Abstract
Objective:
To evaluate the function of long non-coding RNA ANRIL (CDKN2B-AS1) in laryngeal squamous cell cancer (LSCC), and to explore the underlying mechanism.
Methods:
The expression levels of CDKN2B-AS1 in LSCC tissues and cell lines (Tu177, HN4, AMC-HN-8 and NP69) were determined by reverse transcription quantitative PCR (RT-qPCR). AMC-HN-8 cells were then transfected with siRNAs of CDKN2B-AS1. The effects of CDKN2B-AS1 on cell proliferation, cell cycle, and apoptotic protein were determined by CCK-8 assay, flow cytometry analysis, and western blot, respectively. Dual luciferase reporter assay and RNA immunoprecipitation (RIP) assay were employed to verify the targets of CDKN2B-AS1. The miR-324-5p mimics or miR-324-5p inhibitor and ROCK1 over-expression plasmids were also transfected into AMC-HN-8 cells for further analysis.
Results:
CDKN2B-AS1 was upregulated in LSCC tissues, and the upregulation of CDKN2B-AS1 was correlated with overall survival, advanced clinical stage, and lymph node metastasis. In AMC-HN-8 cells, the knockdown of CDKN2B-AS1 by siRNA inhibited cell viability, blocked cell cycle in G1 phase, and increased the expression levels of cyclin-dependent kinase inhibitor 1A (p21), cleaved caspase3, and cleaved PPoly (ADP-Ribose) polymerase 1. Results of dual luciferase reporter assay showed that miR-324-5p could bind to CDKN2B-AS1 or Rho-associated coiled-coil containing protein kinase 1 (ROCK1). Finally, over-expression of ROCK1 in AMC-HN-8 cells revised the inhibitory effect of CDKN2B-AS1 siRNA on cell growth.
Discussion:
The upregulation of CDKN2B-AS1 was correlated with overall survival, advanced clinical stage, and lymph node metastasis and promoted LSCC cell growth via miR-324-5p/ROCK1 axis.
Introduction
Larynx squamous cell carcinoma (LSCC) is one of the most prevalent malignancy types of squamous cell carcinoma of the head and neck (SCCHN), which originates from the larynx. 1 The incidence of LSCC is about 2.4% each year worldwide, and the 5-year survival rate is relatively low due to recurrence or metastasis. 2 Although cisplatin therapy has achieved great progress in the treatment of LSCC, the outcomes for patients were still not good. 3 Considering the important role of the larynx in respiration and phonation, the quality of the life of patients is very poor. 4 Thus, there is an urgent need to explorer the potential therapy targets for LSCC.
The aberrant expression of genes plays key roles in tumorigenesis, and the discovery of competitive endogenous RNAs (ceRNAs) hypothesis has expanded the regulatory mechanism of gene expression. 5 The long non-coding RNA (lncRNA) and targets of micro RNA (miRNA) competitively bind with the same miRNA through the similar binding sites, which are called the miRNA response element (MRE). 5 Fang et al. 6 found there were 82 miRNAs, 1197 mRNAs, and 1041 lncRNAs differentially expressed in head and neck squamous cell carcinoma (HNSCC) from the data of the Cancer Genome Atlas. Among them, 8 miRNAs, 71 lncRNAs, and 16 messenger RNAs (mRNAs) were predicted to construct ceRNAs.
lncRNA ANRIL (CDKN2B-AS1) is an antisense of the cyclin-dependent kinase inhibitor 2B (CDKN2B), which plays important roles in coronary artery disease, ischemic stroke, chronic obstructive pulmonary disease, inflammatory regulation, and various cancers.7-11 Several miRNAs have been identified as targets of CDKN2B-AS1. For example, in glioma cells, CDKN2B-AS1 could promote cell growth, migration, and invasion by regulating miR-34a. 12 However, the role of CDKN2B-AS1 in LSCC has not been studied yet. Rho-associated coiled-coil containing protein kinase 1 (ROCK1) is a serine/threonine kinase, which is a major downstream effector of small GTPase rhodopsin (Rho). 13 Many previous studies have shown that ROCK1 plays an important role in cell proliferation, cell apoptosis, cell adhesion, and other fundamental cellular functions in normal cells or cancers. 14 However, its role in LSCC is still unknown.
In this study, we determined the role of CDKN2B-AS1 in LSCC and explored the potential mechanism. Our results provide a potential biomarker and therapy target for LSCC.
Material and methods
Patients
The research was approved by the Ethics Committee of Beijing Tongren Hospital, Capital Medical University, and followed the World Medical Association Declaration of Helsinki (Approval no. TRECKY2019-086). Written informed consents were obtained from all patients. A total of 60 pairs of LSCC tissues, adjacent normal tissues, and laryngeal papilloma (LP) tissues were collected from LSCC patients, who hadn’t received chemotherapy or radiotherapy from April 2016 to June 2018. The collected tissues were stored in liquid nitrogen until use. The clinicopathological characteristics of LSCC patients are recorded in Table 1.
Relationship between lncRNA CDKN2B-AS1 and clinico-pathological parameters.
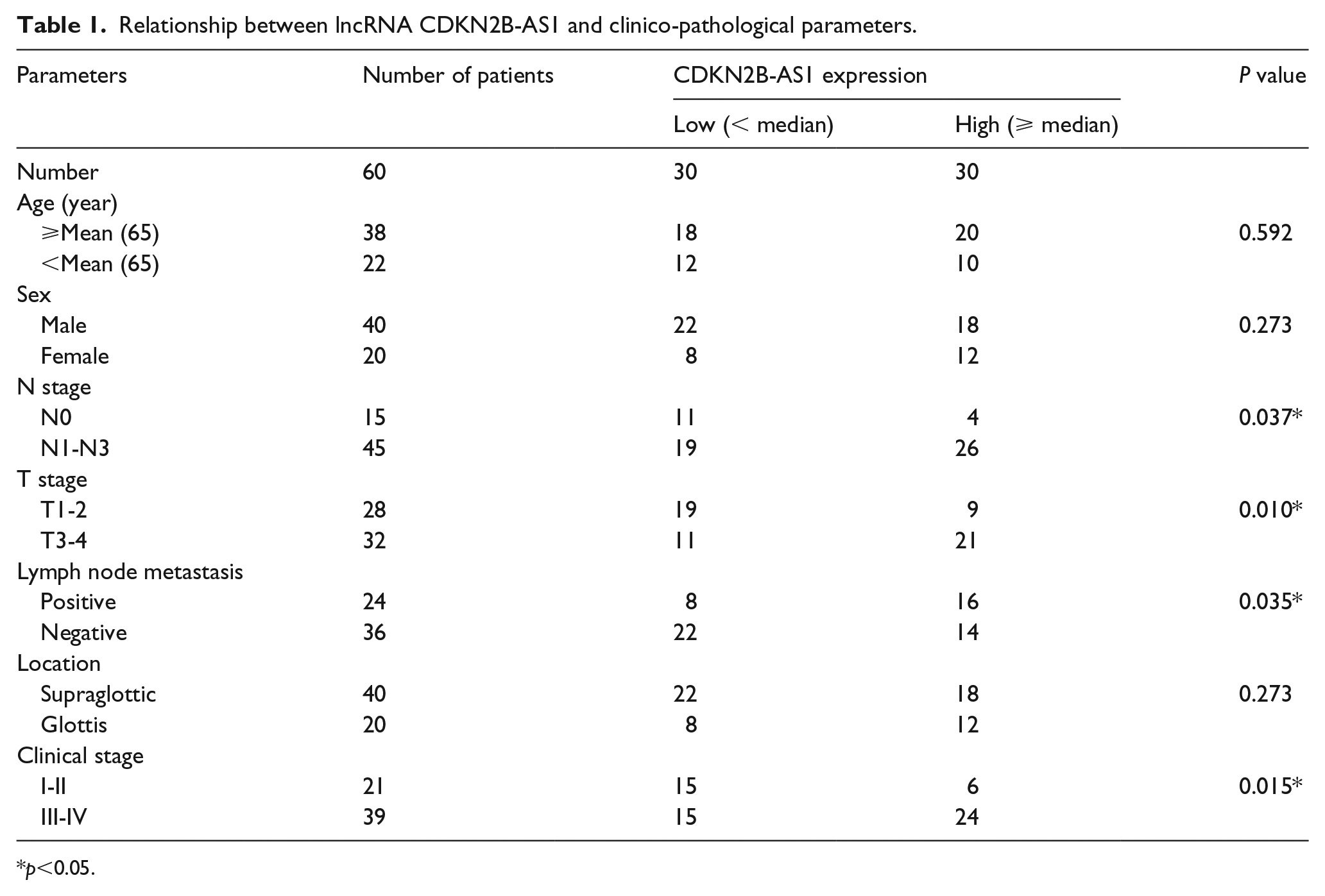
p <0.05.
Cell culture
Normal nasopharynx keratinocytes cell lines carrying SV-40T (NP69, BNCC338439), LSCC cell lines (TU177, BNCC341383; AMC-HN-8, BNCC338377) were purchased from Bena culture collection Co., Ltd. (Beijing, China) and LSCC cell line (HN4, YS448C) was purchased from YaJi biological Co., Ltd. (Shanghai, China). Cells were cultured in Dulbecco’s Modified Eagle Medium (Gibco, Waltham, MA, USA) containing 10% fetal bovine serum (FBS; Invitrogen, Waltham, MA, USA) and 1% penicillin/streptomycin (Invitrogen) in a 37°C humidified incubator with 5% CO2.
siRNAs, mimics and inhibitor of miR-324-5p
siRNAs of CDKN2B-AS1, mimics and inhibitor of miR-324-5p, and their negative controls were purchased from GenePharma (Shanghai GenePharma Co., Ltd., Shanghai, China). siRNAs of CDKN2B-AS1 (si-CDKN2B-AS1 #1, si-CDKN2B-AS1 #2) and its negative control (si-NC) were transfected into AMC-HN-8 cells by using Lipofectamine 3000 (L3000008, Thermo Fisher Scientific (China) Co., Ltd., Shanghai, China). Mimics of miR-324-5p and its negative control (miR-NC), and the inhibitor of miR-324-5p and its negative controls (NC inh) were transfected into AMC-HN-8 cells by using Lipofectamine 3000 (L3000008, Thermo Fisher Scientific (China) Co., Ltd.).
CCK-8 assay
103 AMC-HN-8 cells per well were seeded in a 96-well plate and transfected siRNAs or plasmids. At indicated hours, a CCK-8 reagent (Dojindo Laboratories, Dojindo Molecular Technologies (Shanghai), Inc., Shanghai, China) was added into wells and incubated for 1 hour. Then, optical density (OD) value at 450 nm was measured. Means ±SD were calculated from three independent experiments.
Cell cycle analysis
AMC-HN-8 cells (105 per well) were seeded in a 24-well plate and transfected with siRNAs or plasmids. Forty-eight hours later, the cells were fixed and covered with PBS containing 0.5% Triton X-100, 250 μg/mL RNase A and 10 μg/mL propidium iodide for 0.5 h at room temperature in the dark. Then, the cells were subjected into an Accuri C6 flow cytometer (BD Accuri Cytometers, Ann Arbor, MI, USA) and cells with different DNA contents were counted. Means ±SD were calculated from three independent experiments.
Reverse transcription quantitative PCR (RT-qPCR)
For RT-qPCR, total RNAs from cells or tissues were isolated using TRIzol™ Reagent (15596026, Invitrogen™, Thermo Fisher Scientific (China) Co., Ltd.) according to the manufacturer’s protocol. For miR-324-5p, synthesis of cDNA and quantitative PCR was performed using Mir-X™ miRNA qRT-PCR TB Green® Kit (638314, Takara Bio USA, Inc., CA, USA). For ROCK1 and CDKN2B-AS1, cDNA was reversely transcripted with PrimeScript™ RT reagent Kit with gDNA Eraser (RR047A, Takara, Takara Biomedical Technology (Beijing) Co., Ltd., Beijing, China). qRT-PCR was performed with TB Green® Premix Ex Taq™ II (RR820A, Takara, Takara Biomedical Technology (Beijing) Co., Ltd.). The primers used for qRT-PCR are listed in Table 2. The relative expression levels were calculated using the 2−ΔΔCt methods.
Sequences of primers used for qRT-PCR.
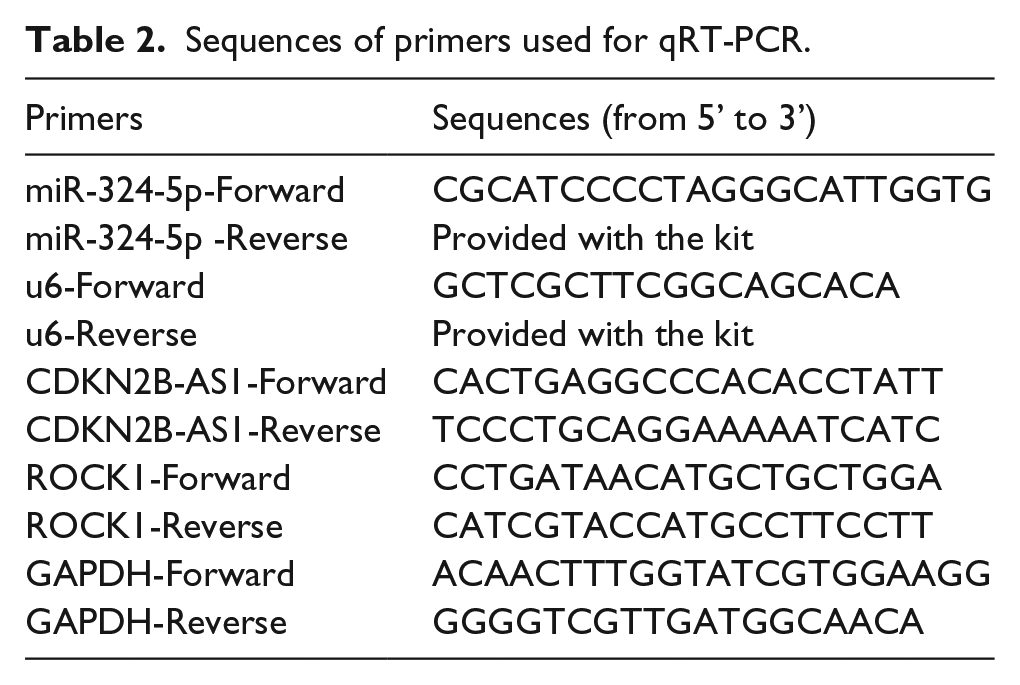
Western blot
Proteins from cells or tissues were extracted with ice-cold RIPA buffer (98065, CST, Danvers, MA, USA) containing PMSF (8553, CST) with protease inhibitor cocktail (5871, CST). Concentrations of isolated proteins were determined by BCA protein assay kit (7780, CST). A total of 10 μg proteins per lane were loaded and separated with sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to a polyvinylidene fluoride (PVDF) membrane. Then, the membrane was blocked using 5% BSA in PBS for 1 h at 37°C, then incubated with primary antibodies. After washing, the membrane was subsequently incubated with HRP-conjugated secondary antibodies and visualized with Pierce™ ECL Plus Western Blotting Substrate (32134, Thermo Fisher Scientific, Waltham, MA, USA). Finally, Image-J software (version 1.8.0_112, Bethesda, National Institutes of Health, MD, USA) was employed to determine the grey value of the bands. The listed primary antibodies were used in this study: anti-PCNA (1:1000, 2586, CST); anti-p21 (1:1000, 2946, CST); anti-cleaved caspase-3 (1:1000, 9661, CST); anti-cleaved PARP (1:1000, 5625, CST); anti-ROCK1 (1:1000, 4035, CST), and anti-GAPDH (1:1000, 5174, CST).
Dual luciferase reporter assay
To assess the interaction between miR-324-5p and CDKN2B-AS1, miR-324-5p and ROCK1, wild type and mutants of CDKN2B-AS1 and 3′-UTR of ROCK1 in pGL3 vector were designed from GenePharma Co., Ltd. (Shanghai, China). The mimics of miR-324-5p or miR-NC combined with synthesized pGL3 plasmids were co-transfected into 293T cells (BNCC100530, Bena Culture Collection Co., Ltd., Beijing, China). Forty-eight hours later, luciferase activity was measured using Dual-Luciferase® Reporter Assay System (E1910, Promega, Madison, WI, USA). The experiments were performed three times.
Statistical analysis
Data were presented as the means ± SEM. Statistical analysis was calculated with GraphPad Prism 6 (GraphPad Software, Inc., La Jolla, CA, USA). Student’s t-test or Tukey’s multiple comparisons test after the ANOVA test were employed for two groups or for three or more groups. Statistical significance was accepted when P < 0.05.
Results
High expression of CDKN2B-AS1 was related to clinicopathological characteristics of LSCC
To determine the function of CDKN2B-AS1 in LSCC, the expression level of CDKN2B-AS1 was firstly determined by RT-qPCR. The results showed that the expression level of CDKN2B-AS1 laryngeal papilloma tissues and LSCC tissues were consecutively significantly increased compared to that in normal adjacent tissues (Figure 1(a)). According to the median expression levels of CDKN2B-AS1, the LSCC patients were divided into two groups: the high expression group and the low expression group. As listed in Table 1, high expression of CDKN2B-AS1 was distinctly associated with poor overall survival, advanced clinical stage, and lymph node metastasis (Figure 1(b) and Table 1). Moreover, expression levels of CDKN2B-AS1 in LSCC cell lines (Tu177, HN4, and AMC-HN-8) were determined. When compared with NP69 cells, the expression levels of CDKN2B-AS1 in HN4, Tu177, and AMC-HN-8 were consecutively significantly increased (Figure 1(c)). The AMC-HN-8 cell line had the highest expression levels, which was thus chosen in further studies.
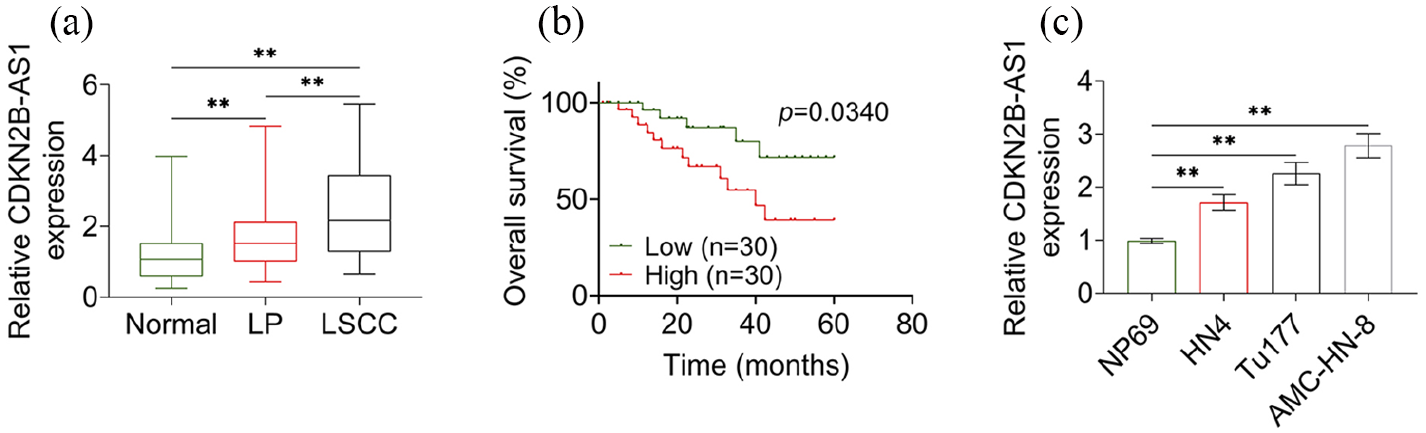
High expression of CDKN2B-AS1 was related to overall survival of LSCC patients. (a) Expression levels of CDKN2B-AS1 in adjacent normal tissues (normal), laryngeal papilloma tissues (LP) and LSCC tissues were determined by RT-qPCR. (b) Overall survival of LSCC patients with high or low expression level of CDKN2B-AS1. (c) Expression level of CDKN2B-AS1 in different cell lines were determined by RT-qPCR.
Knockdown of CDKN2B-AS1 inhibited cell growth of AMC-HN–8
siRNAs were used to knock down the expression level of CDKN2B-AS1 in AMC-HN-8 cells. Two siRNAs (si-CDKN2B-AS1 #1 and si-CDKN2B-AS1 #2) targeting different regions of CDKN2B-AS1 and the negative control (si-NC) were transfected into AMC-HN-8 cells and expression level of CDKN2B-AS1 was determined by RT-qPCR. The results showed that both the two siRNAs decreased the expression levels of CDKN2B-AS1 in AMC-HN-8 cells (Figure 2(a)). Then, the effect of CDKN2B-AS1 on cell growth and cycle was determined. CCK-8 assay showed that knockdown of CDKN2B-AS1 significantly decreased the viability of AMC-HN-8 cells (Figure 2(b)). Cell cycle analysis using propidium iodide staining showed that knockdown of CDKN2B-AS1 arrested cells in the G1 phase and decreased cell numbers in the S phase (Figure 2(c)). Moreover, results of western blot showed that the expression level of PCNA—an indicator of cell proliferation—in CDKN2B-AS1 knock-down cells was significantly decreased, whereas p21, cleaved caspase3, and cleaved PARP (indicators of cell apoptosis) were markedly increased (Figure 2(d)). Taken together, these results indicated that knockdown of CDKN2B-AS1 inhibited viability and induced apoptosis of AMC-HN-8 cells.
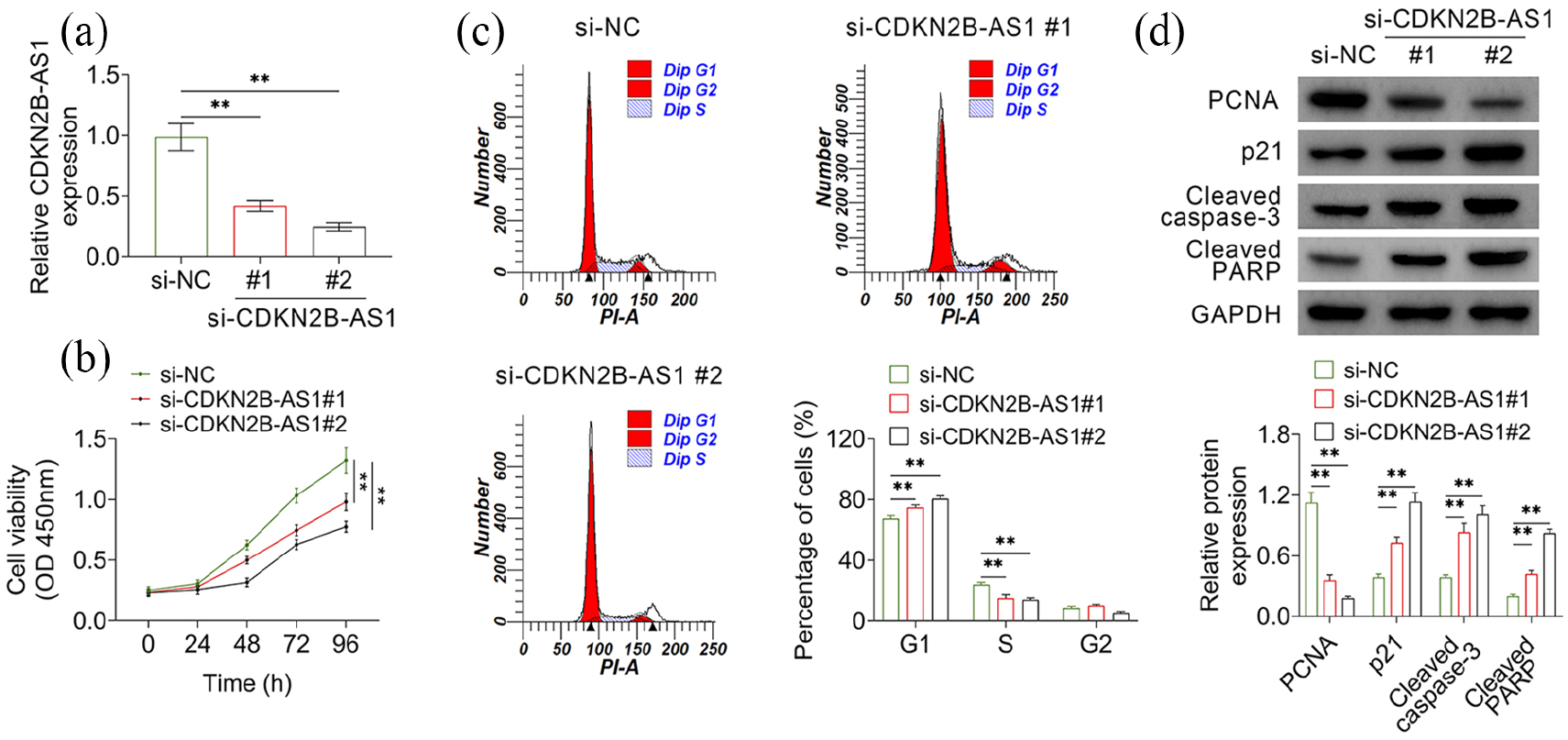
CDKN2B-AS1 regulated expression levels of miR-324-5p in AMC-HN-8 cells
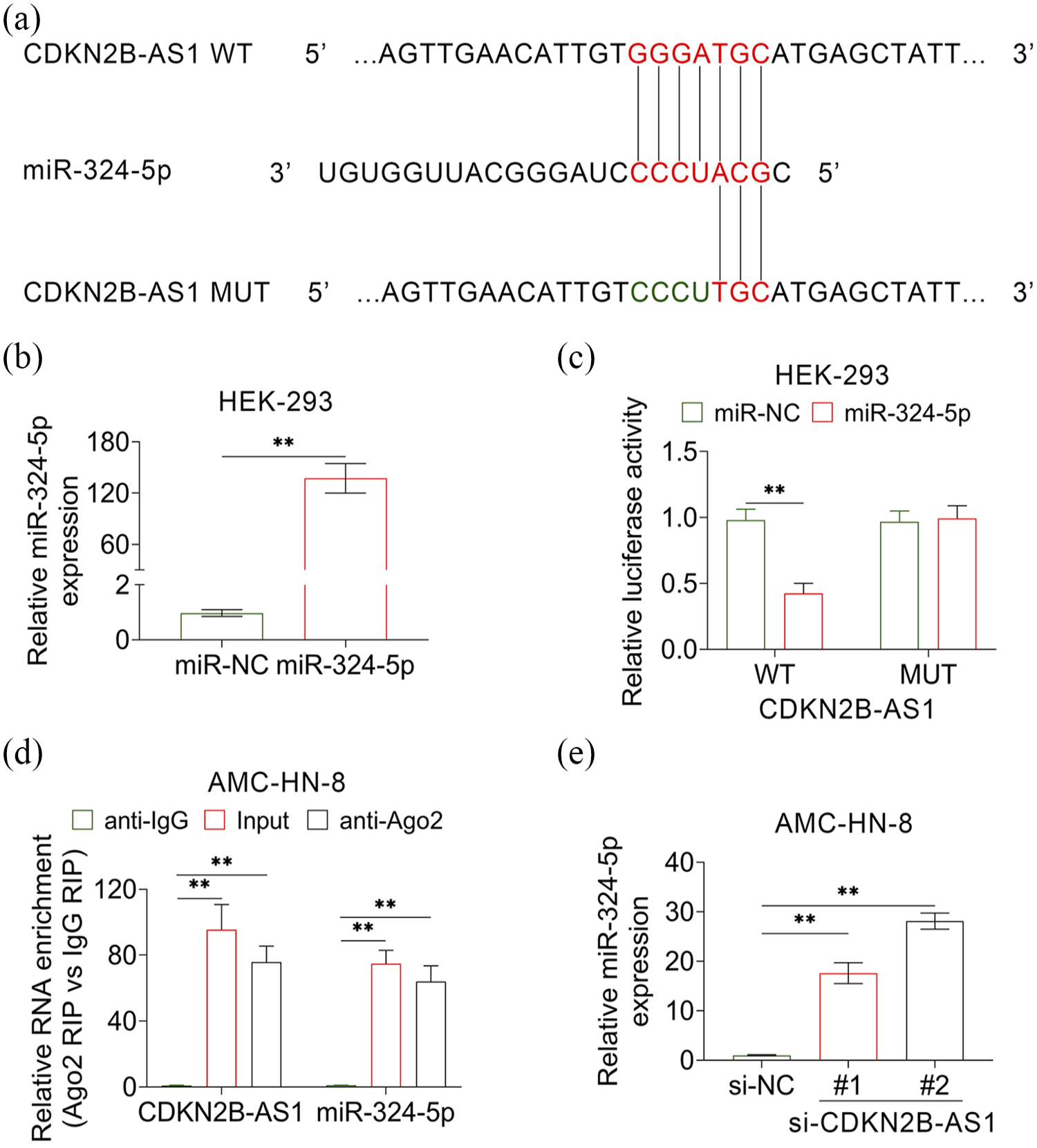
The miRDB (http://mirdb.org/) was used to predict the miRNAs that regulated by CDKN2B-AS1 and the results indicated miR-324-5p was one of the predicted miRNAs (Figure 3(a)). For further verification, mimics of miR-324-5p were transfected into AMC-HN-8 cells and dual luciferase report assay was employed. The expression level of miR-324-5p was increased hundreds of times in cells that transfected with miR-324-5p mimics, determined by RT-qPCR (Figure 3(b)). Then, cells were transfected with mimics of miR-324-5p combined with luciferase report plasmids containing wild type or mutants of CDKN2B-AS1. The results showed that the relative luciferase activity was only decreased in the cells transfected with mimics of miR-324-5p and luciferase report plasmids containing wild type CDKN2B-AS1 (Figure 3(c)). Moreover, the expression levels of CDKN2B-AS1 and miR-324-5p were determined in the ago-RNA complex, which was immunoprecipitation by anti-Ago. Both the expression levels of CDKN2B-AS1 and miR-324-5p were highly enriched in the complex immuno-precipitated with anti-Ago compared with anti-IgG (Figure 3(d)). These results indicated that CDKN2B-AS1 could bind to miR-324-5p in AMC-HN-8 cells. Also, the expression level of miR-324-5p was markedly down-regulated in AMC-HN-8 cells that transfected with siRNAs of CDKN2B-AS1, determined by qRT-PCR (Figure 3(e)). The results showed that lncRNA CDKN2B-AS1 function as sponge for negatively regulating expression levels of miR-324-5p.
CDKN2B-AS1 regulated expression levels of ROCK1 through miR-324-5p in AMC-HN-8 cells
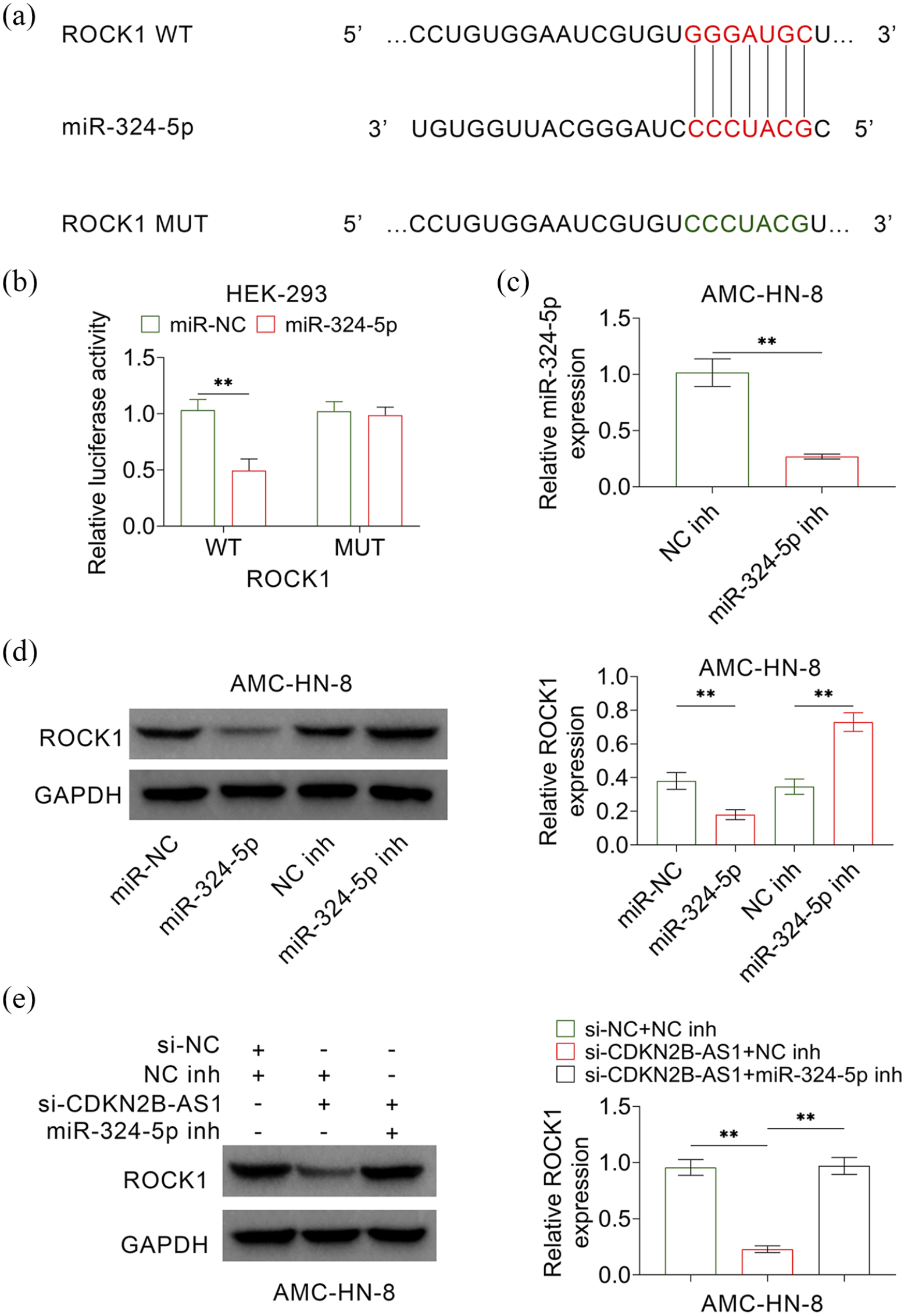
The Targetscan (http://www.targetscan.org/vert_71/) was used to predict the targets of miR-324-5p and the results showed ROCK1 was one of the predicted targets (Figure 4(a)). A dual luciferase report assay was further employed to verify the binding of 3′UTR of ROCK1 and miR-324-5p. The results showed that the relative luciferase activity was only decreased in the cells transfected with miR-324-5p mimics and luciferase report plasmids containing wild type 3′UTR of ROCK1 (Figure 4(b)). Then, the inhibitor of miR-324-5p was synthesized and expression level of miR-324-5p in the cells transfected with miR-324-5p inhibitor was significantly lower than that transfected with negative control (NC inh) (Figure 4(c)). Expression levels of ROCK1 were significantly increased in cells that transfected with miR-324-5p inhibitor, whereas was decreased in cells that transfected with miR-324-5p mimics (Figure 4(d)). These results suggested that miR-324-5p regulated the expression level of ROCK1 in LSCC cells. Finally, the results showed that decreased expression of ROCK1 caused by CDKN2B-AS1 knockdown was reversed by co-transfected with miR-324-5P inhibitor (Figure 4(e)). Thus, the upregulation of CDKN2B-AS1 promoted the expression of ROCK1 by decreasing the expression of miR-324-5p.
Over-expression of ROCK1 reversed the inhibitory effects of CDKN2B-AS1 siRNA on growth of AMC-HN-8 cells
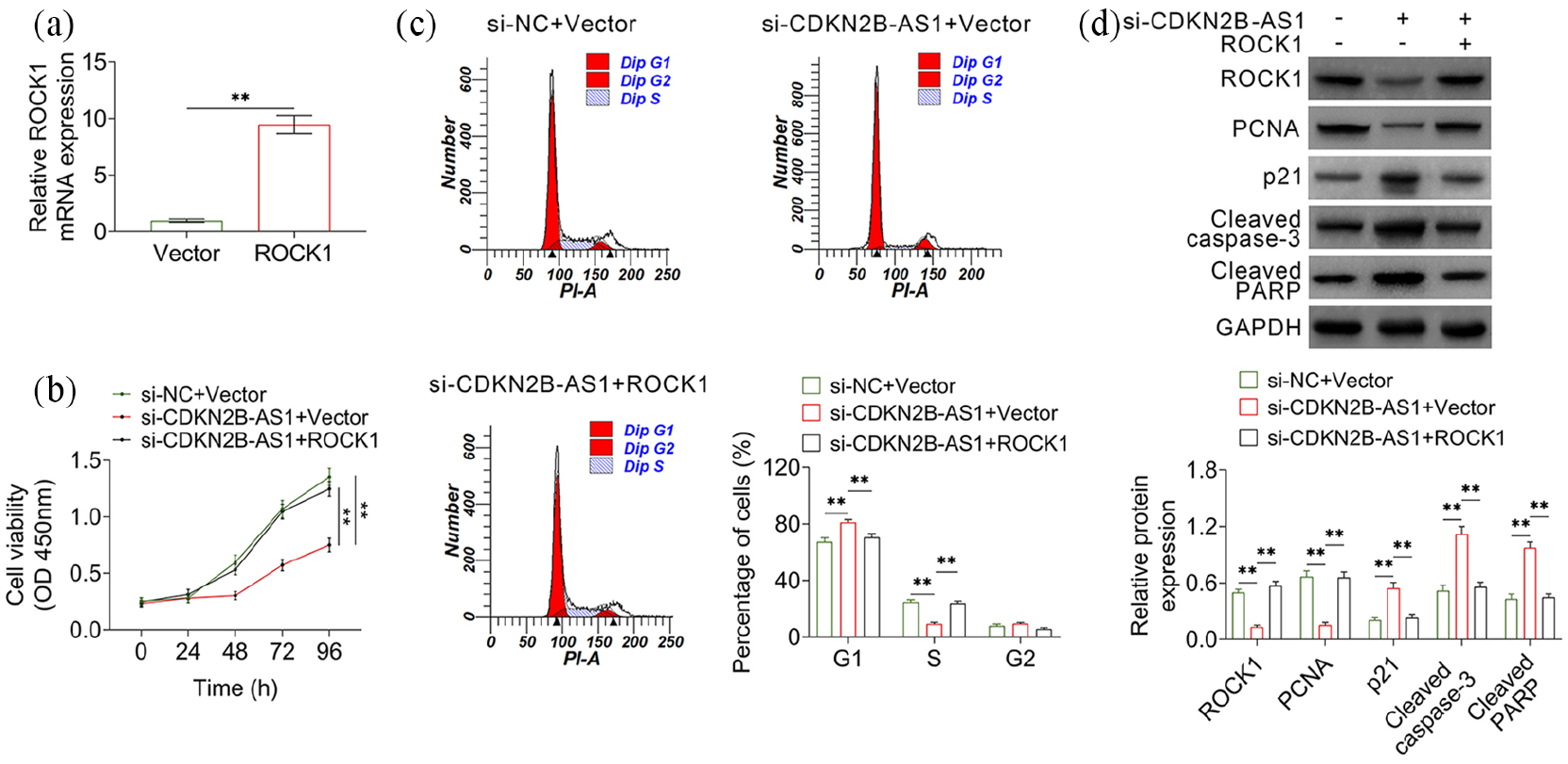
Expression level of ROCK1 was increased more than 10 times by transfection with ROCK1 over-expression plasmids in AMC-HN-8 cells (Figure 5(a)). Then, cell viability, cycle, and apoptosis were determined. Knock down of CDKN2B-AS1 decreased cell viability, whereas co-transfection with ROCK1 over-expression plasmids repaired the inhibitory effect of CDKN2B-AS1 knockdown (Figure 5(b)). Co-transfection with ROCK1 over-expression plasmids also restored the cell cycle distribution (Figure 5(c)). Moreover, enforced expression of ROCK1 in CDKN2B-AS1 knock-down cells also restored the expression levels of PCNA, p21, cleaved caspase3, and cleaved PARP (Figure 5(d)). These results indicated that CDKN2B-AS1 regulated cell viability, cycle, and apoptosis through miR-324-5p/ROCK1.
Discussion
The discovery of ceRNA has broadened our knowledge about the mechanism of gene expression regulation and tumorigenesis. 15 The finding of several ceRNAs in SCCHN from silicon-based analysis of aberrant expression of lncRNAs, miRNAs, and mRNAs suggested that ceRNA also played an important role in the tumorigenesis of LSCC. 6 Although the global lncRNA expression profile in LSCCs was screened using lncRNA microarray 5 years ago, only a few lncRNAs have been studied. 16 Chen et al. 17 found that low expression level of lncRNA AC008440.10 was correlated with increased risk of metastasis, poor prognosis and patient survival, which could be a biomarker for LSCC. High expression of the small nucleolar RNA host gene 20 (SNHG20) was associated with advanced tumor staging and undesirable prognosis, and promoted malignant progression of LSCC by regulating microRNA-140 (miR-140). 18 In this study, we confirmed that the expression of CDKN2B-AS1 was aberrantly upregulated in LSCC, and that the high expression levels of CDKN2B-AS1 was correlated with tumor staging and metastasis. Previous studies have shown that the upregulation of CDKN2B-AS1 was observed in various cancers, including gastric, cervical, glioma cells, hepatocellular carcinoma, prostate, oral, and other cancers. 19 Taken together, these results indicated that lncRNA CDKN2B-AS1 might function as an oncogene in LSCC and it could be a potential biomarker for LSCC.
Further, we found that CDKN2B-AS1 regulated cell viability, cycle, and apoptosis via miR-324-5p/ROCK1 axis. Several miRNAs including miR-96a, miR-449a, miR-199a, miR-323, miR-125a, miR-122-5p, as well as miR-191 have been identified as targets of CDKN2B-AS1.20-22 As far as we know, miR-324-5p/ROCK1 axis is a new target of CDKN2B-AS1. MiR-324-5p played very important roles in various biological processes. For instance, it regulated stemness, pathogenesis, and drug sensitivity in multiple myeloma cells via hedgehog signaling and the suppression of H5N1 virus replication by targeting the viral Protein PB1-F2 (PB1) and host CUE domain containing 2 (CUEDC2).23,24 Also, it could regulate cell proliferation, migration, invasion, and apoptosis in cancers by targeting tetraspanin 8 (TSPAN8), SMAD family member 7 (SMAD7), E26 transformation-specific 1 (ETS1), or specificity protein 1 (SP1).25-27 Here, we proved that miR-324-5p could also decrease the expression levels of ROCK1 by targeting its 3′UTR.
As one of the Rho regulated serine/threonine kinase, ROCK1 is essential for cell cycle progression, senescence, and tumorigenesis by the phosphorylation od various downstream effectors, which is considered an oncogene in cancers. 28 Many inhibitors of ROCK1 are developed in the hope of enhancing the delivery and efficacy of chemotherapy drugs and the effectiveness of radiotherapy for cancers. 29 Here, we also confirmed that the CDKN2B-AS1/ miR-324-5p/ROCK1 pathway regulated cell viability and cell apoptosis in LSCC. Several ceRNAs on ROCK1 have been discovered in cancers. Deng et al. 31 found that the lncRNA small nucleolar RNA host gene 1 (NHG1)/miRNA-101-3p/ROCK1 pathway regulated the proliferation, migration, invasion, and apoptosis of osteosarcoma cells. lncRNA differentiation antagonizing non-protein coding RNA (DANCR)/miR-335-5p and the miR-1972 /ROCK1 pathway regulated proliferation and metastasis in osteosarcoma.30,31 These findings deepen our understanding of ROCK1 in tumorigenesis.
In conclusion, this study revealed that the upregulated expression of lncRNA CDKN2B-AS1 promoted LSCC cell proliferation by miR-324-5p/ROCK1 axis. Also, the upregulation of lncRNA CDKN2B-AS1 could be a potential biomarker and a potential therapy target for LSCC.
Footnotes
Authors’ contributions
JW and FFL conceived and designed the experiments; YX analyzed and interpreted the results of the experiments; and LJM performed the experiments.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by National Key Research and Development Program of China (Grant No. 2017YFC0112500), Beijing Natural Science Foundation (Grant No. 7182036) and the Special Fund of the Pediatric Medical Coordinated Development Center of Beijing Hospitals Authority (Grant No. XTCX201823).
Ethics approval and consent to participate
The study was conducted according to the principles of the Declaration of Helsinki, and it was approved by the Ethics Committee of Beijing Tongren Hospital, Capital Medical University (Approval no. TRECKY2019-086). Informed consent was obtained from each patient before the study’s protocol was implemented.
Patient consent for publication
Not Applicable
Availability of data and materials
All data generated or analyzed during this study are included in this published article.
