Abstract
Background:
Prostate and bladder cancers continue to be the first and fourth most common cancers in men worldwide; thus there is an urgent need for more accurate biomarkers that can detect these types of cancer in a non-invasive way. Liquid biopsy is a new non-invasive tool for diagnosis and with a virtually unlimited supply urine is even more attractive resource since urinary exosomes have been discovered to contain RNAs that are hallmarks of cancer. It is challenging to assay those secreting lower amounts of molecules.
Methods:
This review, based on articles identified through a PubMed/MEDLINE search, comprehensively summarizes state of the art approaches used in the discovery and validation of exosomal RNA biomarkers purified from the urine for lower urinary tract cancer.
Results:
The combination of PCA3 and ERG has shown a relatively good improvement in diagnostic performance; examples of other potential biomarkers and the methods utilized in their discovery are also discussed in this review.
Conclusions:
Of these last markers, to date there are still few data to implement these for routine diagnosis.
Introduction
Almost a decade ago, the term “liquid biopsy” was coined 1 to describe the utility of circulating tumor cells (CTCs) as candidate tumor biomarkers in breast cancer. The existence of CTCs is not new, as their presence in the blood stream was postulated over a century ago. 2 Currently, the term “liquid biopsy” has acquired a new meaning because it also involves the analysis of circulating subcellular structures, as exosomes, nucleic acids (mainly microRNA (miRNA) and cell-free DNA (cfDNA)), of particular relevance in the field of cancer research and clinics. 3 All these subcellular structures obtained from biological fluids express characteristics peculiar to the tissue they originate from. However, the laboratory approaches utilized for their analysis are very different depending on the biomarker and the biological question to be investigated. Moreover, fluids other than blood (e.g. urine, saliva, ascites fluids, pleural effusions) also have been considered to realize biological-functional studies on these subcellular structures.
Exosomes are small vesicles with a size between 30 and 100 nm comparable to viruses, and are secreted by all cells through an ATP-dependent mechanism. 4 The term “exosomes” was coined in 1981 to describe microvesicles with 5’-nucleotidase activity secreted by tumor cell lines. 5 Soon after, the endocytic origin of exosomes was described in cultured reticulocytes. 6 Since then, many papers have shown that exosome production is an ubiquitous cellular process. In fact, tumor cells are particularly active in secreting exosomes in plasma during cancer progression. 7 Exosomes derive from the internal vesicles of the multivesicular bodies and contain nucleic acids such as DNA, miRNA, single-stranded RNA, long non-coding RNA (lncRNA), proteins, and lipids. Typically, exosomes contain the endosome-specific tetraspanins (CD9, CD63, and CD81); accessory endosomal sorting complexes required for transport (ESCRT) pathway proteins (programmed cell death 6-interacting protein, PDCD6IP) and the tumor susceptibility gene 101 protein (TSG101); proteins involved in membrane transport and fusion (Ras-associated binding proteins and annexins); cytoskeletal proteins (actin and tubulin); and G proteins and adhesion molecules (integrins). Some of these proteins reflect the endocytic origin of exosomes and are used as exosome markers (mainly CD63, PDCD6IP, or TSG101). 8 In addition, exosomes contain other proteins that could serve to identify the cellular source, such as CD34 for progenitor cells or CD105 for endothelial cells. 9
Extracellular vesicles, and predominantly exosomes, were identified as important carriers of non-coding RNAs, including circular RNAs lncRNAs, and miRNAs, which were also to be protected from nuclease-mediated degradation in the extracellular space and in body fluid.10,11 MiRNAs can either function as tumor-suppressors or oncogenes depending on their target genes; for example, in particular miR-21 has been considered an oncogene as it has a higher expression level in non-small cell lung cancer patients. 12 Moreover, tumor-derived exosomes can express death ligands (Fas antigen ligand and tumor necrosis factor-related apoptosis-inducing ligand) that could distinguish them from other circulating exosomes released by non-malignant cells. 13
Exosomes interact with recipient cells through several mechanisms: (i) binding to ligand/receptor in the cell surface: (ii) fusion with the plasma membrane; or (iii) uptake by endocytosis, micropinocytosis, or phagocytosis. 14 After exosome engulfing, some of the RNA and the protein transported in exosomes can be delivered to recipient cells serving as a communication mechanism between cells.
Cancer is a disease in which the tumor genetic characteristics dynamically evolve not only as a natural consequence of the tumor genetic instability, but also because of the strong selection pressure due to antitumor treatment, which induces a polyclonal evolution to more resistant clones. 15 Consequently, the identification of the different pre- and post-treatment mutation patterns could provide guidance for therapy and prognosis. 16 Furthermore, studies on the initial biomolecular pattern of the tumor can potentially provide information that is useful for prevention, plus differential and/or early diagnosis of a cancer. Therefore, liquid biopsies are emerging as an attractive solution in which non-invasive samples, such as urine, 17 can be utilized for multiple clinical endpoints and, importantly, they can substitute information obtainable with traditional solid tumor biopsies.
As exosomes are available in body fluids, such as blood and urine, they have been proposed as a biomarker in the diagnosis, prognosis, and follow-up of treatment efficiency; however, there is a strong need to greatly enrich these populations from body fluids, ideally by developing selective and sensitive technology to count a significant number of exosomes. 18 Several examples of such applications for exosomes are available in the literature.
1. Exosome levels have been shown to be elevated in patients with different type of cancers.19,20 Szajnik et al. 19 showed that plasma exosomes from patients with ovarian cancer contained higher levels of proteins compared to those isolated from the plasma of patients with benign tumors or healthy controls; furthermore, the protein content in exosomes from patients with a more advanced stage of the disease is significantly higher than in early stages.
2. Exosome levels variably changed during/after chemotherapy, and correlations between the changes in exosomal protein levels and clinical data suggested that the protein content of exosomes might be useful in predicting response to therapy and prognosis in ovarian cancer.
Survivin, a protein implicated in the inhibition of apoptosis, has been detected at higher levels in blood-derived exosomes from prostate cancer patients compared with healthy controls. Also, Khan et al. 21 showed that exosome survivin splice variants were packaged into exosomes in breast cancer patients’ sera, mimicking the survivin splice variant pattern detected in breast cancer tissues, suggesting that exosomal-survivin analysis may serve as a “liquid biopsy” in early breast cancer patients. 21
Moreover, Raimondo et al. 22 revealed differential protein profiling of renal cell carcinoma urinary exosomes compared with healthy subjects. Approximately 44% of total identified proteins were present only in healthy subjects, while about 22% were detected only in renal cell carcinoma (RCC) urinary exosomes. 22 Also, a transcriptomic analysis showed a change of mRNA content in urinary exosomes. Three transcripts (glutathione S-transferase alpha 1, CCAAT enhancer binding protein alpha, and pterin-4 alpha-carbinolamine dehydratase 1) are reduced in extracellular vesicles derived from clear cell RCC patients with respect to healthy subjects and patients with other types of RCC. These alterations are specific and disappear after nephrectomy. 23
The aim of this manuscript is to quickly review the literature on the clinical and biological role that urinary exosomal nucleic acids can have in urological cancers, paying particular attention to information available for prostate and bladder cancer (Tables 1 and 2).
Potential diagnostic use of urinary exoRNA as prostate cancer biomarkers.
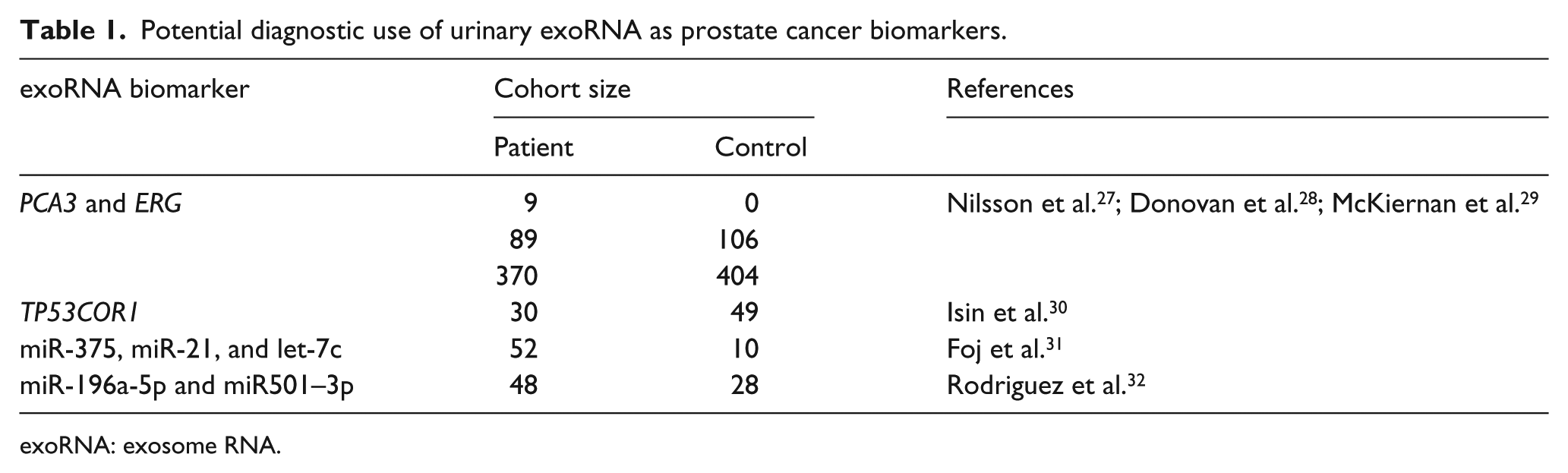
exoRNA: exosome RNA.
Potential diagnostic use of urinary exoRNA as bladder cancer biomarkers.
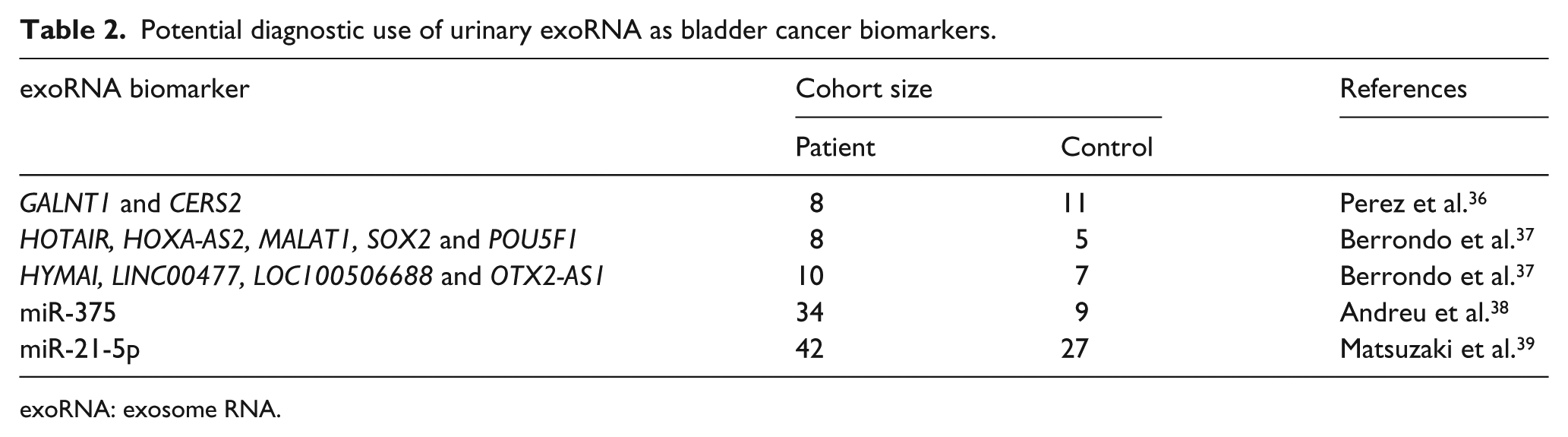
exoRNA: exosome RNA.
Material and methods
A comprehensive literature search of the electronic PubMed/MEDLINE database was conducted to find relevant studies that had assessed the diagnostic value of both urinary and blood exoRNA and exoDNA in prostate and bladder cancer. The literature search was performed from year 2009 to November, 2018, to include the most updated data. To expand our search, references of the retrieved articles were also screened for additional data. For each study included, information was collected concerning basic data (authors, year of publication, country of origin, type of study), methods (isolation and characterization of exosomes, number of patients included, urine volume) and results (exoRNA and exoDNA names, area under the curve (AUC) of the receiver operating characteristic (ROC) curve).
Results Prostate cancer diagnosis potential biomarkers
Digital rectal examination and/or serum prostate specific antigen (PSA) levels are generally utilized by the clinician to suppose the presence of prostate cancer; successively diagnosis depends on histopathological confirmation of adenocarcinoma in prostate needle biopsies. However, PSA is associated with significant false positives 24 and this results in unnecessary biopsies. Therefore, biomarkers including non-invasive methods, such as urinary exosome transcriptome studies, might be useful for the management of patients with prostate cancer. Urine samples—a biological fluid of direct pertinence of urogenital apparatus, which is easy to obtain through non-invasive methods—contains exosomes secreted by the cells lining the urinary tract 25 ; thus this sample is open to a deeper analysis of prostate biomarker discovery. The literature is rich in data on this topic with a plethora of biomarkers considered; however, diagnostic exosomal biomarker performance review is needed for evaluation of the usefulness of these candidate biomarkers.
Nilsson et al. 27 elucidated the potential contribution of these extracellular small vesicles as a source of non-invasive biomarkers. They evaluated a novel approach to find predictive markers for prostate cancer, and analyzed RNA in urine exosomes from nine prostate cancer patients, which indicated that mild prostate massage increased the exosomal secretion into the urethra and subsequently into the collected urine fraction. The exosomal fraction was isolated by differential centrifugation and the exosomal nature was confirmed by immunoelectron microscopy after anti-CD63 gold staining. They demonstrated that urinary exosomes from prostate cancer patients carried the biomarkers TMPRSS2:ERG gene fusion, resulting from a chromosomal rearrangement of ERG to the androgen responsive gene TMPRSS2, and prostate cancer antigen 3 (PCA3). PCA3 is a lncRNA and is one of the most specific prostate cancer biomarkers. It was discovered in 1999 26 and its expression is highly restricted to prostate tissue tumors compared with non-neoplastic prostatic tissue of the same patients. Moreover, its expression was not detected in other tumor types or cell lines. However, although promising, these results have been obtained in a limited number of patients and should be validated in large cohorts. 27
Donovan et al. 28 performed an observational study in men enrolled in a country-wide US study. The “intended use” population was comprised only of men who were undergoing their initial biopsy and had equivocal “gray zone” serum PSA levels (>2 and <10 ng/mL). Exosome RNA (exoRNA) derived from first-catch, non-digital rectal examination urine samples were normalized for RNA levels with SPDEF (SAM-pointed domain-containing Ets transcription factor) to derive “ERG or PCA3 RNA copy number/SPDEF mRNA copy number.” This gene signature building EXO106—an algorithm that associates PCA3 and ERG (including TMPRSS2:ERG) exosomal mRNA levels normalized with SPDEF—demonstrated good clinical performance in predicting high-grade disease (Gleason score ⩾7) with an area under the curve (AUC) of 0.764, in 195 urine samples. The AUC reached 0.803 when combined with clinical parameters (PSA, age, race, and family history). However, a few limitations should be noted. First, the lack of a central pathology review; second, the exosomal ERG RNA copy number is low (average of 1 copy/reaction), suggesting that performance of this gene may be impacted by urine volume; third, shipping conditions and properties of the urine sample including acidity, protein content, and so on. To address the potential impact of urine volume and a dilutional effect on exosome isolation, analyses were limited to samples of ⩽ 40 mL. 28
Exosome Diagnostics Inc. (Cambridge, MA, US), conducted a US national prospective study enrolling men older than 50 years undergoing prostate biopsy with PSA levels (>2 and <10 ng/mL) and analyses limited to ⩽ 49 mL non-digital rectal examination urine samples. The end point of the study was to validate EXO106 under the name ExoDx Prostate Intelliscore (expressed as a risk score ranging from 0 to 100). A total of 774 patients were studied with a “training” cohort of 255 patients and a validation cohort of 519 patients. The association of the exosome-gene expression with clinical parameters (PSA, age, race, and family history) resulted in better discriminative power between insignificant and aggressive disease (AUC = 0.73) compared with the parameters alone (AUC = 0.63). McKiernan and colleagues 29 suggest that this three-gene expression signature assay and similar tests could provide the way to reduce the treatment of low-risk prostate cancer, thus decreasing the number of unnecessary prostate biopsies. Both Donovan et al. 28 and McKiernan et al. 29 filtered urine samples through a 0.8-μm filter and isolated exosomes by ultracentrifugation using the Urine Clinical Sample Concentrator Kit (Exosome Diagnostics).
Isin et al. 30 evaluated exosomal levels of two tumor suppressive lncRNAs: (i) growth arrest specific 5 (GAS5); and (ii) tumor protein p53 pathway corepressor 1 (TP53COR1). These were evaluated in the urine samples after digital rectal examination of 30 patients with prostate cancer, and 49 patients diagnosed with benign prostatic hyperplasia. exoRNA was isolated directly from urine samples using the Urine Exosome RNA Isolation Kit (Norgen Biotek, Thorold, ON, Canada). TP53COR1 was more abundant in prostate cancer patients than in patients with benign prostate hyperplasia (AUC: 0.663). No correlation between the clinical stage (Gleason score) and exosomal lncRNA levels was observed in the prostate cancer group. The sensitivity and specificity of TP53COR1 for predicting prostate cancer were calculated to be 67% and 63%, respectively. The specificity increased to 94% when TP53COR1 was considered in combination with PSA. TP53COR1 provides a promising marker with therapeutic potential for the detection and stratification of prostate cancer. Further studies with larger patient groups are needed to validate the utility of exosomal TP53COR1 levels in urine. 30
Foj et al. 31 analyzed five miRNAs commonly deregulated in prostate cancer in tumor tissues, in serum/plasma, or in exosomes from freshly voided urine samples (30–50 mL) after a prostate massage of 52 patients with prostate cancer, and in 10 healthy volunteers. Exosome were isolated by differential centrifugation and their presence was confirmed by electronic microscopy. They found that miR-375, miR-21, and let-7c were significantly upregulated in the prostate cancer group compared with the healthy group (AUC were 0.799, 0.713, and 0.679, respectively). In addition to the study on exosomes, they demonstrated that a panel of miRNAs from a urinary pellet, mainly consisting of miR-21 and miR-375, can be used to distinguish between healthy individuals and patients with prostate cancer (AUC 0.872). Severe limitations concerning this study have to be considered relative to the healthy control group; for example, the small sample size and the absence of biopsy. 31
Rodriguez et al. 32 reported quantification of exosomal miRNAs by RNA-seq and by quantitative real-time polymerase chain reaction (PCR) analysis from the urine of prostate cancer patients. After isolation of exosomes in urine specimens by differential centrifugation at room temperature, the sequencing of miRNAs in exosomes isolated from urine of 20 prostate cancer patients and 9 healthy male donors revealed that microRNAs (miR-196a-5p, miR34a-5p, miR-143-3p, miR-501-3p, and miR-92a-1-5p) were significantly decreased in prostate cancer patients compared with healthy men. In an independent cohort of 28 prostate cancer patients and 19 healthy men, miR-196a-5p and miR501-3p were significantly decreased in the urine from the prostate cancer patients by quantitative PCR (qPCR), and the AUCs of miR196a-5p and miR-501-3p were 0.73 and 0.69, respectively, for the detection of cancer.
To date, the expression of prostate cancer exoRNAs in the bloodstream exosomes proposed as a diagnostic tool has been considered only by a few authors.
Bryant et al. 33 used a quantitative reverse transcriptase-PCR (qRT-PCR) array to identify differentially expressed miRNAs in the plasma-derived exosomes of 78 prostate cancer patients and 28 controls. The exosomal populations were enriched by filtering the plasma through a 1.2-μm filter to remove cells and cellular debris, and concentrated on a filter concentrator with a 150-kDa molecular weight cutoff. This study showed that miR-107 and miR-574-3p were increased in the circulating exosomes of men with non-metastatic prostate cancer compared with the normal control individuals.
In a study of 20 patients with prostate cancer, 20 patients with benign prostate hyperplasia, and 20 control volunteers, Li et al. 34 showed by qRT-PCR that the level of serum exosomal miR-141 was significantly higher in prostate cancer patients than in benign prostatic hyperplasia patients or healthy controls. Exosomes were extracted from serum samples using ExoQuick exosome precipitation solution (System Biosciences, Mountain View, CA, US), characterized morphologically by electron microscopy analysis and for the expression of CD63 both by western blot and flow cytometric analysis. Moreover, ROC curve analysis showed that serum exosomal miR-141 yielded an AUC of 0.8694, with 80% sensitivity and 87.1% specificity in discriminating patients with metastatic prostate cancer from the patients with localized prostate cancer.
For many years it has been known that circulating DNA can be extracted from serum and plasma. As expected, part of this cell-free genomic DNA is present in exosomes 35 even after being incubated with DNase just to avoid external DNA contamination. Prostate cancer DNA aberrations, such as phosphatase and tensin homolog (PTEN), and tumor protein p53 (TP53) mutations, can therefore be measured in DNA extracted from exosomes present in urine. The profiling of these exoDNA mutations complements exoRNA expression, providing a comprehensive map of intra-exosome changes in prostate cancer using urine-based liquid biopsies.
Bladder cancer diagnosis potential biomarkers
The diagnosis of bladder cancer depends on cystoscopic examination of the bladder and histological evaluation of sampled tissue; however, these procedures are highly invasive and accompanied by undesirable side effects such as hematuria, dysuria, pain over the bladder, and difficulty in voiding. Cytology is helpful in an adjunct to cystoscopy; however, negative cytology does not exclude the presence of a tumor. It is commonly accepted that none of the available molecular marker tests can replace cystoscopy. However, biomarkers can be used as an adjunct to cystoscopy to detect invisible tumors.
Perez et al. 36 conducted a comparative study of the transcriptomes carried within vesicles obtained from urine collected prior to cystoscopy in five bladder cancer and six non-cancer patients using microarray technology. They filtered urine samples through a 0.22 μm pore membrane, isolated exosomes by differential centrifugation, and characterized them by electron microscopy and nanoparticle tracking analysis (NTA). Then they validate the array results by PCR on a new set of samples: three cancer and five control subjects and found polypeptide N-acetylgalactosaminyltransferase 1 (GALNT1) and ceramide synthase 2 (CERS2) mRNAs only in exosomes obtained from cancer patients. Given the reduced number of analyzed samples in this pilot study, the presented results only illustrated the potential of the method to identify candidate RNA that could be useful as biomarkers for the diagnosis of bladder cancer. 36
Berrondo et al. 37 demonstrated by qRT-PCR on exosomes derived from urine collected from eight patients with high-grade muscle invasive urothelial bladder cancer after the induction of general anesthesia and five healthy volunteers, that some lncRNAs as HOX transcript antisense RNA (HOTAIR), HOXA cluster antisense RNA 2 (HOXA-AS2), metastasis associated lung adenocarcinoma transcript 1 (MALAT1), mRNAs as SRY-box 2 (SOX2), and POU class 5 homeobox 1 (POU5F1), which were shown to be expressed in urothelial bladder cancer patients’ tumors, were also enriched in their urinary exosomes relative to healthy volunteers. Exosomes were characterized by western blotting, demonstrating the presence of PDCD6IP and by electron microscopy. exoRNA was isolated from urine samples using the Urine Exosome RNA Isolation Kit (Norgen Biotek). In addition, they searched for lncRNAs using RNA-seq and confirmed by qRT-PCR in 10 patients that their urinary exosomes were enriched in lncRNAs hydatidiform mole associated and imprinted (HYMAI), long intergenic non-protein coding RNA 477 (LINC00477), LOC100506688, and OTX2 antisense RNA 1 (OTX2-AS1) compared to 7 healthy volunteers. Therefore, these lncRNAs may serve as biomarkers for muscle-invasive bladder cancer, though validation in a larger appropriate patient population and comparison with urinary exosomes isolated from patients with non-muscle-invasive disease is necessary. 37
Andreu et al. 38 applied real-time qPCR to validate some of miRNAs selected in microarray detection in exosomes from 34 first morning urine samples collected from patients with bladder cancer (18 high-grade and 16 low-grade) prior to surgery and from 9 healthy volunteer donors. They isolated exosomes by differential centrifugation at 4°C and characterized them by electron microscopy and NTA. However, only miR-375 was significantly lower in high-grade bladder cancer patients compared to healthy volunteers, and miR-146a was significantly upregulated in exosomes from low-grade bladder cancer patients compared to high-grade.
Matsuzaki et al. 39 analyzed, using miRNA microarray, urinary exosomes collected from six patients having urothelial carcinoma (between admission and surgery) and three healthy volunteers. They isolated exosomes by differential centrifugation and characterized them by NTA and by the presence of CD9 using immunoelectron microscopy, western blotting, and ELISA. Then they used qRT-PCR to validate a higher expression for five candidate miRNAs in urinary exosomes of 36 patients compared to the 24 controls (donors for kidney transplantation, healthy volunteers, and postoperative patients of urothelial carcinoma). The results revealed that miR-21-5p was the most potent biomarker for the ability to detect urothelial carcinoma with an AUC of 0.900 (sensitivity and specificity were 75.0% and 95.8%, respectively), though the conclusions drawn are limited by the small sample population.
Baumgart et al. analyzed exosomes recovered using the Total Exosome Isolation reagent (Life Technologies, Carlsbad, California) from urine of 21 bladder tumor patients by qPCR for expression of selected miRNAs, already revealed to be differentially expressed in tumor tissues by microarray analysis but they showed no expression differences between muscle-invasive tumor patients and non-muscle-invasive tumor patients. 40
To date there are some pilot transcriptomics studies on urinary exosomes, but the data are too limited to implement these markers into routine screening programs.
While the studies above have focused on the utilization of exoRNA, only Lee et al. 41 investigated urinary exoDNA in nine patients who underwent radical cystectomy for urinary bladder cancer by targeted deep sequencing of nine genes frequently mutated in urinary bladder cancer: AT-rich interaction domain 1A (ARID1A), phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha (PIK3CA), fibroblast growth factor receptor 3 (FGFR3), HRas proto-oncogene (HRAS), lysine methyltransferase 2D (KMT2D), RB transcriptional corepressor 1 (RB1), TP53, lysine demethylase 6A (KDM6A), and stromal antigen 2 (STAG2). 41 For the isolation of urinary exosomes they used ExoQuick-TC (System Biosciences) reagent, and their presence was examined by electronic microscopy, characterized by western blot demonstrating the presence of PDCD6IP and TSG101, and by NTA. The average allele frequency of exoDNA in the analysis of somatic mutation was 65.6%, indicating an enriched burden of tumor DNA. The same samples presented a similar pattern of copy number aberrations with tumor samples.
Conclusions
In the present review, we summarized current urinary exosome studies to discuss the potential of exosomal shuttle RNAs in the diagnosis of lower urinary tract cancers. As the cargo of biological molecules reflects the patients’ situation, exosomes can be used as new diagnostic tools to serve as a non-invasive liquid biopsy. Supplementary research and development are mandatory for clinical applications, but the clinical utility of exosomes is promising. We actively look forward to the clinical use of exosomes, which will contribute to urological cancer management in the near future.
Footnotes
Declaration of conflicting interest
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
