Abstract
Background:
RP11-108K3.2 was recently identified as a novel long non-coding RNA (lncRNA) transcript, and several single nucleotide polymorphisms (SNPs) have been identified in its coding region. This study aimed to explore the associations of tagSNPs in RP11-108K3.2 with the risk of colorectal cancer and their effects on its expression.
Methods:
A total of 821 colorectal cancer cases and 857 healthy controls were enrolled into this two-stage case-control study. Demographic characteristics and lifestyle information were collected by a validated questionnaire. Six tagSNPs were genotyped by using Sequenom MassARRAY platform. A total of 71 additional colorectal cancer cases were recruited, of which the genotypes of potential polymorphisms and the RP11-108K3.2 expression levels were determined.
Results:
In the discovery set, only the rs2470151 C/T polymorphism was found to have a promising association with the risk of colorectal cancer, and this polymorphism was further replicated in the validation set with a significantly decreased risk of colorectal cancer (adjusted odds ratio 0.73; 95% confidence interval 0.55, 0.97). Combined discovery set and validation set together, this negative association was found both in the heterozygote codominant model and the dominant model. Furthermore, colorectal cancer patients carrying rs2470151 CT/TT genotypes had a marginally lower RNA expression of RP11-108K3.2 than those carrying the CC genotype. Stratified analyses showed the association between rs2470151 and the risk of colorectal cancer were influenced by family history of cancer, smoking, alcohol consumption, and tea drinking.
Conclusions:
These findings suggest that RP11-108K3.2 rs2470151 had a significant association with the risk of colorectal cancer; this may help to predict the susceptibility of colorectal cancer in Chinese populations.
Introduction
Colorectal cancer (CRC) is the third most common cancer and ranks fourth in cancer deaths globally, accounting for roughly 1.2 million new cases and 600,000 deaths every year. 1 Since CRC is a significant threat to public health, a substantial number of studies have investigated its etiology. Apart from age and sex, the following factors have been identified and established to be associated with the risk of CRC in epidemiological studies: family history of CRC, inflammatory bowel disease, smoking, excessive alcohol consumption, high consumption of red and processed meat, obesity, and diabetes.
A substantial number of genetic components play important roles in CRC susceptibility. Elevated genome-wide association studies have indicated that single nucleotide polymorphisms (SNPs) were involved.2-4 Among the numerous SNPs demonstrated to be related to CRC, variants located on the coding region of long non-coding RNAs (lncRNAs) have drawn attention. For example, a two-stage case-control study suggests that genetic variants of HOTAIR rs7958904 may act as a potential biomarker for predicting the risk of CRC. 5
lncRNAs are a large category of transcripts without protein-coding potential, which are >200 nt in length. Although the definite biological roles of most lncRNAs are still unknown, a growing number of literature suggests that they have a wide variety of roles in chromatin remodeling, transcriptional activation, transcriptional repressor, and RNA degradation.6-8 The genetic polymorphisms on lncRNA coding regions may have implications for many human diseases, including cancer, by changing their expression levels. 9
RP11-108K3.2 was recently identified as a novel lncRNA, and its coding region maps on the 15q21 chromosome, which is overlapped by the cytochrome P450 family 19 subfamily A member 1 (CYP19A1) gene. CYP19A1 influences estrogen-metabolizing enzymes and may change the CRC risk through an inflammation-related mechanism. A previous population-based, case-control study suggests that several polymorphisms in the CYP19A1 gene are associated with the risk of CRC, 10 which may have a stronger effect in women due to the influences of sex hormone. 11
In this study, we hypothesized that tagSNPs located on RP11-108K3.2 may be involved in the susceptibility to CRC. To test this hypothesis, we conducted a two-stage, case-control study to explore the association between the tagSNPs of RP11-108K3.2 and the risk of CRC in a Chinese population. We further analyzed the expression levels of RP11-108K3.2 among different genotype carriers if the polymorphism has been proven to be associated with the risk of CRC.
Materials and methods
Study subjects
This study was performed in Jiashan County, Zhejiang Province, China, based on a population screening program. All CRC cases were histopathologically confirmed by the Jiashan Institute of Cancer Prevention and Treatment since 1989. Healthy controls were recruited in parallel from the same population and matched to the cases by age (± 5 years), gender, and residential area. Subjects were excluded if they self-reported a history of familial adenomatous polyposis, ulcerative colitis, or Crohn disease. The recruitment details of the study population have been described previously. 12 Briefly, this two-stage, case-control study ultimately enrolled 320 CRC cases and 319 healthy controls in the discovery set; and 501 CRC cases and 538 healthy controls in the validation set. A face-to-face interview was conducted by trained interviewers using a structured questionnaire to collect the information of demographic characteristics (e.g. age, gender, and body mass index (BMI)), family history of cancer and lifestyle factors (e.g. smoking, alcohol consumption, and tea drinking). The definition of a family history of cancer was a first- or second-degree relative having had cancer at any site. Smokers were defined as those who smoked at least one cigarette per day for more than 1 year, or >300 cigarettes in less than 3 months. Alcohol drinkers were defined as those who consumed at least one alcohol drink per day for more than 3 months. Tea drinkers were defined as those who consumed more than two times of tea drink per week for more than 2 months. Blood sample of each participant was collected and stored at −80°C for DNA isolation.
This study also recruited 71 CRC tissues from patients who underwent surgical operations at the Department of Gastrointestinal Surgery, Hangzhou First People’s Hospital from July 2013 to December 2013. The tissues were preserved in the RNA Later® Stabilization Solution (Invitrogen, Carlsbad, CA, USA) and stored at −80°C.
The written informed consent was obtained from all subjects and this study was approved by the Medical Ethical Committee of Zhejiang University School of Medicine.
SNP selection and genotyping
We selected the tagSNPs that were located on the 2000bp upstream region of RP11-108K3.2 with pairwise correlation r2>0.80 and minor allele frequencies (MAF) > 0.10. Genomic DNA was extracted from peripheral blood by using the modified salting-out procedure. Genotyping was performed by the MassARRAY molecular weight array analysis system (BioMiao Biological Technology Co., Beijing, China). A negative control was included in each 96-well plate for quality control. The 5% of samples were randomly selected for repeated detection and the concordance rate was >99%. Primer information of SNPs of RP11-108K3.2 is shown in Supplementary Table 1.
RNA isolation and quantitative RT-PCR
Total RNA was isolated using a homogenizer (IKA® Works Guangzhou, China) and TRIzol reagent (Invitrogen, Carlsbad, CA, USA) and then purified using the RNeasy Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s protocol. A reverse transcription real-time PCR (RT-PCR) was conducted with StepOnePlus instrument (Applied Biosystems, Foster City, CA, USA) to synthesize cDNA. We performed quantitative real-time PCR in StepOnePlus System (Applied Biosystems, Foster City, CA, USA) using SYBR Green Master Mix (Vazyme, Nanjing, China) to quantify relative RP11-108K3.2 expression levels. The specific forward primer and reverse primer were 5′-GCTGGAACCCAATGGATGA-3′ and 5′-CCTGGAGAATGTAGCAATGGAA-3′. The β-actin was selected as the endogenous control. The specific forward primer and reverse primer were 5′-GTGGCCGAGGACTTTGATTG-3′ and 5′-CCTGTAACAACGCATCTCATATT-3′. All procedures were carried out in triplicate. For quantitative results, the expression of RP11-108K3.2 was calculated according to fold change using the 2−ΔCt method.
Statistical analysis
The Student’s t-test and Pearson’s χ2 test were applied to evaluate the differences of characteristics between cases and controls. Hardy–Weinberg equilibrium was assessed by goodness-of-fitχ2 test. The association between tagSNPs and the risk of CRC was estimated by calculating the adjusted odds ratios (ORs) and corresponding 95% confidence intervals (CIs) in multivariate logistic regression model. Those tagSNPs with a P-value less than 0.20 in the association study of discovery set were further analyzed in the validation set. Pooled analyses were also conducted to estimate the combined effect of these two sets. Different expression levels of lncRNA in samples of different genotypes were analyzed using ANOVA analysis or t-test.
All analyses were performed with SAS software (version 9.2; SAS Institute, Inc., Cary, NC, USA) with two-sided P-values. P < 0.05 was considered statistically significant.
Results
Characteristics of the study population
In the discovery set, the mean age was 65.8 years (range, 35–87 years) for the cases, and 65.3 years (range, 34–89 years) for the controls. In the validation set, the mean age was 62.7 years (range, 32–95 years) for the cases, and 62.3 years (range, 33–92 years) for the controls (Supplementary Table 2). CRC patients had a higher percentage of a family history of cancer (P < 0.001 in the discovery set, P = 0.039 in the validation set) and a lower proportion of tea drinking (P = 0.030 in the validation set) than controls. There was no significant difference in other demographic characteristics (age, gender, BMI) or lifestyle factors (smoking, alcohol drinking) between cases and controls.
Association between RP11-108K3.2 tagSNPs and the risk of colorectal cancer
As shown in Table 1, six tagSNPs were included in the discovery set (rs2470151, rs3751591, rs3751592, rs77348531, rs17602308, and rs75106900), all of which conformed to the request of Hardy–Weinberg equilibrium. However, only the CT genotype of rs2470151 found a promising association after adjusting for age, gender, BMI, family history of cancer, smoking, alcohol consumption, and tea drinking (adjusted OR 0.74; 95% CI 0.51, 1.07; P = 0.106). The other tagSNPs were observed to have no meaningful association with the risk of CRC (P > 0.20).
The association between tagSNPs of RP11-108K3.2 and the risk of colorectal cancer in the discovery set.
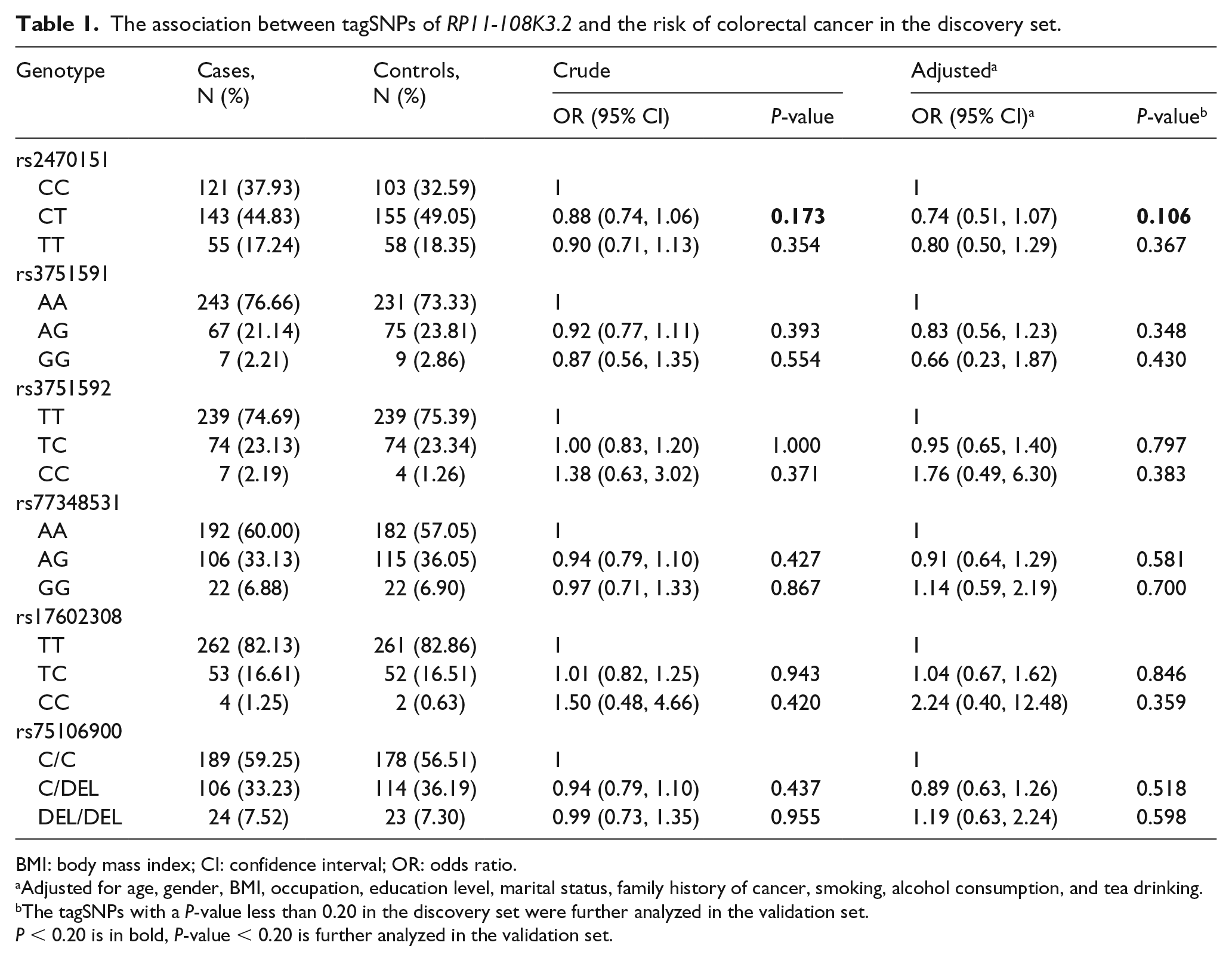
BMI: body mass index; CI: confidence interval; OR: odds ratio.
Adjusted for age, gender, BMI, occupation, education level, marital status, family history of cancer, smoking, alcohol consumption, and tea drinking.
The tagSNPs with a P-value less than 0.20 in the discovery set were further analyzed in the validation set.
P < 0.20 is in bold, P-value < 0.20 is further analyzed in the validation set.
The validation set carried on with the RP11-108K3.2 rs2470151 polymorphism, which was observed to be significantly associated with the risk of CRC in the validation set and the combined set (Table 2). In the validation set, subjects with the CT genotype had a significantly decreased risk of CRC compared with subjects with the CC genotype by a co-dominant model (adjusted OR 0.73; 95% CI 0.55, 0.97; P = 0.030), and this association was confirmed in the combined set (adjusted OR 0.73; 95% CI 0.58, 0.92; P = 0.007). However, the rs2470151 TT genotype did not show a significant difference, which may be due to the low genotype frequency. Also, we applied the dominant model to evaluate the effect of rs2470151 on the CRC risk; it still indicated the CT/TT genotype as being a protective factor for CRC in the combined set (adjusted OR 0.79; 95% CI 0.64, 0.97; P = 0.025). A subgroup analysis by gender is shown in Supplementary Table 3, which indicates that the association was stronger in females than males.
Significant association between RP11-108K3.2 rs2470151 and the risk of colorectal cancer.
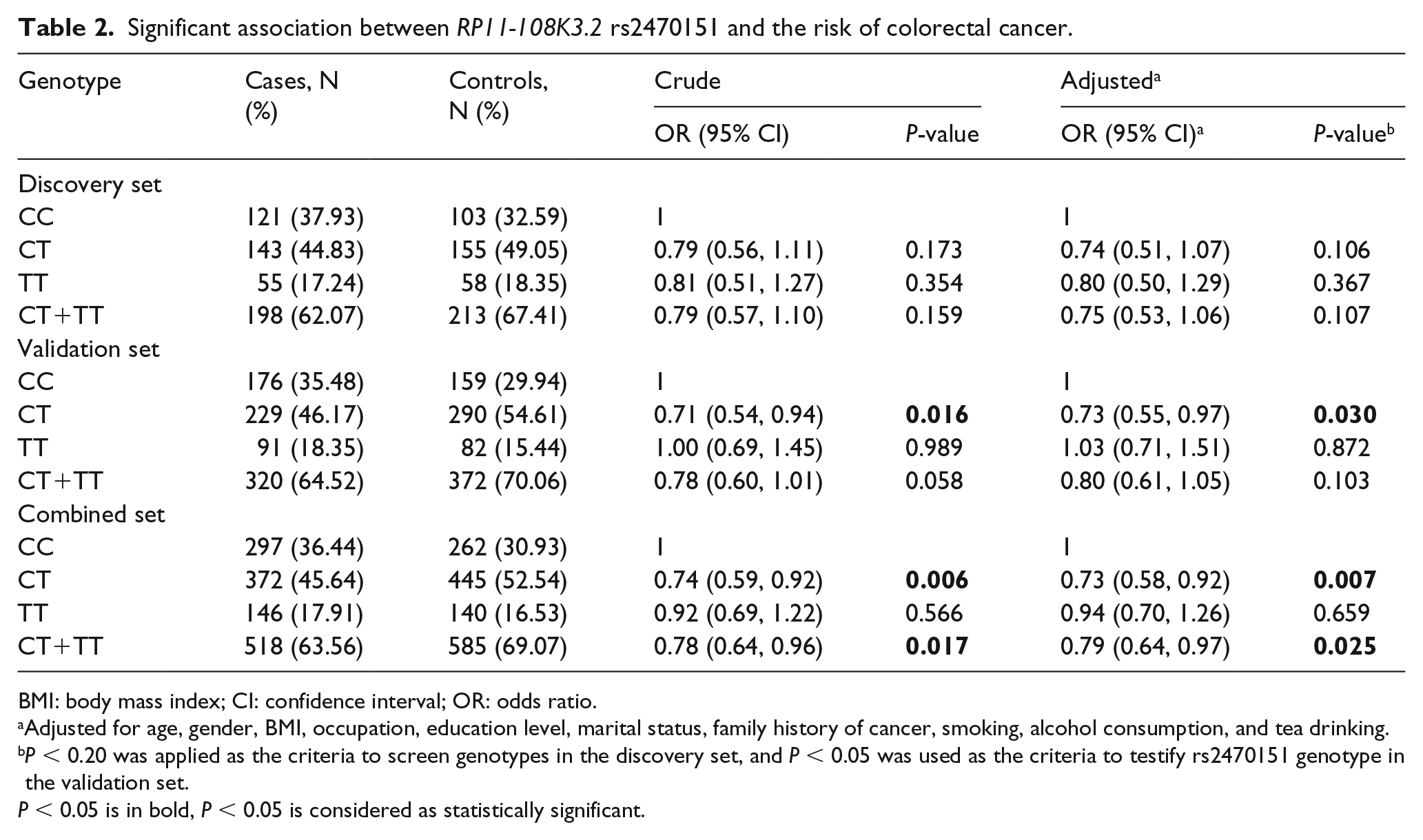
BMI: body mass index; CI: confidence interval; OR: odds ratio.
Adjusted for age, gender, BMI, occupation, education level, marital status, family history of cancer, smoking, alcohol consumption, and tea drinking.
P < 0.20 was applied as the criteria to screen genotypes in the discovery set, and P < 0.05 was used as the criteria to testify rs2470151 genotype in the validation set.
P < 0.05 is in bold, P < 0.05 is considered as statistically significant.
Correlation between rs2470151 genotypes and RP11-108K3.2 expression
Of the 71 CRC tissue samples, CC genotypes were identified in 18 cases (25.4%), CT genotypes in 40 cases (56.3%), and TT genotypes in 13 cases (18.3%). As shown in Figure 1, the relative expression level of RP11-108K3.2 was marginally higher in individuals carrying CC genotypes than in those carrying CT/TT genotypes (fold change = 1.765). The association between RP11-108K3.2 expression levels and clinic pathological characteristics was not statistically significant (Supplementary Table 4).
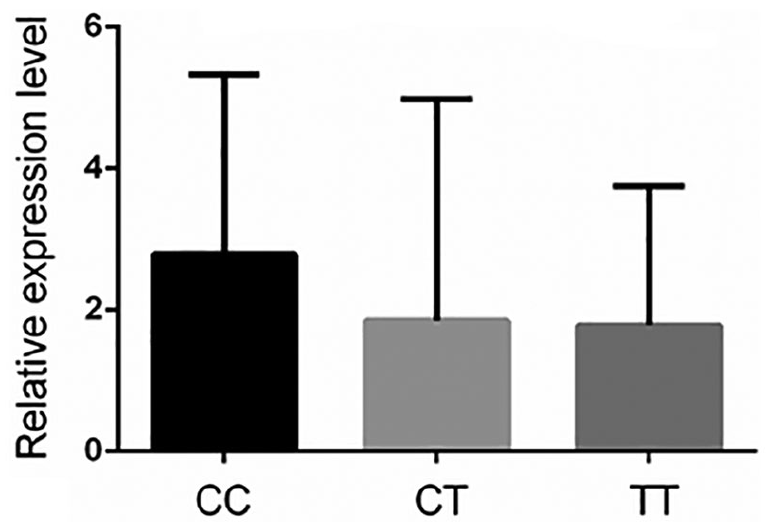
Relative expression level of RP11-108K3.2 mRNA grouped by rs2470151 genotypes in 71 colorectal cancer tissues.
Stratified analysis by selected variables between rs2470151 variant and the risk of CRC
Stratified analyses by selected variables were conducted for the RP11-108K3.2 rs2470151 variant (Table 3). In the subjects without a family history of cancer, CT/TT carriers of rs2470151 had a lower risk of CRC compared with CC carriers (adjusted OR 0.77; 95% CI 0.62, 0.98; P = 0.029). In the combined set, smokers with the CT/TT genotypes had a lower risk of CRC than those with the CC genotype (adjusted OR 0.64; 95% CI 0.45, 0.92; P = 0.014). Non-drinkers with the CT/TT genotypes had a significantly decreased risk of CRC compared with those with the CC genotype (adjusted OR 0.77; 95% CI 0.61, 0.99; P = 0.039). In the non-tea drinkers, carrying the CT/TT genotypes seemed to be a protective factor for CRC (adjusted OR 0.70; 95% CI 0.53, 0.92; P = 0.011).
Risk of colorectal cancer associated with rs2470151 stratified by family history of cancer, smoking, alcohol consumption, and tea drinking.
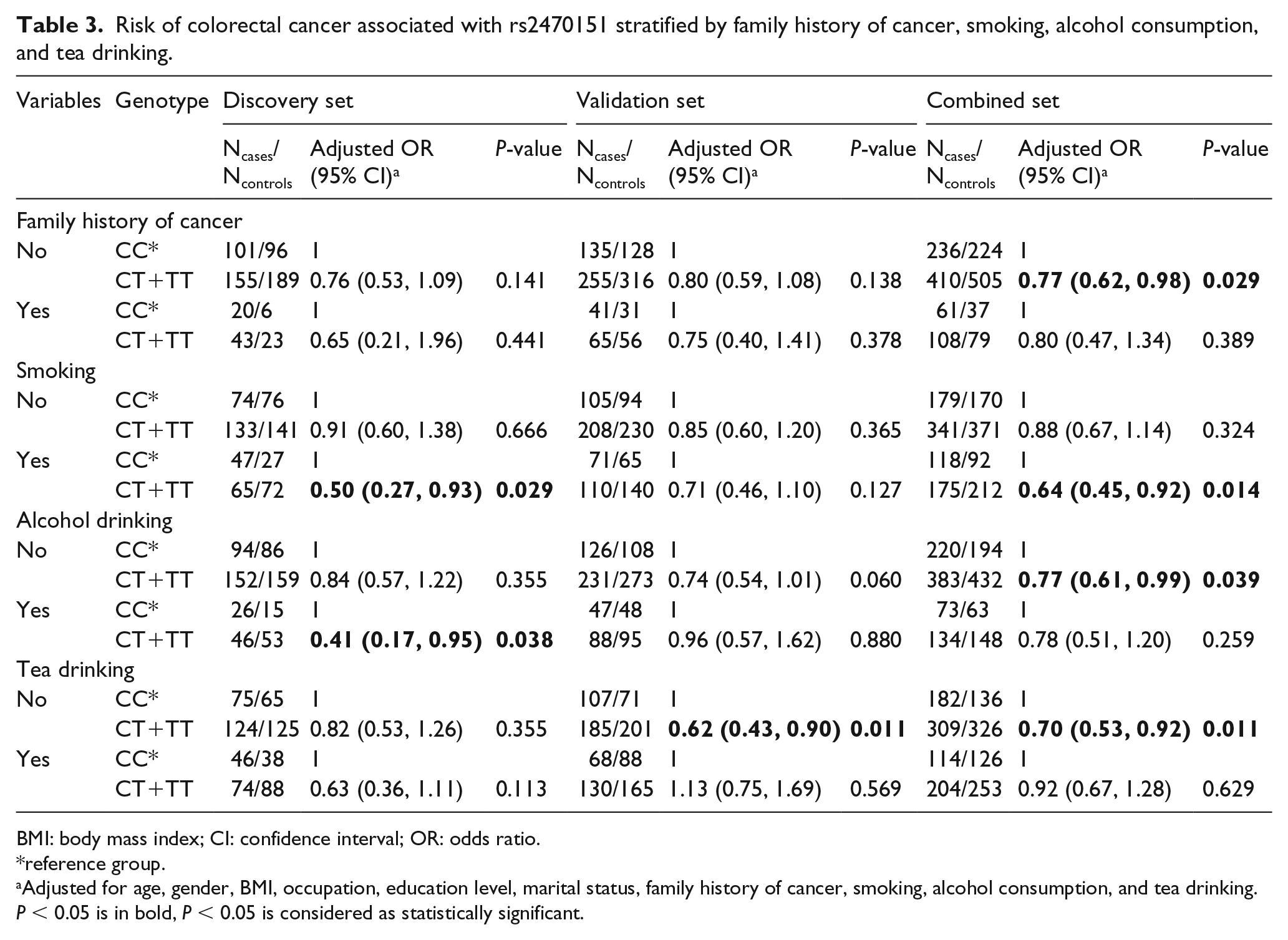
BMI: body mass index; CI: confidence interval; OR: odds ratio.
reference group.
Adjusted for age, gender, BMI, occupation, education level, marital status, family history of cancer, smoking, alcohol consumption, and tea drinking.
P < 0.05 is in bold, P < 0.05 is considered as statistically significant.
Discussion
CRC is a major cause of morbidity and mortality worldwide. 13 One of the fundamental processes driving the initiation and progression of CRC is the accumulation of a variety of genetic and epigenetic changes in the colorectum. 8 Here, we explored the associations between the tagSNPs on RP11-108K3.2 and the risk of CRC, and identified rs2470151 as a potential biomarker for CRC through a two-stage, case-control study. This association between rs2470151 and the risk of CRC was influenced by selected variables including a family history of cancer, smoking, alcohol consumption, and tea drinking. Moreover, carrying the CC genotype of rs2470151 may increase RP11-108K3.2 expression levels compared to the CT/TT carriers. To the best of our knowledge, this is the first study to explore the effect of RP11-108K3.2 on the risk of CRC.
lncRNAs are implicated in serial steps of cancer development, interacting with DNA, RNA, protein molecules, and/or their combinations, acting as an essential regulator in chromatin organization, and in transcriptional and post-transcriptional regulation. 14 There is increasing evidence revealing that SNPs on lncRNAs may be correlated with cancer, which may influence the expression of RNA or the stability of RNA conformation, resulting in the modification of its interacting partners. SNP variants on lncRNAs were observed to function in different cancer types, including gastric cancer,15,16 lung cancer,17,18 prostate cancer,19,20 and breast cancer.21,22 Several studies also have reported that SNPs on lncRNAs are related to the risk of CRC. For example, Ling et al. 23 reported that a novel lncRNA CCAT2 was highly overexpressed in microsatellite-stable CRC, and the SNP status affected its expression, and the risk allele G of rs6983267 produced more CCAT2 transcript. Li et al. 24 demonstrated that rs13252298 and rs1456315 on the lncRNAPRNCR1 were significantly associated with the decreased risk of CRC. Haiman et al. 25 revealed that rs6983267 mapping on the 8q24 chromosomal region was also related to the risk of CRC. Based on the results of current studies, we hypothesized that the SNP rs2470151 may influence the expression of RP11-108K3.2, resulting in its subsequent biological processes. In our study, we observed that the CT/TT genotype carriers of rs2470151 polymorphism (C>T) had a significantly decreased risk of CRC compared with the subjects with CC genotype. Furthermore, different genotype carriers had different expression levels of RP11-108K3.2 in CRC tissues; however, the difference was not significant.
In previous epidemiological studies, there was quite a lot of evidence supporting the association between polymorphisms and different lifestyle factors, including smoking, alcohol, body fatness, diet, physical activity, and medications, in CRC.26-29 Bailie et al. 30 also described several lifestyle factors associated with higher colorectal disease risk, most notably smoking and alcohol. Besides the factors above, family history is a well-established risk factor for CRC. 31 In our study, we also demonstrated that the association between the risk of CRC and RP11-108K3.2 rs2470151 may be influenced by selected variables, including a family history of cancer, alcohol consumption, and tea drinking through stratified analyses.
There are several limitations in this study. First, although we observed a significant association between rs2470151 and the risk of CRC, how the genetic variability at this locus influences RP11-108K3.2 to function still remains to be fully elucidated. Second, the recall bias of lifestyle factors was inevitable in a case-control study. Third, most of the clinical information of the CRC cases was absent due to the population-based study. Fourth, the associations might have insufficient statistical power due to the relatively moderate sample size, and further studies with larger samples are expected to confirm our findings, especially in the subgroup analysis and correlation study between rs2470151 genotypes and RP11-108K3.2 expression levels.
In summary, we have provided initial evidence that subjects carrying the rs2470151 CT/TT genotype had significantly lower CRC risk than those with the CC genotype. We also indicated that the CT/TT genotype of rs2470151 may have an effect on the lower expression level for RP11-108K3.2. This association between rs2470151 polymorphism and the risk of CRC may be influenced by a family history of cancer, alcohol consumption, and tea drinking. These results suggest that the SNP rs2470151 on RP11-108K3.2 may help to predict the CRC risk in Chinese populations.
Supplemental Material
Supplementary_Table_1 – Supplemental material for Polymorphisms of a novel long non-coding RNA RP11-108K3.2 with colorectal cancer susceptibility and their effects on its expression
Supplemental material, Supplementary_Table_1 for Polymorphisms of a novel long non-coding RNA RP11-108K3.2 with colorectal cancer susceptibility and their effects on its expression by Danjie Jiang, Mingjuan Jin, Ding Ye, Yingjun Li, Fangyuan Jing, Xiaocong Zhang, Qilong Li and Kun Chen in The International Journal of Biological Markers
Footnotes
Author contributions
Danjie Jiang and Mingjuan Jin contributed equally to this work.
Declaration of conflicting interest
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the National Natural Science Foundation of China (No.81673262 and 81373087), the National Key Basic Research Program (973 program No.2015CB554003), the Natural Science Foundation of Zhejiang Province, China (No.LQ15H260001), and the Zhejiang Chinese Medical University Foundation (No.2019ZR03).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
