Abstract
Objective:
Abundant evidence has illustrated that long non-coding RNA (lncRNA) plays a vital role in the regulation of tumor development and progression. Ectopic expression of a novel lncRNA, termed lnc-AGER-1, has been discovered in cancers, and this lncRNA was reported to exert an anti-tumor effect. However, its biological mechanism remains unelucidated in colorectal cancer.
Methods:
A total of 159 paired colorectal cancer specimens and adjacent tissues was applied to detect the expression of lnc-AGER-1 by the quantitative Real-time PCR (qRT-PCR), and a series of functional assays was executed to uncover the role of this lncRNA on colorectal cancer.
Results:
We found that the expression of lnc-AGER-1 in the tumor tissues was significantly down-regulated, while compared with adjacent normal tissues (0.0115 ± 0.0718 vs. 0.0347 ± 0.157; P < 0.0001). Also, lnc-AGER-1 was observably associated with clinical T status (r = −0.184, P = 0.024). Patients with advanced T status exerted a significantly lower level of lnc-AGER-1 than those with early T status (20.0% vs. 40.7%, P = 0.021). Over-expression of lnc-AGER-1 inhibited cell proliferation and migration efficiency, and induced cell cycle arrest at the G0/G1 phase, and promoted cell apoptosis. Further research proved that lnc-AGER-1 altered the expression of its neighbor gene, AGER, through acting as a competing endogenous RNA for miR-182 in colorectal cancer.
Conclusion:
lnc-AGER-1 has a suppressive role in colorectal cancer development via modulating AGER, which may serve as a target for colorectal cancer diagnosis and treatment.
Keywords
Introduction
Colorectal cancer (CRC) is the third most common malignant tumor and the second leading cause of cancer-related death worldwide.1,2 Although improvements are achieved on therapeutic strategies, the 5-year survival rate of patients with CRC remains lackluster. 3 Until now, the complicated molecular mechanisms involved in CRC development and metastasis have not been clearly illuminated. 4 Thus, a better understanding of the mechanisms and the identification of therapeutic biomarkers are priorities for improving CRC treatment.
Long non-coding RNAs (lncRNAs)—non-protein-coding transcripts with the length longer than 200 nucleotides—have been considered as special regulators in recent studies.5,6 lncRNAs are expressed in a highly context-specific pattern and are implicated in diverse biological processes, such as cellular proliferation, apoptosis, differentiation, migration, and/or invasion.7-9 Recently, dysregulation of lncRNAs has been reported to be linked to various human diseases, including cancer.10,11 Accumulating evidence suggests that lncRNAs might play vital roles in regulating cancer development and progression. 12 lncRNAs could regulate downstream gene expression through multifarious mechanisms. A considerable amount of literature has been published, documenting that lncRNAs can sponge specific miRNAs and indirectly influence gene expression at the post-transcriptional level.13,14 For instance, one study indicated that lncRNA H19 is upregulated in pancreatic cancer and contributes to cancer cell proliferation and migration through PFTK1 downstream Wnt signaling. 15 Another study presented that SNHG14 is serviced as a facilitator in CRC metastasis through targeting EZH2-regulated EPHA7. 16 lncRNA ZNFX1-AS1 promotes the CRC progression and metastasis by sponging miR-144 to regulate EZH2 expression. 17
Our research has established that lnc-AGER-1 could function as a biomarker to monitor lung cancer development. 18 This lncRNA modulates its nearby gene, AGER, through competitively binding to miRNA-185 and thus contributes to lung cancer progression. However, until now, there has been much less information about the effects of lnc-AGER-1 in CRC. The biological functions and underlying detailed mechanisms to this lncRNA in CRC remain unclear. Therefore, in this study, we aim to ascertain the potential role of lnc-AGER-1 in CRC development and progression; thus, a series of experiments were executed to demonstrate the way that lnc-AGER-1 affected CRC progression.
Materials and methods
Clinical samples
A total of 159 CRC tissues and their adjacent tissues (2 cm away from the tumor border) were obtained from patients who had undergone radical resection and were diagnosed with primary CRC in the Second Affiliated Hospital of Guangzhou Medical University from May 2005 to March 2012. All fresh tissues were collected then immediately frozen in liquid nitrogen and stored at −80°C until further analysis. None of the patients received any chemotherapy or radiotherapy before resection. The clinical characteristics of the patients were recorded after receiving their written informed consent. The study was approved by the Institutional Review Board at Guangzhou Medical University.
Cell culture
The human embryonic kidney (HEK) 293T cells and CRC cell lines, including HCT15, HCT116, DLD-1, SW480, and LOVO, were purchased from Cell Bank of Type Culture Collection of the Chinese Academy of Science (Shanghai Institute of Cell Biology, Shanghai, China). CRC cells were propagated in RPMI 1640 medium (Gibco Life Technologies, Carlsbad, CA, USA) and 293T cells were cultured in DMEM (Gibco). All cells were supplemented with 10% fetal bovine serum (FBS, Gibco) and 1% penicillin-streptomycin. Cells were incubated in a humidified atmosphere (SANYO Electric Co., Ltd., Japan) at 37°C with 5% CO2. The CRC cells were used for analyzing the expression of lnc-AGER-1, and 293T cells were utilized for lentivirus packaging.
Gene expression detection
Total RNA from tissues and cells was extracted using Bio pureTM isolation reagent (Bio Scientific, Austin, TX, USA) according to the manufacturer’s instructions, and was then quantified and reversely transcribed to cDNA by using the cDNA Synthesis Kit (Takara Bio Inc., Shiga, Japan). Quantitative real-time PCR (qRT-PCR) was conducted using TaqMan® Universal PCR Master Mix (Applied Biosystems, Foster City, CA, USA) on the ABI 7500 fast real-time PCR System (Applied Biosystems) to detect the expression level of a gene of interest. Relative expression of genes normalized to β-actin was calculated using the 2−ΔCt method. The primers for lnc-AGER-1 and β-actin were listed as follows: lnc-AGER-1: 5′-AACCAGGAGGAAGAGGAGGA-3′(forward), 5′-TTGGCAAGG TGGGGTTATAC-3′(reverse); β-actin: 5′-GGCGGCACCACCATGTACCCT-3′(forward), 5′-AGG GGC CGG ACT CGT CAT ACT-3′(reverse).
Lentivirus packaging and cell infection
The cDNA of lnc-AGER-1 was synthesized by I Gene Bio Co, Ltd (Guangzhou, China). The cDNA was then cloned into the lentivirus vector pEZ-Lv201. The lentivirus system was applied to introduce pEZ-Lv201-lnc-AGER-1 and empty pEZ-Lv201 into the HCT116, SW480, LOVO, HCT15, and DLD-1 cells. Overall, after the transfection of lentivirus particles, the cells expressing stable levels of lnc-AGER-1 and cells with empty control were used for further studies.
Cell proliferation assay
Cell proliferation was measured by the cell counting kit-8 (CCK8, Engreen, Beijing, China) assay according to the manufacture’s recommendation. Briefly, HCT15 and DLD-1 cells were seeded in triplicates in 96-well plates at 5000 cells/well. After being incubated for 24, 48, 72, and 96 h, a 10 ul CCK-8 solution in 100 ul fresh media cells in each well were added and further incubated for 2 h. The absorbance at 450 nm was measured on a microplate reader. The mean value of the wells with media alone was used as a background and was subtracted from the absorbances of the wells containing cells.
Cell migration and invasion assay
Transwell chambers (8.0 um pore size; Costar; Corning Incorporated, NY, USA), with or without Matrigel coating, were used to measure cell migration and invasion. Briefly, 1.0×105 transfected cells cultured in 200 ul RPIM 1640 without serum were plated on the upper surface of the transwell chambers while 500 ul RPIM 1640 with 10% FBS were added in the lower surface of the chambers. After being incubated in a humidified atmosphere at 37°C with 5% CO2 for 24 h, migration and invasion cells on the opposite side of the membrane were fixed with 4% paraformaldehyde (Asegene) for 30 min and stained with 1% crystal violet (Sigma, Guangzhou, China), whereas non-migrated and non-invaded cells were erased with cotton swabs. The number of remaining cells, as an index for evaluating migration and invasion ability, was counted under the microscope (Olympus, Tokyo, Japan) and analyzed with Image J software.
Flow cytometry assay
The flow cytometry analysis was used to establish whether lnc-AGER-1 influenced cell cycle and apoptosis. For cell apoptosis analysis, HCT15 and DLD-1 cell lines were performed with Annexin V-APC/7AAD Apoptosis Detection Kit (Abbexa, Cambridge, UK) according to the manufacturer’s instructions. For analysis of the cell cycle, cells were harvested and washed with ice-cold PBS, fixed with 70% cold ethanol overnight, then stationed with propidium iodide (PI) solution for 30 min on ice. The distribution of the number in each phase of the cell cycle was analyzed by flow cytometry (FACSCalibur; BD Biosciences).
MicroRNA mimics treatment and luciferase assay
Online bioinformatics analysis (http://www.mirbase.org/index.shtml, http://www.microrna.org/microrna/getMirnaForm.do) indicated that lnc-AGER-1 may function as a competing endogenous RNA (ceRNA) to modulate AGER expression by binding to microRNAs miRNA-4688-5p, miRNA-3171, miRNA-185, and miRNA-182.
Luciferase constructs containing the 3′-UTR of AGER and the mimics or inhibitors of these microRNAs were synthesized. Briefly, cells were seeded into 24-well plates and incubated until the cell confluence reached appropriate 80%. Then, miRNA mimics or inhibitors corresponding plasmids were co-transfected into the cells using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s protocols. Dual-Luciferase Reporter Assay System (Promega, Beijing, China) was applied to determine the luciferase activity according to the manufacturer’s instructions. Independent triplicate experiments were performed using each microRNA.
Statistical analysis
All data in this study were analyzed with SPSS 16.0 software. A comparison between groups was assessed by two-paired t-test, Chi-square test, or Wilcoxon test, as appropriate. The Spearman correlation analysis was performed to evaluate the correlation between two of the qualitative data. All P-values were two-sided, and P < 0.05 was statistically significant.
Results
lnc-AGER-1 is down-regulated in CRC
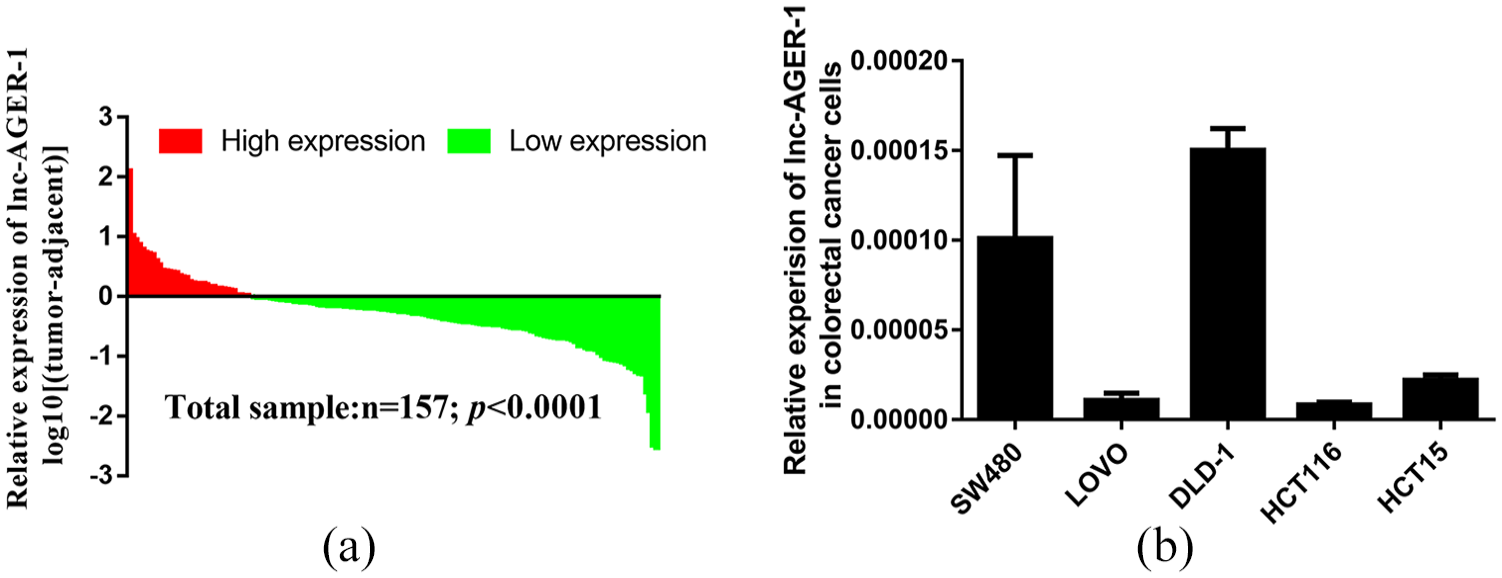
As Figure 1(a) shows, the expression of lnc-AGER-1 was significantly down-regulated in CRC tissues when compared to adjacent normal tissues (mean ± standard deviation: 0.0115 ± 0.0718 vs. 0.0347 ± 0.157; P < 0.0001). Among them, 76.4% (120/157) of CRC patients showed a lower expression level of lnc-AGER-1.
lnc-AGER-1 expression is associated with clinical T status
The demographics and clinical features of studied subjects are listed in Table 1. The expression status of lnc-AGER-1 was classified as “High” or “Low” based on whether the expression of lnc-AGER-1 in CRC tissues is higher or lower than that in adjacent normal tissues. From Table 2, we can see that the expression of lnc-AGER-1 was associated with the risk of clinical T status (r = −0.184, P = 0.024). Patients with advanced T status exerted significantly down-regulation status of lnc-AGER-1 than those with early T status (20.0% vs. 40.7%, P = 0.021). However, there was no notable association between lnc-AGER-1 expression and other clinicopathological characteristics (all P >0.05).
Baseline characteristic in the CRC population.
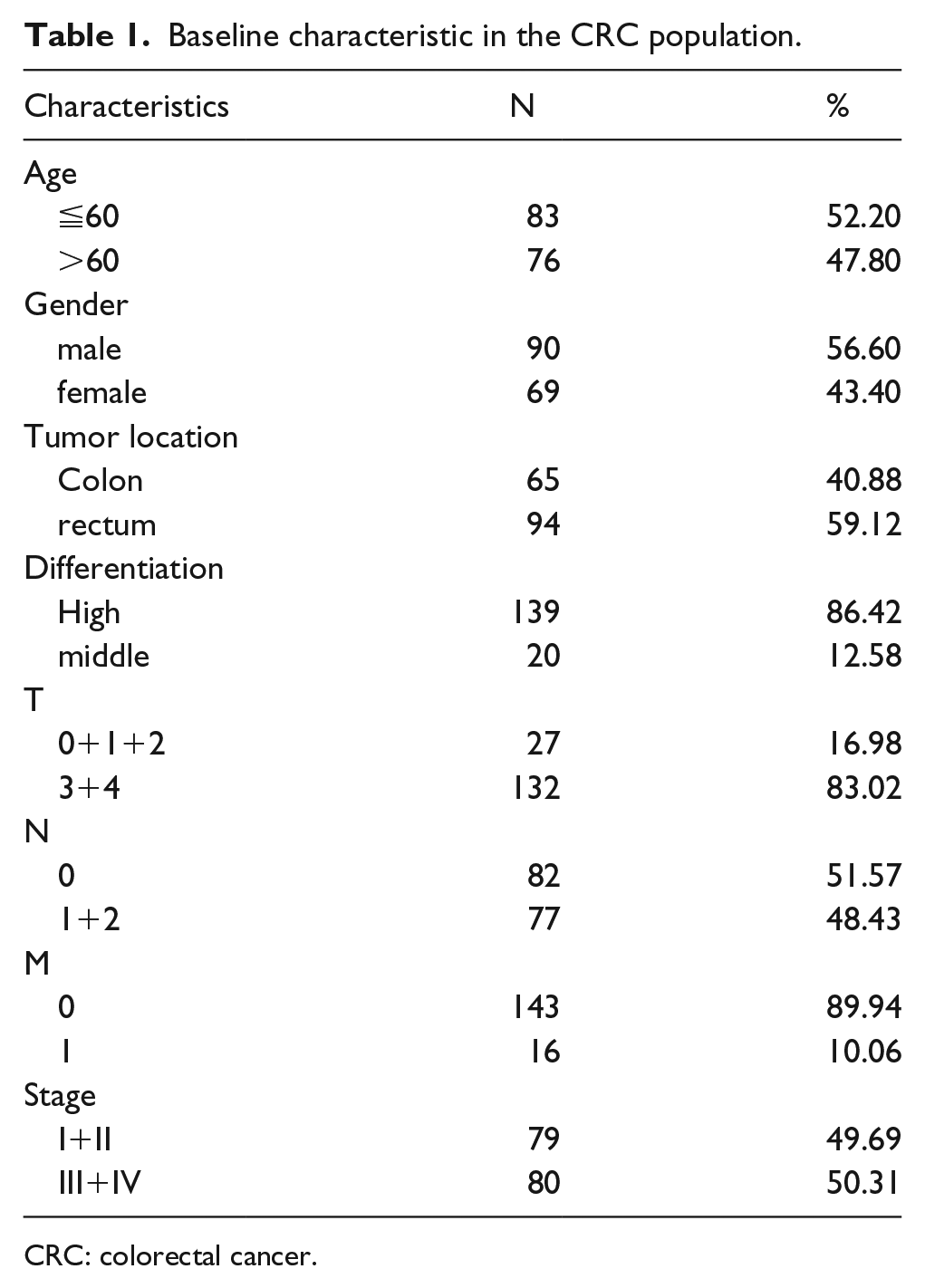
CRC: colorectal cancer.
Correlation between lnc-AGER-1 expression and clinicopathological features in CRC patients.
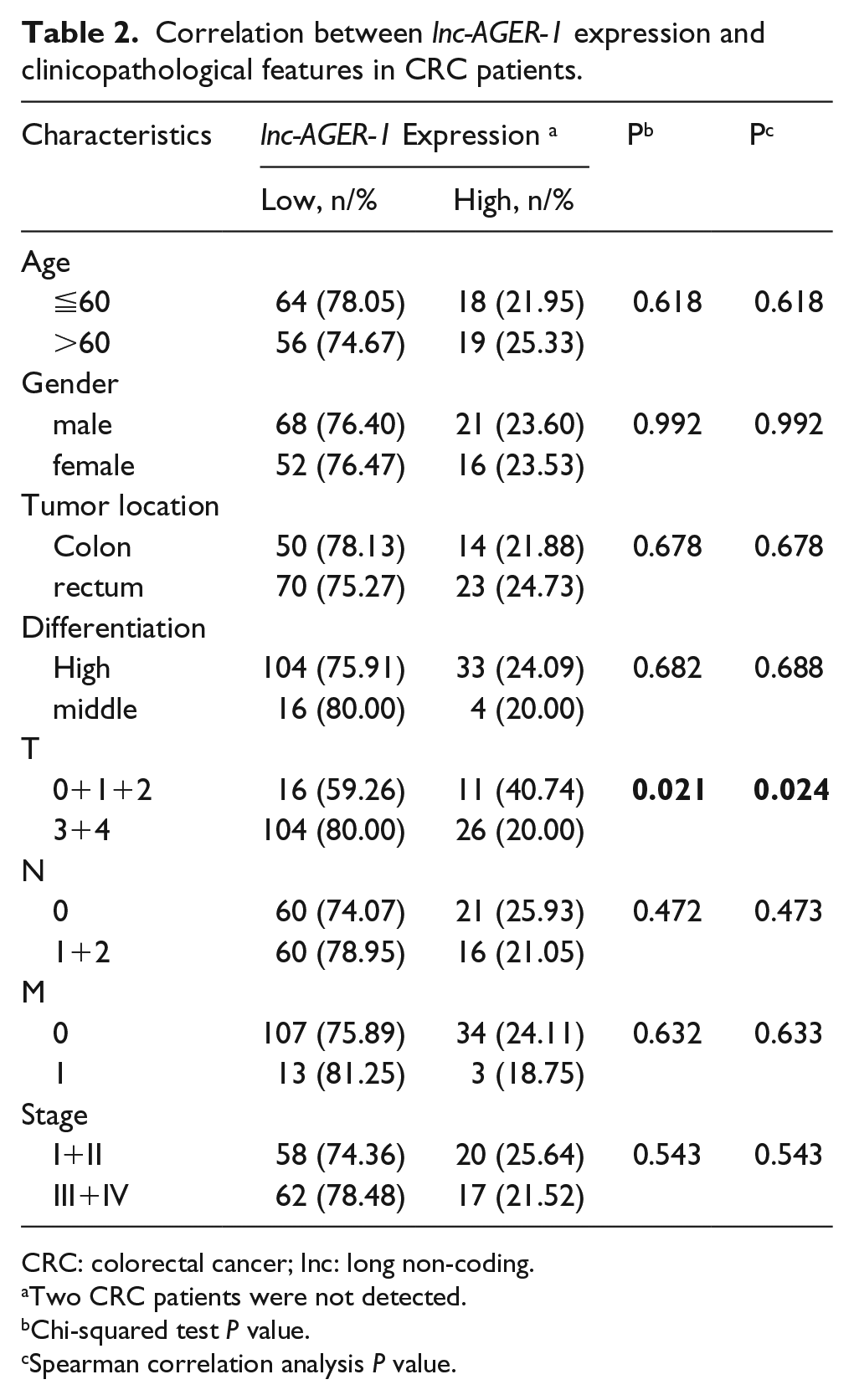
CRC: colorectal cancer; lnc: long non-coding.
Two CRC patients were not detected.
Chi-squared test P value.
Spearman correlation analysis P value.
lnc-AGER-1 inhibits cell proliferation
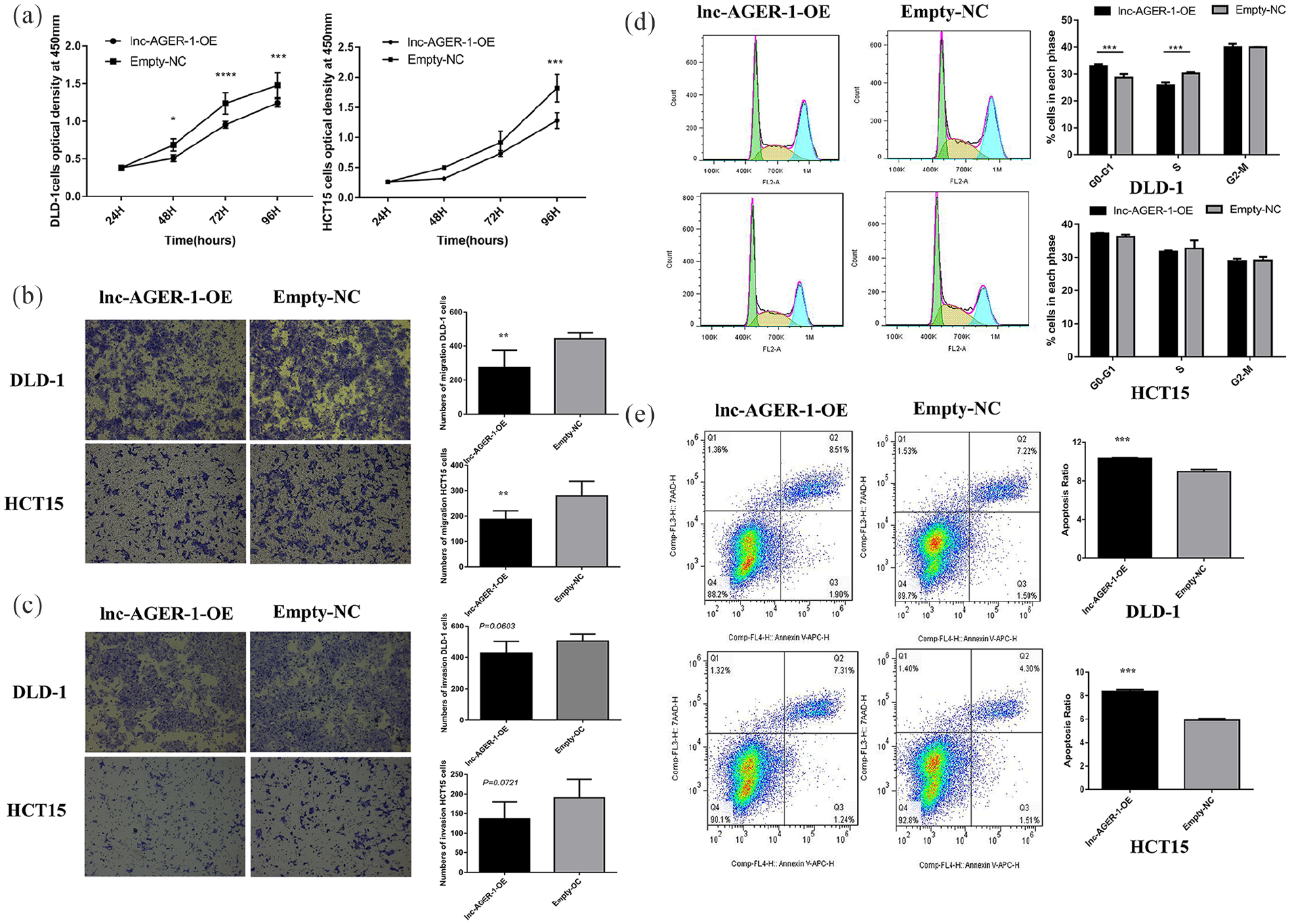
We assessed lnc-AGER-1 initial expression in five CRC cell lines and found that a relatively high expression level of lnc-AGER-1 in DLD-1 and SW480 cells, while a relatively low expression level was found in LOVO, HCT116, and HCT15 cells (Figure 1(b)). Also, after transfection of the lentivirus particles, HCT15 and DLD-1 over-expression cells and all control cells could be observed with green fluorescence, while SW480, LOVO, and HCT116 over-expression cells did not show green fluorescence (Figure S1(c)). Further verification by qRT-PCR also showed that the expression level of lnc-AGER-1 was significantly up-regulated in HCT15 and DLD-1 cells transfected with pEZ-Lv201-lnc-AGER-1 vectors. However, the expression level of lnc-AGER-1 showed no significant difference in SW480, LOVO, HCT116 cells transfected with pEZ-Lv201-lnc-AGER-1 vectors, when compared to cells transfected with empty pEZ-Lv201 vectors (Figure S1(b)). Therefore, DLD-1 and HCT15 cells were used to execute the lnc-AGER-1 gain-of-function studies in CRC. The stably expressed lnc-AGER-1 cells were successfully constructed as presented in Figure S1(a). CCK8 assays showed that the proliferation of cells expressing lnc-AGER-1 was significantly lower than the control groups both in DLD-1 and HCT15 cell lines (P <0.05, DLD-1; P <0.001, HCT15; Figure 2(a)).
lnc-AGER-1 suppresses cell migration, but not the invasion
To determine whether lnc-AGER-1 regulates cell migration and invasion, we carried out the transwell assays. As shown in Figure 2(b), the ability of DLD-1 and HCT15 cells to migrate was repressed in cells with over-expressed lnc-AGER-1 (P <0.01, DLD-1; P <0.01, HCT15; Figure 2(b)). However, the invasion ability through Matrigel™ did not show a significant difference between the two groups (P = 0.0603, DLD-1; P = 0.0721, HCT15; Figure 2(c)).
lnc-AGER-1 affects cell cycle and apoptosis
As shown in Figure 2(d), DLD-1 cells with lnc-AGER-1 highly expressed exhibited an accumulation at the G0/G1 phase (33.0% ± 0.729% vs. 28.7% ± 1.25%, P = 0.0002) and a notably concomitant reduction at the S phase(25.8% ± 1.01% vs. 30.3% ± 0.391%, P = 0.0001), indicating that the inducible expression of lnc-AGER-1 caused G0/G1 cell cycle arrest. However, none of these differences statistically occurred in HCT15 cell lines (G0/G1: 37.3% ± 0.156% vs. 36.2% ± 0.742%, P = 0.608; S: 31.7% ± 0.320% vs. 32.7% ± 2.468%, P = 0.679). Furthermore, compared to empty-control cells, overexpressed lnc-AGER-1 caused a noteworthy high rate of apoptosis both in DLD-1 and HCT15 cells (10.4% ± 0.0603% vs. 8.93% ± 0.252%, P = 0.0007, DLD-1; 8.37% ± 0.161% vs. 5.92% ± 0.115%, P <0.0001, HCT15; Figure 2(e)).
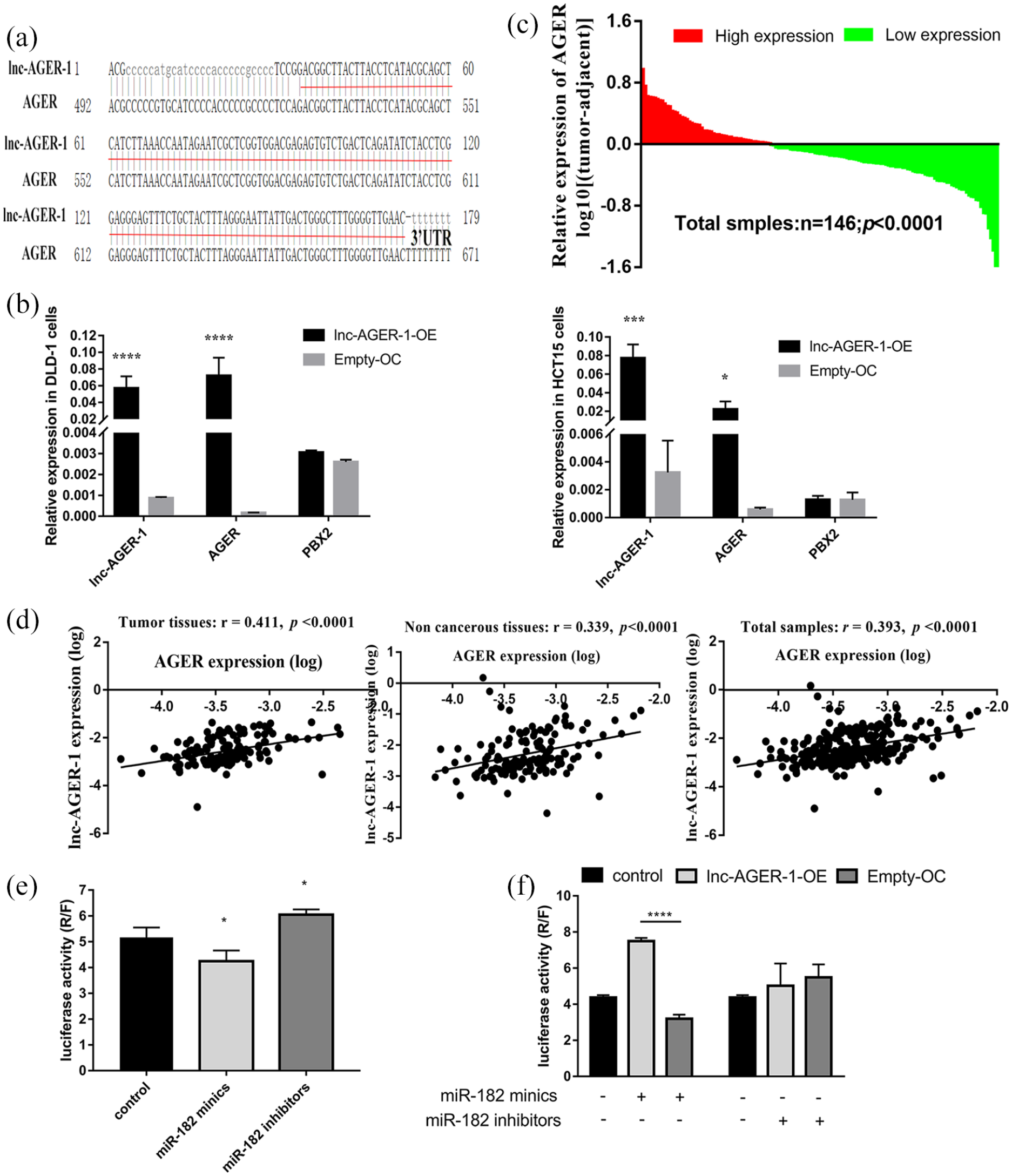
lnc-AGER-1 modulates AGER expression via sponging miR-182
To explore the potential mechanism of lnc-AGER-1, we blasted the sequences of lnc-AGER-1 and the AGER gene online. As seen in Figure 3(a), the same sequence was bound in the lnc-AGER-1 and the 3′UTR of AGER. Accumulating evidence suggests that lncRNA could play a cis-acting role to regulate their neighbor genes. Therefore, we evaluated the potential neighboring targets of lnc-AGER-1 both in transfected cells and CRC tissues and found that the cells that over-expressed lnc-AGER-1 altered the expression of AGER but not PBX2, compared to those control cells (Figure 3(b)). Then, we detected the AGER expression in CRC tissues by qPCR. In Figure 3(c) there is a clear trend of decreasing AGER expression in CRC tissues than that in adjacent normal tissues (mean ± SD: 0.000555 ± 0.000651 vs. 0.000763 ± 0.000898; P =0.0002). Also, the expression of lnc-AGER-1 was prominently correlated with that of AGER in CRC tissues (r = 0.411, P <0.0001), normal tissues (r =0.339, P <0.0001), and in total tissues (r =0.393, P <0.0001, Figure 3(d)). Further experiments revealed that lnc-AGER-1 may regulate AGER expression by binding to miRNA-182. Luciferase reporter assays showed that miR-182 (Figure 3(e)), but not miR-185, miR-3171, or miR-4688-5p (Figure S1) could bind to 3′-UTR of AGER because only miR-182 mimics significantly reduced the luciferase activity, and miR-182 inhibitors enhanced it (P <0.05 for all). The miR-182 mimics caused significantly less reduction of luciferase activity in cells that overexpressed lnc-AGER-1 than control cells, but the inhibitors had no obvious antileptic effect between the up-regulated and control cell lines (Figure 3(f)).
Discussion
Prior studies have noted the importance of lncRNAs in the development and progression of various malignancies, including CRC.19,20 Hence, a better understanding of the functions and biomechanism of lncRNAs may improve the diagnosis and treatment of patients with CRC. In this current study, we observed that lnc-AGER-1 was significantly downregulated in CRC tissues compared with the adjacent non-CRC tissues. Also, lnc-AGER-1 was notably related to clinical T status. The gain of function assays indicated that lnc-AGER-1 inhibited cancer cell proliferation and migration, and promoted cell apoptosis. Mechanistically, lnc-AGER-1 acted as a molecular sponge by competing for miR-182 and indirectly modulating the expression of its target gene AGER.
CRC is one of the most common fatal diseases worldwide and has a poor prognosis because of local tumor recurrence or distant metastasis. Choosing targeted therapy accompanied by conventional therapeutic approaches may eradicate residual micro-metastasis, then ameliorate the prognosis in CRC patients. 21 Although several reports have shown that immunotherapy has largely failed to exhibit clinically meaningful effects for the vast majority of CRC patients, little is well-elucidated regarding the complex immunological interplays in this solid tumor microenvironment. 22 Recently, many publications have reported that the innate immune system is closely linked to CRC progression.23,24 In the previous study, we identified that an innate immune-related lncRNA, termed lnc-AGER-1, exerted an inhibitory action on lung carcinogenesis. 18 Similarly, in the current study, we found this lncRNA acted as a suppressor of CRC development.lnc-AGER-1 showed down-regulated expression in CRC tissues, and the low-expression of lnc-AGER-1 was correlated with advanced T status. The over-expression of lnc-AGER-1 could significantly inhibit CRC cell proliferation. Lnc-AGER-1 also altered cell cycle arrest at the G0/G1 phase with fewer cells in the S phase, suggesting that lnc-AGER-1 impairs cell division. Also, we found that lnc-AGER-1 promoted cell apoptosis. Notably, abnormal lnc-AGER-1 expression is sufficient to inhibit migration but not the invasion of CRC cells. Unlike migration, cell invasion needs gene expression changes causing proteolysis and degradation of the matrix, which fail to achieve by lnc-AGER-1. 25 Cell invasion is close to cancer metastasis and needs synthesis and secretion of degradation enzymes. This may be why lnc-AGER-1 expression correlates with T but not N and M in patients. Therefore, these findings suggest that lnc-AGER-1 plays an anti-colorectal oncogenesis, stimulating a new research direction and therapeutic option.
It is well known that ceRNA is a common mechanism of lncRNAs, among which lncRNA modulates a targeted mRNA via sponging a certain miRNA.14,26 After bioinformatics analysis and mechanism detection, we determined that lnc-AGER-1 acts as an miR-182 sequestrate and thereby increases the expression of its neighbor AGER. Accumulated evidence has shown that miR-182 was upregulated in CRC and exerted its oncogenic role on cell proliferation, invasion, and tumor growth.27,28 lnc-AGER-1 is located on chromosome 6p21.32 that was the susceptible region for cancers,29,30 and its upstream 417 bp is the neighbor AGER gene. AGER is a pattern recognition receptor that binds numerous endogenous and exogenous ligands and plays a critical role in regulating the innate immune response. 31
Studies have illustrated that AGER plays an important role in cancer origin, invasion, and metastasis. 32 Interestingly, dysregulation of AGER had been related to CRC. A study reported that compared to paired normal mucosa, the expression levels of AGER in tumor epithelia were significantly lower. 33 In this study, we also found that AGER was down-regulated in CRC patients. Chronic inflammation is one of the hallmarks for cancers, which fuels and sustains disease progression and neoplastic transformation.34,35 For CRC, inflammatory bowel disease has been identified as a significant risk for malignant transformation. 36 Cells of the innate immune system can be the primary effector in this inflammatory process, including persistent infections and sterile inflammation. 37 While AGER is expressed in innate immune cells, it has enabled the recognition of allergens and thus activated innate immune responses. Together, the host’s own immune system may eradicate the tumor cells that rooted in the continuous inflammatory signals. Based on previous evidence, and combined with our experimental results, it is plausible to speculate that lnc-AGER-1 acted as a ceRNA to regulate AGER, and thereby exerted a suppressor role in colorectal oncogenesis.
There are some limitations in the present study, which are of concern. A follow-up study was not executed, and the potential prognostic role of lnc-AGER-1 needs to be evaluated in further studies. Although we have identified miR-182 as a target miRNA of lnc-AGER-1, lnc-AGER-1 may have other potential regulatory mechanisms. Further research is warranted to study how lnc-AGER-1 affects colorectal oncogenesis and progression. Despite these drawbacks, our study highlights the potential role of lnc-AGER-1 in CRC development.
In summary, the findings from the current study have demonstrated that lnc-AGER-1 is down-regulated in CRC by modulating gene AGER. The over-expression of lnc-AGER-1 inhibits CRC proliferation and migration, induces cell cycle arrest at the G0/G1 phase, and promotes apoptosis. Thus, lnc-AGER-1 could be a promising therapeutic target and novel molecular biomarker for CRC.
Supplemental Material
figure_S1-R1 – Supplemental material for Long non-coding RNA AGER-1 inhibits colorectal cancer progression through sponging miR-182
Supplemental material, figure_S1-R1 for Long non-coding RNA AGER-1 inhibits colorectal cancer progression through sponging miR-182 by Mingzhu Lin, Yinyan Li, Jianfeng Xian, Jinbin Chen, Yingyi Feng, Chun Mao, Yujie Pan, Zhi Li, Yuyuan Zeng, Lei Yang, Jiachun Lu, Yisheng Wei and Fuman Qiu in The International Journal of Biological Markers
Footnotes
Author contributions
FMQ and WYS designed the study. MZL and YYL performed most of the experiments. JFX and JBC revised the paper writing. YYF analyzed the qRT-PCR results. CM, YJP, ZL, and YYZ collected the tissue samples. LY and JCL provided technical support. FMQ and MZL wrote the paper. MZL and YYL contributed equally to this work. All authors read and approved the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the National Natural Science Foundation of China Grants 81602289, 81872127 (F. Qiu); 81872694, 81673267, 81473040 (J. Lu); 81402753, 81672303, 81871876 (L. Yang); the National Key R&D Projects (2016YFC0903700), Local Innovative and Research Teams Project of Guangdong Pearl River Talents Program 2017BT01S155 (J. Lu); Yangcheng Scholar Grants 1201541589 (J. Lu); Guangzhou Science Research Program General Project Grant 201707010123 (F. Qiu); Guangzhou Municipal Scientific Research Project Grant 1201630073 (F. Qiu); Guangdong High School Young Innovative Talents Project Grant 2015KQNCX136 (F. Qiu).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
