Abstract
Keywords
Introduction
Behçet syndrome (BS), also known as Behçet disease, is a chronic, relapsing, multisystemic inflammatory disorder of unknown etiology. 1 Initially described by Dr Hulusi Behçet as a “triple symptom complex” of oral and genital ulcers and uveitis, BS is currently classified as a complex autoinflammatory disease characterized by a variable vessel vasculitis pathology.2–4 In addition to the hallmark mucocutaneous manifestations of BS, the syndrome may also present with diverse ocular, musculoskeletal, gastrointestinal, pulmonary, cardiovascular, and central nervous system lesions.1,4,5
Since its first definition in 1937, some important characteristics of BS have drawn considerable attention, albeit with no clear explanations. 2 The classical high prevalence geography of BS coinciding with the ancient trade route “Silk Road,” the discordant, sometimes paradoxical immunological findings present in BS studies, the divergent clinical phenotypes of the syndrome observed in different geographic regions, and the occurrence of predefined disease clusters with shared clinical manifestations among BS patients are some of these characteristics waiting for explanations.4,6–8 Still another interesting observation is that, even though BS occurs all around the globe, except for its characteristic “Silk Road” geography, BS tends to follow a milder clinical course.9,10
Previously, Oguz et al. were able to document significant gene expression and molecular disease mechanism differences among BS clinical subsets/phenotypes in two different populations (i.e., Portuguese and Turkish).11,12 The presence of such remarkable discrepancies, even among BS patients of the same population, brings to mind the case of BS patients of different ethnicities.
The present study contrasted the gene expression profiling data sets of BS patients of two populations of different ethnicities (i.e., Portuguese and Turkish), first documenting the differentially expressed gene (DEG) sets and then analyzing these DEG sets further with multiple bioinformatics tools. Our aim was to display molecular level similarities and/or discrepancies between Portuguese and Turkish BS patients, which may account for the observed clinical heterogeneity of BS between these two populations.
Methods
The design of the study (secondary analysis of publicly available transcriptome datasets) enabled the researchers to proceed without receiving an ethics committee approval. By clearly depicting the identity of the source database (i.e., Gene Expression Omnibus), the dates of accession, the accession numbers, and the authors of the borrowed datasets, the authors have fulfilled all the obligatory ethical prerequisites for a secondary analysis research paper. The study continued between March 2024 and July 2024. The flowchart summarizing the study is presented in Figure 1. The flowchart of the study. doi.org/10.1007/s00109-013-1022-4,
13
doi.org/10.1002/iid3.836
12
BS: Behçet syndrome; DOI: Digital Object Identifier; GEO: Gene Expression Omnibus; NCBI: National Center for Biotechnology Information; PB: the group inclusive of all Portuguese BS patients; PC: Portuguese control group; PMB: Portuguese mucocutaneous BS group; POB: Portuguese ocular BS group; PVB: Portuguese vascular BS group; RMA: Robust Multiarray Average; TB: the group inclusive of all Turkish BS patients; TC: Turkish control group; TDB: Turkish complex phenotype BS group; TMB: Turkish mucocutaneous BS group; TOB: Turkish ocular BS group; TVB: Turkish vascular BS group.
The gene expression profiling studies by Xavier et al. and Oguz et al
Although their designs showed certain differences, the profiling studies by Xavier et al. and Oguz et al. both included BS patients diagnosed according to the internationally recognized criteria, used GeneChip® Human Genome U133 Plus 2.0 (Affymetrix, Santa Clara, CA, USA) microarrays for hybridization, and were conducted and reported in accordance with the Minimum Information About Microarray Experiment (MIAME) guidelines.12–15
Retrieval of the expression data
The raw microarray data of the study by Xavier et al. was previously retrieved from Gene Expression Omnibus (GEO) data repository using the GEO Accession GSE17114 on August 25th, 2015.13,16 The metadata, raw, and final/normalized data of the study by Oguz et al. were deposited to GEO with the accession GSE209567 on July 22nd, 2022.12,16
The demographic and clinical data of the study population and the definition of the clinical subsets of patients with Behçet syndrome
Clinical characteristics defining individual Behçet syndrome clinical phenotypes.
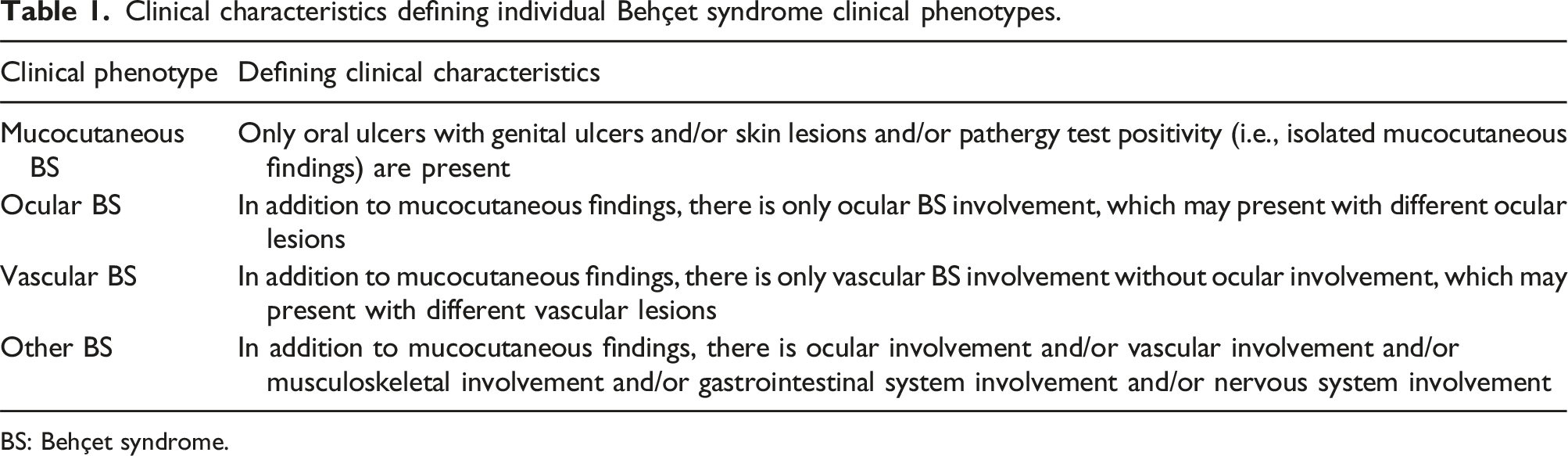

BS: Behçet syndrome.
Excluded samples/cases
Following Oguz et al.’s two studies, the sample GSM428038 from the series GSE17114 (as an outlier in the PVB group) and the samples GSM6376933, GSM6376934, GSM6376935, GSM6376936, GSM6376937, GSM6376938, and GSM6376939 from the series GSE209567 (as complex phenotype BS patients in the TDB group) were excluded from the study (Figure 1).11,12 Regarding the sample GSM428038, according to the information contained in Oguz et al.’s article, while this sample phenotypically belonged to a vascular BS patient, during clustering analysis it consistently clustered with mucocutaneous BS patients. 11 Additionally, the two rationales underlying the exclusion of Turkish other BS (TDB) patients were their significantly low counts in each individual clinical phenotype group and the absence of their corresponding Portuguese counterparts.
Statistical analysis of the demographic characteristics
As the demographic data of the study population did not follow a normal distribution, the median as a measure of central tendency, and the minimum and the maximum values were presented. For the comparisons between two independent groups, the Independent-Samples Median Test and the Chi-Square tests were used. The p values of the analyses were presented with their absolute numerical values and a p value of ≤0.05 was considered statistically significant. Statistical analyses of the demographic data were performed using “SPSS for Windows, Version 16.0” software (SPSS Inc., Chicago, USA).
Pre-processing of the microarray data
Before continuing with the downstream analyses, pre-processing of the microarray data was performed using BRB-ArrayTools Version 4.6.2 Stable Release developed by Dr Richard Simon and BRB-ArrayTools Development Team (Figure 1). 22 The gene expression data present as individual raw .cel files was collated by the data import function of BRB-ArrayTools. The Robust Multiarray Average (RMA) algorithm was implemented for the normalization of the microarray data. 23 Following the normalization step, the replicate spots within each array were averaged and finally, gene filters were used which helped to exclude genes that were not likely to be informative. For this purpose, a “minimum fold-change filter” in addition to a “percent missing filter” were used, which were ready-to-use functions of the BRB-ArrayTools software. 22 As described here, the collation step of the study samples, their collective RMA normalization, the averaging of the replicate spots, and the filtering of uninformative genes were all executed with BRB-ArrayTools’ built-in functions and all the default settings were preserved. 22
Bioinformatics analyses of the gene expression data
Class comparison analyses were performed using BRB-ArrayTools (Version 4.6.2 Stable Release). 22 Two-sample t-test with a random variance model was implemented for class comparison analysis of two classes. Differentially expressed genes were selected using a p value of ≤0.05 and fold changes (FC) of ≥1.5 and ≥2.
Comparisons of the DEG sets with Venn diagrams were performed using Venny 2.1 by Juan Carlos Oliveros. 24 For Venn diagram analyses, DEGs with p ≤ 0.05 and FC ≥2 were used.
BRB-ArrayTools’ (Version 4.6.2 Stable Release) built-in clustering tools, Cluster 3.0 (developed by Michael Eisen and the Stanford group and modified by Michiel de Hoon), and Java Treeview softwares were used for clustering analyses.22,25,26 A hierarchical clustering algorithm using Euclidean distance metric and average linkage was implemented and both the samples/cases and the genes were clustered.
Functional enrichment analyses of the DEG sets were performed with the WEB-Based Gene Set Analysis Toolkit (WebGestalt) and the chosen method of Over-Representation Analysis (ORA) specifically focused on the functional database of gene ontology biological process. 27 In order to provide sufficient number of genes with the uploaded gene lists to WebGestalt, DEG lists with p ≤ 0.05 and FC ≥1.5 were used. For the reference set, “affy hg u133 plus 2” was chosen.
Results
Demographics of the study population
Key demographics of the study population.
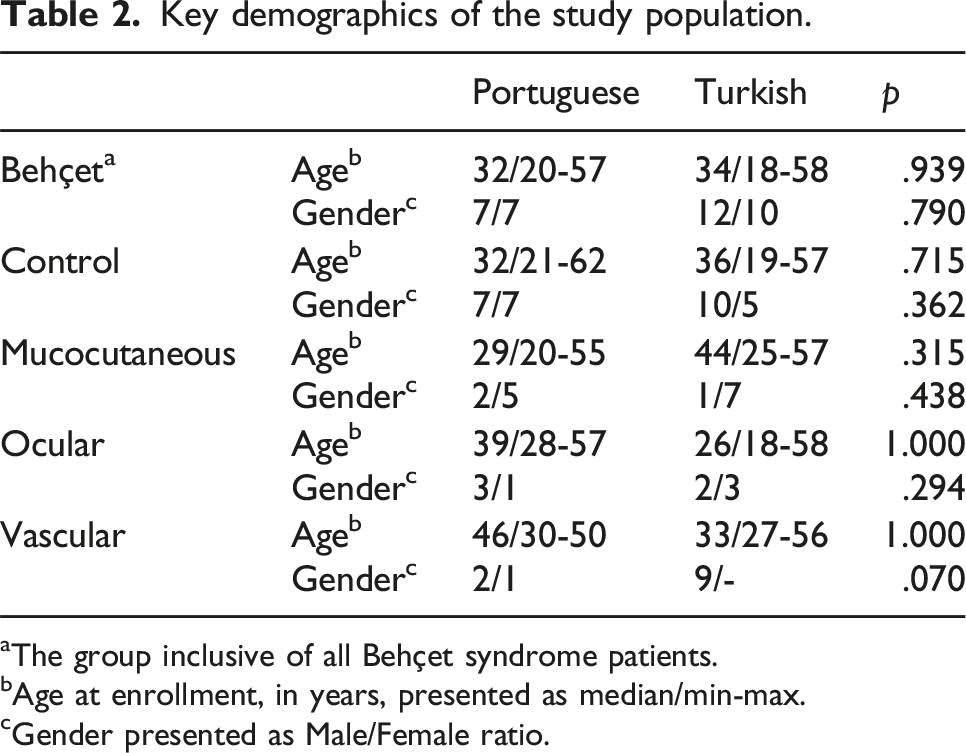
aThe group inclusive of all Behçet syndrome patients.
bAge at enrollment, in years, presented as median/min-max.
cGender presented as Male/Female ratio.
Pre-processing results of the microarray data
The box-and-whisker plots of the normalized microarray expression data are presented in Figure 2. The box-and-whisker plots of the gene expression data normalized using RMA. Samples/cases are along the horizontal axis while log2 intensity values for each gene are plotted on the vertical axis. The black line in the middle of each box plot is the median value for that sample.
Following the pre-processing steps performed on the collated 65 .cel files, 16060 probe sets out of the 54675 present on the GeneChip® Human Genome U133 Plus 2.0 microarrays passed the adjusted filters and were used during the subsequent bioinformatics analyses.
Class comparison analyses
The number of the DEGs obtained during the country specific class comparison analyses.

aIncreased expression in the first class (e.g., B) compared to the second class (e.g., C).
bDecreased expression in the first class (e.g., B) compared to the second class (e.g., C).
B: the group inclusive of all Behçet syndrome patients; C: the control group; FC: fold change; M: mucocutaneous Behçet syndrome group; O: ocular Behçet syndrome group; V: vascular Behçet syndrome group.
The number of the DEGs obtained during the class comparison analyses between the two (Portuguese and Turkish) study populations.

aIncreased expression in the first class (e.g., PC) compared to the second class (e.g., TC).
bDecreased expression in the first class (e.g., PC) compared to the second class (e.g., TC).
FC: fold change; PB: the group inclusive of all Portuguese Behçet syndrome patients; PC: Portuguese control group; PMB: Portuguese mucocutaneous Behçet syndrome group; POB: Portuguese ocular Behçet syndrome group; PVB: Portuguese vascular Behçet syndrome group; TB: the group inclusive of all Turkish Behçet syndrome patients; TC: Turkish control group; TMB: Turkish mucocutaneous Behçet syndrome group; TOB: Turkish ocular Behçet syndrome group; TVB: Turkish vascular Behçet syndrome group.
aThe table displays the top 5 genes with increased and the top 5 genes with decreased expressions for each class comparison.
bThe gene lists exclude gonosomal genes occurring as a result of Male/Female ratio disparities between the compared classes.
cIncreased and decreased expression of genes are always in the patient group with respect to the control group.
B: the group inclusive of all Behçet syndrome patients; C: the control group; FC: fold change; M: mucocutaneous Behçet syndrome group; O: ocular Behçet syndrome group; V: vascular Behçet syndrome group.
The top 10 most differentially expressed genes observed during the class comparison analyses between the two (Portuguese and Turkish) study populations a .

aThe table displays the top 10 genes with increased and the top 10 genes with decreased expressions for each class comparison.
bIncreased expression in the first class (e.g., PB) compared to the second class (e.g., TB).
cDecreased expression in the first class (e.g., PB) compared to the second class (e.g., TB).
FC: fold change; PB: the group inclusive of all Portuguese Behçet syndrome patients; PMB: Portuguese mucocutaneous Behçet syndrome group; POB: Portuguese ocular Behçet syndrome group; PVB: Portuguese vascular Behçet syndrome group; TB: the group inclusive of all Turkish Behçet syndrome patients; TMB: Turkish mucocutaneous Behçet syndrome group; TOB: Turkish ocular Behçet syndrome group; TVB: Turkish vascular Behçet syndrome group.
Venn diagram analyses
The illustrations of the PB versus PC ∩ TB versus TC, PMB versus PC ∩ TMB versus TC, POB versus PC ∩ TOB versus TC, and PVB versus PC ∩ TVB versus TC Venn diagram analyses are presented in Figure 3 (Supplemental Material Table S3). It was striking to see the PB versus PC ∩ TB versus TC and the PMB versus PC ∩ TMB versus TC intersection sets empty (i.e., no shared genes) and the POB versus PC ∩ TOB versus TC intersection set with only 2 genes (Figure 3). The diagrams of the: (A) PB versus PC ∩ TB versus TC, (B) PMB versus PC ∩ TMB versus TC, (C) POB versus PC ∩ TOB versus TC, and (D) PVB versus PC ∩ TVB versus TC Venn diagram analyses (genes in POB vs PC ∩ TOB vs TC: CA1, DEFA4, and in PVB vs PC ∩ TVB vs TC: DCAF16, SLC16A7, IL18R1, EBLN3P, UBA2, PM20D2, HSPA13, SESN3, TMEM267, GFM1, SLC6A8, SLAIN1, PRSS23, TTC37, SNCA, SRSF10, GTF2H3, ETNK1, MEX3C, SHPRH, PYHIN1, TRIM13, MDFIC, IFT80, PRNP, UFL1, EIF1AX, ATF1, PHAX, ATP2B1, HBD, AHSP).
Cluster analyses
The dendrogram and heatmap representations of the results of the cluster analyses of Portuguese and Turkish BS patients and patient subsets are presented in Figure 4 (Supplemental Material Table S6). As shown in Figure 4, the gene sets used and the algorithm implemented successfully clustered both Portuguese and Turkish BS patients and Portuguese and Turkish BS clinical subsets. Important to note, the gene sets used for clustering purposes were constructed from the top 10 most differentially expressed genes observed during the population specific class comparison analyses. The dendrogram and heatmap representations of the results of the cluster analyses: (A) PB & TB, (B) PMB & TMB, (C) POB & TOB, and (D) PVB & TVB. Both the patients (columns) and the genes (rows) are clustered and for ease of demonstration, only the dendrograms for clustering of the patients are shown on the heatmaps. The algorithm chosen and the gene sets implemented (Supplemental Material Table S6) successfully clustered Portuguese and Turkish BS patients.
Functional enrichment analyses
The results of the population specific functional enrichment analyses performed individually with DEG sets with increased and decresed expression are schematically presented in Figure 5 (DEGs with increased expression) and Figure 6 (DEGs with decreased expression) and given in detail in Supplemental Material Table S7. Strikingly and unexpectedly, not even a single gene ontology biological process term was common in Portuguese and Turkish populations’ functional enrichment analyses top 15 terms (Figures 5 and 6). Schematic presentation of the functional enrichment analyses of the individual Portuguese and Turkish class comparison DEG sets with increased expression levels. PB: the group inclusive of all Portuguese Behçet syndrome patients; PC: Portuguese control group; PMB: Portuguese mucocutaneous Behçet syndrome group; POB: Portuguese ocular Behçet syndrome group; PVB: Portuguese vascular Behçet syndrome group; TB: the group inclusive of all Turkish Behçet syndrome patients; TC: Turkish control group; TMB: Turkish mucocutaneous Behçet syndrome group; TOB: Turkish ocular Behçet syndrome group; TVB: Turkish vascular Behçet syndrome group. Schematic presentation of the functional enrichment analyses of the individual Portuguese and Turkish class comparison DEG sets with decreased expression levels. PB: the group inclusive of all Portuguese Behçet syndrome patients; PC: Portuguese control group; PMB: Portuguese mucocutaneous Behçet syndrome group; POB: Portuguese ocular Behçet syndrome group; PVB: Portuguese vascular Behçet syndrome group; TB: the group inclusive of all Turkish Behçet syndrome patients; TC: Turkish control group; TMB: Turkish mucocutaneous Behçet syndrome group; TOB: Turkish ocular Behçet syndrome group; TVB: Turkish vascular Behçet syndrome group.

Discussion
This study sought to investigate the molecular immunopathogenetic similarities and differences underlying BS across two different ethnic populations, Portuguese and Turkish. As Oguz et al.’s two previous studies pointed to the potential pathogenetic role of certain innate immunity genes (i.e., CD69, TNFAIP3, CLEC12A, CLC, and IFI27) in BS, specific focus was on BS’s fundamental innate immunity pathology.11,12 Our main findings highlighted significant gene expression and molecular disease mechanism differences between Portuguese and Turkish BS patients, particularly regarding the anti-inflammatory and the pro-inflammatory genes. While decreased expression of anti-inflammatory genes was a common theme in BS patients of these two populations, the identities of these genes were distinct in each population. Additionally, for Turkish BS patients, increased expression of certain pro-inflammatory genes (i.e., IFI27) might be responsible for the relatively severe disease manifestations and the disease course in this population. As the authors we comment that, the anti-inflammatory and the pro-inflammatory genes are the key to understand the immunopathogenesis of BS 12 and the documented expression differences among the two different ethnicities in this study, may account for the observed discordant, sometimes paradoxical immunological findings present in BS studies and also the divergent clinical phenotypes of the syndrome observed in different geographic regions.7,8 Additionally, the distinct DEGs and the different molecular mechanisms observed across ethnicities, reinforces the concept of BS as a syndrome rather than a single disease entity.
Analysis of demographic characteristics revealed that BS and control groups, as well as the three BS clinical phenotypes (i.e., mucocutaneous, ocular, and vascular), were comparable in terms of age and gender distribution across the two different ethnicities (Table 2). While Turkish BS patients showed a slight male predominance, Portuguese BS patients had an equal Male/Female ratio, which aligns well with the existing literature.4,8,10 Furthermore, a gender-based difference in disease severity was observed among Turkish BS patients. Males were more prominent in the vascular BS group, while the mucocutaneous subset showed a Male/Female ratio of 1/7 (Table 2). This pattern was absent in Portuguese population, consistent with the literature indicating a male predominance in more severe BS phenotypes.4,8
Class comparison analyses revealed notable differences between the two populations. When comparing each population’s BS patients to its respective control, the DEG counts ranged from single to triple digits (min.: 4, max.: 549, p ≤ 0.05, FC ≥2.0) (Table 3). In contrast, comparisons between Portuguese and the Turkish BS groups revealed DEG counts in the four-digit range (min.: 7466, max.: 8642, p ≤ 0.05, FC ≥2.0) (Table 4). Subtracting the Portuguese Control versus Turkish Control class comparison’s DEGs (6549, p ≤ 0.05, FC ≥2.0) did not substantially lower these DEG counts. This finding suggests that, the gene expression differences observed in BS groups are not solely attributable to ethnic background. These substantial molecular discrepancies between the BS groups reinforce the idea that distinct pathways underlie BS across different populations. 1 Comparison of the top 10 DEGs demonstrated minimal overlap between Portuguese and Turkish BS patients, with only CA1 shared among the two populations (Table 5). Notably, previously identified innate immunity genes with potential pathogenetic contribution to BS (i.e., CD69, IFI27, CLEC12A, and CLC) were not shared in the top 10 DEG lists of the two populations (Table 5).11,12 This finding indicates that while gene functions may be similar, the exact molecular drivers differ across populations in BS. The same finding also aligns with our hypothesis that the gene expression and molecular mechanisms involved in BS likely vary by ethnicity, further supporting the notion of BS as a heterogeneous syndrome.
In Portuguese BS patients, CD69 showed increased expression across patient sub-groups relative to their Turkish counterparts (Table 6). The CD69 gene, previously associated with milder disease expression in Portuguese BS patients by Oguz et al., suggests a protective, anti-inflammatory role that might contribute to milder BS manifestations in Portuguese patients.11,28–30 Conversely, the involvement of genes such as IFI27, CLEC12A, and CLC in Turkish BS patients suggests a more severe clinical presentation in this population (Table 5). 12 Together, these findings support a model where distinct genetic profiles influence BS severity across ethnicities, potentially providing a molecular basis for population-specific therapeutic strategies in future.
Venn diagram analyses revealed no shared DEGs in the intersections of PB versus PC ∩ TB versus TC and PMB versus PC ∩ TMB versus TC (Figure 3). It was striking, especially not to see even one shared gene in the PB versus PC ∩ TB versus TC set. When the relatively more severe forms of BS (i.e., ocular and vascular BS) were analyzed, the number of shared DEGs increased (2 for POB vs PC ∩ TOB vs TC [out of 166 genes] and 32 for PVB vs PC ∩ TVB vs TC [out of 733 genes]) (Figure 3). This may be due to common gene expression modifications occuring secondary to prolonged and severe tissue damage and/or therapeutic effects in these more severe BS phenotypes. In cases of severe BS, especially in those patients with organ and/or life threatening disease, common immunosuppressive or immunomodulatory treatments are often used to limit the disease activity with potential consequences in gene expression profiles. 31
Clustering analyses successfully distinguished Portuguese and Turkish BS patients and their respective clinical phenotypes using only around 20 genes per analysis (Figure 4). Notably, IFI27 emerged as the only shared gene across clustering DEG lists, underlining its potential importance as a key pro-inflammatory gene in Turkish BS patients. 12 This finding may (1) support the view that IFI27 may play a pivotal role in distinguishing BS patients across ethnicities, (2) emphasize its relevance in BS pathogenesis in Turkish population, and (3) further underline innate immunity and inflammatory mechanisms as central in BS pathogenesis. Also, as the authors we hypothesize that, given the role of IFI27 in driving pro-inflammatory pathways, it may represent a key therapeutic target for BS management, at least in Turkish population.12,32,33
The functional enrichment results further confirmed significant molecular differences between Portuguese and Turkish BS patients. No shared enriched gene ontology terms were identified among the top 15 in either population whether using DEGs with increased or decreased expression (Figures 5 and 6). Functional enrichments differed significantly between Portuguese and Turkish BS patients, with Portuguese patients showing pathways related to neutrophil degranulation and immune response, while Turkish patients displayed enrichment in Type I interferon signaling and T-cell receptor pathways (Figures 5 and 6). This lack of common functional enrichments between the two populations again supports the hypothesis that BS’s molecular mechanisms vary by ethnicity.
Both innate and adaptive immune responses are held responsible in BS pathogenesis.34–38 Recently, a pro-inflammatory and mainly Th1/Th17 type cytokine profile was also documented in BS patients.39–41 It was suggested by Direskeneli that, BS might be linked to a primary immune alteration resulting from a genetic variation, effecting an adhesion molecule, a proinflammatory cytokine or chemokine or a transcription/regulatory factor, that predisposes the individual to intense neutrophil and T cell responses. 42 As the authors, we believe that the swift and exquisite immune response observed in BS patients, may well be secondary to supressed anti-inflammatory activity as well as enhanced pro-inflammatory activities. Therefore, taking into account the findings of our study, we may suggest that CD69 and CLEC12A, both C-type lectin domain containing inhibitory receptors expressed in various leukocytes including neutrophils, NK, and T cells, may be added to Direskeneli’s list. 42 On the other hand, CLC (also known as GAL10), recently shown to be a cytoplasm located protein, seems to play a vital role for regulatory T cells’ immune homeostasis functions.43,44 So, at least these three genes (i.e., CD69, CLEC12A, and CLC) with their important anti-inflammatory functions and decreased expressions, may be contributing to the pro-inflammatory milieu observed in BS. In addition to the Pathergy reaction characteristically observed in BS patients, “skin urate” reaction believed to occur as a result of increased neutrophil responses to urate crystals was also demonstrated in BS patients. 45 It is interesting that CLEC12A, which is an anti-inflammatory brake functioning in neutrophils, is also a uric acid crystal receptor that attenuates inflammation in response to cell death. 46 A hypothesis article by Oguz et al. tried to explain the links between CLEC12A, BS, gout, and the therapeutic agent colchicum. 47 IFI27 was another noteworthy gene in our study. Well-known for its interferon-stimulated gene property, IFI27 is presented as an emerging key player in immune responses. 48 While there is only one article touching on IFI27’s potential contribution to BS pathogenesis, IFI27 has been studied in several inflammatory conditions with significant differential expression, suggesting a possible involvement in immune dysregulation, disease severity, and pathogenesis.12,48
Limitations
We must acknowledge several limitations regarding the study. First, this in-silico study was relying on microarray data, which, while valuable, lacks the advantages of next-generation sequencing methods. As is well known, RNA sequencing could capture rare transcripts and identify genetic variants which are not detectable in micro-arrays, potentially expanding our understanding of BS pathogenesis. 49 While single nucleotide polymorphisms, noncoding RNAs with regulatory functions, and novel transcripts may carry important implications for BS pathogenesis, these can only be documented with next generation sequencing data. On the contrary, microarrays harbor important design bias and are not able to display such important genomic and transcriptomic details. Furthermore, this study was limited to Portuguese and Turkish populations, which clearly is not able to represent BS heterogeneity globally. Additionally, treatment effects on gene expression cannot be ruled out as BS patients enrolled in the study were known to continue their pharmacological therapies.12,13 Finally, the present study did not include BS patients with complex clinical phenotypes involving multiple organs/organ systems. Future research employing next generation sequencing on larger treatment-naive BS patients, across more diverse geographic regions and ethnicities, and including also the more complex disease phenotypes, could provide further insights into BS’s immunopathogenesis and immunopathogenetic diversity. Also, in addition to using a clinical phenotype based clustering of BS patients, an approach implementing a disease severity score based grouping of these patients may further add valuable insights to BS molecular level heterogeneity present across ethnicities.
Conclusion
In conclusion, this study demonstrated significant gene expression and molecular disease mechanism differences between Portuguese and Turkish BS patients. This finding strongly supported the complex autoinflammatory disorder classification of BS and added that, in addition to its complex genetic nature, BS is also a heterogeneous syndrome. The observed interethnic differences of gene expression and molecular mechanisms may account for the inconsistent, sometimes contrasting immunological findings observed in BS research and the distinct clinical phenotypes of BS cumulating in certain geographic regions of the globe. Naturally, future research should take into account the molecular level heterogeneity of BS across ethnicities.
Decreased expression of anti-inflammatory genes was a common theme documented in both Portuguese and Turkish BS patients, albeit the identities of these genes differed across populations. This finding occurred to such an extent that, important anti-inflammatory genes with potential pathogenetic roles in one population were not detected in the other. Additionally, if present, pro-inflammatory gene expression differences seemed to enhance disease severity. Future research should explore the anti-inflammatory pathways, especially involving innate immunity proteins like CD69, CLEC12A, and CLC, as potential therapeutic targets to reduce inflammation in BS. These anti-inflammatory genes may also potentially help with the development of an animal model of BS which is currently lacking.
Taken together, these insights signal the forthcoming era of personalized therapeutics based on molecular profiles for BS patients. Finally, while this in-silico study was able to document important findings, undoubtedly, there is an absolute need for a precisely designed in vitro validation study which will try to replicate our findings. For this future work, the patient group should be carefully selected, taking into consideration of the specific limitations of the present study.
Supplemental Material
Supplemental Material - Comparative expression profiling of Portuguese and Turkish Behçet syndrome patients: Are we looking at the same Behçet?
Supplemental Material for Comparative expression profiling of Portuguese and Turkish Behçet syndrome patients: Are we looking at the same Behçet? by Tulin Isik, Seda Tasir, Abdulfettah Zolkhfan, Cagdas Sahap Oygur and Ali Kemal Oguz in European Journal of Inflammation
Supplemental Material
Supplemental Material - Comparative expression profiling of Portuguese and Turkish Behçet syndrome patients: Are we looking at the same Behçet?
Supplemental Material for Comparative expression profiling of Portuguese and Turkish Behçet syndrome patients: Are we looking at the same Behçet? by Tulin Isik, Seda Tasir, Abdulfettah Zolkhfan, Cagdas Sahap Oygur and Ali Kemal Oguz in European Journal of Inflammation
Supplemental Material
Supplemental Material - Comparative expression profiling of Portuguese and Turkish Behçet syndrome patients: Are we looking at the same Behçet?
Supplemental Material for Comparative expression profiling of Portuguese and Turkish Behçet syndrome patients: Are we looking at the same Behçet? by Tulin Isik, Seda Tasir, Abdulfettah Zolkhfan, Cagdas Sahap Oygur and Ali Kemal Oguz in European Journal of Inflammation
Supplemental Material
Supplemental Material - Comparative expression profiling of Portuguese and Turkish Behçet syndrome patients: Are we looking at the same Behçet?
Supplemental Material for Comparative expression profiling of Portuguese and Turkish Behçet syndrome patients: Are we looking at the same Behçet? by Tulin Isik, Seda Tasir, Abdulfettah Zolkhfan, Cagdas Sahap Oygur and Ali Kemal Oguz in European Journal of Inflammation
Supplemental Material
Supplemental Material - Comparative expression profiling of Portuguese and Turkish Behçet syndrome patients: Are we looking at the same Behçet?
Supplemental Material for Comparative expression profiling of Portuguese and Turkish Behçet syndrome patients: Are we looking at the same Behçet? by Tulin Isik, Seda Tasir, Abdulfettah Zolkhfan, Cagdas Sahap Oygur and Ali Kemal Oguz in European Journal of Inflammation
Supplemental Material
Supplemental Material - Comparative expression profiling of Portuguese and Turkish Behçet syndrome patients: Are we looking at the same Behçet?
Supplemental Material for Comparative expression profiling of Portuguese and Turkish Behçet syndrome patients: Are we looking at the same Behçet? by Tulin Isik, Seda Tasir, Abdulfettah Zolkhfan, Cagdas Sahap Oygur and Ali Kemal Oguz in European Journal of Inflammation
Supplemental Material
Supplemental Material - Comparative expression profiling of Portuguese and Turkish Behçet syndrome patients: Are we looking at the same Behçet?
Supplemental Material for Comparative expression profiling of Portuguese and Turkish Behçet syndrome patients: Are we looking at the same Behçet? by Tulin Isik, Seda Tasir, Abdulfettah Zolkhfan, Cagdas Sahap Oygur and Ali Kemal Oguz in European Journal of Inflammation
Footnotes
Acknowledgements
The authors express their sincere gratitude to all the scientists who share their valuable research data with their colleagues for the benefit of humanity.
Ethical statement
Author contributions
Seda Tasir and Ali Kemal Oguz conceptualized the study; Ali Kemal Oguz designed the study; Seda Tasir performed the formal analysis; Tulin Isik and Abdulfettah Zolkhfan performed the investigation; Tulin Isik and Abdulfettah Zolkhfan wrote the original draft; Cagdas Sahap Oygur and Ali Kemal Oguz reviewed and edited the manuscript. All authors have read and agreed to the final submitted version of the manuscript.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Data Availability Statement
All the bioinformatics data supporting the findings of this study are either presented within the manuscript or with the Supplemental Material files accompanying the manuscript. The microarray expression data of the studies by Xavier et al. and Oguz et al. can be obtained through Gene Expression Omnibus data repository with the accession numbers GSE17114 and GSE209567, respectively.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.

