Abstract
Myalgic encephalomyelitis/chronic fatigue syndrome (ME/CFS) is a poorly understood disease affecting 0.2%–2% of the global population. To gain insight into the pathophysiology of ME/CFS in New Zealand, we examined the transcriptomes of peripheral blood mononuclear cells by RNA-seq analysis in a small well-characterized patient group (10 patients), with age/gender-matched healthy controls (10 control subjects). Twenty-seven gene transcripts were increased 1.5- to sixfold and six decreased three- to sixfold in the patient group (
Keywords
Introduction
Myalgic encephalomyelitis/chronic fatigue syndrome (ME/CFS) affects approximately 0.2%–2% of the global population. 1 All age groups and socioeconomic strata are susceptible to the syndrome, but proportionally more women are affected.1,2 As few as 5% return to a state of well-being. 1 ME/CFS is a multifactorial and multisystem illness, impacting the neurological, immune, endocrine and musculoskeletal pathways, and often the gastrointestinal system. 1 Often ME/CFS onset is described as ‘sudden’, following an acute infectious period, but gradual onset is also reported. 2 Initiating factors such as chronic viral infection, immune activation, exposure to chemicals, stress, hypotension, lymphocyte abnormalities and neuroendocrine dysfunction have all been proposed, 2 but none validated as a single unifying cause. A formal medical diagnosis of ME/CFS is made after excluding all other recognized illnesses with a similar presentation, and by self-reported symptoms that fit within one of several broad sets of clinical criteria available. 1 They highlight as a key symptom disabling physical and mental fatigue, lasting over 6 months that is exacerbated by exercise, or mental or emotional exertion.2,3 While ME/CFS pathophysiology is still poorly understood, it is hypothesized that profound dysregulation of the central nervous system and immune system, dysfunction of cellular energy metabolism and cardiovascular abnormalities may play important roles in the pathophysiology of the illness. 2
A number of ME/CFS immune cell gene expression studies have been published. Collectively they have identified disturbances in pathways related to immune, neuronal, mitochondrial and metabolic function, but there is little consensus on the genes that are specifically differentially expressed. High-throughput RNA sequencing technology (RNA-seq) has only recently been utilized to investigate the ME/CFS transcriptome.
4
Our RNA-seq study described here focuses on peripheral blood mononuclear cells (PBMCs) in a well-characterized cohort of ME/CFS patients. PBMCs are reportedly stable, accurate reflectors of biological differences in healthy and unwell subjects, particularly for immunological or inflammatory pathways. We identified 27 gene transcripts significantly increased in the ME/CFS group when compared to healthy controls, and 6 gene transcripts significantly decreased (
Methods
PBMC collection and isolation
The study had approvals from Health and Disability Ethics Committees in New Zealand (NTY/12/06/055 and 15426). A total of 10 ME/CFS patients, diagnosed according to the canadian consensus criteria (CCC) guidelines
3
by an experienced ME/CFS clinician, were recruited along with matched controls. All participants gave their informed signed consent to participate. The control group had no history of significant illness, injury or fatigue-related disorders. ME/CFS patients completed a detailed questionnaire, developed in-house, providing their clinical history and current health status (Supplementary Table S1). Within 4 h of collection, whole blood samples were diluted 1:1 with filter-sterilized phosphate-buffered saline (PBS), layered onto Ficoll-Paque PLUS (Sigma-Aldrich) and centrifuged at 400
Transcriptome analysis
The transcriptome analysis was performed in collaboration with New Zealand Genomics Ltd. RNA was isolated from PBMC samples using a MirVana™ RNA Isolation Kit (Thermo Fisher Scientific) following the manufacturer’s instructions. All samples passed quality control RNA integrity analysis by Fragment Analyzer (RIN ⩾7). Library preparation used an Illumina TruSeq Stranded total RNA library kit (with RiboZero Human/Rat/Mouse). Total RNA libraries were pooled and sequenced by Illumina HiSeq 2x125 bp paired-end sequencing. Raw FASTQ sequences had an average per base mean quality score (Q score) of 35.84. A Q score of >30 indicates >99.9% base call accuracy and is the quality bench mark for next-generation sequencing. Raw FASTQ files were mapped and aligned to the human genome (hg38) with TopHat (tophat2) and Bowtie (bowtie2), resulting in 13,593 reads mapped to coding regions. The data were analysed in R using EdgeR (bioconductor release 3.4). A generalized linear model fit was used to normalize the data. A t-test identified significantly changed gene transcripts between the ME/CFS patients and controls, identified on the UCSC genome browser (https://genome.ucsc.edu/). The false discovery rate (FDR) reduces false positives, but at the inverse cost of increasing the number of false negatives so that the likelihood of the data to show real effects, particularly with small patient cohorts, can be easily reduced. In light of this, statistical analysis was achieved by setting the change in gene expression threshold to an un-adjusted
RT-qPCR validation
Taqman gene expression (RT-qPCR) assay validation was performed for the three most significantly changed gene transcripts
FNA
Differentially expressed gene transcripts (n = 33,
IPA
A pool of 125 upregulated and 40 downregulated gene transcripts (
Results and discussion
The concepts of Precision Medicine with appropriate statistical analysis have been applied to this small clinically well-characterized patient cohort in this RNA-seq transcriptome analysis. This approach has been successfully applied to cohorts of rare diseases, where available patients are small in number. In addition, the RNA-seq analysis itself has appropriate sequencing depth to compensate for the smaller sample size.
RNA-seq analysis revealed an altered PBMC transcriptome
A t-test of normalized RNA-seq data found 27 significantly increased gene transcripts and 6 significantly decreased gene transcripts in the ME/CFS cohort (
Transcripts with altered gene expression in the ME/CFS group compared to healthy controls (
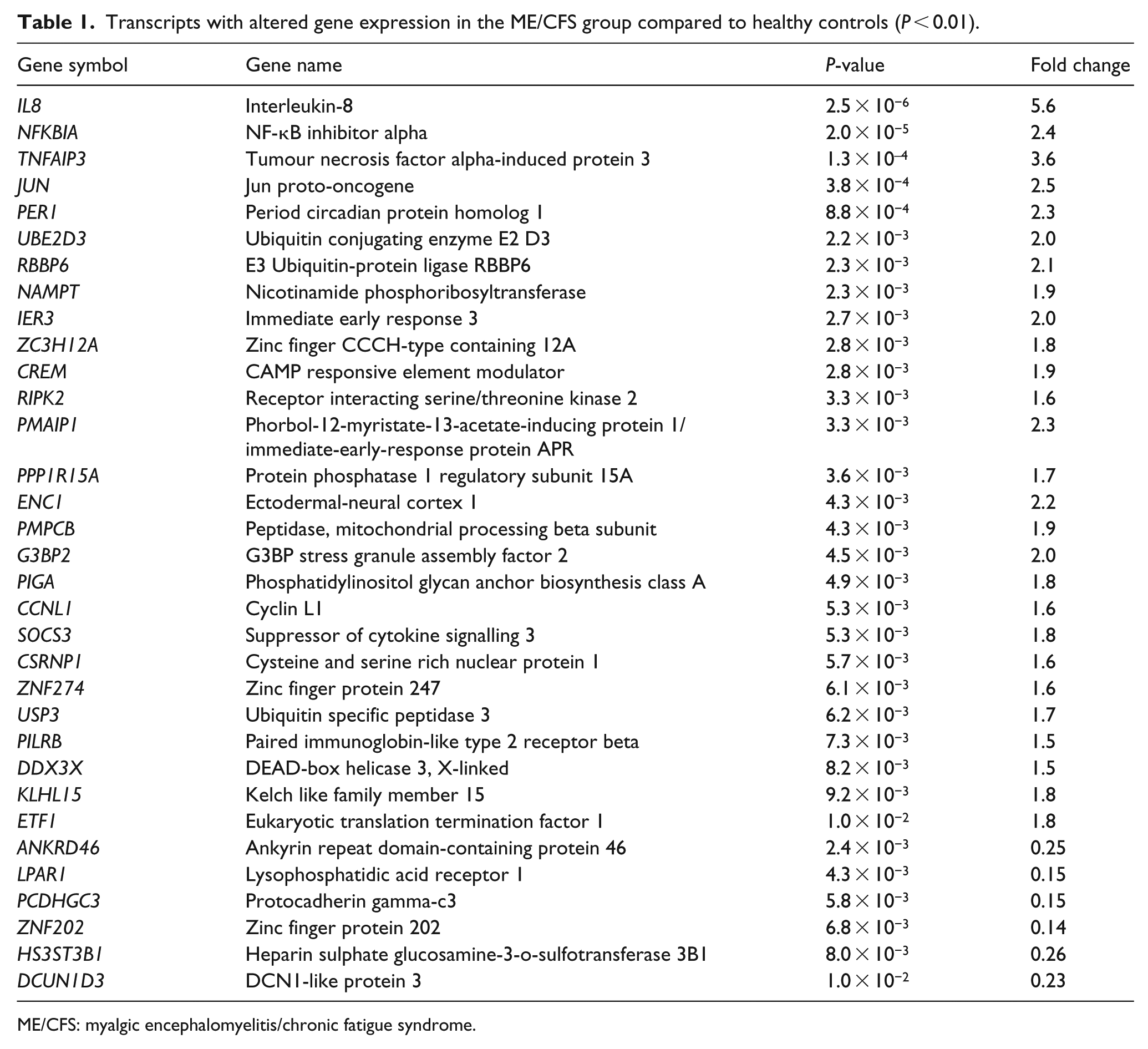
ME/CFS: myalgic encephalomyelitis/chronic fatigue syndrome.
qPCR validation of changes in gene expression
The top three significantly changed gene transcripts in the ME/CFS group were evaluated by RT-qPCR on the study group. The results were consistent with the RNA-seq data for
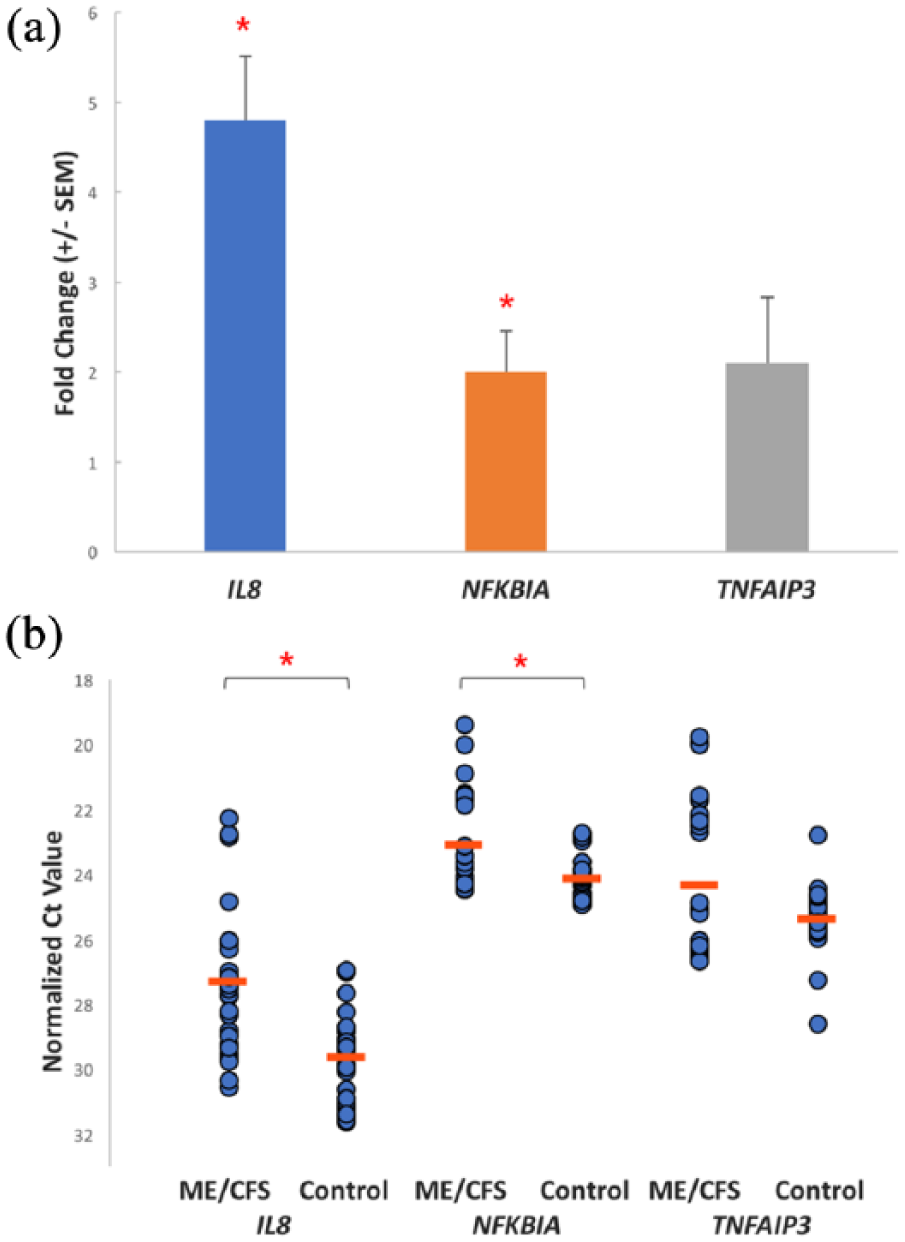
(a) Mean fold-change in expression between ME/CFS and controls of
Analysis of the transcriptome data
FNA and IPA examined the enriched functional interactions and the biochemical and physiological pathways derived from the analysis of the differentially expressed gene transcripts in ME/CFS.
FNA
The functional association networks of proteins encoded by differentially expressed gene transcripts (n = 33,

STRING analysis of significantly altered gene transcripts in the ME/CFS cohort (n = 33,
IPA shows inflammation and oxidative stress in ME/CFS
IPA was carried out on the transcriptome dataset with significance set at
Anti-inflammatory responders to TNFα and NF-κB
The three top upregulated genes in our study,
Significant biological pathways enriched in the ME/CFS group, identified by FNA and IPA
The circadian rhythm
Several elevated gene transcripts in the ME/CFS group encode core proteins that regulate the circadian clock (Supplementary Table S2), the central mechanism that drives our 24-h circadian rhythm. A disturbed circadian rhythm can be linked directly to many ME/CFS symptoms, such as pain, fatigue, sleep disturbance, flu-like symptoms, cognitive impairment, post-exertional malaise and dysfunction of metabolic and immune systems. 6 These results suggest that disruption of the circadian rhythm is likely in ME/CFS and could be a plausible candidate for sustaining both the symptoms and severity of ME/CFS.
Mitochondrial dysfunction
Mitochondria are essential for cellular energy production, metabolism and regulating stress responses. Results from FNA and IPA indicate mitochondrial dysfunction, characterized by signs of oxidative stress and downregulation of metabolic pathways (Supplementary Tables S3 and S4). Three significantly increased gene transcripts in the ME/CFS cohort (
Chronic inflammation
Sustained inflammation is emphasized in the ME/CFS transcriptome, with NF-κB and TNFα central to two primary canonical pathways predicted to be enriched by IPA (Supplementary Table S4). As mentioned previously,
These results reinforce the involvement of inflammation, disturbed circadian rhythm, metabolic dysregulation and oxidative stress in ME/CFS.
Previous ME/CFS transcriptome studies support inflammation and mitochondrial involvement
To the best of our knowledge, this is the first RNA-seq that has analysed PBMCs of ME/CFS patients. This technology was, however, used in a recent whole blood study analysing adolescent ME/CFS participants by gene set enrichment.
4
Our analysis was consistent with their findings, suggesting impairment of B cell differentiation and survival, enhanced innate antiviral responses and inflammation. Co-expression patterns and single gene transcripts were associated with neuroendocrine markers of altered HPA axis and autonomic nervous activity, plasma cortisol, blood monocyte and eosinophil counts. We have found significant molecular changes in our ME/CFS cohort that are consistent with those reported in larger complementary but not identical studies. Previous microarray and differential display studies of gene expression in ME/CFS have indicated disturbances in immune pathways, mitochondrial function, cell stress and apoptosis. A microarray investigation of PBMCs from a small group of post-infective ME/CFS males by Gow et al.
10
identified differentially expressed genes with roles in immune modulation, oxidative stress and apoptosis and, of particular interest, also found increased
Our exploratory approach has enabled us to obtain a rich differentially expressed gene dataset to identify changed biology in ME/CFS. We have identified the circadian rhythm dysregulation pathway as a new possible underlying cause of the unrefreshing sleep, fatigue and metabolic abnormalities seen in ME/CFS. Furthermore, impaired mitochondrial function and resulting oxidative stress, coupled with chronic immune-inflammatory signalling, provides a compelling explanation for the fatigue, cognitive dysfunction and post-exertion malaise experienced in ME/CFS.
Therefore, this study is a further step towards gaining an understanding of the disease process and identifying putative biomarkers to support clinical diagnosis. The biological pathways identified offer a rational explanation of the complex and often multi-systemic nature of ME/CFS.
Supplemental Material
Supplementary_Material – Supplemental material for Changes in the transcriptome of circulating immune cells of a New Zealand cohort with myalgic encephalomyelitis/chronic fatigue syndrome
Supplemental material, Supplementary_Material for Changes in the transcriptome of circulating immune cells of a New Zealand cohort with myalgic encephalomyelitis/chronic fatigue syndrome by Eiren Sweetman, Margaret Ryan, Christina Edgar, Angus MacKay, Rosamund Vallings and Warren Tate in International Journal of Immunopathology and Pharmacology
Supplemental Material
Supplementary_references_used_to_support_statements_in_the_text – Supplemental material for Changes in the transcriptome of circulating immune cells of a New Zealand cohort with myalgic encephalomyelitis/chronic fatigue syndrome
Supplemental material, Supplementary_references_used_to_support_statements_in_the_text for Changes in the transcriptome of circulating immune cells of a New Zealand cohort with myalgic encephalomyelitis/chronic fatigue syndrome by Eiren Sweetman, Margaret Ryan, Christina Edgar, Angus MacKay, Rosamund Vallings and Warren Tate in International Journal of Immunopathology and Pharmacology
Footnotes
Acknowledgements
We express our gratitude for the support of our ME/CFS patient and control cohorts. We thank the National ME/CFS Disease Association – Associated New Zealand ME Society (ANZMES) – for their ongoing support. We especially wish to thank the patients and healthy control subjects who willingly took part in our study.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship and/or publication of this article: We gratefully acknowledge the financial contributions and grants from the ME/CFS Association of New Zealand (ANZMES), the HS & JC Anderson Charitable Trust, NZ Lottery Health, Otago Medical Research Foundation and generous private bequests towards this study.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
