Abstract
Objective
MicroRNA-related single-nucleotide polymorphisms (miR-SNPs) can alter microRNA (miRNA) expression profiles, thereby influencing the risk of rheumatic diseases. Herrin a case control study, six miR-SNPs in miRNA processing machinery genes, namely RAN (rs14035), XPO5 (rs11077), Dicer (rs3742330), GEMIN3 (rs197412), GEMIN4 (rs2740348), and TNRC6B (rs9623117), were genotyped to assess their correlation with the risk of systemic lupus erythematosus (SLE).
Methods
We included 119 patients with SLE and 130 healthy controls. The genotypes of the six miR-SNPs were determined using polymerase chain reaction (PCR). Serum cytokine levels were assessed using a cytometric bead array, and fluorescent probe technology was used to determine plasma reactive oxygen species (ROS) levels.
Results
The AA genotype of Dicer was correlated with a 0.566-fold decreased risk of SLE compared with that of the AG + GG genotype (odds ratio, 0.566; 95% CI, 0.342–0.935; p = .026), and the rs3742330 A allele was associated with a significantly decreased risk of SLE (p = .035) compared with that of the rs3742330G allele. Additionally, AA genotype carriers exhibited lower levels of interleukin-6 (IL-6) in the blood (p = .013). Subsequent analysis revealed increased ROS production in patients with SLE than that in the controls (621.042 ± 425.285 vs 499.966 ± 302.273, p = .011).
Conclusion
Our findings suggest that ROS generation participates in SLE pathogenesis. The identification of Dicer gene SNP rs3742330 as a potential modifier of SLE risk via mediating IL-6 overproduction suggests a potential avenue for targeted interventions to manage SLE and its associated immune dysregulation.
Keywords
Introduction
Systemic lupus erythematosus (SLE) is a complex and chronic autoimmune disorder characterized by the production of autoantibodies targeting cytoplasmic and nuclear antigens, leading to diverse immunological and clinical manifestations.1,2 In China, the prevalence of SLE ranges from 40 to 70 per 1000 people, with a ratio of 1:9 for males and females respectively. 3 Several factors, including environmental influences, cytokines, genetic variations, and oxidative stress, contribute to the onset and progression of SLE; however, the precise etiology remains unclear.4–6
MicroRNAs (miRNAs), non-coding ssRNA molecules of 19–22 nucleotides, participate in crucial biological processes, including metabolism, proliferation, apoptosis, differentiation, stress responses, and hormonal secretion.7–9 Growing evidence suggests the involvement of miRNAs in rheumatic disorders, including rheumatoid arthritis (RA), SLE, and Sjögren’s syndrome.10–12 During the processing of miRNA to produce a 21-base-pair dsRNA, processing of the long primary miRNA transcripts takes place in the nucleus via RNase III Drosha generating pre-miRNA (precursor miRNA), and then with both RAN-GTPase (RAN) and exportin-5 (XPO5), the pre-miRNA becomes located in the cytoplasm, where the processing of the pre-miRNA takes place by RNase III Dicer and the transactivation-responsive RNA-binding protein. This results in the formation of miRNA duplex that decompresses releasing mature mi-RNA. The RNA-induced silencing complex, containing GEMIN3/4, directs mature miRNA to its target messenger RNA sites.13–18 miRNA can inhibit the expression of proteins by either degradation of mRNA or inhibition of translation. Single-nucleotide polymorphisms in miRNA-related genes (miR-SNPs), including miRNA processing machinery and binding sites, can modify miRNA expression and influence the risk of rheumatic diseases, such as rheumatoid arthritis, osteoarthritic chondrocytes, and polymyositis.18–20
Cytokines and oxidative stress play significant roles in SLE pathogenesis. An imbalance between anti- and pro-inflammatory cytokines contributes to SLE inflammation and tissue damage. 21 Additionally, reactive oxygen species (ROS)-modified DNA generates new antigen epitopes, forming immunological complexes with circulating anti-DNA antibodies and initiating SLE pathogenesis. 22
In this case-control study, genotyping of six miR-SNPs in miRNA processing machinery genes—RAN (rs14035), XPO5 (rs11077), Dicer (rs3742330), GEMIN3 (rs197412), GEMIN4 (rs2740348), and TNRC6B (rs9623117)—was performed to assess their correlation with SLE risk. Cytokine and ROS roles in SLE were also investigated.
Methods
Study population
In accordance with the 1997 classification criteria of the American College of Rheumatology, 119 women with SLE treated between May 2017 and October 2023 at the Second Hospital of Hebei Medical University were recruited as the cases group. Additionally, 130 age-matched healthy women, devoid of any autoimmune disease history, were included in this study. The clinical characteristics of the 119 patients, including 67 cases with rashes, 9 with ulcers, 40 with arthritis, 12 with serositis, 57 with anti-dsDNA positivity, 39 with lupus nephritis (LN), 22 with neuropsychiatric SLE (NPSLE), and 65 with hematological abnormalities, were retrospectively collected (Table 2). The clinical disease activity of SLE was assessed using the Systemic Lupus Erythematosus Disease Activity Index 2000 (SLEDAI-2K). All individuals provided written informed consent, and the study was conducted in adherence to the Declaration of Helsinki under the supervision and approval of the Human Tissue Research Committee of the Second Hospital of Hebei Medical University (2022-R730).
Genotyping of miR-SNPs
Primers and probes.

Measurement of cytokine levels
A filter plate V02 (BioLegend, San Diego, CA) equipped with a human TH1/2 panel (8-Plex) was used for measuring the levels of cytokines, such as interleukin (IL)-2/4/5/6/10/13, interferon (IFN)-γ, and tumor necrosis factor (TNF)-α. The procedure was conducted in darkness at room temperature, involving the addition of 25 μL of the serum sample (two-fold dilution using an assay buffer) to an equivalent volume of antibodies for each cytokine. Subsequently, the mixture was incubated, combined with beads, and shaken for 1 h at approximately 800 r/min, followed by the addition of streptavidin–phycoerythrin (SA-PE) into the tubes, with a subsequent 30-min shaking period at approximately 800 r/min. The quantification of the phycoerythrin fluorescence signal in the analyte-specific bead region and the determination of specific analyte concentrations were performed using the MACSQuant Analyzer 10 flow cytometer (Miltenyi Biotec, Bergisch Gladbach, Germany) and data analysis software from BioLegend (San Diego, CA), respectively.
ROS measurement
ROS levels were monitored using the BBOXiProbe® Plasma Active Oxygen Detection Kit (BestBio Technology, Shanghai, China). Plasma (100 μL) was incubated with 10 μL of O12 probes at 37°C in darkness for 30 min, and fluorescence absorbance was measured at 488 nm (excitation) and 520 nm (emission) using a fluorescence plate reader (BIOTEK, Winooski, VT, USA).
Statistical analysis
Statistical analyses were conducted using SPSS software (version 21.0; SPSS Inc., Chicago, IL, USA). For continuous variables, we employed either the Student’s t-test or rank sum test, whereas the chi-square test or Fisher’s exact test was used to evaluate qualitative variables. We used either Spearman’s or Spearman’s rank correlation analysis to assess the correlation between variables. A sample size of 117 yielded 90% power to detect a medium effect size (W) of 0.30, using a one-degree-of-freedom chi-square test with a significance level (alpha) of 0.05 for PASS 15. A p-value (two-sided) < .05 indicated a significant difference.
Results
Genotyping and patient characteristics
Clinical data of SLE patients and controls.

SLE: systemic lupus erythematosus; Anti-dsDNA: anti-double stranded deoxyribonucleic acid; SLEDAI: systemic lupus erythematosus disease activity index; χ2: chi-square.
Genetic models and Dicer gene-located SNP analysis
Associations of the six SNPs with systemic lupus erythematosus risk.

SLE: systemic lupus erythematosus; χ2: chi-square; OR: odds ratio; CI: confidence interval.
In the dominant model, the Dicer gene-located SNP rs3742330 revealed that the genotype frequencies for AA and AG + GG were 42.0% and 58.0% in patients with SLE, whereas in controls, they were 56.2% and 43.8%, respectively. The AA genotype was correlated with a 0.566-fold decreased risk of SLE compared with that of the AG + GG genotype (odds ratio, 0.566; 95% CI, 0.342-0.935; p = .026, Table 3). Additionally, the analysis revealed that the rs3742330 A allele was associated with a significantly decreased risk of SLE (p = .035) compared with that of the rs3742330 G allele.
Cytokine levels, clinical parameters, and their association with ROS levels
Assessment of the IL-2/4/5/6/10/13, IFN-γ, and TNF-α cytokine levels served for the purpose of evaluating their association with this SLE risk-correlated miR-SNPs by Wilcoxon rank sum test (Figure 1). The AA rs3742330 genotype displayed lower IL-6 expression than that of the AG + GG genotype (p = .013), suggesting that the Dicer SNP may participate in SLE pathogenesis by alteration of IL6 expression. The association of rs3742330 genotype with cytokines.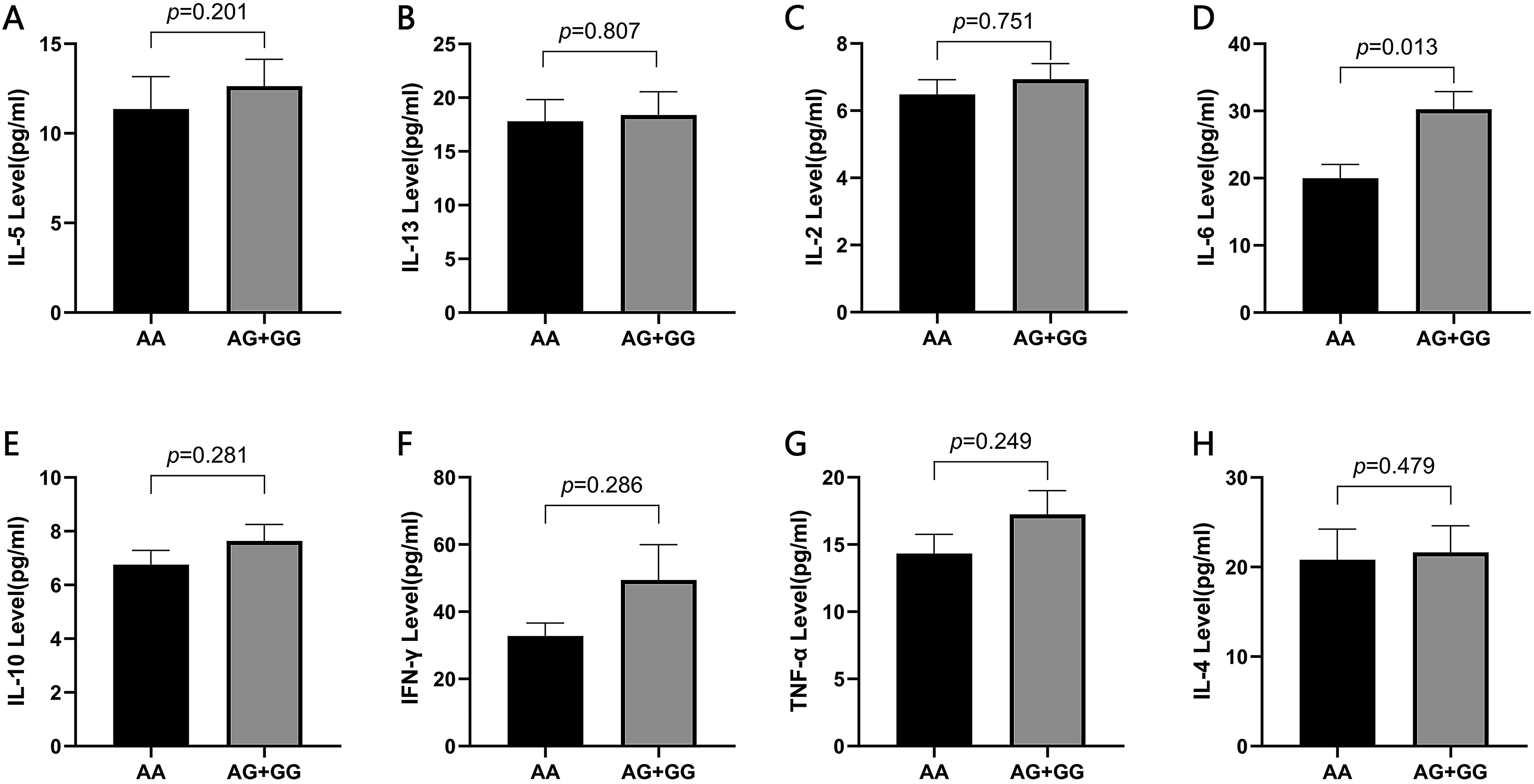
Comparison of different clinical parameters, including SLEDAI-2K, lupus nephritis, and neuropsychiatric SLE, with cytokine levels is depicted in Figures 2–4. However, no significant differences were observed between groups. An assessment of different clinical classifications, including lupus nephritis, neuropsychiatric SLE, and hematological abnormalities, in relation to SLE risk-associated miR-SNPs is presented in Table S1. However, no significant differences were observed between groups. The association of SLEDAI-2K with cytokines. The association of lupus nephritis with cytokines. The association of neuropsychiatric SLE with cytokines.


A subsequent investigation into SLE risk-associated miR-SNPs and ROS levels revealed no significant difference (AA: 615.080 ± 408.763 vs AG + GG: 625.362 ± 439.787, p = .896). Additional analysis revealed that in comparison to the control, ROS levels in the SLE group exhibited higher expression than control group and this was statistically significant (621.042 ± 425.285 versus 499.966 ± 302.273, p = .011, Figure 5). Spearman’s correlation analysis indicated no association between IL-6 and ROS levels. ROS level in SLE patients. (A) The ROS levels in AA and AG + GG genotypes of rs3742330. (B) The ROS levels in SLE patients and controls.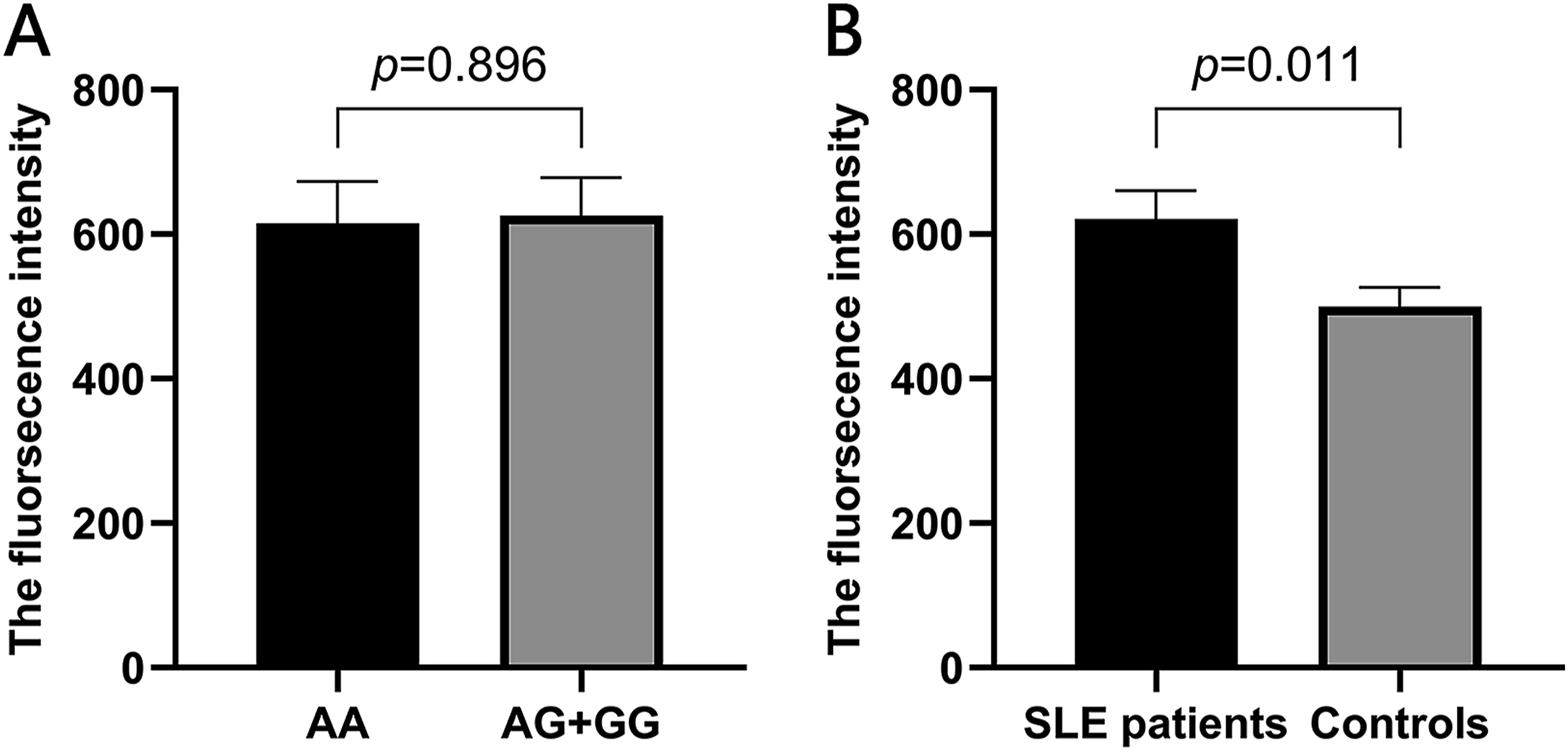
Discussion
In this study, we genotyped six miR-SNPs in miRNA processing machinery genes—RAN (rs14035), XPO5 (rs11077), Dicer (rs3742330), GEMIN3 (rs197412), GEMIN4 (rs2740348), and TNRC6B (rs9623117)—to explore their correlation with SLE risk. Our findings revealed an association between the Dicer gene-located SNP rs3742330 and SLE risk, with the rs3742330 A allele significantly linked to a decreased risk of SLE compared with that of the rs3742330 G allele.
Dicer, a member of the RNase III family, plays a crucial role in cleaving dsRNAs to generate small interfering RNAs and miRNAs, thus suppressing the target gene expression.23,24 Dicer-controlled miRNA pathways are involved in the function and immune tolerance of Treg cells.25–27 Altered Dicer expression in Tregs from lupus-prone MRL-Faslpr/lpr mice exhibited a diminished suppressive capacity and an altered phenotype characterized by expressional changes of CD62L and CD69. The altered miRNA profiles mediated by reduced Dicer expression may be implicated in the functional alterations in the Tregs, potentially initiating the SLE development. 28 The miRNA profile changes influenced by the Dicer gene 3′-untranslated region (UTR)-located miR-SNP rs3742330 could modulate the SLE process by influencing miRNA expression profiles; however, the precise target miRNAs involved in this process remain to be identified.
Our observation of an association between the Dicer AA genotype and decreased IL-6 levels aligns with previous research suggesting that reduced Dicer expression promotes IL-6 expression in inflamed colon tissues. 29 IL-6, a multifunctional pro-inflammatory cytokine, modulates the inflammatory response, influences B cell differentiation into plasma cells that secret immunoglobulin, 30 and contributes to autoantibody production. Moreover, IL-6-knockout mice exhibit delayed lupus nephritis onset and prolong survival.31,32 Concordantly, IL-6 expression is also significantly increased in patients with SLE in addition to correlating with disease activity. 33 This implies that Dicer may regulate IL-6 expression by modulating miRNA expression, thereby influencing the progression of SLE.
Consistent with prior studies, our data indicated higher ROS levels in patients with SLE compared with those in healthy controls. Aberrant ROS levels can induce oxidative damage to intracellular components, including lipids, DNA, proteins, and phospholipids, potentially triggering cellular dysfunction or autoimmune responses. 34 Additionally, ROS can initiate the formation of pro-inflammatory cytokines through the activation of the nuclear factor-κB axis and contribute to the generation of SLE autoantigens, including the apoptotic bleb and neutrophil extracellular trap. 35
However, our study has certain limitations, including its single-center nature and a relatively small sample size. Further investigations involving data from multiple centers and a larger participant pool are warranted. Additionally, the exact mechanism by which the identified SNP contributes to the risk of developing SLE remains unknown and requires further investigation.
Conclusion
Our findings suggest that the SNP rs3742330 located in Dicer may modify SLE risk by mediating IL-6 overproduction. ROS generation also participates in SLE pathogenesis.
Supplemental Material
Supplemental Material - Single nucleotide polymorphisms in the Dicer gene modify the risk of systemic lupus erythematosus
Supplemental Material for Single nucleotide polymorphisms in the Dicer gene modify the risk of systemic lupus erythematosus by Jingjing Zhang, Yufei Zhao, Song Wang, Shasha Zhang, Xiaoyun Zhang, Chenxing Peng and Qingyi Liu in European Journal of Inflammation.
Footnotes
Acknowledgements
We would like to thank all patients for their participation.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Key Science and Technology Research Program from Health Commission of Hebei Province (grant number 20230133).
Ethical statement
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
