Abstract
Introduction
The concept of liquid biopsy was developed for circulating tumor cell (CTC) more than 10 years ago.1,2 Liquid biopsy has become a new method of collecting biomarkers from body fluids, for instance, blood, urine, and cerebrospinal fluid. Shortly afterward, researchers promptly extended this technique to circulating tumor DNA (ctDNA), 3 and other tumor-associated products, such as circulating cell-free RNA,4,5 besides tumor-educated platelets. 6 Liquid biopsy has emerged as a prominent research focus in molecular biology due to its significant advantages as a minimally invasive diagnostic modality with high analytical accuracy. This innovative approach enables real-time monitoring of circulating biomarkers through advanced genomic and proteomic profiling techniques, offering substantial clinical potential for early disease detection and personalized therapeutic interventions.
Different types of nanoscale sized exosomes that are continuously released by viable cell which belong to EVs in various ways, for instance, exosomes can actively releasing through plasma membrane fusion through extracellular interactions with vesicular vesicles, or by the cell surface germinated micro cytoplasmic protrusion. The average diameter of an exosome is between 30 and 200 nm, making them a nanoparticle.7–9 Various therapeutically relevant diagnostic options exist due to the rich chemistry of exosomes, which include proteins, nucleic acids, lipids, oligosaccharides, and metabolites. 10 There is a real possibility that exosomes may be used for cancer screening and early diagnosis because of the information they disclosed about viable tumor cells. A cancer cell may release approximately 20 000 of these vesicles within about 48 hours. 11 The transportation mechanism has the potential to carry a diverse range of biomarkers that originate from the tumor or tumorigenic process. Exosomes have the capability to expedite the proliferation of tumor cells, stimulate the formation of new blood vessels (angiogenesis), facilitate the migration of tumor cells, impede the immune response, and facilitate the development of metastases. These aforementioned characteristics are recognized as key indicators of malignant cellular behavior. 12 These findings lead to the orientation of becoming a new approach for monitoring cancer biomarkers.
Using the existing medical technology to identify optimal tools for early-stage tumor diagnosis while reducing diagnostic harm is a crucial challenge.13,14 With superior sensitivity for early disease detection and exceptional biomarker stability, exosome-based liquid biopsy emerges as an ideal approach satisfying these critical diagnostic needs.
15
This review describes tumor-related exosomal biomarkers in several common cancers, including lung cancer, breast cancer (BC), kidney cancer, prostate cancer (PCa), and colorectal cancer (CRC), using exosome-based liquid biopsy techniques (Figure 1). We further explore their clinical prospects and summarize the technologies for isolating exosomes and detecting exosomal-based biomarkers. Finally, the limitations and prospects of exosome-based liquid biopsy in the early screening and diagnosis of cancers are discussed. Exosome-Based Liquid Biopsy for Several Common Cancers. Tumor-Related Exosomal Biomarkers can be Detected in Body Fluid (Plasma, Serum, Pleural Fluid, Semen, Urine) Using Exosome-Based Liquid Biopsy Techniques.
Exosomal Biomarkers in Cancers
Lung Cancer
According to global statistics, the frequency of lung cancer is the highest amongst cancer subtypes.16,17 In the year 2022, lung cancer constitutes 12% of newly diagnosed cancer cases. Furthermore, it is noteworthy that lung cancer has the highest fatality rate among both males and females, accounting for 21% of all cancer-related deaths. 18 Earlier detection and diagnosis of lung cancer in high-risk populations is crucial and has a non-negligible effect on lung cancer outcomes. The implementation of timely identification and assessment leads to a notable enhancement in the 5-year relative survival rates. Specifically, the survival rate escalates from a mere 6% for cases diagnosed at an advanced stage to 33% for those detected at a regional stage, and further rises to 60% for those diagnosed with localized-stage illness. 18 This intervention has the potential to greatly enhance the likelihood of survival in individuals diagnosed with lung cancer. Recent research has shown that exosomal biomarkers play a pivotal role in facilitating the transmission of information between cells affected by lung cancer and neighboring cells. 19 Exosomes can thus be used for early diagnosis. Understanding exosomes and their biomarkers may be the key to understanding lung cancer. Exosomes and their biomarkers play an important role as mediators for intercellular information exchange between lung cancer cells and other cells, and may become important tools for the detection and early diagnosis of lung cancer. 20
Lung Cancer Potential Biomarkers in Clinical Practice.

Lung cancer is mainly divided into two types: small-cell lung cancer, or another types of non-small cell lung cancer (NSCLC), 24 which as an important protagonist and with the accounts for approximately 85% of all lung cancer types. 25 A study analyzing the dynamic profiles of plasma exosomal miRNAs in advanced NSCLC patients treated with nivolumab identified that baseline miR-181a-5p and miR-574-5p, combined with performance status, effectively stratified patients into distinct prognostic groups (median overall survival of 4 months vs over 9 months), establishing a clinical prognostic stratification model incorporating exosomal miRNAs. Further investigation revealed six dynamically regulated exosomal miRNAs (including miR-19a-3p, miR-1260a, miR-1260b, and others) in responding patients, which exhibited significant downregulation during early treatment cycles and sustained suppression for over 6 months, suggesting their potential as real-time diagnostic and therapeutic monitoring biomarkers. 26 A noteworthy study focusing on early‐stage NSCLC and its histologic heterogeneity recently reported that four exosomal miRNAs (miR‐let‐7b‐5p, miR‐let‐7e‐5p, miR‐23a‐3p, and miR‐486‐5p) were promising diagnostic markers of stage I NSCLC, with sensitivity and specificity values of 80.5% and 92.31%, respectively. 27 At present, therapeutic drugs show positive effects on NSCLC patients with mutant epidermal growth factor receptor (EGFR) expression, whereas a lower beneficial effect is generally noted on NSCLC patients expressing wild-type EGFR.28,29 Therefore, new detection methods must be identified for the accurate clinical diagnosis of NSCLC. Similar to exosomal miRNAs, exosomal circRNAs are expected to act as effective non-invasive biomarkers for liquid biopsy-based early diagnosis of NSCLC. CircSATB2 was highly expressed in NSCLC cells and serum exosomes from lung cancer patients and exhibited high sensitivity and specificity for clinical detection, which indicated that circSATB2 may be a potential biomarker for NSCLC diagnosis. 30
Both lung adenocarcinoma (LUAD) and lung squamous cell carcinoma (SCC) manifest as distinct molecular subtypes within the heterogeneous spectrum of NSCLC, exhibiting divergent genomic landscapes and therapeutic vulnerabilities.24,31 Clinically actionable extracellular miRNAs, particularly those differentially packaged in tumor-derived exosomes, function as subtype-specific biomarkers in discriminating LUAD and SCC. Further studies have conducts research and analysis to identify plasma exosomal miRNAs (miR-19b-3p, miR-21-5p, miR-221-3p) were up-regulated with the expression profiling was performed using exiqon panels, which means they can identified to the detection of the potential biomarkers for LUAD diagnosis. 32 Using qRT-PCR in the screening phase and followed by verification (30 LUAD VS. 10 healthy controls (HCs). Meanwhile, in another study, tumor-derived exosomes were isolated from the plasma of early-stage NSCLC patients (46 stage I NSCLC patients and 42 HCs) to identify and validate miRNAs which is LUAD- or SCC-specific. According to the results, plasma exosomal miRNAs (miR-30a-3p, miR-30e-3p, miR-181-5p, and miR-361-5p) were LUAD-specific, while plasma miRNAs (miR-10b-5p, miR-15b-5p, and miR-320b) were SCC-specific. 33 These findings indicated that the detection of exosomal biomarkers, especially exosomal nucleic acids, can be a new early screening entry for the lung cancer. Exosome-based liquid biopsy has the potential to assist in diagnosis of the lung cancer pathological classification, and provide assistance for non-invasive personalized diagnosis and treatment.
In lung cancer, tumor cells can transmit oncogenic signaling molecules through exosomes, which reflect the actively secreted molecular profiles of tumor cells and reveal real-time biological behaviors. The core challenge in the clinical translation of exosome-based liquid biopsy for lung cancer lies in the subpopulation heterogeneity of tumor-derived exosomes, necessitating the development of multi-marker panels to address variability. Machine learning models (such as random forest) can enhance classification accuracy by integrating exosomal miRNA, proteomic data, and clinical parameters. Additionally, comprehensive exosome-based integrative analyses, like combining exosomal RNA, ctDNA, and CTCs, may enable the construction of molecular subtyping models of lung cancer diagnosis.
Breast Cancer
Globally, BC remains one of the leading causes of cancer-related mortality among women, based on 2024 epidemiological data. 16 Female BC incidence rates have been increasing at a low rate of approximately 0.5% per year since the mid-2000s. 18 Regarding earlier detection and diagnosis of BC, mammography is the established gold standard for BC screening in clinical practice. 34 While debates persist regarding the hypothesized association between mammography screening and reported increases in BC incidence rates, this contention has prompted emerging research efforts to identify exosomal biomarkers capable of improving early detection of BC.
Breast Cancer Potential Biomarkers in Clinical Practice.

An elevation in the levels of serum exosomal miRNAs (miR-21-5p, miR-23a-3p) were founded to distinguish early and advanced BC cases, which are useful for detecting early-stage BC. 38 The study results suggest that exosomal miRNAs can be used as potential biomarkers for early BC diagnosis, thereby overcoming the limitations of mammography when used as a diagnostic technique auxiliary to mammography.
To date, exosomal miRNAs significant for BC diagnosis, including for the differentiation of tumor subtypes, have been established. One investigation showed that miR-93 can be used for the early diagnosis of ductal carcinoma in situ (DCIS),39,40 as patients diagnosed preoperatively with DCIS of BC may develop invasive ductal carcinoma. Thus, applying exosomal miRNAs as BC malignancy diagnostic indicators is challenging and needs further exploration. Additionally, one study employed untargeted lipidomic profiling to reveal that breast cancer-derived EVs are specifically enriched in sphingolipids and glycerophospholipids, enabling accurate discrimination between cancerous/normal cells and their subtypes (100% accuracy in vitro models) through lipid composition analysis. 41 Validation using plasma EVs demonstrated high-accuracy classification of breast cancer patients vs healthy individuals (93.1%), as well as primary vs metastatic cancers (89.5%), validating the potential of EVs (such as exosomes) lipids as non-invasive diagnostic biomarkers for breast cancer. This proof-of-concept study establishes a methodological framework and translational pathway for developing exosomes lipidomics-driven biomarker discovery strategies.
Exosomes carry breast cancer-specific biomolecules like HER2 protein or miRNA, driving tumor growth, metastasis, and drug resistance by transmitting oncogenic signals. Their phospholipid membrane enhances biomarker stability compared to ctDNA, enabling noninvasive dynamic monitoring. Key clinical translation challenges include exosomal subpopulation heterogeneity like triple-negative, lack of standardized isolation protocols, and high costs. Proposed solutions involve multi-marker panels (like HER2+ exosomal miRNA), machine learning integration of multi-omics data, and automated microfluidic detection. Future efforts require multicenter validation of sensitivity (>85%) and specificity (>90%) to promote their application in early screening, treatment response prediction, and recurrence monitoring, advancing precision oncology.
Kidney Cancer
Global epidemiological data reveal a concerning upward trajectory in kidney cancer incidence, with this malignancy now demonstrating a sustained annual growth rate of approximately 2%, a trend exceeding population aging projections and positioning renal cell carcinoma among the fastest-rising genitourinary cancers worldwide.16–18 It is noteworthy that over 30% of individuals diagnosed with kidney cancer experience the development of metastasis. Furthermore, a majority of these instances of distant metastasis occur during the early stages of the disease. 42 Global epidemiological data reveal a concerning upward trajectory in kidney cancer incidence, with this malignancy now demonstrating a sustained annual growth rate of approximately 2% – a trend exceeding population aging projections and positioning renal cell carcinoma among the fastest-rising genitourinary cancers worldwide. Emerging translational studies substantiate that exosomal cargo components like miRNAs, lncRNAs, and tumor-specific surface proteins demonstrate diagnostic utility in renal cell carcinoma through their roles in intercellular oncogenic signaling and immune microenvironment modulation, achieving 82-91% sensitivity in multicenter validation cohorts. 43 This positions exosome-based liquid biopsies as clinically actionable noninvasive biomarkers for early detection, molecular subtyping, and therapeutic response monitoring in kidney cancer management.
Kidney Cancer Potential Biomarkers in Clinical Practice.

RCC constitutes over 90% of primary renal malignancies, with clear-cell RCC (ccRCC) representing 80-90% of cases characterized by distinct histopathological features (cytoplasmic clearing and nested architecture) and canonical genetic alterations (VHL gene inactivation). This predominant subtype’s molecular signature, including chromosome 3p loss and HIF pathway activation, underpins both its diagnostic classification and therapeutic vulnerability to targeted therapies. 42 In one study, early-stage ccRCC patients, healthy individuals, and patients with various urinary system malignancies were compared using NGS-identified differentially expressed urinary exosomal miRNAs. The research showed that miR-30c-5p in urinary exosomes can function as a diagnostic biomarker for ccRCC at an early stage and may be able to regulate the production of HSPA5, a recombinant heat shock protein that is linked to the development of ccRCC. 48 Based on ccRCC exosomal miRNA expression profiles and the literature, a study analyzed the expression of selected six exosomal miRNAs (miR-210, miR-224, miR-452, miR-155, miR-21, and miR-34a) in tissues and sera and hypoxia-induced (with CoCl2) renal cell lines through qRT-PCR. Only exosomal miR-210 was significantly upregulated in patients with ccRCC than in HCs, so serum exosomal miR-210 exhibits potential as a diagnostic for ccRCC. 49 The combinatorial analysis of circulating miRNAs reveals that dual-miRNA signatures (miR-126-3p/miR-449a, AUC = 0.84; miR-126-3p/miR-34b-5p, AUC = 0.79) demonstrate significant diagnostic potential for distinguishing ccRCC from healthy controls, 50 with particular clinical utility in differentiating small renal masses and benign lesions (AUC = 0.79-0.85), while postoperative miRNA normalization patterns and exosomal transfer mechanisms (evidenced by cancer cell-secreted miR-126-3p/miR-17-5p/miR-21-3p/miR-25-3p internalization) suggest their dynamic role in tumor microenvironment crosstalk and potential as liquid biopsy biomarkers. Emerging liquid biopsy platforms demonstrate transformative potential in early-stage renal cancer management, as evidenced by tumor-derived exosomal biomarkers (e.g., miRNAs, proteins) enabling ultrasensitive detection of subclinical tumors through multi-analyte signatures (superior sensitivity: 85-92% vs conventional imaging). Crucially, exosomal payloads maintain tumor-specific molecular fingerprints - including therapy-resistant clones and metastatic drivers - permitting longitudinal monitoring of minimal residual disease undetectable by standard radiographic modalities.
Exosomes in renal cancer carry tumor-specific molecules that drive progression by regulating HIF-1α and immune suppression. Their stability and enrichment enable superior early detection and recurrence monitoring compared to traditional biomarkers. Clinical translation challenges include subtype heterogeneity and low-abundance biomarker sensitivity. Solutions involve multi-omics integration (exosomal RNA/metabolites + ctDNA), AI models (combined with radiomics), and microfluidic enrichment. Future multicenter validation of biomarker panels with AUC >0.88 is critical to advance applications in early screening and targeted therapy monitoring, enhancing precision oncology.
Prostate Cancer
Prostate cancer (PCa) represents the most frequently diagnosed malignancy in the male population, accounting for approximately 27% of all incident cancer cases. Globally, this neoplasm stands as the second-leading cause of cancer-related mortality among men, demonstrating significant epidemiological impact through both its high incidence rates and substantial disease burden. 18 Obviously, the demand for more effective diagnosis and treatment for PCa is explicit. Currently, concerns exist regarding the precision of clinical risk stratification models such as prostate-specific antigen (PSA) levels, as well as the diagnostic and safety yield of PCa biopsy. Exosome-based liquid biopsies offer significant advantages in prostate cancer management by enabling non-invasive detection of tumor-derived biomarkers, including nucleic acids and proteins, with enhanced sensitivity for early-stage disease identification. This approach circumvents the need for invasive tissue biopsies. Exosomal biomarkers may have a potential application in the early screening and diagnosis of PCa through a fast and high-throughput method.
Prostate Cancer Potential Biomarkers in Clinical Practice.

Survivin expression has been detected both in PCa tissues and plasma-derived exosomes across distinct clinical cohorts, including HCs, BPH patients, and PCa patients, suggesting its potential dual role as a tissue-specific oncoprotein and circulating biomarker. 57 The relative amounts of exosomal survivin in the plasma of PCa patients were significantly higher than those in the pre-inflammatory plasma of BPH patients and HCs. Exosomal survivin was differentially expressed in both patients with newly diagnosed and advanced PCa with low- or high-grade cancers. Plasma exosomal survivin levels may act as a convenient diagnostic tool for PCa. Because exosomal survivin levels are elevated in low as well as high Gleason-scored samples, these survivin levels may be used for early PCa detection. 58 Overall, the liquid biopsy diagnosis of PCa has a variety of exosomal biomarkers, and the combination of prostate-specific markers and exosomal biomarkers provided novel strategy for the early screening and diagnosis of PCa.
Exosomes carry prostate cancer-specific molecules (e.g., miR-141), driving tumor progression by regulating androgen receptor signaling and promoting bone metastasis. Their phospholipid membrane enhances biomarker stability compared to ctDNA, enabling noninvasive dynamic monitoring. Key clinical translation challenges include subtype heterogeneity (hormone-sensitive vs castration-resistant subtypes), lack of standardized isolation protocols, and low biomarker abundance. Solutions involve developing multi-marker panels like PSA + exosomal miRNA, AI-driven integration of multi-omics data including exosomal RNA, ctDNA, radiomics, and automated microfluidic detection. Future multicenter validation of sensitivity and specificity is critical to advance their application in early diagnosis and precision oncology.
Colorectal Cancer
CRC constitutes a major global health burden, epidemiologically significant as the second most prevalent malignancy and the second-leading cause of cancer-associated mortality worldwide, underscoring its substantial disease burden and urgent need for improved diagnostic and therapeutic strategies.18,59 While surgical resection remains the diagnostic gold standard for tumor assessment, conventional tissue biopsy methodologies present inherent clinical limitations due to their invasive nature and inability to comprehensively characterize tumor heterogeneity, primarily stemming from sampling bias in localized specimen acquisition that fails to capture spatiotemporal tumor evolution. 59 Hence, the examination of exosomal biomarkers assumes significant importance in addressing the pressing requirement for the discovery of a new diagnostic approach that is minimally invasive or noninvasive, specifically for the early detection of CRC.
Colorectal Cancer Potential Biomarkers in Clinical Practice.
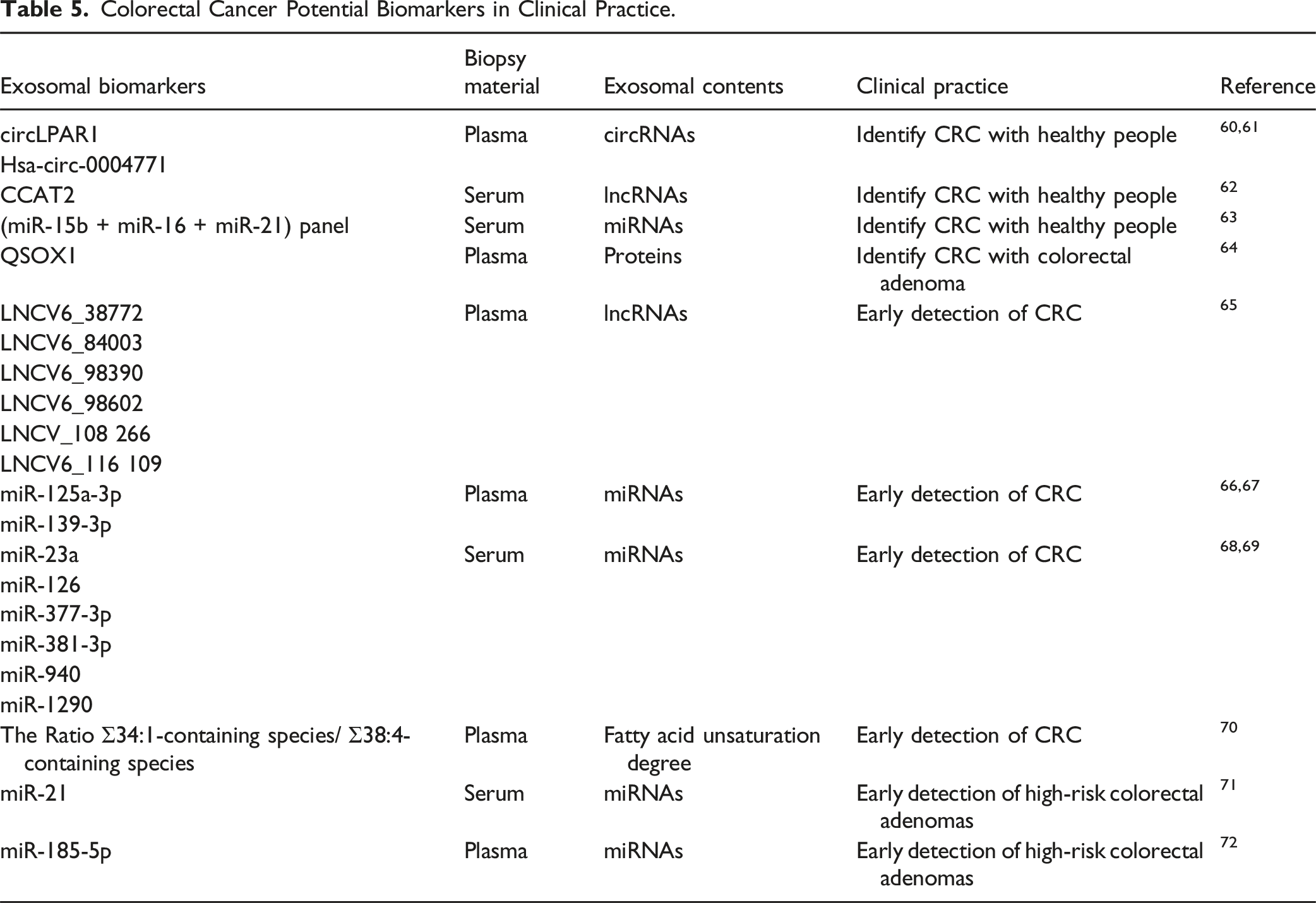
Early CRC detection is critical so that early-stage cancer patients have a survival rate highly than 90%. 18 In a study with an independent test cohort, recombinant quiescin q6 sulfhydryl oxidase 1 (QSOX1) was observed in the plasma exosomes of CRC patients compared with HCs, which has a significantly reduced protein abundance, with the diagnostic ROC curve analysis revealing an AUC of 0.904. 64 This revealed that exo-associated QSOX1 may be a promising novel marker for the early diagnosis of CRC and non-invasive risk stratification in CRC. Notably, the combination of exosomal lncRNAs demonstrated markedly elevated AUCs of 0.736 and 0.758 for CRC and early-stage CRC respectively. Furthermore, a separate study found that the expression of six exosomal lncRNAs (LNCV6-38772, LNCV6-84003, LNCV6-98390, LNCV6-98602, LNCV-108266 and LNCV6-116109) was considerably elevated in the plasma of 50 CRC patients, suggesting that they may serve as possible non-invasive biomarkers for early CRC detection. 65 Researchers have found that miR-125a-3p and miR-139-3p in plasma-derived exosomes can serve as novel biomarkers for early-stage CRC, leading them to focus more attention on circulating exosomal miRNAs in CRC and their function in CRC early detection,66,67 miR-377-3p and miR-381-3p, both of which are found in serum exosomes, have been identified as potential new biomarkers for early-stage CRC. 68 In addition, high diagnostic values to identify CRC patients (TNM stage I) from HCs led to the identification of four serum exosomal miRNAs (miR-126, miR-1290, miR-23a, and miR-940) in CRC patients as possible biomarkers for the early detection of CRC. 69 Furthermore, the data revealed that, independent of internal miRNA expression, CRC cells secrete exosomal miRNAs into the extracellular environment via exosomes.
The level of fatty acid unsaturation in plasma exosomes was studied as a potential exosomal biomarker in colorectal cancer patients. Patients with CRC were categorized as either healthy, hyperplastic polyps, adenomatous polyps, invasive neoplasia CRC, or having hereditary non-polyposis CRC. Plasma exosome lipid fingerprint was shown to be useful in differentiating between HCs and pathological patients based on the ratio of 34:1-containing species to 38:4-containing species, and this study further bolstered the potential of this non-invasive biomarker for therapeutic applications. 70 The ability of exosomal miRNAs to diagnose high-risk cases of colorectal adenomas was demonstrated in another study. Twenty normal colonic mucosa, twenty-seven adenomas, and nineteen CRC tissues were studied for miRNA expression, and the expression of potential exosomal miRNAs was assessed in total serum and exosomes from 26 adenoma patients and 47 HCs. According to the findings, exosomal miR-21 has a strong relationship with both the number and size of adenomas, and it can distinguish between patients with low- and high-risk adenomas. 71 Exosomal miR-185-5p in patient plasma has also been used as a marker for the presence of advanced adenomas and colorectal cancer. 72 These exosomal biomarkers, especially miRNA panel provides the possibility for early detection of CRC in clinical practice.
In colorectal cancer exosomes carry tumor-specific biomarkers (e.g., miR-21) that drive tumorigenesis via Wnt/β-catenin pathway activation and immune evasion. Their phospholipid membrane protects cargo stability, outperforming ctDNA for noninvasive early detection and monitoring. Clinical translation faces challenges like exosomal heterogeneity, isolation standardization, and cost barriers. Solutions include multi-marker panels (CEA + exosomal miRNA + KRAS), AI-driven integration of exosomal RNA, ctDNA, and radiomics data, and microfluidic platforms for automated analysis. Future multicenter validation is essential to advance applications in early screening and personalized therapy guidance, aligning with regulatory pathways for clinical adoption.
Isolation and Detection Technology for Exosome-Based Biomarkers
Methods for the Isolation of Exosomes
Highly purified exosomes are the prerequisite for the identified exosome-related biomarkers and for obtaining reliable data for precise cancer diagnosis. Therefore, the isolation of exosomal biomarkers needs to meet the pre-analysis requirement.73–75 Ultracentrifugation, immunoaffinity isolation, polymeric precipitation isolation, and liquid chromatography are among the prevailing methodologies employed for the extraction of chemicals from human samples, encompassing a range of physical, chemical, and biologically-based approaches (Figure 2). Traditional and Novel Methods for the Isolation of Exosomes.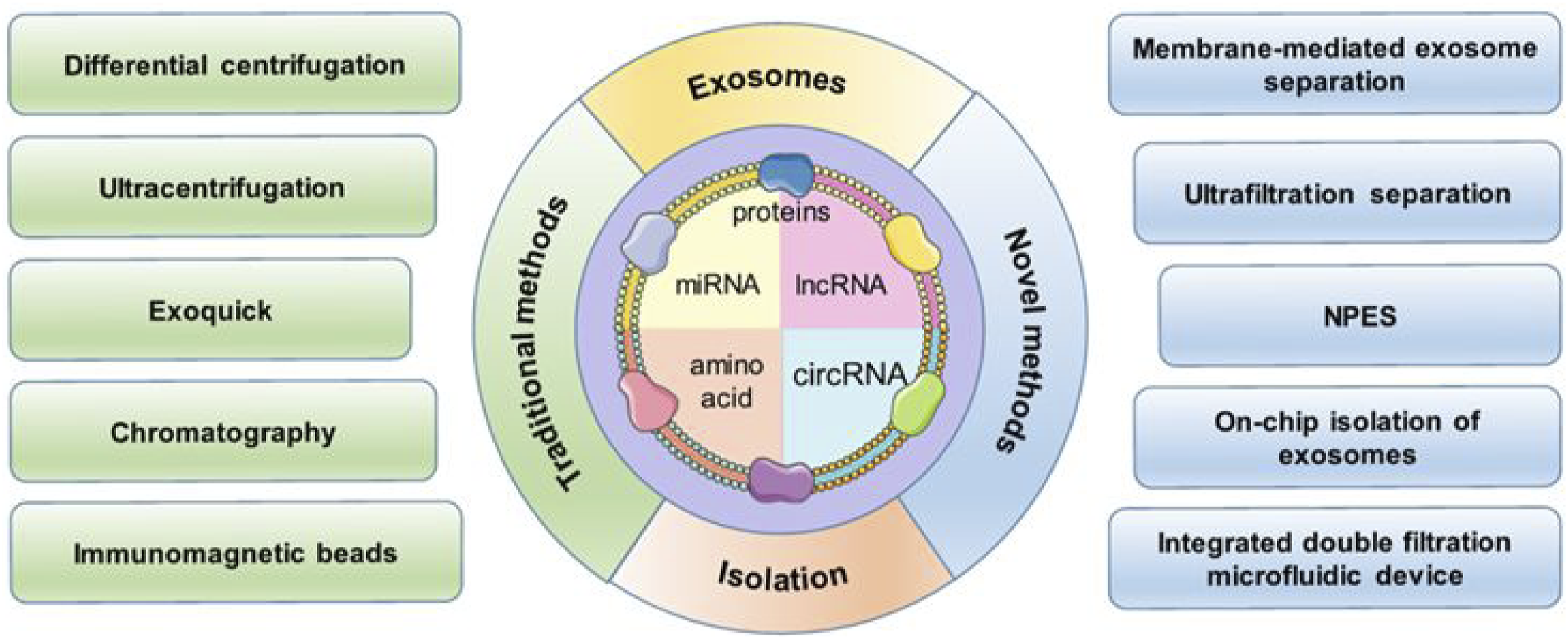
Comparison of Common Exosome Isolation Techniques.

Interestingly, a cascaded microfluidic circuit is designed using electro-hydraulic simulation. For pulsatile filtering of EVs, this circuit combines a cell removal circuit with the exosome isolation method circuit. 86 This pneumatic clock pulse generator-driven microfluidic device can directly separate preprogrammed, non-blocking, mild, high-yield, and high-purity EVs from the blood within 30 min. A polyethylene glycol-enhanced hyperthermia sensor is then used to detect protein markers of EVs isolated from the patient’s blood, thus proving its clinical application. The accuracy rate of this early diagnosis of BC is 91%. Cascaded microfluidic circuits have broad application prospects for the novel isolation of exosomes.
Detection Technology for Exosome-Based Biomarkers
Detection Technology for Exosomal Proteins
Exosome-based membrane proteins may be involved in cancer development and progression. 87 Consequently, exosome-based liquid biopsies can be utilized to target exosomal proteins for the purpose of detecting malignancy. The WB technique and the enzyme-linked immunosorbent assay (ELISA) are the prevailing methodologies employed for the detection of exosomal proteins. 88 Nevertheless, the utilization of these methodologies is sometimes hindered by challenges related to the intricacy of programming and the limited level of sensitivity they offer. Various innovative methods have been explored thus far for the identification of exosomal proteins.
Colorimetry is a scientific discipline that involves the analysis of chromogenic substances by the comparison or measurement of their color intensity, with the aim of determining its constituent composition. A recently developed colorimetric aptasensor has been designed to detect cancer-derived exosomes with simplicity, sensitivity, and selectivity. 89 The biorecognition of target exosomes was facilitated by employing the CD63 aptamer, after the collection of those exosomes using latex beads through aldimine condensation. The process of biotin-streptavidin binding was employed in order to conjugate horseradish peroxidase (HRP) with the aptamer under investigation. The colorimetric detection method required an only 10-minute duration. The colorimetric signal was enhanced by the in situ application of polydopamine (PDA) onto exosome particles. The signal could be visually perceived by individuals. In summary, the utilization of HRP-accelerated DA polymerization and in situ PDA deposition holds great potential for the advancement of point-of-care cancer detection kits in clinical settings.
Fluorescence spectrophotometry identifies and determines the substance content based on the positivity and intensity of the fluorescence spectrum lines. A novel ap-tamer-based method for the competitive detection of exosomes based on a CD63-specific aptamer was developed while ensuring sensitivity cost and reducing difficulty significantly. 90 Another study reported the development of a hybridization chain reaction-based assay for the fluorometric determination of exosomes by using magnetic nanoparticles and both aptamers and antibodies as recognition elements. The aptamer sequence binds to the exosomal surface proteins (from HepG2 cells). 91
Electrochemical detection possesses the capability to detect the analyte’s presence with high sensitivity by monitoring the electro-chemical potential or current of the sample across a wide range of dynamics. 92 A study introduce a novel electro-chemical aptasensor (DeMEA)-equipped detachable microfluidic device for the purpose of real-time evaluation of cancer-associated exosome concentration. The DeMEA method is employed for the quantification of exosomes extracted from plasma samples collected from breast cancer patients at different stages of the disease. Hence, the utilization of microfluidic-integrated biosensors is employed in this methodology for the early detection of cancer biomarkers.
The utilization of optical sensors in surface plasmon resonance (SPR) biosensors involves the application of surface plasmon polaritons, which are a distinct type of electromagnetic radiation, for the purpose of examining the interactions between analyte molecules in a solution and biomolecular recognition components that are attached to the surface of the SPR sensor. 93 SPR technology is employed in the clinical detection of human blood tumor markers to identify a sandwich-type immunological complex. This complex comprises a sensor surface that has been immobilized with a primary antibody, the tumor marker itself, and a secondary antibody. The proposal of a new type of integrated microfluidic device can quickly and hyper sensitively detect extracellular proteins. The results show differential expression of lung cancer specific extracellular protein leucine-2-glycoprotein 1 (LRG1) collected from patient urine samples, this technology exhibits promising potential in the identification of individuals afflicted with early-stage lung cancer, as well as in the comparative analysis of those individuals with a group of healthy controls. The recent research has developed a micro-fluidic device based on dual tangential flow filtration. This method can be employed for the isolation of exosomes from human serum, which has been demonstrated to exhibit heightened sensitivity towards cell membranes. 94
Surface-enhanced Raman spectroscopy (SERS) uses light to interact with nanostructured materials like gold and silver, resulting in a potent SPR effect. The Raman signal of molecules adsorbed on the nanostructure surface can be greatly amplified by this phenomenon, leading to a detailed fingerprint of the sample or Raman probe molecules with high sensitivity.95,96 SERS exhibits considerable promise as a viable technique for application in the fields of life science and clinical testing due to its notable advantages. These advantages encompass non-invasiveness, heightened sensitivity, robust selectivity, expedited processing speed, and less susceptibility to water interference. With the development of plasma-enhanced methods, many research teams have used raman spectroscopy to analyze exosome phenotypes and characteristics. These studies have indicated that EVs in various biological samples, from cell culture media to blood, can be recognized even when surface markers and specific antigens are absent.
A literature analysis indicates that numerous exosomes utilizing SERS have been suggested for qualitative and quantitative investigations of specific targets, employing various recognition, separation, and amplification techniques, 91 cancer screening, and early detection of various cancers, such as ovarian cancer, lung cancer, BC, pancreatic cancer, melanoma, and osteosarcoma. The advancements in various fields, including multivariate statistical methodologies, machine learning, and AI, have shown to be highly productive in enabling accurate and simple identification of these vibration spectra. Researchers from Korea’s Exo PERT company and Korea University have demonstrated a novel liquid biopsy technology that combines AI with SERS to achieve a hybrid approach. 97 Using 520 test samples not used for training, six cancer types (namely lung cancer, BC, CRC, liver cancer, pancreatic cancer, and gastric cancer) were detected. The diagnostic performance of this method has been tested and used for simultaneously diagnosing multiple cancer types through unmarked analysis of plasma exosomes. This study had a retrospective design, wherein AI was used to analyze the SERS spectrum. This study aims to investigate the role of exosomes in the detection of six early malignancies and evaluate their diagnostic accuracy. The extraction of exosomes, the acquisition of SERS data, and the subsequent processing of these signals using deep learning algorithms were undertaken by researchers in order to carry out a comprehensive investigation of the heterogeneity of cancer for diagnostic purposes. Researchers used 520 test samples and obtained an analysis system that could identify the presence of cancer, by having a 0.970 AUC area under the curve ROC analysis. Furthermore, the performance of the system in categorizing the tumor organ types of 278 patients with early-stage cancer was evaluated using the area under the AUC, which yielded a value of 0.945. The ultimate integrated decision-making model demonstrated a sensitivity of 72% for false positives and a specificity of 94% for the detection of tumor organs. Furthermore, as a non-specific analysis of Raman signals was used in this method, its diagnostic range may be expanded to include other disease types for more applications.
Detection Technology for Exosomal Nucleic Acids
Exosomes carry various nucleic acids, with RNA being one of the predominant types, and extensive research on exosomal RNAs has shown their potential as specific biomarkers for cancer diagnosis. 98 The most commonly used detection techniques for quantifying the expression level of extracellular nucleic acid are qRT-PCR, microarray, and NGS. 99 Because of the limitation of qRT-PCR for nucleic acid detection using known sequences and the high cost, large data volume, complex library construction for NGS, and low sensitivity of the microarray, scientists are committed to developing convenient and highly sensitive methods for detecting exosomal nucleic acids.99,100 Herein, we focus on ddPCR and magnetic beads (MB) technology.
The ddPCR method employs a technique whereby the PCR mixture is partitioned into several water-in-oil emulsion droplets, reaching a scale of tens of thousands. The primary means of obtaining genetic material for tumor genomic profiling is through plasma, making it the focus of investigation for liquid biopsy using ddPCR. Researchers are exploring the use of different biological fluids, including cerebrospinal fluid, urine, stool, ocular fluids, sputum, saliva, bronchoalveolar lavage, pleural effusion, mucin, peritoneal fluid, fine needle aspirate, bile, and pancreatic juice. 99 A recent research conducted an optimization of annealing parameters, including annealing temperature, annealing duration, and number of PCR cycles, for the purpose of detecting urinary exosomal miRNAs using ddPCR. Furthermore, the study compared the performance of ddPCR with qPCR. The findings of the study indicated that ddPCR exhibited superior levels of sensitivity, reproducibility, and accuracy in comparison to qPCR. 101 In order to facilitate the timely identification of endometrial cancer in individuals, a technique known as small RNA sequencing was used to ascertain potential exosomal miRNAs (specifically miR-15a-5p) that might serve as diagnostic biomarkers. This was accomplished via the utilization of ddPCR. 102
The hairpin-like oligonucleotide, known as MB, employs fluorescent dyes and quenching agents at both ends of the probe for the purpose of labeling and facilitating the spontaneous hybridization of the target sequence. This hybridization process leads to the disruption of the hairpin ring structure and subsequently induces fluorescence. 99 The project included the construction of a DNA zipper-mediated membrane fusion method to facilitate the quick detection of exosomal miRNA. This was achieved by loading zipper DNA constructs (ZDCs) onto the surface of MB. In the present study, the exosome of interest is subjected to the introduction of cZDCs. When ZDCs (Zinc-Doped Carbon Dots) and CZDCs (Cadmium-Zinc-Doped Carbon Dots) are combined, the interconversion between these two structures leads to the merging of exosome membranes and vesicle probes. This process activates the detection of exosomal miR21 by the encapsulated MB and subsequently results in the emission of fluorescence. This fluorescence may be readily monitored within a time frame of 30 min. Significantly, the use of flow cytometry enables the differentiation between tumor exosomes that overexpress miR-21 and exosomes released by normal cells. This distinction has considerable promise in the field of liquid biopsy, particularly for the timely detection of cancer. 103 MB biosensors have enormous potential in cancer diagnosis through liquid biopsy, with the advantages of simple operation and dispensing with extracellular pre-separation and extraction of nucleic acid.
Limitations and Perspectives
The excellent circulating stability of exosomes in body fluids has been confirmed, and the status of tumor progression, which contains diverse information, can be reflected through exosomes. Different research data can be combined to reflect the enormous potential of exosomal biomarkers as tools for early cancer screening and detection, ultimately contributing to clinical diagnosis.98,104–106 However, because of the high heterogeneity and nanoscale limitations of exosomes, exploring the molecular information and interactions of exosomes has been posed with significant technical challenges. The comprehensive validation of the isolation and characterization of individual exosomes associated with a particular kind of cancer from a substantial cohort of clinical samples need to be extensively established. However, there is still a lack of proven standardized techniques and protocols for the efficient and precise isolation of exosomes from cell culture medium and human body fluids, particularly under high-throughput and high-purity conditions. The ideal scenario is to develop a standardized method for isolating exosomal biomarkers from human samples in the long future. This method should be simple, time-saving, accurate, and affordable. The use of this technology has the potential to facilitate early screening and detection of cancer patients within a substantial population. Additionally, this will facilitate the development of novel uses of exosomes inside clinical laboratory settings.
Exosomes, as nano-scale vesicles carrying bioactive molecules such as nucleic acids, proteins, and lipids, demonstrate significant potential in disease diagnosis and prognostic evaluation. However, the clinical translation faces critical bottlenecks due to the lack of standardized protocols and validation systems. To achieve standardization, comprehensive operational guidelines must be established across the entire workflow: from sample collection (e.g., standardized temperature and time controls for blood/urine processing), exosome isolation (harmonizing methods like ultracentrifugation, size-exclusion chromatography, or commercial kit selection), to biomarker detection (e.g., homologous sequence calibration for RT-qPCR primer design in miRNA quantification). Cross-platform technical standards should be developed to ensure comparability. For clinical validation, adherence to the gold standards of IVD development is essential. Multi-center, prospective cohort studies are required to verify biomarker sensitivity and specificity. For instance, in early lung cancer screening, studies should enroll over 2000 high-risk individuals, comparing exosomal EGFR mutation detection against the gold standard of tissue biopsy, while analyzing ROC curve parameters and negative/positive predictive values. Addressing exosomal heterogeneity, machine learning-based multi-omics integration models should be developed to enhance diagnostic accuracy. Ultimately, navigating regulatory pathways will enable closed-loop translation from laboratory research to clinical implementation in biomolecules like nucleic acids, proteins, metabolites, offer a unique molecular snapshot of tumor heterogeneity, yet their clinical utility can be amplified through integration with complementary liquid biopsy technologies.
Combining exosomal multi-omics with ctDNA mutation analysis or CTC enumeration may enhance diagnostic accuracy by cross-validating tumor-derived signals. For instance, exosomal PD-L1 expression could synergize with ctDNA-based tumor mutational burden to predict immunotherapy response, while exosome-derived glycan patterns might resolve ambiguities in CTC classification. Machine learning frameworks could unify these multimodal datasets to map dynamic tumor evolution, such as detecting minimal residual disease by correlating exosomal KRAS mutations with CTC phenotypic plasticity. Challenges include optimizing bioinformatics pipelines to handle cross-technology data noise and establishing standardized protocols for co-analysis. Addressing these could enable next-generation “liquid biopsy panels” that leverage exosomes’ functional cargo to decode tumor-stroma crosstalk, ultimately advancing precision oncology from early detection to real-time therapeutic monitoring devices.
Conclusion
Exosome-based liquid biopsy, as an emerging non-invasive diagnostic approach, demonstrates high sensitivity and broad application prospects in early cancer diagnosis. The collaborative optimization of traditional and novel extraction techniques has significantly improved exosome isolation efficiency, while breakthroughs in multi-omics detection technologies enable precise identification of biomarkers such as proteins and nucleic acids. Despite challenges in standardization, detection specificity, and clinical translation, multidisciplinary collaborative innovation and large-scale clinical validation may position exosome detection as a core tool for cancer early screening and dynamic monitoring, driving the innovative development of personalized diagnosis and treatment systems.
Footnotes
Author Contributions
Xuying Hao, Zihan Liu contributed equally to this work. Writing of the manuscript: Xuying Hao, Zihan Liu, Feifei Ma, Chengbin Liu, Tuo Li, Ning Wang and Jiebing Guan. Review, editing, and supervising: Ningning He, Jiao Liu, Shujuan Lu, Huijuan Song, Jianguo Li, and Kaixue Wen.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by the Natural Science Foundation of Shandong Province, China, grant number ZR2022MH011.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
