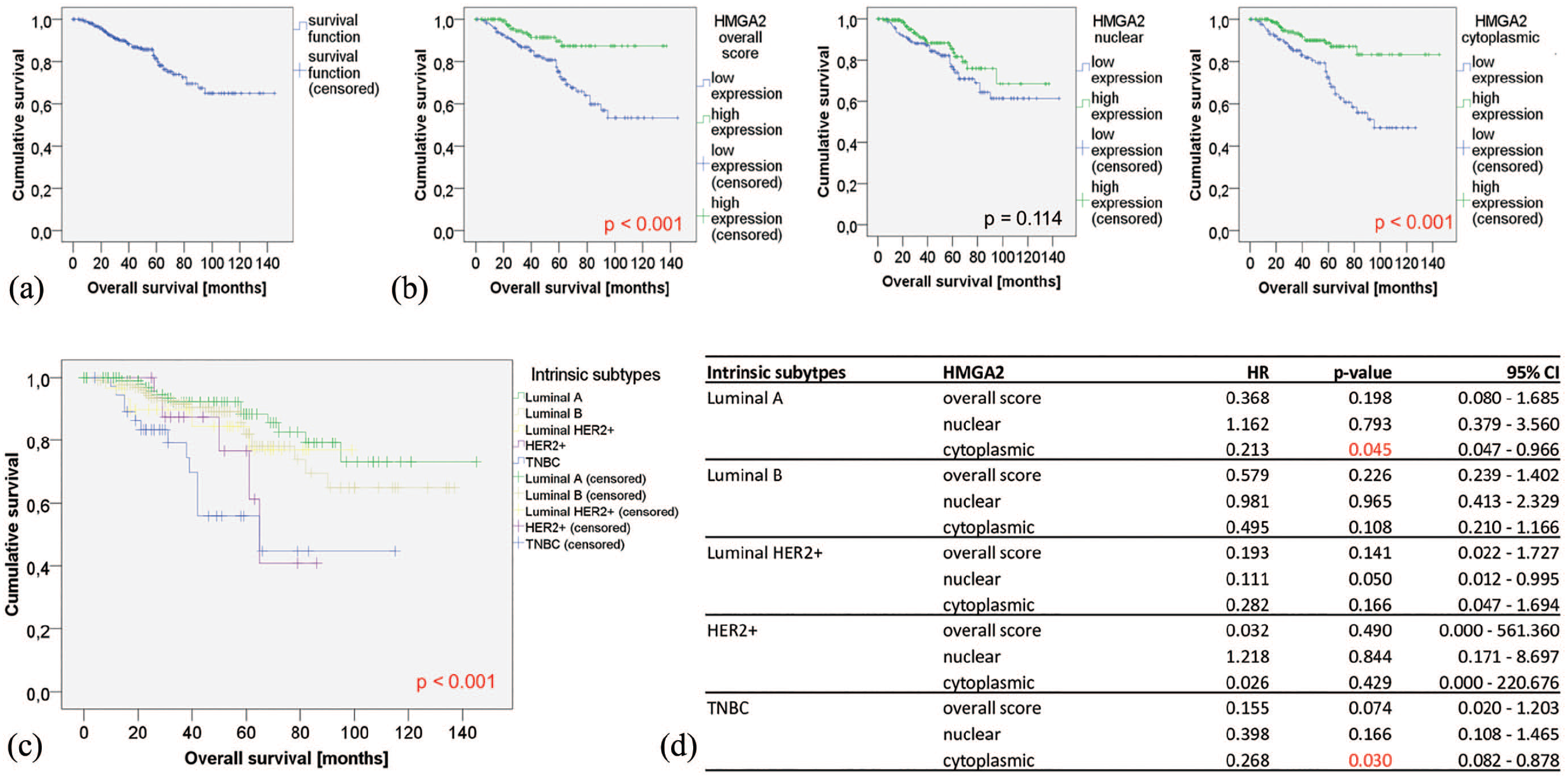
(a)Combined survival analyses (OS) in 342 breast cancer patients in our collective using Kaplan–Meier estimates. (b) OS in the breast cancer collective divided by subcellular levels of HMGA2: Overall and cytoplasmic, but not nuclear, HMGA2 correlated with better survival. Green curves: high HMGA2 scores. Blue curves: low HMGA2 scores. Vertical bars indicate censored patients. (c) Kaplan–Meier curves for all five intrinsic subtypes in our collective of 342 breast cancer patients. Patients of the HER2-positive subgroups did not receive HER2-targeted therapies routinely. (d) Cox regression models showing hazard ratios (HR) of subcellular HMGA2 distribution divided by intrinsic subtypes. Only in Luminal A and TNBC subtypes, high cytoplasmic HMGA2 scores led to better survival prognoses.
HMGA2 levels in Luminal intrinsic subgroups
Next, subgroup analyses were restricted to 256 Luminal A and B subtype patients. Here, neither the overall nor the nuclear HMGA2 levels showed associations with survival, while cytoplasmic HMGA2 was associated with a better overall survival (OS) (HR 0.42; 95% CI 0.21, 0.86; P = 0.017). To further exclude clinically relevant biases, these analyses were additionally restricted to 240 patients with early tumor stages (T1 and T2). Within this subgroup, high overall and cytoplasmic levels were associated with a favorable survival (HR 0.38; 95% CI 0.15, 0.96; P < 0.05; and HR 0.28; 95% CI 0.12, 0.67; P < 0.01, respectively, (not shown)), whereas no prognostic associations were found for nuclear HMGA2 levels. When 174 patients from the Luminal subgroups with a negative axillary node status were evaluated separately, again, only cytoplasmic by trend HMGA2 impacted the survival probability in our collective (HR 0.18; 95% CI 0.02, 1.06; P = 0.057 (not shown)).
HMGA2 levels and prediction of therapy response in Luminal subtypes
In 256 patients with Luminal A and B breast cancer subtypes high overall, nuclear, or cytoplasmic levels of HMGA2 did not statistically significantly impact the survival probability of patients treated with either endocrine therapy alone (n = 144; HR 1.11; 95% CI 0.30, 4.13; P = 0.872) or with chemotherapy followed by endocrine therapy (n = 75; HR 0.50; 95% CI 0.09, 2.71; P = 0.417 (not shown)).
HMGA2 distribution and complementary breast cancer risk stratification
To test the possible strength as a biomarker in breast cancer, the effect of an additional inclusion of HMGA2 scores in a hypothetical model of conventional breast cancer risk parameters was determined. First, a Cox model of established risk parameters (tumor size, node status, grading, and ER positivity) was calculated in our collective (Figure 4, left). Second, utilizing hierarchical Cox models, the subcellular HMGA scores were included in this risk stratification separately. Figure 4 demonstrates that the incorporation of the overall (HR 0.39; 95% CI 0.19, 0.83; P < 0.05) and the cytoplasmic (HR 0.39; 95% CI 0.21, 0.74; P < 0.01)—but not the nuclear (HR 0.69; 95% CI 0.37, 1.27; P > 0.05)—HMGA2 levels significantly sharpened the selectivity of this combined risk stratification of conventional breast cancer risk factors. In other words, only the overall and the cytoplasmic HMGA2 scores led to an improved model fit (P-values at the bottom of Figure 4).
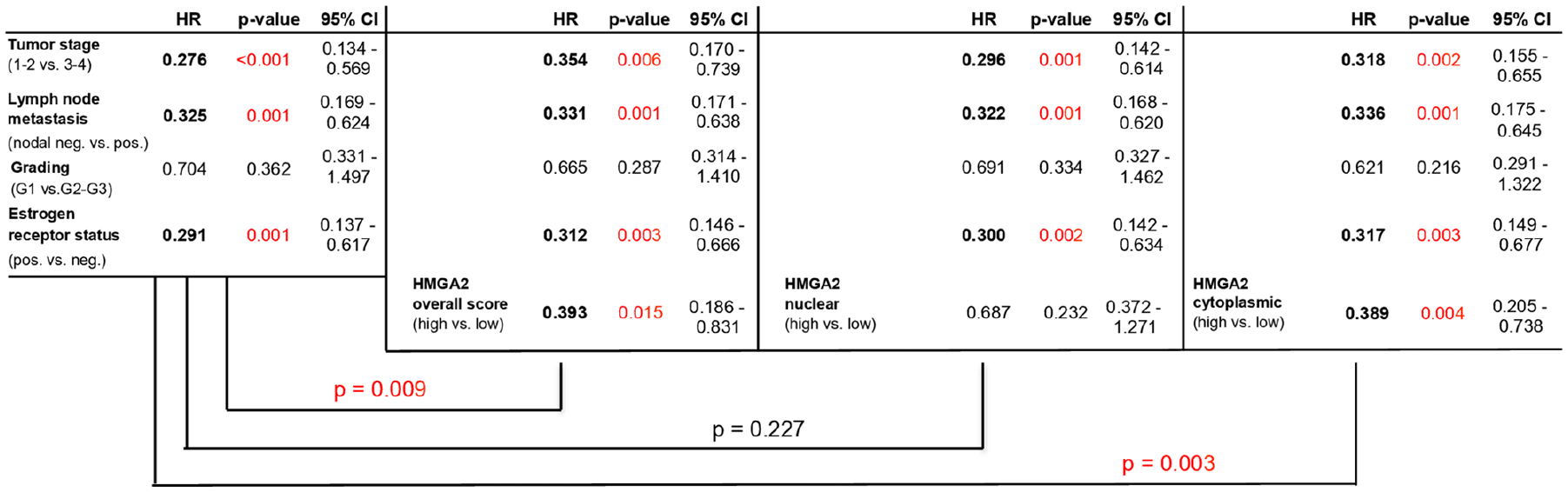
HMGA2 scores and OS: two-step hierarchical Cox regression model for conventional breast cancer risk factors and surrogate markers incorporating subcellular HMGA2 levels. Incorporating overall and cytoplasmic—but not nuclear—HMGA2-based models allowed more accurate risk stratification in our collective (P < 0.001).
Discussion
The prognostic relevance of HMGA proteins in breast and other cancers has been repeatedly tested. Here, we present an unprecedented analysis on the subcellular distribution of HMGA2 in 342 breast cancer specimens. Unexpectedly, high levels of HMGA2 were associated with a favorable OS of breast cancer patients, and this effect was mostly determined by cytoplasmic HMGA2. Implemented in a risk stratification model of conventional risk markers, HMGA2 overall and cytoplasmic—but not nuclear—levels significantly sharpened the prognostic selectivity of each model. We also delineated better survival prognoses upon cytoplasmic HMGA2 levels within joint Luminal A and B subgroups independent of tumor size or axillary lymph node status. Despite a median follow-up of only 48 months, and a limited number of late events in our patients’ collective, this could be of special interest in sharpening prognostic evaluations and tailoring breast cancer therapies in Luminal subgroup of breast cancer patients.
Progressive insights on HMGA biology in breast cancer have been gained. 32 Wu et al. 20 found nuclear HMGA2 expression to associate with advanced tumor stages, gene signatures of mesenchymal phenotypes, and an impaired OS in low-grade and non-TNBC tumors. In contrast, Wend et al. 33 linked HMGA2 expression specifically to TNBC types and described a correlation of HMGA2 with mesenchymal tumor characteristics, a poor recurrence free survival (RFS), and higher rates of metastasis in a patient’s panel who was suffering from TNBC. Interestingly, this study also referred to the nuclear HMGA2. High HMGA2 levels in less differentiated breast cancers were observed by Rogalla et al. 18 and Ryu et al. 19 In our analyses, the protective effects of cytoplasmic HMGA2 were retained in the biologically diverse Luminal A and TNBC subgroups. This ostensible inconsistency might indicate pan-subtype effects of HMGA2 expression or distribution, as HMGA proteins are linked to various cancer hallmarks. 32 For example, Thuault et al. 34 linked HMGA2 to epithelial-to-mesenchymal transition elicited by transforming growth-factor-beta in breast cancer cells in which HMGA2 regulated the expression of the transcription factors Snail, Slug, and Twist as major regulators of tumor invasiveness and metastasis. 34 Much larger analyses, including patients receiving HER2-targeting treatments, are therefore required to systematically analyze the prognostic value of HMGA2 in different intrinsic breast cancer subtypes. In order to align our contrasting prognostic findings of HMGA2 with TCGA data, we utilized the public, mRNA-based Kaplan–Meier Plotter. 35 Within this data set, HMGA2 was associated with an HR of 0.84 (95% CI 0.72, 0.98; P = 0.025) in 1764 breast cancer samples read out for RFS, and an HR of 1.01 (95% CI 0.74, 1.38; P = 0.94) in 626 breast cancer samples read out for OS. These observations are in line with data published by Huang et al., 36 who could not link HMGA2 to poorer survival in analyses based on TCGA breast cancer data sets. More specifically, mRNA-based approaches showed no association of HMGA2 expression with any particular subtype or tumor grading. 32 Therefore, Sgarra et al. 32 proposed that, due to its complex post-transcriptional regulation, HMGA2 scores should be evaluated at the protein level. 32
HMGA proteins have long since been recognized in extranuclear localizations and, specifically, the mitochondria. Dement et al. 37 have described a deregulated transport of HMGA1 from the nucleus to the cytoplasm in malignant cells. In normal cells, HMGA1 is exclusively present in the nucleus except during the late S-phase and G2-phase, when it is transiently found at mitochondrial sites. At these stages of the cell cycle, HMGA1 protein is phosphorylated, which drastically reduces its DNA binding capacity. In malignant cells, HMGA1 is found in the cytoplasm throughout all stages of the cell cycle and binds to AT-rich DNA strands in the regulatory D-loop in the mitochondrial genome, thereby influencing mitochondrial DNA maintenance and organelle function contributing to cancer-related mitochondrial abnormalities. 38 Also, high mobility group box (HMGB) proteins were found in cytosol and mitochondria, where they regulate mitochondrial functions and morphology. 39 To date, prognostic implications of HMGAs remain controversial, especially for different family members. 40 Moreover, concrete anti-tumoral effects of HMGA2 also have been described, linking HMGA2 protein to a cellular stage of senescence, in which—by accumulation on the chromatin—HMGA2 contributes to the repression of genes associated with proliferation. 41 HMGA2 is translocated to the nucleus via Importin-α2, which binds to the nuclear localization signal (NLS) within the second AT-hook. 42 The mechanism of its nuclear export has not been unraveled so far. For the transcription factor p8, which is biochemically related to HMGAs, a cell-cycle-dependent expression pattern has been described. The inhibition of p8-deacetylation promoted its cytoplasmic accumulation, and the nuclear export receptor CRM1 was not involved. 43 In the context of oncologic strategies, the inhibition of nuclear import mechanisms has been proposed for other key proteins also involved in cell growth, such as NFκB or β-catenin. 44 Therefore, it is tempting to speculate that targeting intracellular trafficking might lead to a beneficial accumulation of HMGA2 in the cytoplasm.
To the best of our knowledge, we present the first systematic valuation of the prognostic implications of HMGA2 in breast cancer depending on its subcellular localizations. It remains speculative whether nuclear and cytoplasmic levels of HMGA2 should be valued with an equal emphasis in terms of prediction of survival. A delineation of organelle-specific, anti-tumoral functions of HMGA2 in the cytoplasm from its other potentially oncogenic functions will be a challenge for future studies. Our HMGA2-based risk groups were defined and validated in the same patient set. Therefore, confirmation of our findings should be attempted in a validation set for breast and other cancers. Nonetheless, our findings emphasize the biologic and prognostic relevance of subcellular levels of HMGA2 in breast cancer, thereby adding more complexity to the implications of HMGA2 in cancer biology.
In summary, our findings provide a rationale for the evaluation of subcellular HMGA2 distribution patterns in breast cancer, rather than referring to the total amount of HMGA2. Our set of data does not support the conception of utilizing HMGA2 as an adverse biomarker in breast cancer. In contrast, our findings might stimulate research on extranuclear, novel functions of HMGA2 as well as on mechanisms responsible for HMGA2 shuttling between cellular compartments. Confirmation of our findings in separate breast cancer sets may unveil intracellular HMGA2 distribution as a pan-subtype biomarker in breast cancer.
Footnotes
Acknowledgements
None
Conflict of interest
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Declaration of Helsinki
The authors confirm that this study has been performed in accordance to the ethical standards of the Declaration of Helsinki and with approval of the local ethics committee (Christian-Albrechts-University of Kiel, Germany, Faculty of Medicine: #D450/10).
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
