Abstract
Introduction:
MicroRNA373 was highly expressed in many tumors including esophageal cancer. However, its molecular mechanism is still unclear, especially epigenetic modification, in esophageal squamous cell carcinoma (ESCC).
Methods:
In this study, we investigated serum levels of the miR-371-373 cluster in ESCC patients before and after surgical removal, and further focused on the expression level of miR-373-3p in tumor tissues of ESCC patients and its target genes. In addition, the epigenetic alterations of miR-373-3p promoter was analyzed.
Results:
The expression levels of miR-371-5p and miR-373-3p were significantly increased in preoperative serum of ESCC patients compared with that of healthy volunteers (P<0.01); however, they dropped significantly after surgical removal (P<0.01). Compared with adjacent normal tissues, miR-373-3p also showed significant up-regulation in cancer tissues (P<0.05). The methylation levels of miR-373-3p promoter were 42.86% in ESCC cancer tissue and 66.67% in adjacent normal tissues. The low methylation of the miR-373-3p promoter may promote the expression of miR-373-3p. Large tumor suppressor 2 (LATS2) and oxidation resistance 1(OXR1) are predicted to be targets of miR-373-3p by the bioinformatics method. They are the genes in the Hippo and the p53 signaling pathway, respectively. Their respective upstream genes, neurofibromatosis type 2 (NF2) and Jun Kinase, and the downstream genes, transcriptional co-activator with PDZ-binding motif (TAZ) and caspase 9, were also detected. The expression of all these genes were significantly decreased in ESCC cancer tissues compared with adjacent normal tissues.
Conclusions:
This study shows that DNA epigenetic modification in the miR-373-3p promoter region and the Hippo and p53 signaling pathways play important roles during the miR-373-3p mediating ESCC development process.
Introduction
Esophageal cancer can be divided into two major pathological types, esophageal squamous cell carcinoma (ESCC) and adenocarcinoma. In China, ESCC is the most common pathological type, and about half of the ESCC cases in East Asia occur in China.1,2 In early-stage patients, the cancers are most commonly treated by surgical removal; while patients at advanced stages are often treated by chemotherapy, but the efficacy is poor with an overall 5-year survival rate of only 15%~25%. 3 Therefore, it would be of great significance to clarify the pathogenesis of ESCC and to develop methods for early diagnosis and treatment of this disease.
MicroRNAs (miRNAs) are a group of non-coding small RNAs capable of regulating gene expression at a post-transcriptional level and play an important role in regulating cell differentiation, proliferation, invasion, angiogenesis and cell regulation. Current studies have shown that abnormal expression of miRNAs is closely correlated with a variety of malignancies. MiRNAs are dysregulated (up-regulated, down-regulated, or deleted) in different tumor types and carcinogenic pathways, showing a role similar to that of oncogenes or tumor suppressor genes. For example, miR-21 is highly expressed in gastric cancer and may serve as a potential biomarker 4 ; miR-486-5p is low in colon cancer and may be one of the therapeutic targets 5 ; and miR-34a is low in colorectal cancer and presumably plays a role in its metastasis. 6
The relationship between miRNAs and target genes is complex; one miRNA often regulates multiple target genes, and multiple miRNAs may regulate the same target gene. miRNAs target a large number of genes, which are widely involved in multiple cell signaling transduction pathways, thus forming a complex signaling network that plays a variety of biological roles in life activities and diseases, including cancers. 7
As one of the human embryonic stem cell (ESC)-specific miRNAs, miR-373 is one of research hotspots in recent years, especially in cancer. 8 It belongs to miR-371-373 gene cluster (miR-371, miR-372, and miR-373). In 2006, Voorhoeve et al. 9 first proposed miR-373 as a novel oncogene, confirming its involvement in the progression of testicular spermatogonia tumor by blocking the p53 pathway. Later, other studies proposed that miR-373 promoted liver cancer growth by targeting PPP6C (protein phosphatase 6 catalytic subunit) 10 ; and may be involved in lymph node metastasis of breast cancer. 11 Abnormal expression of miR-373 was also found in ESCC, but Lee et al. 12 only measured the expressions of miR-373 and its target gene LATS2 (large tumor suppressor 2). Recently, Ghasemi et al. 13 observed the up-regulation of the miR-371-373 cluster in the tumor tissues of ESCC patients. However, the changes of the miR-371-373 cluster in the serum of ESCC patients have not been studied, and the molecular mechanism of miR-373 in the progression of ESCC is still unclear. The aim of the present study was to investigate the serum levels of miR-371-373 cluster in ESCC patients before and after surgical removal, and to further clarify the association between miR-373-3p expression and esophageal cancer—both in the serum and the tumor tissues. In addition, epigenetic alterations of the miR-373-3p promoter and the expression levels of its target genes were measured to explore the function of miR-373-3p in ESCC.
Materials and methods
Patients and tissue samples
This study was approved by the Ethics Committee of the Xinxiang Central Hospital. A total of 74 ESCC patients from the Xinxiang Central Hospital and 94 healthy subjects were consecutively recruited to this study between September 2016 and June 2017. All participants were genetically unrelated ethnic Han Chinese from the same geographic region (Henan Province, China). The diagnosis of ESCC was confirmed histologically in all patients. Written informed consent was obtained from all participants prior to the study. ESCC tissues and adjacent non-cancerous esophageal tissues (at least 5 cm away from the tumor) from 23 patients were collected. Three milliliters of peripheral blood was collected from 51 ESCC patients and 94 healthy persons; for ESCC patients, peripheral blood was collected twice, pre-operative and 7 days of post-surgery.
Tissue and serum sample processing and RNA isolation
All the tissue samples that were collected during surgery were immediately snap-frozen in liquid nitrogen and stored at −80°C until RNA extraction was performed. Total RNA was isolated using Trizol (Invitrogen) according to the instructions of the manufacturer.
Peripheral blood was collected in tubes containing a separating gel and a clot activator, then put in water bath for 20 min at 37°C and centrifuged at 3,500 g for 10 min at room temperature. The supernatants were transferred to Eppendorf tubes. A second centrifugation at 12,000 g for 10 min at 4°C was performed to completely remove all cellular components. The serum was then aliquoted and stored at −80°C until RNA extraction was performed. All blood samples were processed within 3 h after they were obtained. Total serum RNA was isolated from 100 μL serum and eluted in 300 μL of RNase-free water using Trizol (Invitrogen) according to the instructions of the manufacturer for blood samples.
Real-time quantitative reverse transcriptase polymerase chain reaction
The expression levels of individual miRNA were ascertained on independent sets of serum or tissue samples using real-time quantitative reverse transcriptase polymerase chain reaction (qRT-PCR). The PrimeScript™ RT Reagent Kit (Perfect Real Time) (TaKaRa) was used to synthesis cDNA. SYBR Green (TaKaRa) quantitative PCR (qPCR) assay was used to detect miRNA expression of miR-373-3p, miR-371a-5p, miR-371b-5p, miR-372-3p, and miR-16-5p; also, mRNA expression of LATS2, neurofibromatosis type 2 (NF2), transcriptional co-activator with PDZ-binding motif (TAZ), oxidation resistance 1 (OXR1), Jun Kinase (JNK), and caspase 9 (CASP9). PCR reaction was performed for 45 cycles (95°C, 10 s; 60°C, 30 s) after an initial denaturation step (95°C, 5 min) on the CFX96 system of Bio-Rad. The expression levels of miRNA and mRNA were quantified using the 2-ΔΔCT method; miR-16-5p and GAPDH were used as the internal control for miRNA and mRNA, respectively. All reactions were done in triplicate. Primers were synthesized by Applied Biosystems (Shanghai, China) and their sequences were listed in Supplementary Table 1.
Prediction of miR-373-3p target genes
The prediction of miR-373-3p target genes was carried out using TargetScan (http://www.targetscan.org/) and PicTar (http://pictar.mdc-berlin.de/).
Genomic DNA isolation and methylation analysis
Genomic DNA was extracted using an EZ DNA Methylation-Gold™ Kit (Qiagen, Valencia, CA, US). The methylation status of the miR-373-3p promoter region was determined using methylation-specific PCR (MSP) on bisulfate-treated genomic DNA according to the manufacturer’s protocol. The specific primers for either unmethylated or methylated alleles are listed in Supplementary Table 1. As an internal control, all purified genomic DNA samples were successfully tested by PCR with TaKaRa EpiTaqTM HS kit (for bisulfite-treated DNA). Normal human peripheral lymphocyte methylated and unmethylated DNA was used as a positive control for the methylated genes, and as a negative control for unmethylated genes, respectively. PCR products were analyzed on 1% agarose gel, stained with ethidium bromide, and visualized under ultraviolet light. Each MSP was repeated at least once to confirm the results.
Statistical analysis
Data were reported as mean ± SD for quantitative variables. The difference in mRNA or miRNA expression levels between paired tissue samples was calculated using the Wilcoxon matched-pairs test. Correlations between independent samplings and qRT-PCR of LATS2, NF2, TAZ OXR1, JNK, CASP9 and miRNAs were determined by the Spearman correlation test. The Mann–Whitney test was performed to determine the significance of serum miRNA levels. The area under the curve (AUC) for tissue and serum microRNAs was determined using Receiver Operator Characteristic (ROC) analysis. P values <0.05 were considered as statistically significant. The statistical analysis was performed with software SPSS version 17.0.
Results
Expression levels of miR-371-373 cluster in ESCC
To investigate the role of miR-371-373 cluster in ESCC, we first examined the serum levels of miR-371a-5p, miR-371b-5p, miR-372-3p, and miR-373-3p in ESCC before and after surgical removal, and in normal subjects. Serum levels of miR-371a-5p, miR-371b-5p, and miR-373-3p were significantly higher (P<0.01) in ESCC patients before surgery than in normal subjects, by fold change 3.6, 2.7, and 3.9, respectively, while their levels were significantly (P<0.01) reduced after surgical removal, falling to similar levels to the normal subjects (Figure 1(a), (b), and (d)). However, for miR-372-3p, there was no significant difference in ESCC patients before and after surgical removal compared with normal subjects (Figure 1(c)). Further study mainly focused on miR-373-3P in ESCC.
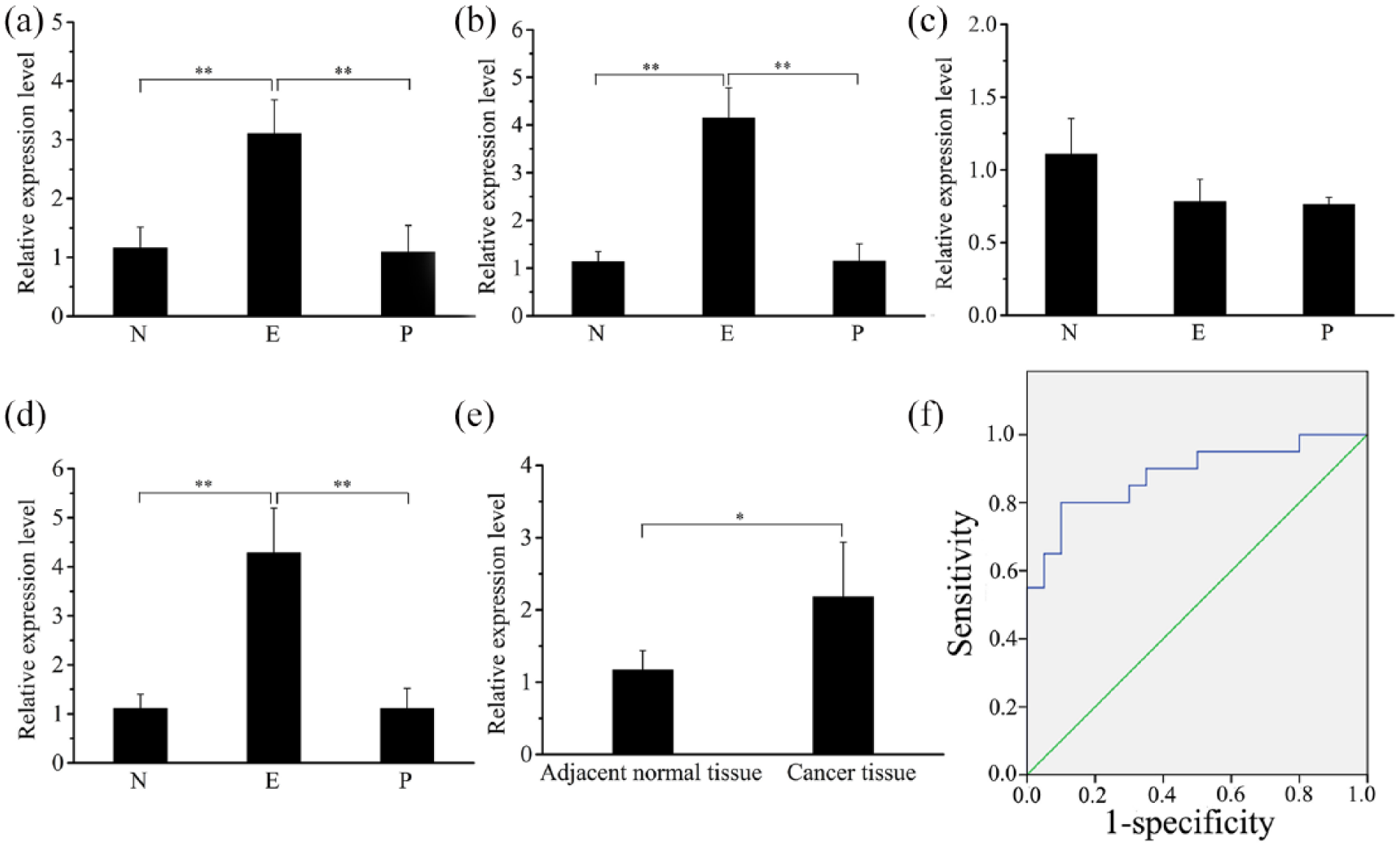
Expression levels of miR-371-373 cluster and ROC curve analysis of miR-373-3p. (a) miR-371a-5p expression in serum of ESCC patients by qRT-PCR. (b) miR-371b-5p expression in serum of ESCC patients by qRT-PCR. (c) miR-372-3p expression in serum of ESCC patients by qRT-PCR. (d) miR-373-3p expression in serum of ESCC patients by qRT-PCR. (e) miR-373-3p expression in tissues from 23 ESCC patients by qRT-PCR. (f) ROC curve analysis of miR-373-3p. N, E and P represents the serum from 94 normal subjects, 51 esophageal squamous cell carcinoma patients and postoperative patients, respectively.
The low level of miR-373-3p at post-surgical removal indicated that a higher serum level might be derived from cancer tissue. To test this hypothesis, we compared the expression level of miR-373-3p between cancer tissues and adjacent non-cancerous tissues. As shown in Figure 1(e), the expression level of miR-373-3p was significantly higher (P<0.01) in cancer tissues than in adjacent non-cancerous tissues.
The significant change of serum level of miR-373-3p indicates that this miRNA has a potential diagnostic role for ESCC. To evaluate the diagnostic value of serum miR-373-3p, we analyzed its ROC curve and AUC value. ROC curve analysis showed that the AUC value of miR-373-3p was 0.883 (95% confidence interval (CI): 0.776, 0.989). When the cutoff value was set at 0.7, a sensitivity of 80% and specificity of 90% was achieved (Figure 1(f)). AUC can be used to determine the diagnostic value of diagnostic methods. The higher the AUC (up to 1), the more accurate and higher the diagnostic value; therefore, the serum level of miR-373-3p could be used as an ESCC diagnostic marker.
Analysis of miR-373-3p promoter region methylation
DNA methylation is an inheritable feature that regulates gene expression without changes in DNA sequence. It is the most common epigenetic change observed in tumors and plays a role in the development and progression of tumors by regulating the expression of oncogenes or tumor suppressor genes. 14 Therefore, we detected the methylation level of the miR-373 promoter region in the cancerous tissue and peri-cancerous tissue of ESCC patients. To check whether the miR-373-3p expression level is related to methylation of its promoter region, we examined the methylation of the promotor of miR-373-3p in cancerous tissue and peri-cancerous tissue. It was found that the average methylation level of the miR-373-3p promoter was 42.86% in cancerous tissue, which was significantly lower than that in peri-cancerous tissue (66.67%; P < 0.01; Figure 2), indicating that the low methylation level of the miR-373-3p promoter may promote the expression of miR-373-3p.
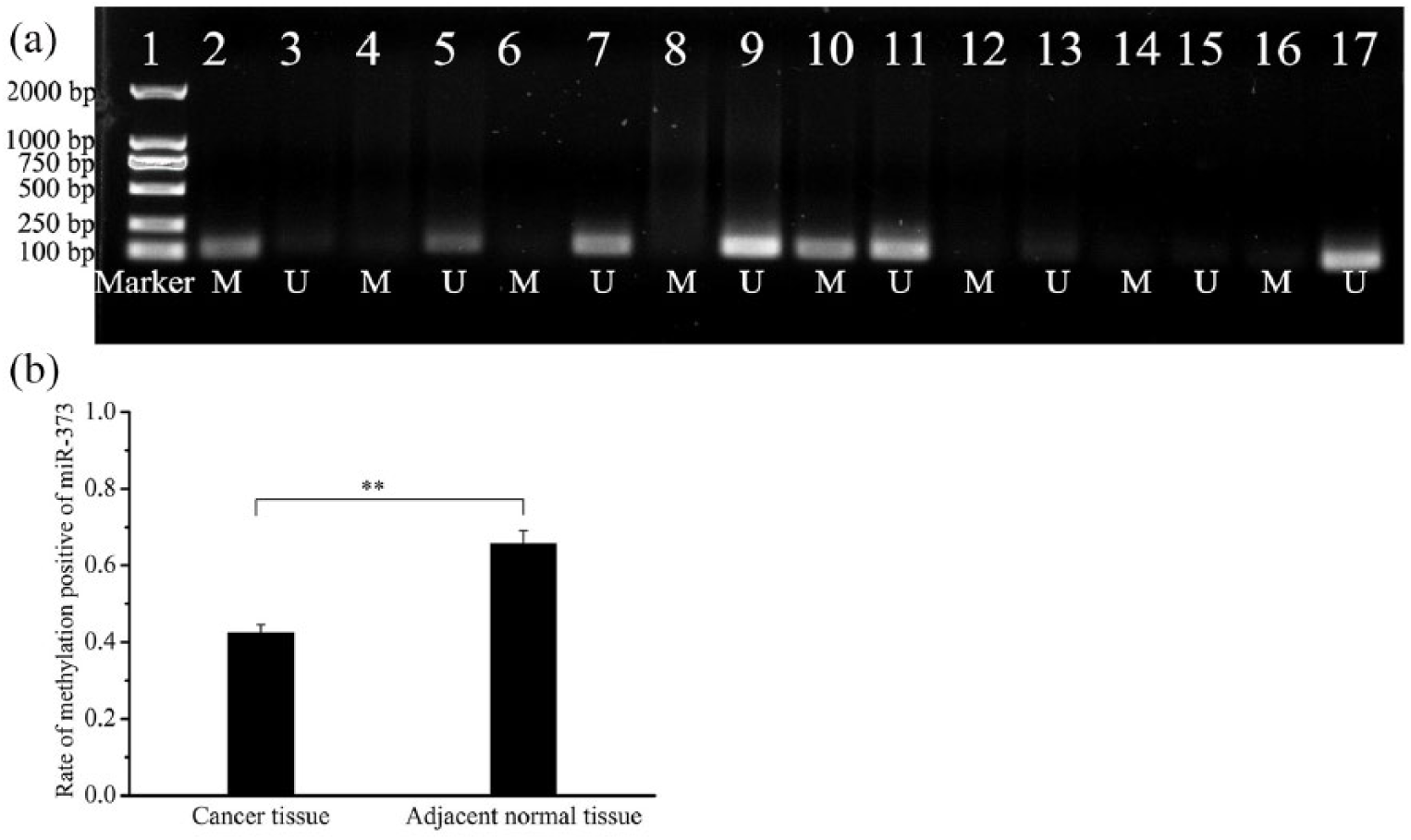
Detection of the methylation status of the miR-373-3p gene promoter region. (a) 1: DL2000 marker; 2,6,10,14 methylation of cancer tissue; 3,7,11,15 non-methylation of cancer tissue; 4,8,12,16 methylation of adjacent normal tissue; 5,9,13,17 non-methylation of adjacent normal tissue. (b) The average methylation level of the miR-373-3p promoter in cancer tissue and adjacent normal tissue. **P<0.01.
Negative correlation between miR-373-3p and OXR1 and LATS2
In order to illustrate the mechanism of miR-373-3p in ESCC development, we further predicted the target genes of miR-373-3p using bioinformatics software, such as TargetScan and PicTar. Among hundreds of putative target genes, we focused on two targets: LATS2 and OXR1 (Figure S1). LATS2 is one of the central components in the Hippo signaling pathway, which plays an important role in controlling organ size and balancing cell proliferation, and has been proven to be involved in the tumor development of multiple organs.15,16 OXR1, an antioxidant gene, regulates the network of reactive oxygen species, and regulates cell cycle and apoptosis via the p53 signaling pathway. The p53 signaling pathway has a role in inducing cell cycle arrest, promoting apoptosis and DNA repair, and is thus involved in the prevention of DNA damage accumulation, the maintenance of genomic stability, and the regulation of cell differentiation and aging.17,18 The role of miR-373-3p in ESCC might be due to miR-373-3p-mediated regulation on OXR1 and LATS2, which further affects Hippo and p53 signaling pathways.
We analyzed the expression level of the Hippo-signaling-pathway-related genes, which included LATS2 and its upstream gene NF2, and the downstream gene TAZ. The results demonstrated that the expression levels of the three genes in the cancer tissues were significantly lower compared with the adjacent non-cancerous tissues (P<0.05) (Figure 3). The Pearson correlation coefficient analysis revealed a negative correlation between the expression levels of miR-373-3p and LATS2 (r = −0.704; P<0.01).
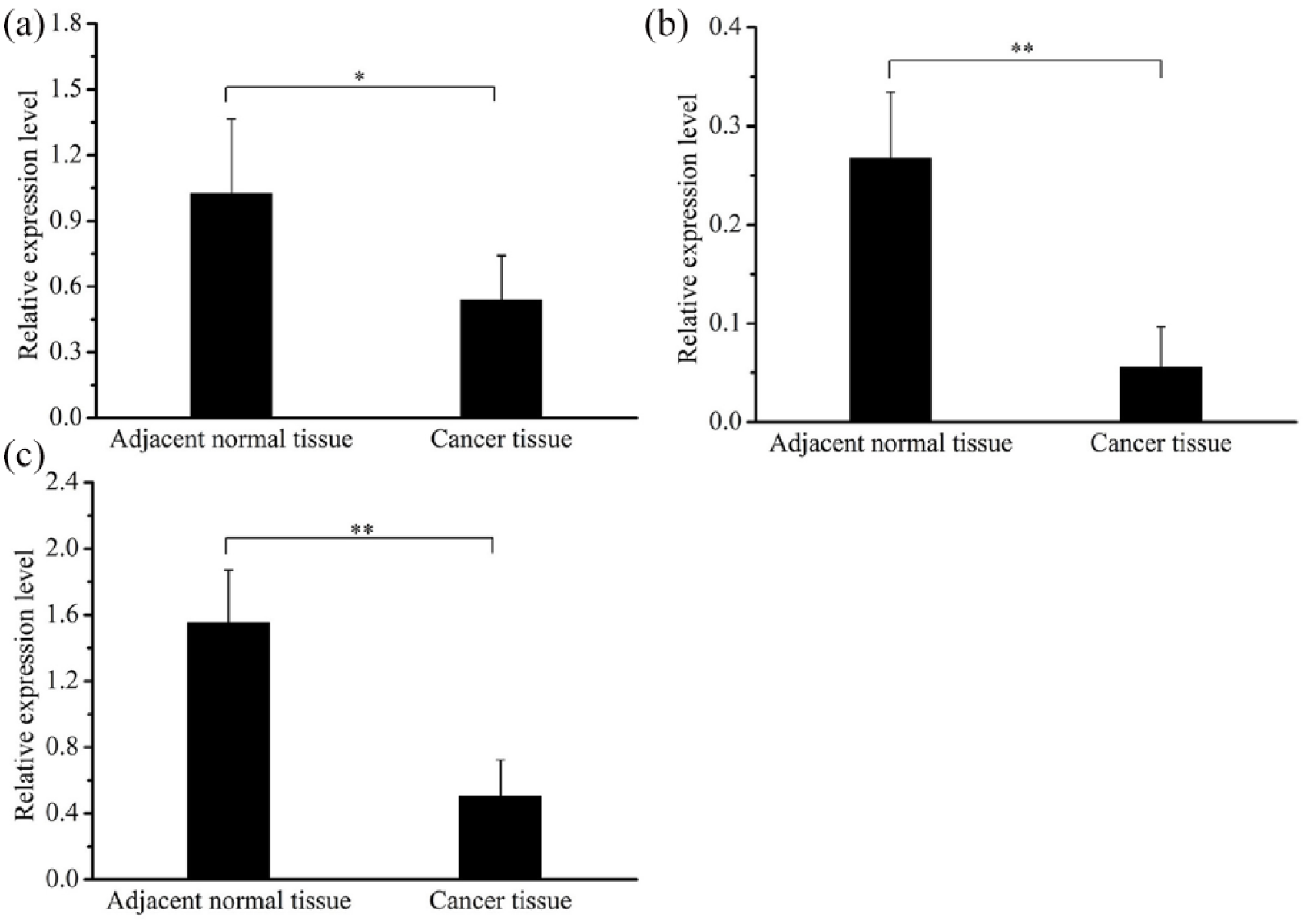
Expression levels of Hippo-signaling-pathway-related genes in ESCC. (a) The expression level of NF2 in the cancer tissue of 23 ESCC patients. (b) The expression level of LATS2 in the cancer tissue of 23 ESCC patients. (c) The expression level of TAZ in ESCC patients. GAPDH was used as the reference gene to normalize the expression levels.
We also analyzed the expression level of the p53-signaling-pathway-related genes, OXR1 and its upstream gene JNK, and the downstream gene CASP9. The results demonstrated that the expression levels of the three genes in the cancer tissues were significantly lower compared with the adjacent non-cancerous tissues (P<0.05) (Figure 4). The Pearson correlation coefficient analysis demonstrated that the expression levels of miR-373-3p and OXR1 were negatively correlated (r = −0.699; P<0.05).
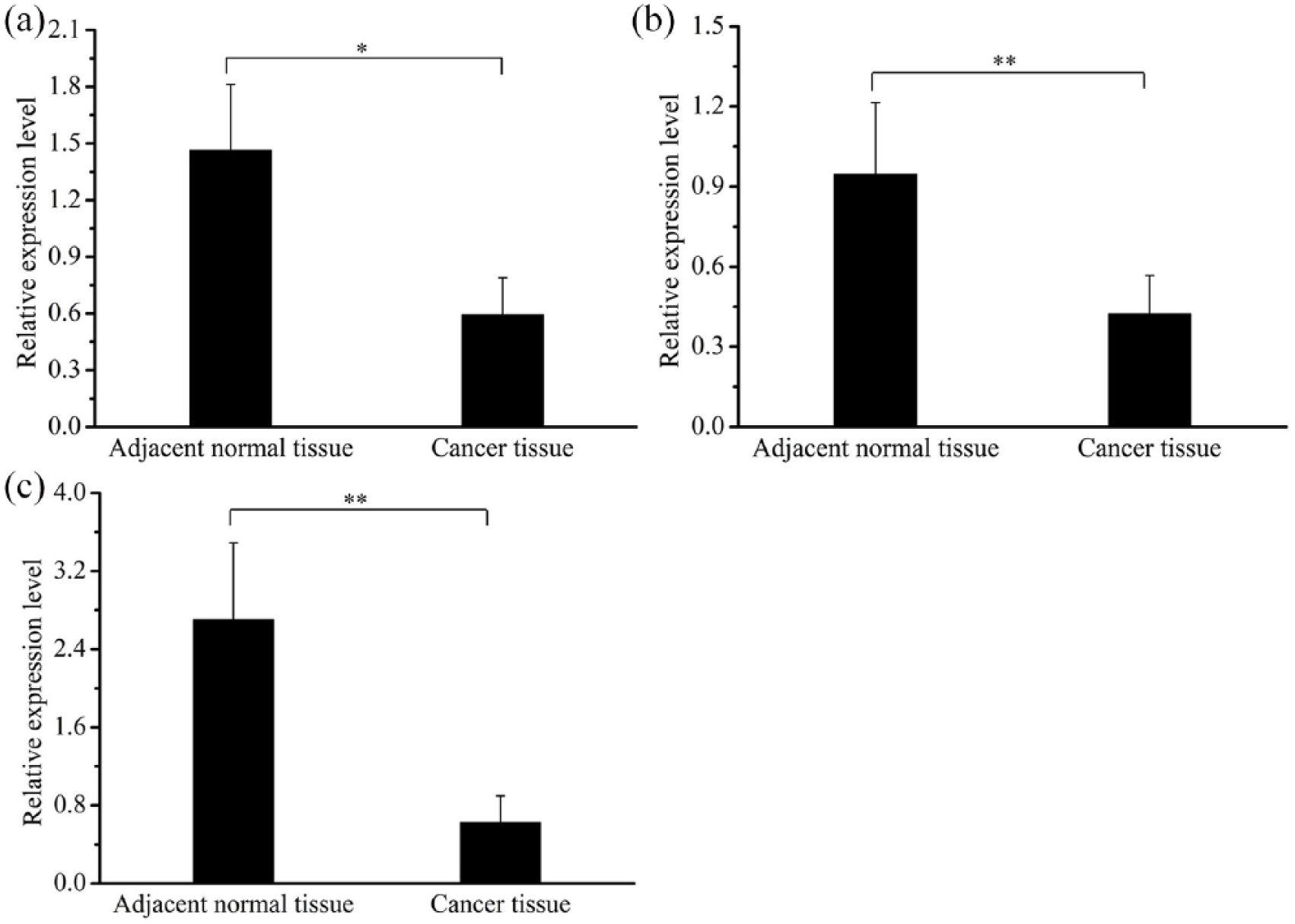
Expression levels of p53-signaling-pathway-related genes in ESCC. (a) The expression level of JNK in the cancer tissue of 23 ESCC patients. (b) The expression level of OXR1 in the cancer tissue of 23 ESCC patients. (c) The expression level of CASP9 in the cancer tissue of 23 ESCC patients. GAPDH was used as reference gene to normalize the expression levels.
Discussion
MicroRNAs from the miR-371-373 cluster have been reported to either act as tumor suppressors or behave like oncomirs, depending on the specific cellular context. 8 High expression of the miR-371-373 cluster was observed in the formalin-fixed paraffin-embedded tumor tissues of ESCC patients. 13 The current study is the first to evaluate the serum level of the miR-371-373 cluster in ESCC patients before and after surgical removal. Our results showed that serum levels of miR-371 and miR-373 were significantly high in ESCC patients, and dropped significantly after surgical removal, reaching similar levels to that of normal subjects (Figure 1). However, miR-372 from the miR-371-373 cluster showed no significant differences in the serum of ESCC patients before and after surgical removal compared with normal subjects. miR-371 and miR-373 could be used as candidate serum markers for ESCC patients’ diagnoses, with miR-373 showing the greater up-regulation.
The increased expression of miR-373 has been observed in several cancer cell lines (including breast cancer, testicular seminoma, and esophageal cancer, etc.),9,11,12 and high-level miR-373 may have a direct effect on CD44 (a signal molecule involved in cell growth and adhesion) and may promote tumor cell growth. 19 Liu et al. 19 found that the miR-373 expression level in the plasma and tumor tissues of ESCC patients was closely correlated with differentiation, tumor status, and lymph node metastasis (P<0.05), but not with gender, age, tumor node metastasis stage, smoking, and alcohol intake (P>0.05). In this study, significant high-level miR-373 was observed in both the serum and cancerous tissue of ESCC patients (Figure 1), suggesting that miR-373 was produced by the ESCC cells and secreted into circulating blood. The results were in line with our previous reports about the expression of miR-483-5p in ESCC, supporting the statement that serum miRNAs are mostly derived from cancerous tissue. 20 Therefore, miR-373 acts as an oncomir in ESCC.
The role of epigenetic modification in gene expression is one of the current research hotspots. miRNA may be generated from an independent gene or from the introns of coding genes. We have previously reported that the expression of miR-483-5p (located in an intron of IGF2 gene) was affected by methylation of the IGF2 promoter region. 20 miR-373 is derived from an independent gene; our results showed that the miR-373 promoter region was hypomethylated in the cancerous tissue of ESCC patients (Figure 2). Therefore, promoter methylation also regulates the expression of independent miRNAs. Promoter hypomethylation is one of the reasons that led to the high expression of either the independent miRNAs or those within the intron of other genes.
Bioinformatic analysis predicted hundreds of potential targets of miR-373, among which the tissue inhibitor of metalloproteinases-3 (TIMP3) and LATS2 have been validated as targets of miR-373 in esophageal cancer.12,21 miR-373 may promote the migration and invasion of esophageal cancer by targeting TIMP3. In this study, we found a negative correlation between miR-373 and LATS2 in the cancerous tissue of ESCC patients compared with peri-cancerous tissue, which is in accordance with previous reports that high levels of miR-373 significantly correlate with low levels of LATS2 protein. 12 NF2 and TAZ locate upstream and downstream of LATS2, respectively, in the Hippo signaling pathway. We also found that the expression level of NF2 and TAZ was lower in the esophageal cancer tissue than in the adjacent normal tissue. The lower expression levels of all these genes in esophageal cancer tissue compared with normal tissue indicate the suppression of the Hippo signaling pathway in esophageal cancer. LATS2 was decreased by several folds, showing a much greater decrease compared to the other two genes (Figure 4), suggesting a key role of miR-373 in the suppression of the Hippo signaling pathway. There is a positive feedback loop between the p53 and LATS2 tumor suppressors. 22 LATS2 can activate p53, another tumor suppressor gene, while p53 can also up-regulate LATS2 expression. 22 Therefore the LATS2 mRNA level can be used as a sensitive marker for tumor suppression activity. The suppression of LATS2 mRNA translation by miR-373 may weaken the LATS2-p53 interaction and promote tumor growth.
The pathogenesis of ESCC is not yet fully understood, although the imbalance between oxidant and antioxidant has been proposed as one of the causes. 23 In this study, we investigated the expression level of OXR1, another target of miR-373, as well as its upstream gene JNK and downstream gene CASP9. The expression levels of these three genes were lower in esophageal cancer than in adjacent normal tissues. JNK is a member of the mitogen-activated protein kinase (MAPK) superfamily; the JNK signaling pathway can be activated by various factors, such as cytokines, growth factors, and stresses. 24 Studies have shown that JNK acts upstream of OXR1 and can significantly up-regulate the expression of OXR1. 25 Silencing JNK reduced the expression of OXR1 in female mosquitoes. 26 Therefore, the decreased expression of OXR1 may be due not only to the high level of miR-373 but also to the low level of JNK in ESCC (Figure 4). CASP9 plays an important role in the p53-dependent mitochondrial apoptosis pathway under the regulation of OXR1. 21 OXR1 depletion caused the up-regulation of CASP9 in HeLa cells. 27 In this study, the expression of CASP9 showed a significant decrease, although OXR1 was suppressed in ESCC. The decreased mRNA levels of CASP9 were found in different cancer cells, such as hepatocellular carcinoma, breast carcinomas, and colorectal cancer,28-30 whereas the up-regulation of CASP9 was observed when the proliferation of tumors was inhibited. Some studies have reported that CASP9 can be down-regulated by multiple miRNAs in cancer cells.28,31 Therefore, the development and proliferation of ESCC also may be needed in the inhibition of p53-dependent apoptosis by down-regulating CASP9.
Conclusion
This study is the first to investigate the serum levels of the miR-371-373 cluster in ESCC patients before and after surgical removal. The high level of miR-373 in both serum and tumor tissues before surgical removal, which fell to a regular level after surgical removal, further confirmed it as an oncomir in ESCC. Importantly, the expression of the direct target genes of miR-373 and its upstream/downstream genes were simultaneously reduced, which revealed that the targets of miRNA also can be modulated by its upstream genes and affect it’s the level of its downstream genes. The suppression of Hippo and p53 signaling pathways may promote the development and progression of ESCC. However, there are still some limitations in our study. Validation of the new target OXR1 of miR-373 in vitro needs to be performed. In addition, the protein levels of NF2, TAZ, OXR1, JNK, and CASP9 need further investigation by inhibiting and overexpressing miR-373 in the esophageal cancer cell line.
Supplemental Material
Supplemental_Material – Supplemental material for MicroRNA-373 promotes the development of esophageal squamous cell carcinoma by targeting LATS2 and OXR1
Supplemental material, Supplemental_Material for MicroRNA-373 promotes the development of esophageal squamous cell carcinoma by targeting LATS2 and OXR1 by Li Wang, Lifeng Wang, Weidong Chang, Yongfang Li and Linsong Wang in The International Journal of Biological Markers
Footnotes
Author contributions
Li Wang and Lifeng Wang contributed equally to this study.
Declaration of conflicting interest
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Key Projects of Science and Technology in Henan Province (Grant No.172102310407) and the Key Research Projects of Henan Higher Education Institutions (Grant No.16A180028).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
