Abstract
Purpose:
Bone metastases from non-small cell lung cancer (NSCLC) are common, and current prognostic stratification methods are challenging to predict outcomes. The aim of the study is to examine circulating tumor DNA as a potential biomarker to gauge overall survival.
Methods:
Late stage NSCLC patients associated with bone metastasis were recruited for the study. Circulating tumor DNA was quantified using droplet digital polymerase chain reaction, a sensitive assay that is capable to pick up low mutant DNA frequencies. KRAS and EGFR genes were serially profiled at monthly intervals to ascertain their variations in different patients during monitoring. These were correlated to overall survival as an endpoint measure.
Results:
Analysis of circulating tumor DNA and tumor tissues samples yielded a 94.7% overall agreement for KRAS and EGFR mutations. Higher circulating tumor DNA quantities of more than 1.4-fold were also detected in patients with bone metastases. To gauge the prognostic utility of circulating tumor DNA for these patient groups, circulating tumor DNA quantities as well as the patients’ genotypes were used to stratify patients in the survival analysis. Hazard ratios for patient groups with and without bone metastasis was 1.63. Patients groups separated by circulating tumor DNA quantity and molecular profile were 1.51 and 1.58, respectively. The results showed that circulating tumor DNA was a useful marker to identify patients with better survival outcomes.
Conclusion:
The good overall concordance in genetic profiling and circulating tumor DNA measurements associated with NSCLC patients with bone metastases supports plasma-based testing. This study presents exploratory evidence of the prognostic role of circulating tumor DNA that can be of value in the management of the disease.
Introduction
Cancer is one of the major causes of mortality worldwide (1) and warrants critical attention. Among the different types, lung cancer is a leading form and accounts for more than 1.8 million new cases yearly (2). The lethality of the disease is attributed to the difficulty in early detection and highly metastatic potential (3). In many cases, patients have progressed to the advanced stage during the first diagnosis and metastasis has occurred (4). Bone metastases from lung cancer are common, affecting about 30%–40% of the advanced stage patient population (5). The usual symptoms are considerable pain, occasionally fractures, or interference with daily activities (6).
Clinically, studies have shown that besides reducing the patient’s quality of life, bone metastases as a result of lung cancer is associated with poor prognosis (7). With the progress in therapy and alternative treatment options (8), early detection and monitoring of disease progression are important to enable the best care for patients in a timely fashion. Current detection modalities for bone metastases rely heavily on imaging methods (9), such as x-ray, computed tomography scans, and magnetic resonance imaging (MRI). These techniques are useful but do not provide the molecular profiles of patients that will be needed for therapy intervention. Alternative probing methods, such as circulating tumor DNA (ctDNA), have been demonstrated to be useful in detecting and monitoring clinical progression (10, 11). CtDNA that is present in blood as a result of necrotic or apoptotic tumor cells shedding genetic materials into the circulation (12) is highly correlative to lung cancer (13, 14). For instance, it has been demonstrated that ctDNA extracted from patients carried the same critical mutations associated with the disease (15). Also, monitoring these changes during the treatment cycles has been valuable for patient stratification with the endpoint measures for disease progression and overall survival (15). Its benefits also include being responsive to potential cancer treatment (16-18). This is useful to address the rapidly changing dynamics of the disease (19, 20). The most significant benefit for measuring disease changes using ctDNA is the ease in sample retrieval and extraction. Unlike invasive techniques, such as needle biopsy and surgical resection, tumor materials can be accessed via a straightforward peripheral blood draw. In other studies, it was determined that ctDNA could better epitomize the disease, as tumor tissues via subsampling might not be representative of the heterogeneous nature of the entire tumor mass and its secondary metastases (21).
In the current study, this addresses the non-small cell lung cancer (NSCLC) patients who had developed bone metastases. The primary aim is to correlate ctDNA levels with overall survival. The secondary study aims are to estimate KRAS and EGFR mutations, to compare ctDNA levels in late-stage NSCLC patients with and without bone metastases, and to estimate the agreement between ctDNA and primary tumor samples. This exploratory study will measure the clinical utility and prognostic ability of ctDNA for these patient groups.
Methods
NSCLC patient characteristics
The present study recruited 150 late-stage NSCLC patients. The participants of the trial gave their informed consent. The procedures used for sample collection strictly followed the guidelines approved by the institutional review board. Patients in the trial were consecutively enrolled to reduce selection bias. The study was divided into two groups: (a) patients with bone metastasis; and (b) a small proportion without. The patients were recruited from a single center. The adjudication process was based on MRI findings. Patients were treatment naïve at baseline and had complete molecular profiling performed for KRAS and EGFR during recruitment. Those without quality or insufficient tissues specimens were excluded, as were patients with cancer of unknown origin. Patients with existing cardiovascular diseases were also excluded. These patients’ cell-free DNA quantity may have been affected and may have prevented bias in the results. The detailed breakdown of the patient group is shown in Table I. For these patients, they were serially tracked and measurement was based on overall survival. This refers to the length of time from the date of first late-stage diagnosis of each patient until death from any cause.
NSCLC patient characteristics
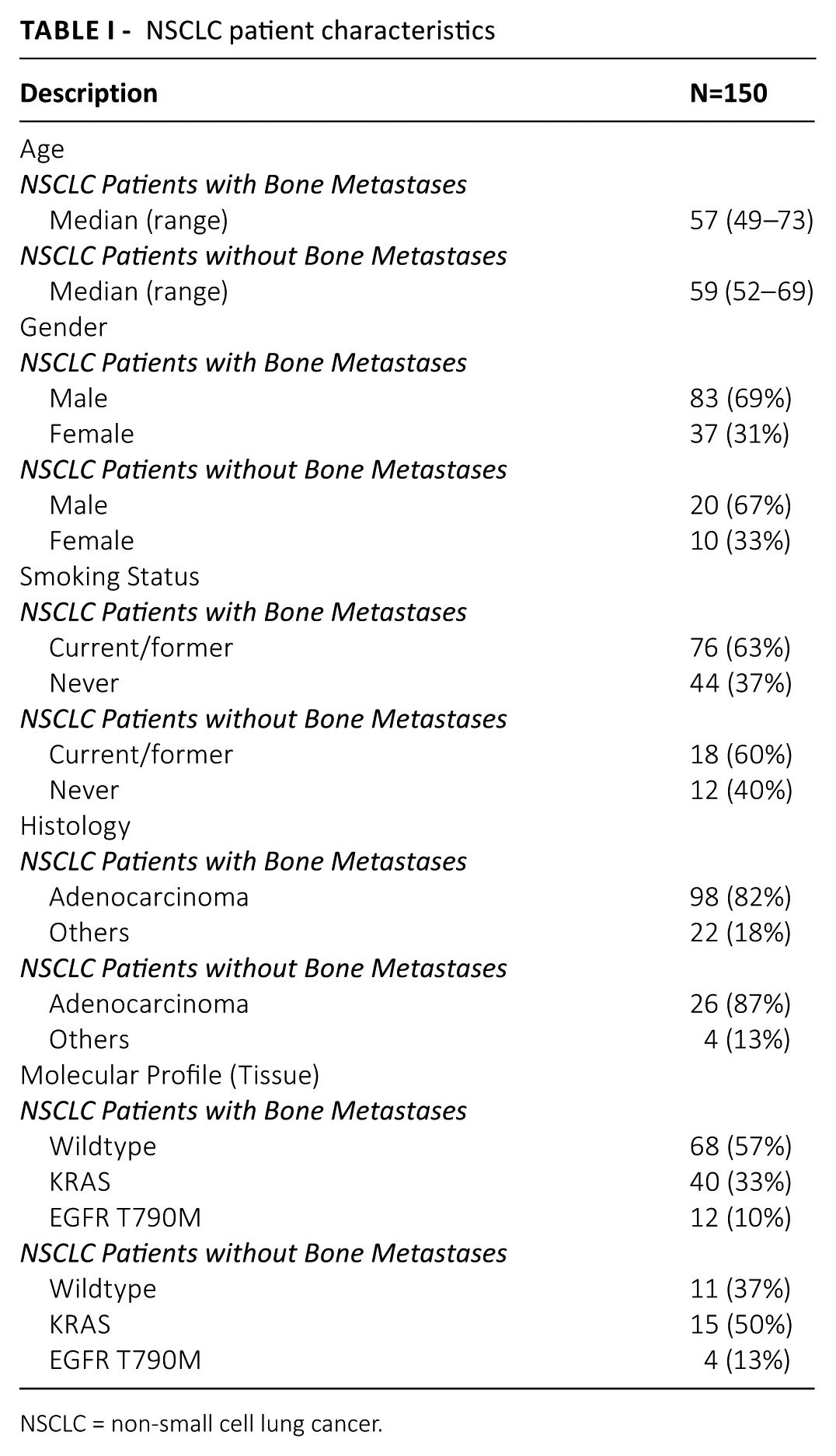
NSCLC = non-small cell lung cancer.
Specimen collection and ctDNA purification
Venous blood extraction was performed multiple times on each patient at different time points. This allowed serial comparisons and variations analysis to establish clinical correlations. Blood samples of 9–10 mL were drawn each time, collected in ethylenediaminetetraacetic acid (EDTA) tubes and processed within two hours. The first step for sample processing was to separate the blood plasma from whole blood. To achieve this, the blood specimen was spun down at 1000 g for 10 minutes, 4
Digital droplet polymerase chain reaction for mutation detection in ctDNA
Identification of key mutations prevalent in NSCLC was conducted using digital droplet polymerase chain reaction (ddPCR). Purified DNA was put through the QX100 ddPCR system (Bio-Rad Lab Inc., Hercules, CA, USA). Detection for KRAS and T790M was performed for each ctDNA sample. Primers and probes were purchased from Bio-Rad’s PrimePCR™ ddPCR Mutation Assays (Bio-Rad Lab Inc.). We performed assay validation runs to define the sensitivity of the assay. For this, spiked controls using KRAS and T790M mutant plasmids were used. For molecular profiling, the procedures followed the manufacturer’s recommendations. In brief, 20 uL reactions were prepared followed by PCR amplification. Thermocycling conditions were set as follows: 1.95
Statistical Analysis
Comparison of ctDNA quantities among different groups of NSCLC patients was done using the unpaired Student t test. Concordance with primary tissue analysis and ctDNA was correlated using the Cohen’s Kappa. The Kaplan-Meier method was used to estimate survival functions, and the hazard ratio was calculated using the Cox regression model to estimate the relative risk of event (i.e. death) between groups of patients. For all statistical analysis, the Prism software (Graphpad Inc., La Jolla, CA, USA) was used to compute the results. For all tests, a P value of less than 0.05 was considered statistically significant.
Results
Study design and ddPCR assay validation
In this current study, we aim to address how ctDNA variations affects the prognostication for late stage NSCLC patients, particularly for patients with bone metastasis. Our study recruited 150 patients and aimed to follow them through during their treatment process. Of the study cohort, 120 had confirmed bone metastasis, which was established from MRI analysis. Another 30 patients without bone metastasis were also recruited as the experimental control. The clinical staging was determined as per guidelines on the tumor nodes metastasis (TNM) classification (22). These patients were enrolled between January 2014 and March 2015. The median age for the entire study cohort was 58 years old with mainly adenocarcinoma histology. Baseline molecular profiles were obtained using tumor tissue samples extracted during the initial diagnosis. Patients had mixed profiles of KRAS and EGFR mutations mostly. The rest had wildtype characteristics. The proportion of patients with mutant KRAS and EGFR mutations determined using tumor tissues is shown in Figure 1A. For patients with bone metastases, 10% had T790M mutation (n=12), 33% were KRAS positive (n=40) and the rest were of wildtype genotype (n=68). For the control group without bone metastases, 13% had T790M mutation and 50% were KRAS positive. The rest were of wildtype genotype.
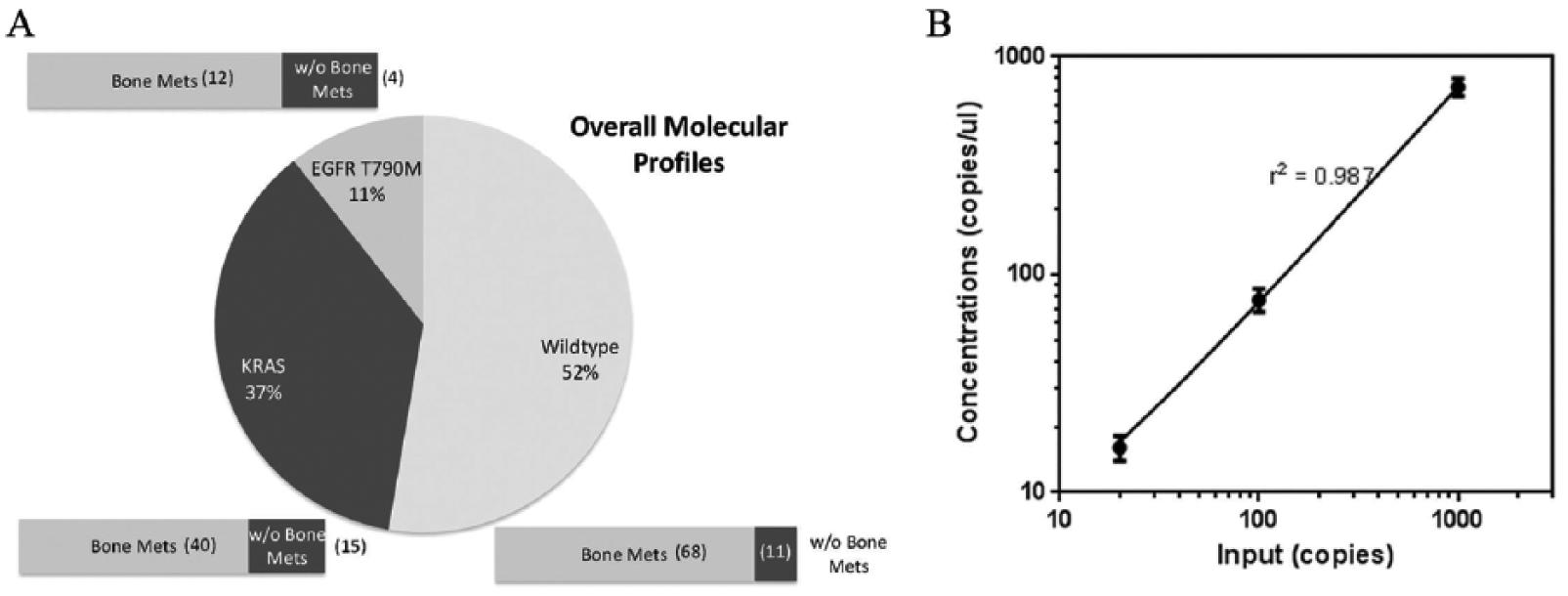
Molecular profiles of trial participants and detection assay validation. A) Tissue sample analysis for KRAS and EGFR T790M for patients with and without bone metastases. B) Linear regression analysis for spiked controls demonstrating good sensitivity and linearity.
It is important that the detection assay is sensitive as past literature has demonstrated that mutant signatures within ctDNA are of very low frequency (23). To validate the ddPCR assay, we included spiked conditions of 20, 100 and 1000 copies into wildtype genomic DNA. Figure 1b shows the results from the ddPCR analysis. Each setting was independently repeated five times. We observed good linearity in the results and were able to successfully detect samples with as little as 20 copies of mutant DNA. Fitting a linear model using regression analysis showed an r2 value of 0.987.
Baseline ctDNA and correlation to NSCLC with bone metastases
In our baseline measurements, we determined the association of ctDNA with different groups of NSCLC patients. The quantity of DNA extracted from blood plasma is shown in Figure 2A. As clearly demonstrated, the amount of ctDNA extracted from NSCLC patients with bone metastases is much higher than the group without bone metastases. We observed that the difference in mean quantities was 1.41-fold higher. Further analysis of the subtypes of patients within this patient group with bone metastases in terms of their molecular profiles showed other interesting trends. A one-way ANOVA showed significant differences in the mean values within the three different subgroups (Figure 2B, P < .05). We observed that the mean ctDNA quantity for KRAS or T790M-positive patient group, as detected using tissue profiling, was much higher than the wildtype group (unpaired Student t test, P < .05). No significant difference was observed comparing the KRAS and T790M patients (unpaired Student t test, P = .132). This seems to suggest that patients with critical disease mutations have a higher content of ctDNA.
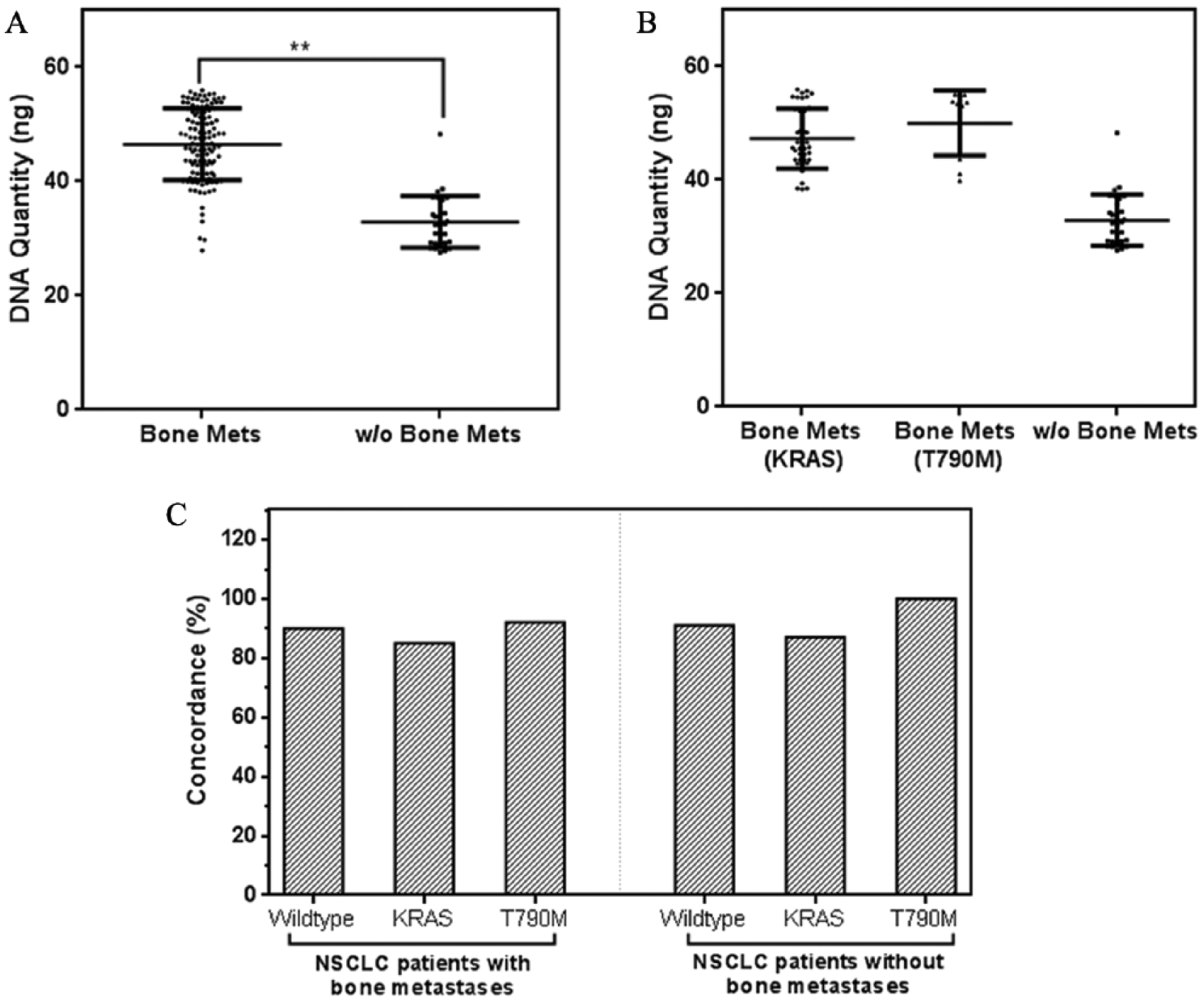
Baseline detection of ctDNA from patients with and without bone metastases. A) Direct ctDNA quantification between different groups of NSCLC patients show significant differences in mean extracted DNA. B) Patients split by their molecular profiles: those with bone metastases compared to those without bone metastases. C) Concordance measure demonstrates good agreement for ctDNA when referenced to tumor tissue samples.
To ascertain if the ctDNA detected is associated with the disease, we performed mutational profiling using ctDNA for KRAS and T790M for all patients. Figure 2C summarizes the results comparing them to tumor tissue profiling. Of the tissue and ctDNA samples, we observed an overall agreement of 88.7% (k = 0.773; 95% confidence interval (CI) 0.671, 0.874). As tabulated in supplementary Table 1, the sensitivity and specificity were both 89.9%. A further breakdown of the subgroups showed that for the KRAS and T790M-positive cohort, the concordance rates were 85.5% and 93.8%, respectively. The wildtype group had 89.9%. We also measured the variability for the concentration of mutant recovered ctDNA from each of the patient groups. Overall CVs for NSCLC patients with and without bone metastases for both KRAS and T790M-positive patients were 13.4% and 13.7%, respectively. KRAS and T790M-positive patients had coefficient of variations (CVs) of 11.1% and 11.4%, respectively. We observe that variations were relatively close for the different mutational profiles. Taken together, this suggested that ctDNA has a significant correlation with the disease, and its variability in DNA concentrations is relatively stable across different patients.
Importance of serial measurements and their association to overall survival
Patients were monitored monthly for six months, where possible, for ctDNA quantity changes as well as mutational profile variations. It was interesting that we observed persistently positive detection in the tumor tissue discordant cases detected at baseline (Figure 3A). We noted that this group of patients tended to have lower ctDNA concentrations compared with the rest of the NSCLC-positive cohort as shown in Figure 3B. We tested the hypothesis of whether lower ctDNA concentrations associated with the discordant cases were statistically significant. A one-tail Student t test comparing the mean detection concentrations showed it was significantly lower (P < .05). Of the mutation-positive discordant cases detected using ctDNA with tumor tissue profiles, we observed intermittent positive signatures at subsequent time point analysis for these nine patients (Figure 3C). Similarly, the retrieved concentration of mutant DNA was significantly lower than the rest of the positive patients (Figure 3D). Including these patients, the overall concordance rate for patients with bone metastases was 94.7% (k = 0.894, 95% CI 0.822, 0.965). Serial monitoring of patients showed several important changes in the disease. We noticed variations to the T790M profile for patients with initial wildtype genotype. As shown in Figure 3€, 24 patients were detected with the emergence of the mutation throughout the six-month period during their treatment phase. More importantly, we also observed that variations in ctDNA quantities had strong links to disease progression. In measuring the ctDNA variations during the monitoring period for an NSCLC patient with bone metastases, we noticed a deterioration in the condition, and a significantly large increase in ctDNA quantity (3.51-fold) at six months (Figure 3F).
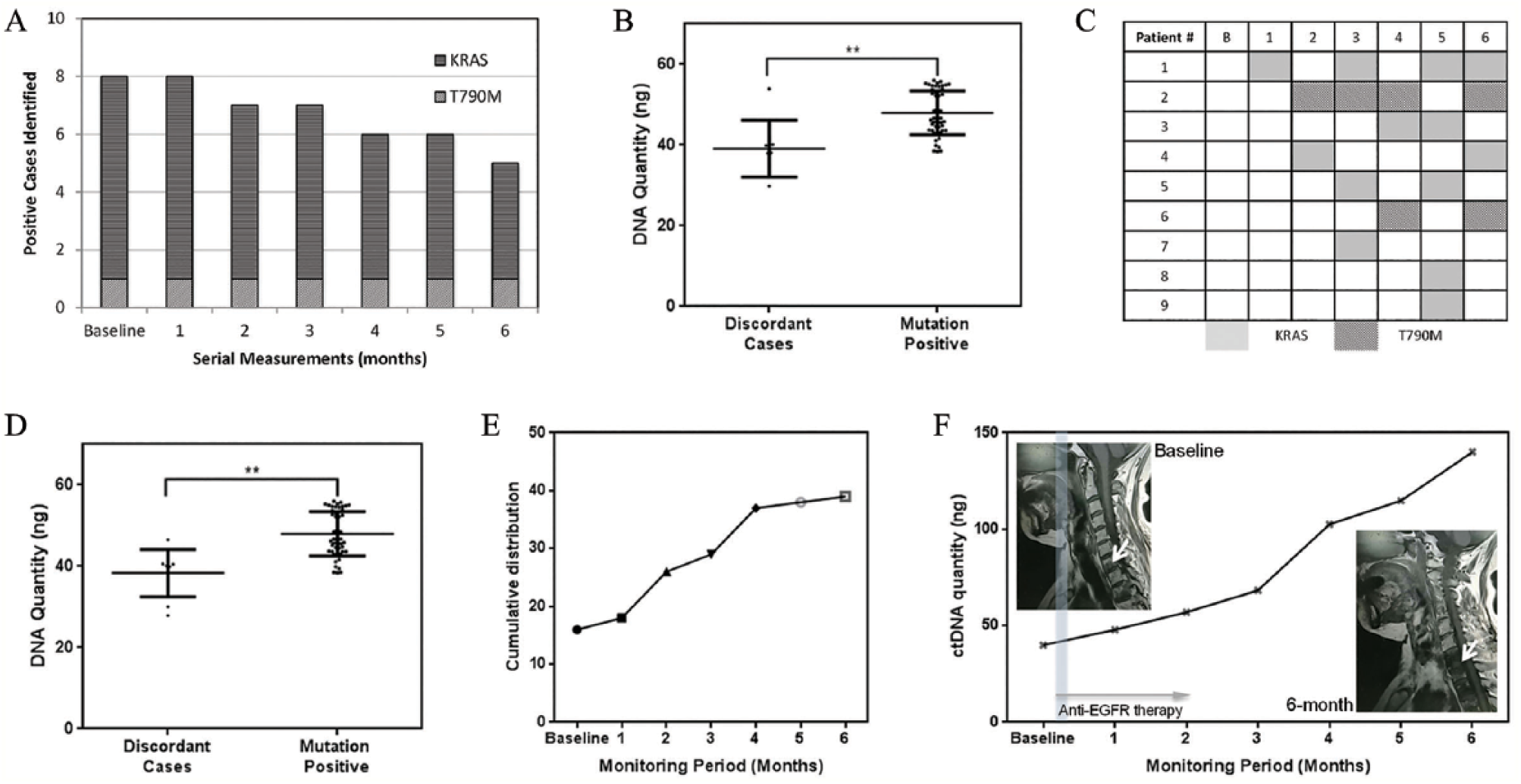
Serial ctDNA measurement and molecular profiling for NSCLC patients. A) Tissue discordant cases demonstrated at baseline show persistently positive detection throughout. Three patients died during this six-month period. B) Comparison of baseline ctDNA quantity for tumor tissue discordant cases. C) Intermittent positive detection for patients with baseline negative ctDNA mutational results. D) ctDNA discordant cases and measurement of DNA quantity. E) Frequency distribution of additional EGFR T790M detection among patients during monitoring. No other patients were detected with KRAS mutation other than the ctDNA discordant cases. F) Case study for serial measurements of ctDNA quantities and corresponding magnetic resonance imaging scans.
To gauge the prognostic utility, we performed a number of subgroup analyses with the endpoint measure of overall survival (OS). We assessed whether the patient groups with and without bone metastases had an effect on OS. As shown in Figure 4A, the hazard ratio determined using the COX regression estimate was 1.63 (95% CI 1.13, 2.49) with a P value of .020, which was statistically significant. Median survival for the NSCLC group with bone metastases was five months less (8 vs. 13 months). Within the NSCLC group for bone metastases, we further divided the cohort into subgroups to assess the effects of ctDNA and molecular profiles. We analyzed whether higher ctDNA quantity within these patients affected the OS, as we hypothesize that this could be closely related to severity. We divided the group by taking a median split of the ctDNA quantity recorded over the entire monitoring period for these patients. Indeed, we observed statistical significance (Figure 4B) using the Kaplan–Meier (KM) analysis (P = .031) and the hazard ratio was 1.51 (95% CI 1.07, 2.33). Further splits using the upper and lower quartile value showed that patients with a higher ctDNA content had a worse outcome (supplementary Figure 1). Median survival for the group with higher ctDNA quantity was 1 month less (eight vs. nine months). In Figure 4C, we compared the groups that were positively identified by ctDNA for KRAS and T790M mutations to the wildtype cohort. We also observed that for the former, the median survival was worse. The hazard ratio was 1.58 (95% CI 1.23, 2.89; P = .014). The results demonstrated the usefulness for ctDNA profiling within NSCLC patients with bone metastases, which will highlight the patients who are at higher risk.
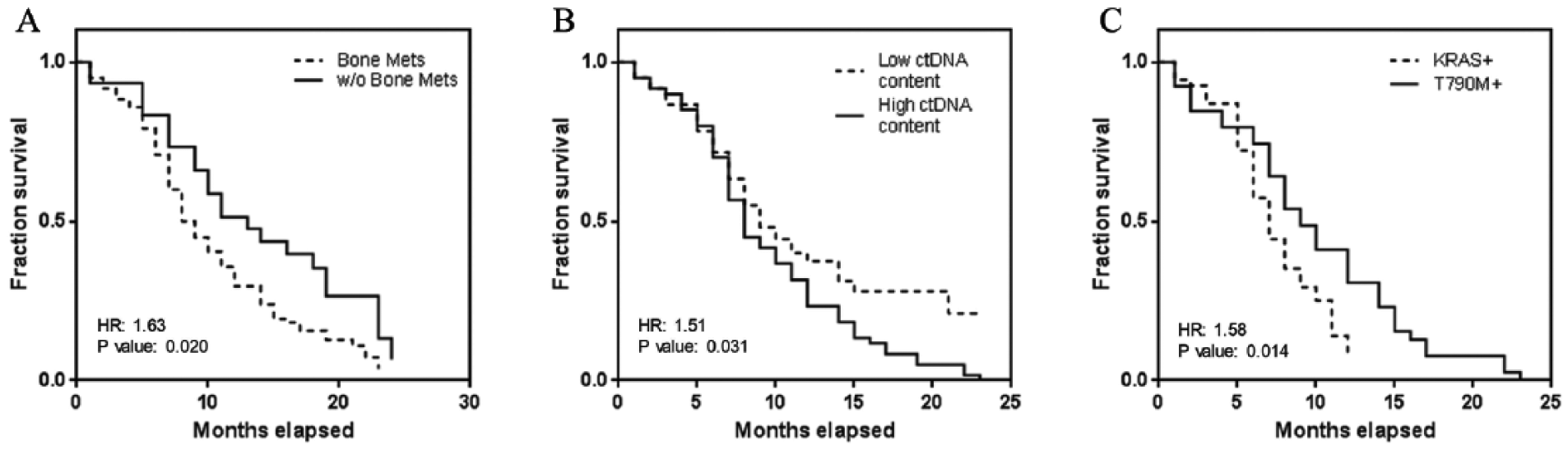
KM estimate for different profiles of NSCLC patients. A) Comparison of survival data for NSCLC patients with and without bone metastases. B) Comparison of survival data for NSCLC patients with bone metastases split by the extracted ctDNA quantity. C) Comparison of survival data for NSCLC patients with bone metastases by EGFR T790M or KRAS.
Discussion
In this study, we evaluated the association of ctDNA in NSCLC patients with and without bone metastases. Primarily, we were interested in understanding if the quantity of ctDNA and the molecular profiles associated with its detection would be useful as a prognostic biomarker. Lung cancer patients with bone metastases rely heavily on imaging modalities for diagnostic applications; however, this provides little benefit in subsequent prognostic utility. We envisioned that ctDNA would better address or supplement current disease management routines to probe the dynamics of the NSCLC, especially in this patient group. The current study was exploratory, and encompassed heterogeneous patient profiles (such as histology, age, and gender), which provided preliminary data for future confirmatory studies with a tighter selection of patient cohorts. The ease of sample retrieval, as shown in this study, makes the assay more attractive than invasive surgical procedures. ctDNA has been the focus of attention on a number of diseases and clinical applications (24). Its applications span from non-invasive prenatal screening tests to diagnostic and prognostic applications in different cancer types, such as NSCLC (25, 26), breast cancer (27), and prostate cancer (28). By using this approach, constant monitoring and detection can be provided, which, hopefully, will lead to better clinical predictions for NSCLC patients with bone metastases.
We observed several interesting trends related to NSCLC patients with bone metastases, and this is one of the first studies to address this group of patients using ctDNA. In our baseline measurements, we noted that NSCLC patients with bone metastases had higher ctDNA content compared with patients without bone metastases. We hypothesized that this could have been largely associated with the disease and its severity. Using ddPCR as a detection technique, we profiled these patients for two critical mutations, which are also fairly prevalent within the NSCLC cohort. The detection of KRAS and EGFR T790M mutations are important as these two mutations are strongly linked to weakened drug response in NSCLC patients (29-31). In a related study (32), variable mutation profiles linking to the EGFR mutation demonstrated key associations to variable responses to treatment. ddPCR is a sensitive method to detect genetic aberrations (33, 34) and was critically needed for our application as studies have shown that mutant ctDNA within blood plasma are of low frequency (35). In our validation experiments, we have shown that samples with as low as 10 mutant copies can be reliably detected. Using the detection method on clinical specimens, we noted good agreements of the molecular profiles of all NSCLC patients compared with their corresponding tumor tissue specimens. This presented evidence of the good clinical relevance of ctDNA to the disease in all different patient groups (Figure 2©). Our results are in good agreement with other studies that have shown the clinical significance of ctDNA (19, 21). For instance, studies have demonstrated that ctDNA represented well the genetic diversity within the disease (21), and its quantity within peripheral blood may be linked to tumor burden (19), or it may be associated with chemotherapy (36). In our study, we strongly noted these two attributes specifically with NSCLC patients with bone metastases. Additionally, our investigations showed that patients with bone metastases had a higher amount of ctDNA within circulation, which could have disease implications.
To be able to track the disease changes, we performed a monitoring trial for each patient for six months. Our results showed that this was important as it captured the dynamic variability of the disease. For instance, it was shown that certain patients had intermittent positive detection for the key driver mutations KRAS and EGFR T790M. We observed that this was likely due to the low concentrations within blood specimens, and serial monitoring of these patients effectively provided the confirmation status. In our trial to detect T790M mutations, we also noted the emergence of this mutation among wildtype patients detected at baseline. The ability to actively pick up patients with changes to their molecular profiles can effectively aid clinical intervention. For instance, NSCLC patients on EGFR first-generation tyrosine kinase inhibitors (TKIs) can switch to third-generation TKIs much earlier as the newer drugs act better on these mutations (37). We noted that several other studies that addressed advance stage NSCLC patients had similar observations for T790M emergence during monitoring using tissue rebiopsy (38, 39). Our technique is, effectively, more patient compliant and allows easier access to tumor materials for genotyping. Additionally, our analysis of these patients’ KRAS and EGFR profiles showed that ctDNA could better represent the disease molecular status and intra-tumoral heterogeneity. A small group of patients within our trial was positively identified for the two different mutations using ctDNA but was absent within tumor tissue tests. We were able to confirm the positive detection through subsequent specimens tests extracted from the same patient. The means to provide accurate genetic profiles for patients with bone metastases is critical to be able to give timely clinical interventions. Additionally, we realized that measuring ctDNA quantities was effective in tracking disease progression (Figure 3(f)). In one case study of our patients, we noted deterioration in the patient disease as confirmed from our MRI scans in a six-month period. This was accompanied by a sharp increase in ctDNA quantity compared with baseline. The benefits of ctDNA in disease management from both genetic profiling and quantification can provide better real-time updates to the disease.
Our main aim in the longitudinal study was to be able to accurately profile NSCLC, especially for patients with bone metastases. In the KM analysis with different patient subgroups, we noted that both ctDNA quantity and mutational status corresponded well to stratify patients with worse outcomes. This could potentially be used clinically to identify patients with a higher likelihood of poorer survival, and to justify the use of alternative treatments or maintenance therapies. In our analysis, patients with higher ctDNA quantities were observed to have a median survival of one month less than the other patients. Using the log rank test, this was statistically significant. We postulate that this was significantly related to the tumor burden, which some studies have shown (19). Our study further demonstrated that this contributed to worse survival outcomes for NSCLC patients with bone metastases. In our trial, we also observed that the survival analysis showed better outcomes for NSCLC patients that did not harbor the two critical mutations that were monitored. Median survival for the mutation positive group was two months less, and the hazard ratio showed a good split within the two patient groups (Figure 4©). This can be likely attributed to the treatment response of the patient groups, as KRAS and EGFR T790M are known to affect the efficacy of NSCLC drugs. We noted that the molecular profiling using ctDNA is effective to separate out the high-risk patient group, and that these patients could benefit highly from constant monitoring of the ctDNA profiles. Our study demonstrated clear clinical associations of ctDNA and its use for NSCLC with bone metastases, which may lead to better clinical prognosis and potential clinical intervention measures.
The main limitation observed in our study for the current methodology was the low mutant DNA frequencies seen in several patients. This presents strong requirements in detection assays to prevent false negative results that may impact prognostic predictions. We noted that this can be compensated by continual serial monitoring of patients’ blood specimens as shown in our study, or by the use of more sensitive molecular assays, such as ICE-COLD PCR (40) or BEAMING (41), which will be able to enhance the purity of mutant DNA. Alternatively, sensitive detection has been used in single circulating tumor cell analysis (42) or microRNAs (43) that coexisted in blood specimens. These methods will complement the current study to reduce false negatives or to provide alternative materials that also may be suitable for disease probing.
The current study was mainly exploratory and the perspectives of using ctDNA in clinical practice are numerous: to monitor the genetic evolution of tumor; to identify earlier stage NSCLC patients at high risk for relapse who may benefit from additional adjuvant treatment; and the potential use to monitor treatment response. Validating the role of plasma genotyping will require other well-planned clinical studies.
Conclusion
ctDNA can be useful in the management of late-stage NSCLC, and the current exploratory study has demonstrated several promising advantages for serial monitoring and prognostication. This forms the basis for future confirmatory studies, which can bring the use of ctDNA closer to routine clinical use. Our results demonstrated good concordance rates to tumor tissue specimens and good clinical relevance. The main advantage of ctDNA measurements is better patient compliance, as sample extraction is minimally invasive. This provides opportunities for long-term study of the disease, which may aid in future treatment monitoring.
Supplemental Material
Supplementary_Materials – Supplemental material for Circulating tumor DNA as prognostic markers for late stage NSCLC with bone metastasis
Supplemental material, Supplementary_Materials for Circulating tumor DNA as prognostic markers for late stage NSCLC with bone metastasis by Jiguang Jia, Bin Huang, Zhengling Zhuang, Sen Chen in The International Journal of Biological Markers
Footnotes
Author contributions
Both authors contributed equally to this study.
Disclosures
Financial support: This study was supported by a research grant provided by the Xiangyang No.1 People’s Hospital, Hubei University of Medicine.
Conflict of interest: None of the authors has financial interest related to this study to disclose.
Ethical approval
All human and animal studies have been approved by the appropriate ethics committee and have therefore been performed in accordance with the ethical standards laid down in the 1964 Declaration of Helsinki and its later amendments.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
