Abstract
BACKGROUND:
ARHGDIB, a Rho GDP dissociation inhibitor protein, has been reported playing critical roles in regulation of multiple biological responses. However, whether ARHGDIB serves as a valuable biomarker in cancer is little known so far, especially in breast cancer.
OBJECTIVE:
In this study, we aimed to investigate the importance of ARHGDIB in breast cancer, including but not limited to biomarker-like role, as well as potential mechanisms.
METHODS:
Total 100 breast cancer samples and 100 benign breast disease samples were enrolled and underwent detailed pathological assessment and IHC analysis. Human breast cancer cell lines and epithelial cell line were subjected to siRNA-mediated knock-down, RT-qPCR, western blot, MTT staining, cell cycle assay, transwell analysis respectively.
RESULTS:
We observed the expression of ARHGDIB is significantly higher in human breast cancer tissues compared with the benign tissues. ARHGDIB expression was positively correlated with tumor size, lymph node metastasis and TNM stage in breast cancer patients. Moreover, ARHGDIB depletion decreased proliferation, migration and invasion of breast cancer cells. Furthermore, we found ARHGDIB mediated epithelial-mesenchymal transition, and MMP2 is the key downstream effector of ARHGDIB.
CONCLUSIONS:
Hence, our results suggested the significance and predictive role of ARHGDIB in breast cancer. High expression of ARHGDIB indicated the poor prognosis for breast cancer patients.
Introduction
Breast cancer, one of the most common malignant diseases, is one of the leading causes of cancer death for woman worldwide. According to the report, from 2005 to 2014, overall breast cancer incidence rates increased among Asian/Pacific Islander (1.7% per year) women [1]. As a result of the increasing longevity, increased urbanization and the adoption of western lifestyles, the incidence of breast cancer is increasing in the developing world [2]. Metastatic dissemination of cancer cells from the primary tumor and their spread to distant sites in the body is the leading cause of mortality in breast cancer patients, which is often associated with epithelial-mesenchymal transition (EMT) mediated metastasis [3, 4]. It has been shown that EMT is regulated by transcription factor Snai1 [5]. Despite kinds of therapies for the clinical treatment of breast cancer, such as radiation, chemotherapy and surgical resection, 41070 women with breast cancer died in the United States in 2017 [6]. However, the molecular mechanisms underlying the recurrence and metastasis of breast cancer remain to be elucidated. Thus, new biomarkers for the diagnosis and prognosis of breast cancer are required.
Rho GDP dissociation inhibitors (RhoGDIs), the pivotal regulators of Rho GTPase function, with the character of forming a complex with Rho GTPase, regulates their nucleotide exchange and membrane association [7, 8]. ARHGDIB is one of the most critical members in RhoGDIs. It is mainly located in hematopoietic, endothelial, and epithelial cells [9] and involved in a variety of cellular functions, including signaling, secretion, cytoskeletal organization and proliferation [10]. Previous studies have reported the dysregulation of ARHGDIB in a variety of human tumors, while the role of ARHGDIB appears as controversial in tumor development and progression. For example, low expression of ARHGDIB was reported in bladder cancer and Hodgkin’s lymphoma and was identified as a presumptive metastatic suppressor [11, 12]. On the other side, overexpression of ARHGDIB was observed in pancreatic cancer and gastric cancer [13, 14]. Furthermore, few studies conducted in the role of ARHGDIB in breast cancer [15]. The clinical significance of the expression of in breast cancer patients and the underlying mechanism remain to be elucidated. In this study, we investigated the role of ARHGDIB in cell proliferation and metastasis in breast cancer and the clinical significance in breast cancer patients. Our study suggest ARHGDIB might play an important role in breast cancer progression.
Results
Expression of ARHGDIB in clinical breast tissue specimens and cell lines
To address the role of ARHGDIB on the development and progression of breast cancer, we first detected the expression of ARHGDIB in clinical breast specimens by IHC. ARHGDIB was predominantly expressed at the cytoplasm of breast epithelium, and the expression of ARHGDIB protein was frequently observed stronger in breast cancer tissues, in comparison to benign breast disease tissues (Fig. 1A). Moreover, ARHGDIB protein expression was observed in 74 of the 100 (74%) breast cancer specimens which was significantly higher than benign breast disease tissue (
Expression of ARHGDIB in breast cancer samples and benign tissues
Expression of ARHGDIB in breast cancer samples and benign tissues
Differential expression of ARHGDIB in breast clinical samples and cell lines. A. Expression of ARHGDIB protein in breast cancer tissues and benign breast disease tissues was detected using IHC. B. Cell lysates, as indicated, were subjected to western blot analysis using antibodies specific for ARHGDIB. 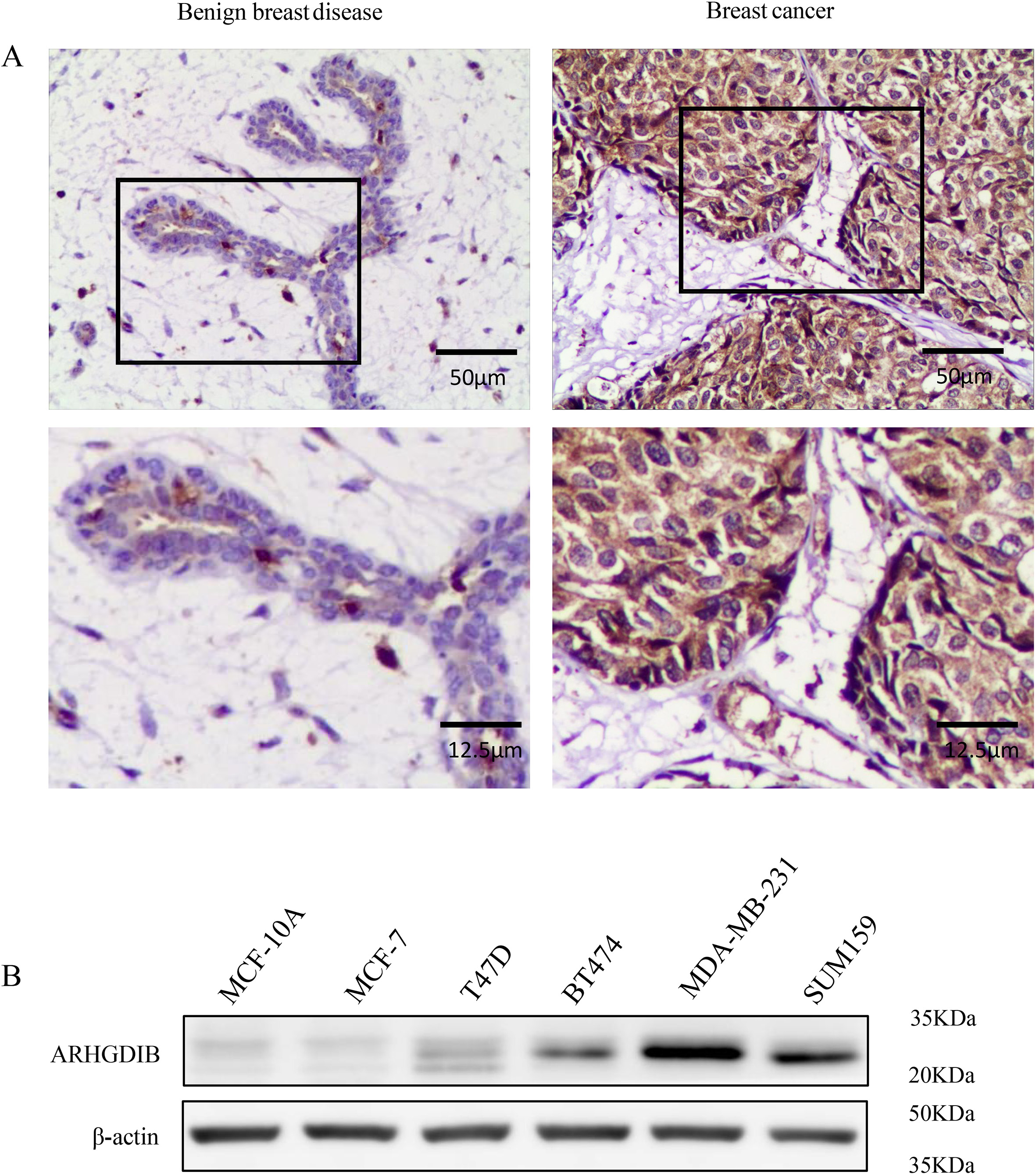
Correlation of ARHGDIB expression and clinicopathological parameters in breast cancer patients
To explore the clinical significance, we next investigated the association of ARHGDIB expression with clinicopathological parameters in breast cancer patients. As shown in the Table 2, the expression of ARHGDIB protein was significantly associated with large tumor size (
Impact of ARHGDIB expression on breast cancer cell proliferation
To assess the functional role of ARHGDIB in breast cancer, we transfected either ARHGDIB specific siRNA (siARHGDIB) or negative control (NC) into MDA-MB-231 cells. The results of western blot and RT-qPCR showed that ARHGDIB was remarkably decreased in the siARHGDIB groups compared to the NC group (Fig. 2A and B). In colony formation assays, the cells with transiently transfected into siARHGDIB formed smaller and fewer colonies per plate than that in the NC cells (Fig. 3A). In MTT assay, we observed that cell viability of the cells with transfection of siARHGDIB was significantly decreased compared with transfection with NC (Fig. 3B). Similarly, the cell numbers were decreased significantly in the group of siARHGDIB transfection compared with cells transfected with NC (Fig. 3C).
Construction of ARHGDIB knockdown system in MDA-MB-231 cells. A. Western blot analysis showed that siARHGDIB markedly reduced the protein expression of ARHGDIB. B. RT-qPCR analysis showed that siARHGDIB markedly reduced the mRNA level of ARHGDIB in MDA-MB-231 cells. 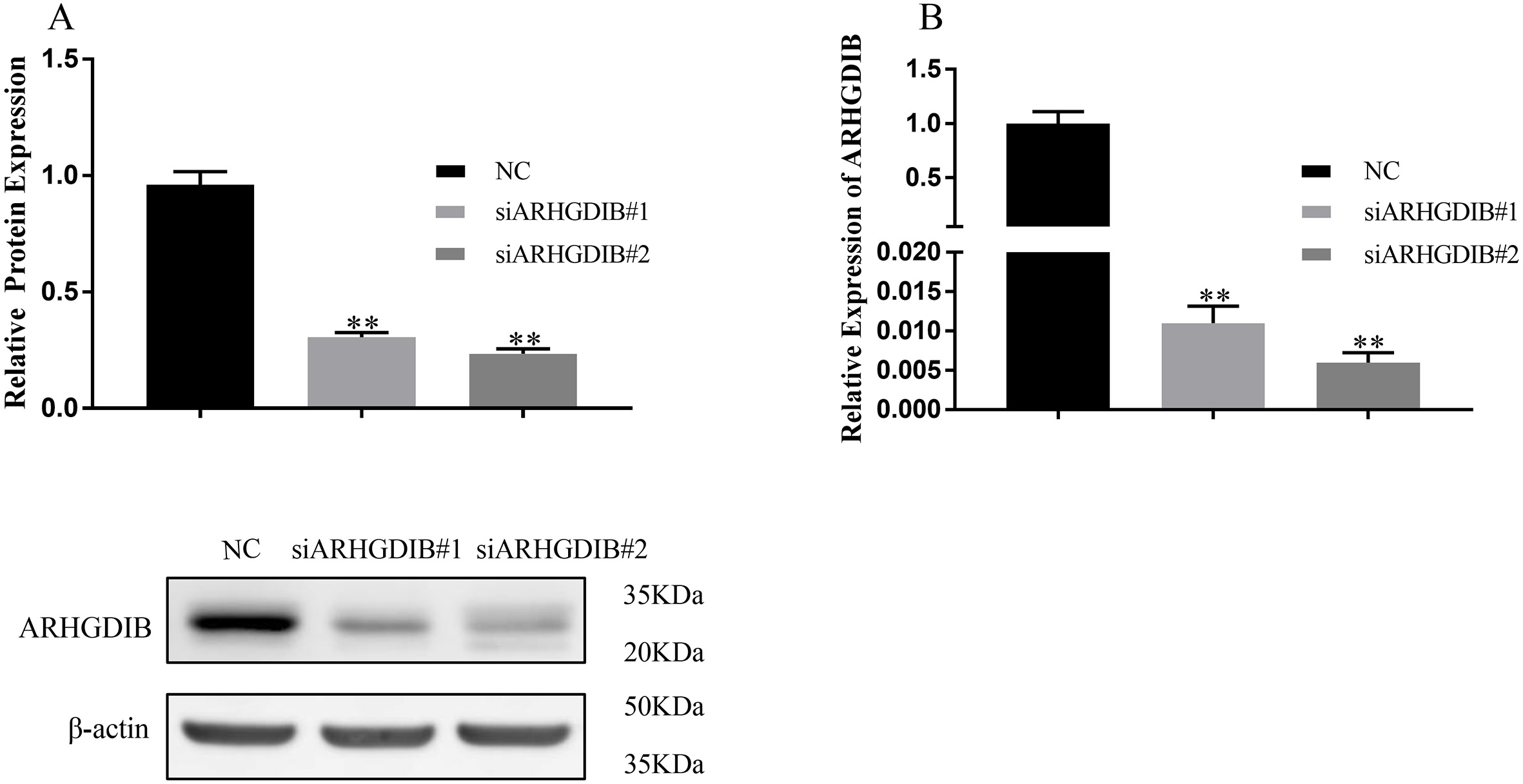
Inhibitory effects of ARHGDIB knockdown on breast cancer cell proliferation and cell cycle distribution. (A) Colony formation assay, (B) MTT assay and (C) cell growth assay indicated that knockdown of ARHGDIB significantly suppressed the proliferation of MDA-MB-231 cells. (D) The cell cycle distribution was determined with flow cytometric analysis. The representative FACS analysis is shown. The percentage of MDA-MB-231 cells in the G0/G1, S, and G2/M phases. The data represent one of three separate experiments. 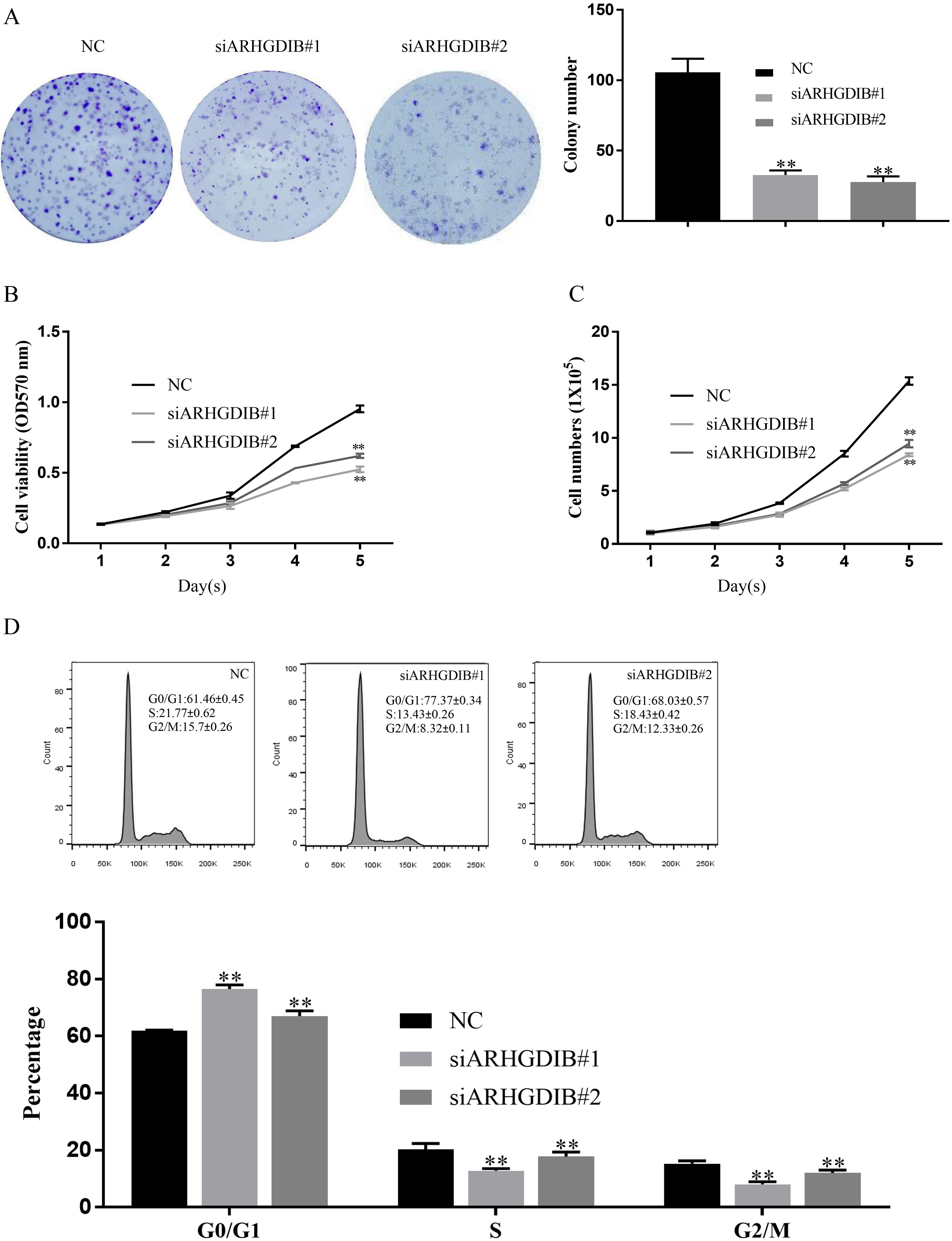
In order to determine whether ARHGDIB knockdown affects cell cycle regulation of breast cancer cells, flow cytometric analysis was performed at 48 h after cell transfection with either siARHGDIB or NC control. As shown in Fig. 3D, the population of both siARHGDIB-transfected MDA-MB-231 cells in the G0/G1 phase increased significantly. Moreover, the increase in G0/G1 phase cell population was accompanied with a concomitant decrease of cell number in S phase and G2/M phase.
Impact of ARHGDIB expression on migration and invasion of breast cancer cells
In order to determine the role of ARHGDIB in migration and invasion of breast cancer, cell migration and invasion assays were individually performed in MDA-MB-231 cells at 48 h after cell transfection with either siARHGDIB or NC control. Compared with NC group, both cell migration (Fig. 4A) and invasion (Fig. 4B) decreased significantly in MDA-MB-231 cells with siARHGDIB, which suggested that ARHGDIB might be involved in breast cancer metastasis.
Knockdown of ARHGDIB inhibits migration and invasion of breast cancer cells. (A) Migration assay and (B) Invasion assay indicated that the knockdown of ARHGDIB by siARHGDIB significantly suppressed the migration and invasion of MDA-MB-231 cells. 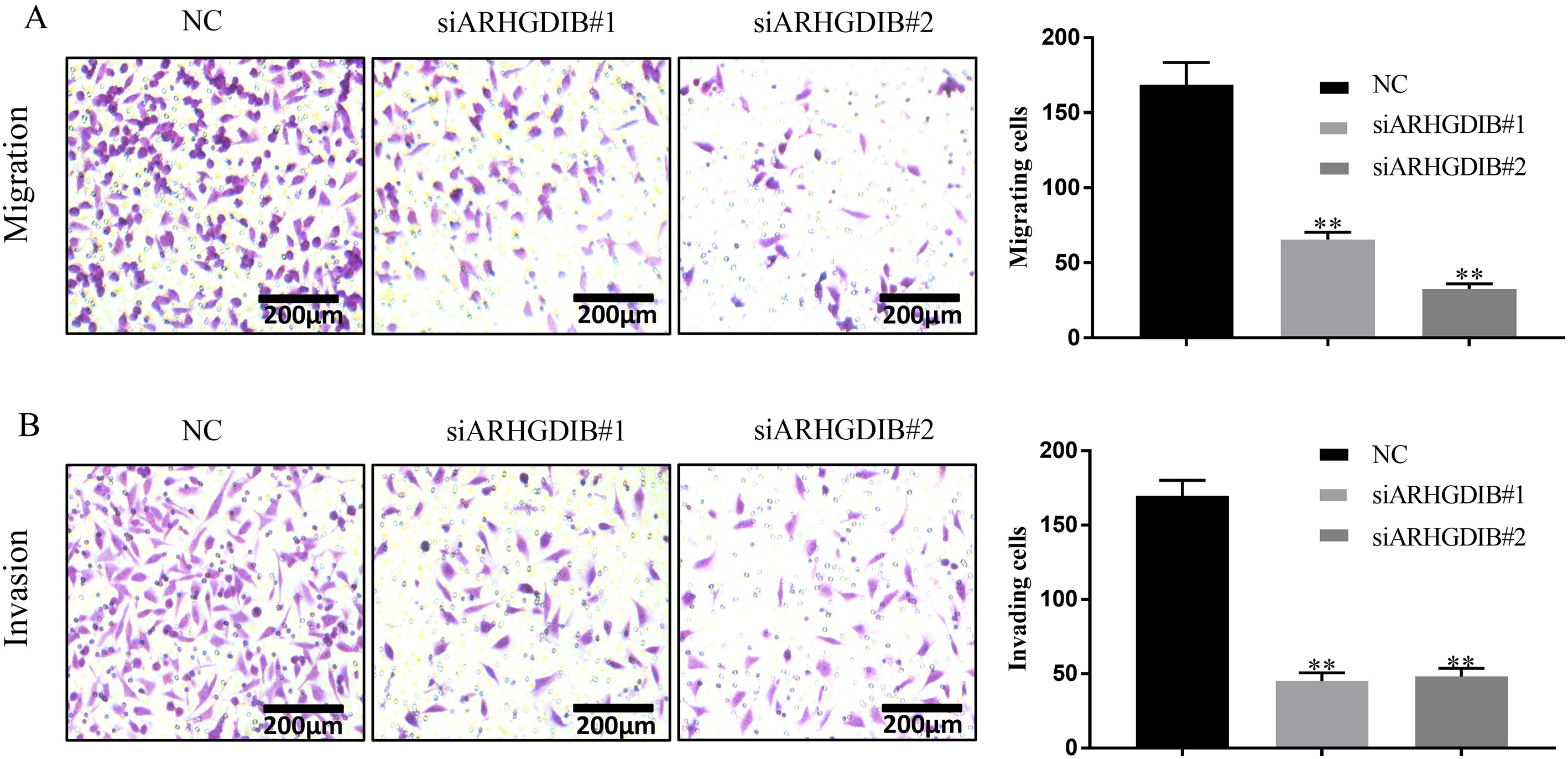
Considered the close connection between cancer migration, invasion and EMT, we further examined the protein levels of several key EMT biomarkers in breast cancer cells transfected with either siARHGDIB or NC control [16, 17]. As expected, protein levels of epithelium marker (Occludin) was increased, while mesenchymal markers (MMP2, Vimentin and Snai1) were decreased significantly in the group of siARHGDIB compared with NC control (Fig. 5).
ARHGDIB depletion suppresses EMT in breast cancers. Representative western blot for epithelial marker Occludin, as well as mesenchymal markers MMP2, Vimentin and Snai1 in siARHGDIB cells and NC control. 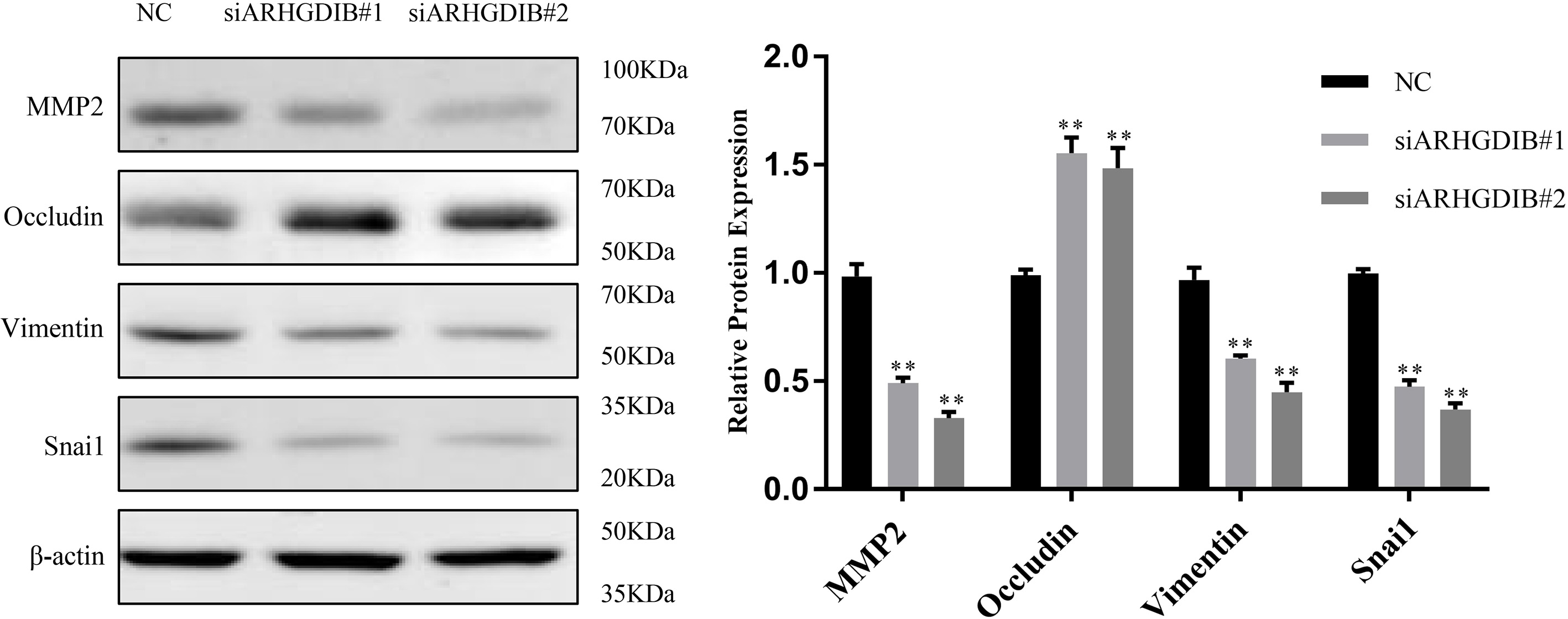
Rho GTPase proteins are involved in cytoskeleton component remodeling, thus influencing cell mobility [18]. Therefore, the regulation of Rho family proteins might have a profound impact on metastasis, a process that involves in cancer cell migration from one location to another to invade the destination tissue/organ [19]. ARHGDIB is a Rho GTPase regulatory protein that binds and holds GDP bound Rho proteins in an inactive non-membrane localized, cytoplasmic compartment [20]. Alterations of ARHGDIB expression levels have been shown in a variety of human tumor types [21]. Yi et al. showed that ARHGDIB promoted pancreatic carcinoma cell invasion and metastasis in vitro [13]. Yang et al. reported that high ARHGDIB expression level promotes the aggressive behavior of hepatocellular carcinoma [22]. These studies suggest an oncogene role of ARHGDIB in cancer progression. However, Gildea et al. reported that ARHGDIB is an invasion and metastasis suppressor gene in human bladder cancer [23]. Herein we systematically observed the role of ARHGDIB in human breast cancer both in vitro and in clinical tissues. The expression of ARHGDIB is frequently increased in breast cancer cell lines and clinical breast tissue specimens. Both the capacity of proliferation and metastasis were dramatically decreased after the knockdown of ARHGDIB in breast cancer cells. This is consistent with previous reports in the literature [15, 24]. In addition, the G0/G1 phase increased while S and G2/M phase decreased after transiently transfected into siARHGDIB in breast cancer cells. It is known that uncontrolled proliferation is a major feature of cancer cells, and these cells have typically acquired a complex defect in cell cycle progression. However, a previous report contradicts our existing conclusions. This study shows Rho-GDI
EMT is an important cellular mechanism during tumor invasion and metastasis development and results in enhanced cell motility [26]. However, it has not been determined whether ARHGDIB expression result in EMT in breast cancer. In this study, the knockdown of ARHGDIB was observed to inhibit EMT by down-regulating mesenchymal-associated markers MMP2, Vimentin and transcription factor Snai1 while up-regulating the epithelial-associated marker Occludin. To our knowledge, this is the first report suggesting that ARHGDIB can induce EMT in breast cancer.
In summary, our study demonstrated that the altered expression of ARHGDIB was exist in breast cancer and ARHGDIB play an important role in breast cancer cell proliferation and metastasis. Our finding suggests a role of ARHGDIB in the molecular etiology of breast cancer and provides a possible target for therapeutic strategies.
Materials and methods
Clinical sample collection
In total, 100 breast cancer samples and 100 benign breast disease samples were enrolled in the present study. These tissue specimens were collected from the patients treated with surgery between January 2013 and December 2014 at the First Affiliated Hospital of Anhui Medical University (Hefei, China). All resected tissues underwent detailed pathological assessment, and all patients had not treated with chemotherapy of radiation therapy before surgery, and no evidence of rheumatic disease, acute infection, human immunodeficiency virus or other type of cancer. The clinicopathological parameters of these patients were collected which were determined based on the 2012 World Health Organization (WHO) classification system [27]. The protocol to use these tissues was approved by the Biomedical Ethics Committee of Anhui Medical University with informed consent of all the involved patients.
Immunohistochemical analysis
For immunohistochemistry (IHC) of ARHGDIB expression in tissue samples, a rabbit anti-ARHGDIB polyclonal antibody was obtained from Proteintech Group (Proteintech Group, Chicago, IL, USA) and was used at a dilution of 1:50 according to our previous studies [28, 29]. Expression of ARHGDIB protein in breast cancer and benign breast disease tissues were observed by a pathologist who was blinded to the patients. The staining intensity was scored as: 0, negative; 1, weak; 2, medium and 3, strong. The extent of staining was scored as: 0, 0%; 1, 1–25%; 2, 26–50%; 3, 51–75% and 4
Cell culture
All human breast cancer cell lines (MCF-7, T47D, BT474, MDA-MB-231 and SUM159) and human breast epithelial cell line (MCF-10A) were purchased from the American Type Culture Collection (ATCC, Manassas, VA). MCF-7, T47D and BT474 cells were maintained in the RPMI-1640 medium supplemented with 10% fetal bovine serum (FBS, Hyclone). MDA-MB-231 cells were cultured in L15 medium supplemented with 10% FBS. SUM159 cells were cultured in the Ham’s F12 medium supplemented with 5% FBS, 10
siRNA transfection and knock-down
Small interfering RNAs (siRNA) for targeting ARHGDIB and negative control (NC) RNA were purchased from GenePharma (Shanghai, China). The sequences of the siRNAs and NC used for transfection are shown in Table 3. The MDA-MB-231 cells were seed in 6-well plates overnight and then transiently transfected with siARHGDIB or NC using Lipofectamine 3000 (Invitrogen, USA) according to the manufacturers’ protocol. Real-time PCR analysis was used to confirm the efficiency of siRNA transfection.
Sequence of the oligonucleotides used for knockdown experiments
Sequence of the oligonucleotides used for knockdown experiments
Total RNA was extracted from MDA-MB-231 cells using Trizol reagent (Invitrogen). The sequences of the primers used for RT-qPCR are shown in Table 4. The reverse transcription reaction was performed for total RNA using the all-in one first-strand cDNA Synthesis Supermix for qPCR (TransGen Biotech, Beijing) and Light Cycler Fast Start DNA Master Plus SYBR Green I kit (TransGen Biotech, Beijing) was used for RT-qPCR according to the manufacturers’ instructions protocol to analyze the mRNA expression of ARHGDIB. The expression level of the target gene was detected using the 2
Specific primers used for RT-qPCR experiments
Specific primers used for RT-qPCR experiments
Total proteins were lysed with RIPA lysis buffer (Beyotime, Shanghai, China) from each group of MDA-MB-231 cells, and total proteins were extracted to measure the concentration by a BCA protein quantification kit (Thermo, USA). Then about 15
Cell proliferation assays
For cell proliferation, methyl-thiazolyl-tetrazolium (MTT) assay, total number assay and colony formation assay were used. In total number assay, 1
Cell cycle assay
MDA-MB-231 cells were seed in 6-well plates overnight and then transiently transfected with siARHGDIB or NC. Cell Cycle and Apoptosis Analysis Kit (Beyotime, Shanghai, China) was used to explore the cell cycle, when the confluence of each group of cells approximately reach 70%. Briefly, the MDA-MB-231 cells were collected, washed with PBS for three times, and fixed with 70% ice-cold ethanol overnight. The ethanol-suspended cells were then washed and stained with propidium iodide (PI) for 30 min in the dark at 37
Migration and invasion assays
MDA-MB-231 cells were seed in 6-well plates overnight and then transiently transfected with siARHGDIB or NC and incubated for 48 hours before the migration and invasion experiments. For the cell migration assay, cells were performed using uncoated (8
Statistical analysis
The association between the expression of ARHGDIB and clinicopathological parameter in clinical samples was analyzed using the chi-square test. In addition, the statistical differences of the experimental data were evaluated by one-way analysis of variance (ANOVA) using GraphPad Prism 7 software package. All experiments were performed at least three times.
Footnotes
Acknowledgments
This work was supported by the National Nature Science Foundation of China (81972472, 31970696 and 81502975), Anhui provincial academic and technical leader reserve candidate (#2016H074), and Key Program of Outstanding Young Talents in Higher Education Institutions of Anhui (#gxyqZD2016046). The research was supported in part by the China Postdoctoral Science Foundation (2016T90413 and 2015M581693).
Conflict of interest
The authors declare that they have no conflicts of interest.
