Abstract
Background
In light of the growing problem of antibiotic resistance, it is imperative to investigate new sources, and plants offer a promising supply of bioactive chemicals. Because of its numerous uses in industry, health, and nutrition as well as its antibacterial qualities, Cannabis sativa (C.sativa) has garnered a lot of study interest. This study sought to determine whether ethanolic extracts from C.sativa leaves have antibacterial properties against six human pathogenic microorganisms.
Methodology
The antibacterial activity of C.sativa ethanolic extract was tested against six bacteria according to design of experiments made by Agar diffusion method accompanied by response surface method (RSM) of Minitab 17 software. The different combinations set were, concentration: 5.0, 7.5, and 10.0, pH: 5.0, 6.5, 8.0 and temperature: 35°C, 37.5°C, 40°C. By using RSM, maximum antibacterial activity has been checked for ethanolic extract of C.sativa against six bacteria by choosing three independent variables, temperature, pH, and concentration. In in-Silico studies, homology, threading approach, structure prediction, ligands designing and docking studies was performed against the antimicrobial target sequences for Beta-Lactamase, GABA Receptor, Lipoteichoic Acid, N-Acetylglucosamine (NAG), Peptidoglycan and Topoisomerase-IV through FASTA format from UniProt for structure prediction.
Results
The results indicated that the three concentrations were effective against tested bacteria. Moreover, effect of pH caused a significant variation in zone of inhibition. The graphs presented in this study indicate the highest zone of inhibition for plant extract; have been achieved at concentration of 10.0, pH 5.1 and temperature 37.5°C. It shows that by keeping the pH low, antibacterial activity will increase. Through the multiple regression analysis on the experimental data, the fitted regression model for the response variable and the test variable x1, x2, x3 are correlated by the second order polymeric equation.
Conclusion
It has been concluded that C.sativa can be considered as an effective drug in curing diseases caused by bacteria. Using the optimized values of temperature and pH analyzed in this experiment.

Keywords
Introduction
The herbaceous plant Cannabis sativa L. (C.sativa) is classified as part of the Cannabaceae family. Many people allude to this plant species by its many colloquial names, such as hemp and marijuana. Despite being indigenous to Central Asia, this plant has spread throughout the world due to its capability to adapt to many environmental variables. 1 Humans have been employing C.sativa since ancient times, and numerous historians have recorded multiple uses of this plant abroad. This plant has been cultivated for religious and recreational purposes, as well as for food, fiber, and oil, according to recorded history. 2 C.sativa is also used therapeutically to treat depression, inflammation, and chronic pain, according to numerous ethnobotanical surveys. 3 An estimated 550 or more molecules of bioactive chemicals are thought to be present in cannabis. These substances are classified as alkaloids, carotenoid, flavonoid, lignanamide, terpenoid, stilbenoid, and cannabinoid. 4
The plant is also an important source of various types of compounds with diverse chemical structures. It have been reported that the leaves of this plant contain C-glycosyl flavones like orientin in L form, vitexin and iso-vitexin and also iso-orientin, lucenin-II, vicenin-II, diosmetin 6-C-beta-glucoside, apigenin, luteolin, kaempferol and chrysoeriol. Thus, multiple cannabinoids make up the complex chemical composition of the psychoactive substance of marijuana or cannabis. Psychoactive effects are exhibited by a class of secondary metabolites called cannabinoids. The two types of cannabinoids are exogenous (made by the C. sativa plant or synthetically) and endogenous (generated by the human body). Cannabinoids are composed of two G-protein coupled receptors, CB1 and CB2. These receptors are an integral part of the human body’s endocannabinoid system. Both endogenous and exogenous cannabinoids adhere to receptors; the most efficacious exogenous ligand is tetrahydrocannabinol (THC). Yet another exogenous cannabinoid that has been shown to mitigate the negative side effects of THC is cannabidiol (CBD). It has anxiolytic, anti-sedative, and anti-psychotic properties while acting as an antagonist for CB1 and CB2 receptors. 5
Since it was reported that the plant can grow in presence of heavy metals, it can be cultivated in polluted areas where it can resist heavy metal toxicities. The scientific name C. sativa has common names as Palo Verde, Horse bean and Jerusalem thorn. However, in Pakistani language it is known as Kabuli kikar and Vilayati kikar. It pertains to Phylum Spermatophyta, family Fabaceae (leguminosae) and genus Paekinsonia. It is inherent to Southern USA, Northern Mexico, Southern Africa, Pakistan and Central America. 6
Given that, it is emerging that infectious diseases have threatened the lives of many people across the world. 7 Besides, microorganisms causing these diseases are now resistant to wide range of antibiotics. For instance, Methicillin-resistance Staphylococcus aureus (S. aureus) (MRSA) is the dangerous, genetically different strain of S. aureus that have become resistant to several antibiotics. Infection caused by it is grueling to treat. Therefore to overcome such issues, medicinal plants are being explored as a source of drugs that is hitherto unprecedented. Although new antimicrobial agents are being synthesized but due to the increasing widespread resistance indicate that they will have relatively short therapeutic life span. 8
It has been a historic norm to combat infectious diseases using medicinal plants. The medicinal value of such plants is due to the fact that they contain secondary metabolites, which affect the physiology of an organism. As stated by World Health Organization (WHO), plants are meant to be the right source of antibacterial drugs. Therapy with plant extracts having antimicrobial properties is worthy of attention. WHO has also indicated that about 20 000 plant species have been known to possess therapeutic potential that could be attributed to the bioactive compound synthesized for the duration of the secondary metabolism of plants. 9 Medical microbiology concerns in preventing and treating disease caused by these microorganisms. Inhibitory chemicals used to kill microorganisms or retard their growth intend to cure diseases. These chemicals are called as antimicrobial agents and are classified with respect to their utilization and spectrum of activity. 10 Antibacterial agents are classified into antibiotics, non-antibiotics chemotherapeutic agents such as disinfectants, antiseptics, preservatives and immunological products. Salicylic is a strong antiseptic and Benzoic acid is used in lotions and ointment as antiseptic. Other than this mandolic acid, ethanol, propanol, chlorine, iodine containing compounds, heavy metals, dyes such as crystal violet, soaps and detergents also to some extent possess antimicrobial activities and are used as disinfectants and antiseptics. Immunological products include vaccines and monoclonal antibodies. 11
Multidrug resistance (MDR) pathogens have been recognized by the WHO as a serious hazard to world health. An estimated 1.27 million fatalities in 2019 were attributed to antibiotic-resistant bacteria (ABR). This highlights the significance of innovative therapy approaches. Therefore, finding and creating novel chemicals is essential to preserving the efficacy of present and upcoming pharmacological interventions for infectious diseases. 12 Hereby, numerous cannabinoids have demonstrated strong antibacterial effects against MRSA and other Gram-positive bacteria. Cannabinoids also may help eliminate dangerous microorganisms from the environment, according to in vitro researches. Additionally, there is proof that some of the chemicals in C. sativa may have antibacterial qualities. 5 This implies that more research is required to fully grasp their potential. With the ongoing emergence of antibiotic resistance, cannabis may offer a novel approach to treating infectious illnesses. This study emphasizes how urgently more investigation and standardized clinical trials are needed to confirm these results and create cannabinoid-based therapies.
Materials and Methods
C. sativa leaves were purchased from the market in Lahore and identified by Prof. Dr Ijaz Rasool, Department of Botany, Agriculture University, Faisalabad, Pakistan. Escherichia coli, Pseudomonas aeruginosa (ATCC-27853), Stephylococcus aureus (ATCC-25922), Bacillus subtilis, Klebsiella pneumonia and Salmonella typhi were collected from Microbiology lab of Institute of Molecular Biology and Biotechnology (IMBB), The University of Lahore, Lahore, Pakistan.
Collection and Preparation of C. sativa Extracts
2.0 g of 80 mesh plant sample was dissolved in 4.0 mL of 80% ethanol, placed overnight at room temperature for 24-h. After filtration, extract was subjected to filtration through Whatmann filter paper 1.0 and evaporated the solvent for overnight to get concentrated filtrate, which was dissolved in DMSO, followed by centrifugation for 10 minutes at 14 000 r/min. This supernatant preparation took 4 days. After that supernatant was saved and stored in refrigerator at 4°C until the preparation of extracts at different concentrations (5.0 µg/mL, 7.5 µg/mL, 10.0 µg/mL) and at different pH (5.0, 6.5, 8.0) by using 1.175% BaCl2 and 1% H2SO4, to use them at different temperatures (35°C, 37.5°C, 40°C) for antibacterial activity by using agar well diffusion method and response surface methodology (RSM) by Minitab 17 software against six bacteria.
Human Pathogenic Bacterial Preparation
Three Gram-positive bacteria were used in the study: Staphylococcus aureus, Bacillus subtilis, and Staphylococcus typhi. Three Gram-negative bacteria were also used, including Pseudomonas aeruginosa, Klebsiella pneumoniae, and Escherichia coli. Their significance as human pathogenic microorganisms led to their selection. A single colony of bacteria was used to cultivate each strain using nutrition broth (NB), which was stirred for 18 hours at 37°C. The bacterial concentration was calibrated to an optical density (OD) of 0.1 at 600 nm before to use.
Agar Disc Diffusion Assay
The Agar disc diffusion assay was used as the first screening technique to determine whether the extracts have antibacterial properties. Six strains of bacteria that are harmful to humans were inoculated. After being cultivated from a single colony in 10 mL of nutrition broth (NB), each bacterial strain was incubated for the entire night at 37°C. After the bacterial cells were incubated, they were centrifuged, and the concentration of cells was adjusted until the optical density (OD) was 0.1 at 600 nm. Each bacterial suspension of 100 μL was then aseptically placed onto nutrient agar (NA) plates. The NA plates were covered with 0.6 mm-diameter sterile paper discs. Ethanolic extracts of different concentrations (5,7.5 & 10 μL) made from the stems and leaves of C.sativa were used to saturate each of these separate discs, along with negative (DMSO) and positive control (ciprofloxacin 5 µg/disc) at different temperatures (35, 37.5 & 40) and pH (5, 6.5 & 8). This process was repeated thrice. After 15 minutes of diffusion, the NA plates were incubated for 24 h at 37°C in a bacterial incubator (Memmert, Willi-Memmert-Straße, Büchenbach). The development of inhibition zones surrounding the paper discs were noted and documented after incubation. Bacterial susceptibility to the extracts was shown by the sizes of these zones, which were expressed in millimetres (mm), where larger zones indicated greater susceptibility. This whole study took one and half months to get the results in vitro.
Response Surface Methodology (RSM) Model
Counter plot is used to study the relationships among the test variable and to check the maximum level of each variable that can give higher zone of inhibition due to antibacterial activity of C. sativa extract.
Data Analysis
The SAS software version 5.0 was used to analyse the inhibitory zone diameter. Each treatment had three replicates, each with five plates, in an experiment that used a completely randomised design (CRD). Analysis of Variance (ANOVA) and mean comparisons between approaches were included in the statistical study. P-values below .05 were regarded as statistically significant.
In silico Analysis
Selection of Ligands and Proteins
Cannabinol, Delta-8-Tetrahydrocannabinol, Tetrahydrocannabinol, Cannaabicyclol, Beta-Eudemsol, Tetrahydrocannabivarin, Nabilone, Cannabitriol and Cannabigerolate of C. sativa leaves were downloaded from the PubChem database in sdf format while Beta-Lactamse, DNA Gyrase inhibitory protein, Lipoteichoic Acid, Pinicillin-Binding Protein and Topoisomerase-IV were downloaded from Protein Data bank in pdb format (https://www.pdb.org/pdb).
Protein Preparation
Receptor proteins were opened in discovery studio software, 2021 (https://discover.3ds.com/discovery-studio-visualizer-download) to remove undesired substances. All the Heta atom, water molecules and bounded ligands were removed while hydrogen atoms were added with proteins and were saved in pdb format.
SDF formats of ligands were uploaded in Open Babel window of PyRx by minimizing energy to convert them into pdb format. 13
Molecular Docking
Molecular docking was run after creating grad box using vina wizard. The grad box dimension for Beta-Lactamse (X: 55.99, Y: 47.56, Z: 43.31), DNA Gyrase inhibitory protein (X: 41.70, Y: 41.67, Z: 54.17), Lipoteichoic acid (X: 57.83, Y: 63.72, Z: 62.80), Pinicillin-Binding Protein (X: 59.05, Y: 80.01, Z: 70.24) and Topoisomerase-IV (X: 52.70, Y: 73.39, Z: 55.43) were followed and Discovery studio 2021 was used for results visualization including 2D structure, hydrogen bonding and other type of bonds. Using the standard precision algorithm (SP) with flexible ligand sampling, the generated library of ligands (phytochemicals) was docked into the target protein’s active site. This was followed by extra precision (XP) with limited refine ligand sampling. The ligand-residue interaction diagram for the target protein’s active site was viewed using the ligand interaction tool.
Drug-Likeness and ADME (Absorption, Distribution, Metabolism and Excretion) Analysis
SwissADME online software (https://www.swissadme.ch/) was used for checking drug-likeness and ADME analysis. 14 Canonical smiles of ligand compounds were retrieved from PubChem and were pasted in swissADME to analyze drug-likeness and ADME parameters.
Toxicity
Canonical smiles of all the ligands were retrieved from PubChem database and were pasted one-by-one in Protox-II online web server (https://tox.charite.de/protox_II) 15 and admetSAR (https://lmmd.ecust.edu.cn/admetsar2) for toxicity determination. Protox-II online and admetSAR was run to check hepatotoxicity, mutagenicity, cytotoxicity, AMES toxicity, carcinogenicity and acute oral toxicity of the compounds.
Bioactivities
Orally administrated drugs need to be compatible with the drug-likeness properties to be finalized as pharmaceutically consistent with their bioactivity score and that’s why Molinspiration Toolkit was used for prediction of selected compounds (https://www.molinspiration.com/cgi-bin/properties).
Experimental Design and Statistical Analysis
The significant terms were seen in the P-values (less than .05). A second order regression equation proposed for the analysis of data was used in experiment.
Here X1 is temperature (35 to 40°C), X2 is pH (5.0 to 8.0), X3 is Concentration (5.0 to 10.0 µg/mL).
Three dimensional response surface and counter presentations were plotted to study the interaction among various factors used and to find out the optimal level of each factor of maximum zone of inhibition. A set of 60 experiments (all factorials and none of the experiment was zero point test) was performed to approximate their error. Counter plot was also used to study the relationships among the test variable and to check the maximum level of each variable that can give higher zone of inhibition due to antibacterial activity of C. sativa extract. While, out of three independent variables, one variable was taken constant and the other two were kept in the x axis and y axis in the counter plot graph. Then the relationship between two variables was examined using the graph for S.aureus.
Results
Molecular Docking
Molecular Docking of Ligands With Different Receptors

Where: Ala, Alanine; Arg, Arginine; Asn, Asparagine; Asp, Aspartic acid; Cys, Cysteine; Glu, Glutamic acid; Gln, Glutamine; Gly, Glycine; His, Histidine; Ile, Isoleucine, Leu, Leucine; Lys, Lysine; Met, Methionine; Phe, Phenylalanine; Pro, Proline; Ser, Serine; Thr, Threonine; Trp, Tryptophan; Tyr, Tyrosine; Val, Valine.
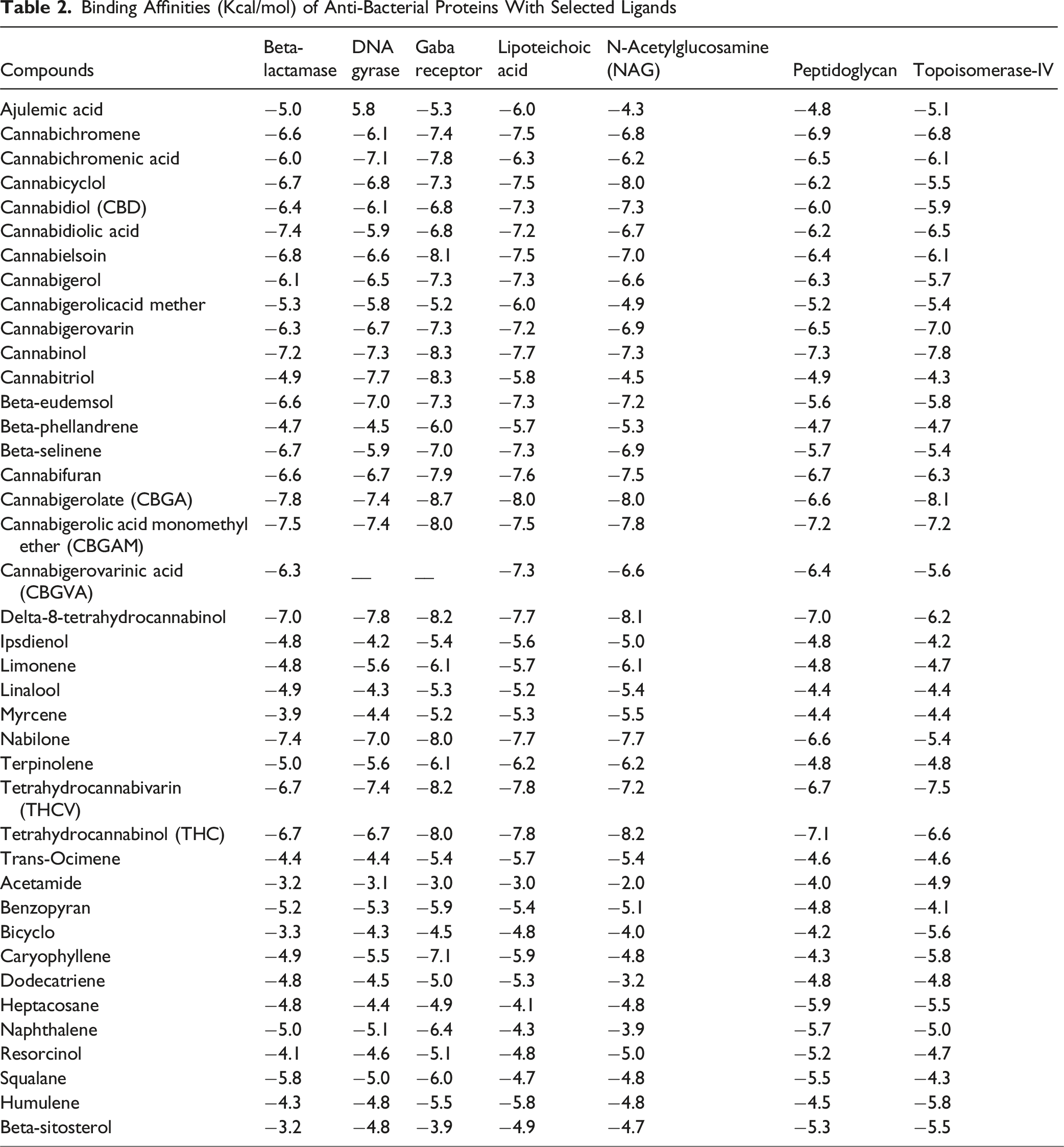
Binding Affinities (Kcal/mol) of Anti-Bacterial Proteins With Selected Ligands
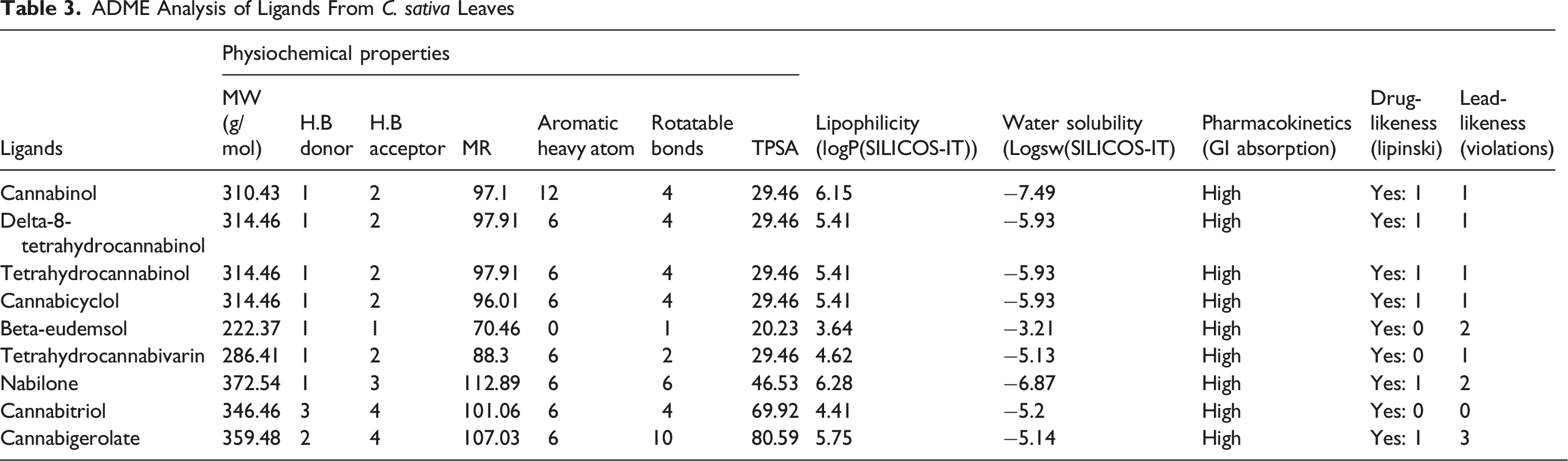
ADME Analysis
ADME Analysis of Ligands From C. sativa Leaves
Protox-II Toxicity Test
Toxicity Analysis for all Selected Compounds Using ADMET SAR

Bioactivities Molinspiration Test
Analysis of Bioactivities for All Selected Compounds
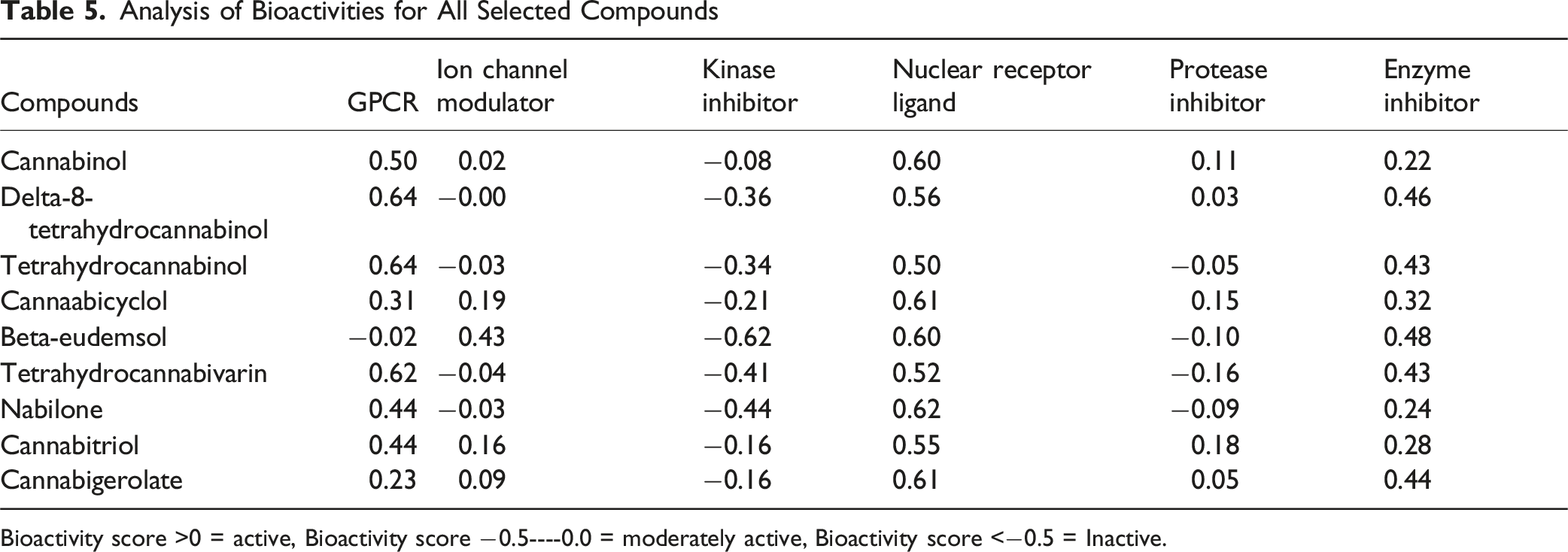
Bioactivity score >0 = active, Bioactivity score −0.5----0.0 = moderately active, Bioactivity score <−0.5 = Inactive.
Antibacterial Activity of Ethanolic Extract of C sativa Leaves at Different Concentrations, pH and Temperatures
Zones of Inhibition at Different Concentrations of C. sativa Leaves, Temperatures and pH Against Different Bacteria


Zone of Inhibition by Ethanolic Extract of C sativa Leaves at Different Concentrations, pH and Temperatures
Response Surface Methodology (RSM) Model
Results of RSM model indicated that the three concentrations were effective against tested bacteria. More over effect of pH caused a significant variation in zone of inhibition. Ethanolic extract of C. sativa has shown an important role in affecting the growth of all bacteria. The model equation, represented on the graphs, shows Response Surface Plots that represent the individual and interactive effects of test variables on the response. P-value and the Response Surface Graphs shows that temperature and pH are the effective parameters for giving highest antibacterial activity against all bacteria as clarified in surface plot with significant interactions between the concentration, pH and temperature. In the current work, the ethanolic extracts are optimized for maximal antibacterial potency against selected bacterial strains that interact with the three variables. The least square estimate of parameters (β) of the final model was calculated following the analysis of response data and treatment combination. It was determined that the pH and temperature of Staphylococcus aureus, Bacillus subtilis, and Staphylococcus typhi, Pseudomonas aeruginosa, Klebsiella pneumoniae, and Escherichia coli were significant among the three interaction terms.

Regression Coefficients for the Optimization of Antibacterial Activity of Cannabis Sativa Against Selected Bacteria
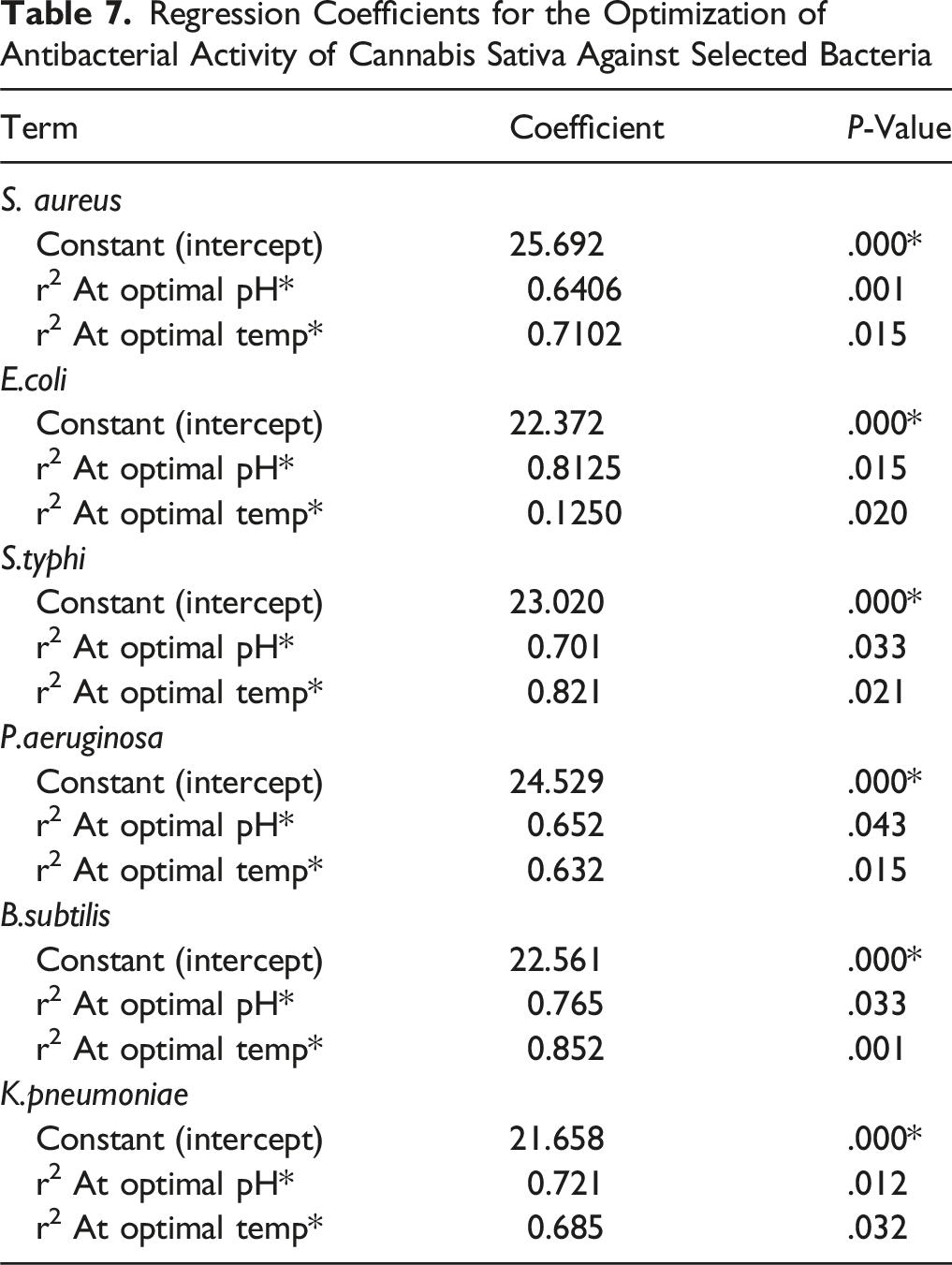
To confirm that the interaction effects were significant, the P-values were employed as a tool. The adjusted r2 and the coefficient of determination (r2) were used to represent the quality of regression model fitness. Our findings consistently showed r2 values in the range of 0 and 1. The response is more accurately anticipated and the model is stronger when the value of r2 approaches 1 (Table 7).
Counter plot is used to study the relationships among the test variable and to check the maximum level of each variable that can give higher zone of inhibition due to antibacterial activity of C. sativa extract. Out of three independent variables, one variable is kept constant and the other two are kept in the x-axis and y-axis in the counter plot graph. Then the relationship between two variables is examined using the graph for bacterial strains.Counter plot between the three variables are given in Figure 1. The graphs in Figure 2 indicates that the highest zone of inhibition of S.typhi by plant extract have been achieved at 10.0 µg/mL, pH 5.1 and temperature 37.5°C. It shows that by keeping the pH low, antibacterial activity will increase. Contour Plots of Different Strains of Bacteria Using RSM
In the graphs of Figure 2 of S.aureus, the growth of bacteria is inhibited by all the given concentrations but when the pH was increased to 6.5, the bacterial growth was inhibited only at the concentration of 10 µg/mL. This shows that pH has a major effect on the growth of S.aureus. However concentration did not show much effect though it has also contributed to the results. All the P-value of temperature, pH and concentration of all six bacteria indicated that the variable, which caused the highest antibacterial activity, is pH and temperature. Each graph showed different pattern. Graph in Figure 2 of E.coli indicates that the combined effect of temperature and pH gives the maximum antibacterial activity. This shows that concentration of plant extract is playing less contribution in effecting the growth of E.coli. The contour plot of P.aeruginosa also indicated that concentration contribute very less in the results, as maximum antibacterial activity was achieved at a pH of 5.0, temperature 35° but any concentration between 5.5 to 9.5 µg/mL. Graph in Figure 2 for B.subtilis shows that major contribution in maximum antibacterial activity was due to pH (5.0), at 35 to 38.9 °C. Graph of K.pneumoniae in Figure 2 also shows less contribution of the concentration in getting maximum antibacterial activity.
Discussion
Infectious diseases continue to become the major cause of mortality and morbidity all over the world. Microorganisms occupy approximately 60% of the total earth’s biomass. In addition to this they have a wide variety in their genetic, metabolic and physiological makeup which makes them more complex for scientist to come up with the cure for the disease they cause. As microorganisms becomes resistance to antibiotics by many ways, making the situation worse. Temperature and pH have affected the results more than the concentration has affected as clearly indicated by contour plot graphs, in this study. The results of this investigation into the antibacterial properties of the ethanolic extract derived from C. sativa leaves are in line with earlier findings by Serventi et al 18 They emphasized the hydroalcoholic extract’s antibacterial properties from the strawberry inflorescences of C. sativa L. cv.19-24 With regard to Escherichia coli, Bacillus subtilis, and Pseudomonas aeruginosa, the extracts’ minimum inhibitory concentrations (MIC) ranged from 4.96 to 39.68 μg/mL, 1.56 to 19.8 μg/mL, 39.68 to 62.99 μg/mL, and 15.74 to 62.99 μg/mL, respectively, according to the results. This was consistent with research by Appendino et al, 24 who found that C. sativa cannabinoids suppressed Staphylococcus aureus growth with MIC values ranging from 0.125 to 128 μg/mL. Additionally, Iseppi et al 25 revealed that essential oils derived from fiber-type C. sativa L. had antibacterial properties against bacterial infections. According to their findings, the minimum inhibitory concentrations (MIC) for Staphylococcus aureus, Staphylococcus epidermidis, Enterococcus faecalis, Bacillus subtilis, and Bacillus cereus ranged from 2-32 μg/mL, 1-16 μg/mL, 0.5-32 μg/mL, and 1-26 μg/mL, respectively. A study on the antibacterial qualities of silver nanoparticles made by combining several C. sativa leaf extracts was carried out by Csakvari et al 26 Their results showed that the combination of C. sativa leaf extracts with silver nanoparticles had strong antibacterial action against Staphylococcus aureus, Escherichia coli, Klebsiella pneumoniae, and Pseudomonas fluorescens. These results implied that the ethanolic extract from C. sativa leaves might prevent or lessen the growth of germs that are harmful to humans, which could be advantageous for the creation of natural antibiotic medications. By simulating the interaction between different elements influencing the antibacterial activity and the response (eg, inhibition zone size), RSM can be used to anticipate and optimise the antibacterial properties of plants. This technique makes it possible to determine the best circumstances for plant extract extraction, processing, or application in order to optimise their antibacterial activity. 27 The primary goal of this study was to use the RSM to maximize the antibacterial activity of C. sativa extract as it is based on a number of statistical and mathematical methods It forecasts the optimization process, lowers simulation costs, and optimizes a process related to optimization. The Box-Behnken-based Design-Expert® (v. 7) software was utilized in the current investigation to predict the best response using RSM and assess the impact of independent variables on response performance. In order to examine the antibacterial activity of C. sativa extract, this approach chose three independent factors: temperature, pH, and concentration with the inhibition rate. As seen by the surface plot with the notable interactions between concentration, pH, and temperature, the contour plots demonstrate that temperature and pH are the most efficient parameters for providing the highest antibacterial activity against all bacteria. This was confirmed with a variety of grading or evaluation methods provided by docking findings. To find the best ligand orientation and conformation needed for binding at the active site, molecular docking techniques typically use an energy-based scoring system. 28 The scoring function separates the incorrect conformations to be sorted from the proper macromolecular-ligand (C. sativa compounds as target drugs) conformations in order to identify legitimate binding conformations with bacterial proteins. 28 While all ligand compounds in our study have the ability to bind with receptor proteins because their binding affinities are greater than −7.0 Kcal/mol (mentioned in Table 1), Tetrahydrocannabinol, Cannabicyclol, Tetrahydrocannabivarin, and Nabilone have premising binding affinities of −8.2 Kcal/mol, −9.4 Kcal/mol, −8.3 Kcal/mol, and −8.0 Kcal/mol, respectively, indicating that these compounds can interact with the targeted proteins more affectively leading to the fact that they can be considered as a components of future drugs. All of the chosen compounds meet the Lipinski rule of five, which states that where cannabinol, delta-8-tetrahydrocannabinol, tetrahydrocannabinol, cannabicyclol, napinol, and cannabigerolate have one infraction each, beta-eudemsol, tetrahydrocannabivarian, and cannabitriol have none. In our study, the in vitro analyses of the contour plots are in correspondence with the docking scores of the target bacterial proteins with the cannabinoids. The compounds cannabicyclol, nabilone, tetrahydrocannabin, beta-eudemsol and tetrahydrocannabivarin have the maximum binding affinity against the proteins beta-lactamase 1SHV, DNA gyrase inhibitor, lipoteichoic acid, penicillin binding protein and topo-isomerase IV respectively, which clearly confirms the anti-bacterial potential of above said cannabinoids against the target proteins at suitable pH and temperature, while concentration has no significant effect on inhibition activity. All of the substances are very soluble, as demonstrated by intestinal absorption. Different categories of solubility are indicated by logs for water solubility, which vary from −10 to 0 (−10, −6, −4, −2, and 0 for insoluble, poorly soluble, soluble, very soluble, and very soluble, respectively).
Limitations
Limitations of this study include that only the particular range of input parameters tested yields reliable predictions from the RSM model. It is unable to accurately predict or interpret results for systems that operate outside of this specific experimental window. Another primary drawback of RSM is that, it only approximates a system mathematically rather than elucidating the biological dynamics at play. It is unable to clarify the way in which specific elements combine to create an antibacterial action, which is essential for comprehending the creation of novel medications. Furthermore, its incapacity to take into consideration the intricacy and interactions of several active chemicals in the extract also the challenge of pinpointing the precise elements causing activity, and the necessity of having a solid grasp of the factors affecting antibacterial qualities beforehand.
Conclusion
This study is the continuation of the research to examine the effectiveness of ethanolic extracts made from C. sativa leaves against harmful microorganisms in humans. The results show that this extract has strong antibacterial activity against a variety of pathogens, such as Pseudomonas aeruginosa, Klebsiella pneumonia, Escherichia coli, Bacillus subtilis, Staphylococcus typhi, and Staphylococcus aureus which is affected more strongly by the pH and temperature variations rather than the concentrations of the extract. Moreover, it is confirmed by the application of the RSM model which indicates its activity. The zones of inhibition produced in the repetitive study has been concluded that C. sativa may be qualified as the drug of the future that can be efficacious for combating bacterial infections. The said plant is of high importance to synthesize a very high potency antibacterial drug by using the optimized ranges of temperature and pH.
Supplemental Material
Supplemental Material - RSM-Based Optimization of Dose Response and Antibacterial Potential of Cannabis sativa (L.) Leaves Using Computational Analysis
Supplemental Material for RSM-Based Optimization of Dose Response and Antibacterial Potential of Cannabis sativa (L.) Leaves Using Computational Analysis by Sara Zahid, Sampath Chinnam, Asma Ahmed, Syed Muzzammil Masaud, Beenish Khurshid, Saira Farman, Sana Rashid, Fatima Zahid in Dose-Response
Footnotes
Acknowledgements
Authors are highly thankful for of all the staff and students of lab-313, The University of Lahore-Pakistan for their valuable contributions in the preparation of this manuscript.
Ethical Considerations
This article has been approved by the Public Health Committee of the Institutional Review Board (IRB/2009-45140) of The University of Lahore, Lahore, Pakistan.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
This article includes all of the available data. No further data is available.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.

