Abstract
Vicilins are major seed storage proteins and show differential binding affinities toward sugar moieties of fungal cell wall and insect gut epithelium. Hence, purpose of study is the thorough in-silico characterization of interactions between vicilin and chitin oligomer followed by fungal and insecticidal bioassays. This work covers the molecular simulation studies explaining the interactions between Pisum sativum vicilin (PsV) and chitin oligomer followed by protein bioassay against different pathogens. LC-MS/MS of purified PsV (∼50 kDa) generated residual data along high pea vicilin homology (UniProtKB ID; P13918). Predicted model (PsV) indicated the characteristic homotrimer joined through head-to-tail association and each monomer is containing a bicupin domain. PsV site map analysis showed a new site (Site 4) into which molecular docking confirmed the strong binding of chitin oligomer (GlcNAc)4. Molecular dynamics simulation data (50 ns) indicated that chitin-binding site was comprised of 8 residues (DKEDRNEN). However, aspartate and glutamate significantly contributed in the stability of ligand binding. Computational findings were further verified via significant growth inhibition of Aspergillus flavus, A. niger, and Fusarium oxysporum against PsV. Additionally, the substantial adult population of Brevicoryne brassicae was reduced and different life stages of Tribolium castaneum also showed significant mortality.
Introduction
Vicilins (150–170 kDa) are trimeric proteins where each subunit is containing two copies of cupin superfamily domain. 1 Vicilins contain low number of sulfur containing amino acids and tryptophan along high content of acidic and basic amino acids. 2 Vicilins have shown toxicity against certain fungal and insect pathogens by the virtue of their inherent ability of binding with sugar moieties that are present inside the fungal cell wall and insect gut epithelium. Similar to other sugar-binding proteins like lectins, chitin-binding proteins and chitinases, vicilins also bind to chitin moieties and inhibit fungal and insect growth.3-5 However, the real-time picture of in-vivo molecular interactions between vicilin and chitin molecules is lacking and remains poorly understood. Chitin is an unbranched polymer of amino sugar occurring in the fungal cell wall, in the exoskeleton of invertebrates, peritrophic membrane (PM) and peritrophic gel of the insect gut.6,7 PM is a semipermeable structure comprised of chitin, protein, glycoprotein, and proteoglycan conjugates. It has been suggested that chitin-binding proteins interact with the chitinous structure, which is inevitable for insect survival.8,9
Recently, the antifungal and insecticidal potential of vicilin has been reported against pathogenic fungi and stored grain insect pests. 10 In developing countries, insect-related economic losses are estimated to be $500 million to $1 billion per year. 11 Tribolium castaneum is one of the most damaging and widespread pest discovered in cereals and processed foods, 12 as well as the representative pest in flour mills. T. castaneum is found in tropical and warm climates around the world, primarily in Africa, South America, and South Asia.13,14 Similarly, the cabbage aphid, Brevicoryne brassicae L. is a destructive pest native to Europe but spread globally, 15 and attacks economically important crops including broccoli, oilseed rape, cauliflower, and black and white mustard. It also acts as a vector for several viral diseases in crucifers, including cauliflower mosaic virus and turnip mosaic virus.16,17 Synthetic chemical insecticides have been used in agricultural practices for controlling destructive pests. However, the indiscriminate application of synthetic pesticides for crop productivity and protection has been linked to carcinogenesis, fertility issues, and mutagenesis in humans.18-20 Plant-derived substances can be used to control pests due to concerns about the usage of synthetic insecticides. 21
This study provides a detailed computational analysis of interactions between vicilin and chitin oligomer ligand, aiming to reveal the putative chitin-binding sites in the modeled vicilin structure and further exploration of the complex stability up to atomic interactions between two molecules via MD simulation followed by the potential assessment of vicilin against fungal and insect pathogens.
Materials and Methods
Isolation and Purification of Pisum sativum Vicilin (PsV)
Pisum sativum seeds (10 g) were soaked and homogenized in 100 mL of phosphate buffer (100 mM, pH 7.0) and centrifuged at 10 000 ×g for 15 minutes. Supernatant was collected and filtered through Whatman filter paper (11 µm). The clear solution was subjected to 60% ammonium sulfate saturation under cold conditions and centrifuged at 3000 ×g for 5 minutes. Supernatant was collected and salts were removed by using a dialysis membrane of 3.5 kDa MWCO (Spectra/Por 3; Catalog no. 132724) in the same extraction buffer. Dialyzed sample was loaded onto anion exchanger-Hi Trap Q FF column with 100 mM phosphate buffer (pH 7.0) at a flow rate of 0.2 mL/min and eluted with NaCl gradient of 0-1 M in the same buffer. Purified vicilin was subjected to Hi-load 16/600 Superdex 200 gel filtration column (GE Healthcare) using the same buffer and a UV detector (280 nm) was used for the recording of eluent absorbance. Fractions with maximum protein quantities were pooled together and were loaded on SDS-PAGE under both reduced and non-reduced conditions to analyze protein banding patterns along the protein ladder (Thermo ScientificTM catalog number 26616). 22 Protein quantification was done by nanodrop (NanoDrop™ 2000/2000c Spectrophotometers).
LC-MS/MS Data
Coomassie-stained protein bands were excised into 2 mm3 pieces and kept in the sterile micro-centrifuged tube. The proteins were reduced by DDT (10 mM, 56°C, 30 minutes.) and cysteine residues amended with iodoacetamide (55 mM, 20 min. in the dark). The gel excised pieces were digested with trypsin (5 ng trypsin/µl, trypsin, Promega, Madison, USA). 23 After complete digestion, peptides were treated with 50% acetonitrile and 5% formic acid and then dried in a vacuum concentrator that was again solubilized in 20 μL of .1% formic acid. LC-MS/MS data were recorded with a nano-liquid chromatography system (nanoACQUITY, Water, Manchester, UK) fixed via ESI to a quadrupole-orbitrap mass spectrometer (Orbitrap QExcactive, Thermo Scientific, Bremen, Germany). LC-MS raw data were subjected to proteome Discoverer 2.0 (Thermo Scientific, Bremen, Germany). MS/MS spectra were hunted with Sequest HT against the plant SwissProt database (www.uniprot.org) and a homemade contaminant database (298 entries). The searches were made using following parameters: precursor mass tolerance was set to 10 ppm and fragment mass tolerance was set to 0.5 Da. Furthermore, two missed cleavages were allowed and a carbamidomethylation on cysteine residues as a fixed modification as well as oxidation of methionine residues as a variable modification. All identifications were validated manually and MS/MS spectra, for which no significant hit was found in spite of high data quality, were analyzed manually.
Molecular Structure Prediction
LC-MS/MS generated residual sequence was BLAST in UniProtKB (https://www.uniprot.org/blast/) 24 and showed a high identity score with vicilin of Pisum sativum. Primary sequence of Pisum sativum vicilin (UniProtKB Accession ID: P13918) was put for model prediction into Swiss-Model online server (http://swissmodel.expasy.org/). 25 The predicted structure of PsV was developed by using crystal coordinates of 7S globulin of adzuki bean (PDB ID: 2EA7) as a template. 26 The generated PDB file was converted into visual cartoon model by using PyMOL. 27 The quality of structure was validated by the Ramachandran plot by using, VERIFY3D, ERRAT and PROCHECK online server.28,29
Site Map Analysis and Molecular Docking
Site map analysis of PsV was done by using Schrodinger site map analysis tool which identified the possible binding sites. 30 Docking studies of chitin oligomer (GlcNAc)4 into identified sites were performed in glide docking panel of Schrodinger. 31 At first, chitin oligomer was prepared for docking study by performing LigPrep which was performed to desalt and make all possible tautomer states at pH 7.0 using Epik; definite chiralities were retained, and ligands were minimized using the OPLS 2005 force field. Similarly, protein is also prepared in Protein Preparation Wizard of Maestro. Protein was prepared after ensuring chemical correctness, assigning bond orders, eliminating water molecules, and adding hydrogens for pH 7.0 using Epik. Prime was used to complete missing side chains and loops, and termini were capped. Using the default constraint of 0.30 Å RMSD and the OPLS 2005 force field, a restrained minimization of the protein structure was performed to complete protein preparation. The binding site was defined around the site points generated by site map analysis, and the receptor grid was prepared around the “site 4.” Molecular docking was performed using the Glide ligand docking module in SP (Standard precision) mode, where the receptor grid defined in the receptor grid generation folder was selected for the docking of chitin oligomer, which was prepared by LigPrep. The binding conformations were examined to identify critical interactions.
Molecular Dynamics Simulation
The most stable dock protein-ligand complex was subjected to molecular dynamics (MD) simulations using Desmond program with fixed OPLS 2005 force field.32,33 The system was prepared for simulation by developing water model (SPC: Single point charge) as solvent in orthorhombic box with specific boundary conditions which determined the shape and size of box as 10 Å × 10 Å × 10 Å distance. The charge of the system was neutralized by adding Na+ and Cl− counter ions and 0.15 M NaCl concentration was set corresponding to the physiological system using the Desmond System Builder panel. The Nose–Hoover thermostat algorithm and Martyna–Tobias–Klein Barostat algorithm were used to maintain the constant temperature (300 K) and pressure 1 atm respectively during simulation.34-36 Molecular simulations of both native protein and docked complex were done for 50 ns and comparative analysis was performed to understand the structural stability of protein and protein-ligand complex, and following quality parameters; Root Mean Square Deviation (RMSD), Root Mean Square Fluctuations (RMSF), Radius of Gyration (RoG), intermolecular hydrogen bonds (H-bonds) and Solvent Accessible Surface Area (SASA), were calculated. PyMOL was used for analyzing the MD simulation results of both bounded and unbounded forms of protein.
Antifungal Activity
Three phytopathogenic fungi [Aspergillus niger (FCBP-PTF-720), Aspergillus flavus (FCBP-PTF-862), and Fusarium oxysporum (FCBP-PTF-866)] were bought from First Culture Bank of Pakistan (FCBP), University of Punjab, Lahore, Pakistan. Spores were collected from fungal cultures by adding 15 mL of pre-chilled distilled water. Fungal cultures were filtered through cheesecloth along with continuous stirring by sterile loop and the filtrate was collected into the sterile falcon tube. The collected spores were counted at 400x magnification of binocular microscope (Ernst Leitz Wetzlar GMBH, Germany) by using the hemocytometer (NeubauerHausser Bright-Line; Catalog No. 3100) and optimized at a standard concentration of 2 × 104 spores/mL. Different concentrations of PsV (30, 40, 50, 60, and 70 µg) were mixed with fungal spores and incubated at 30°C in a 200 µL 96 well microtiter plate. Fungicide (TOPSIN®) and extraction buffer were used as positive and negative controls respectively. The OD600 was recorded after 0, 24, and 48 hours. The experiment was done in triplicate and data were processed by using Microsoft Excel. Another test was performed by using three sterile discs (15 mm) placed at equal distances in Petri plates. Fungicide TOPSIN® 4.5 FL (10 µL) was used as positive control which inhibits the fungal growth and extraction buffer was used as a negative control. The remaining disc was loaded with 60 µg purified PsV to test the antifungal activity. A loop full of fungus culture was then placed in the center and incubated at 30°C. Fungal growth was observed after 48–72 hours of incubation.
Insecticidal Activity Test
The insecticidal activity of PsV was evaluated against two pests Brevicoryne brassicae L. (Cabbage aphid) and Tribolium castaneum Herbst (Red Flour Beetle). Three different concentrations of protein (1, 2 and 3 mg/mL) were prepared and added to feed of pests. All concentrations were sprayed upon fresh-cut cabbage leaves (2 × 2 cm) in 5 replicates and placed in petri dishes (90 mm diameter, 15 mm depth). Ten individuals of Brevicoryne brassicae were placed into each petri dish and allowed to feed the protein contaminated leaves. After 12 hours, dead and alive insects were separated and counted. The data were repeatedly collected up to 72 hours.
Similarly, another experiment was designed for T. castaneum with same concentrations. Each concentration was blended with 150 g of wheat flour to develop dough. 37 Dough was air-dried and ground into a fine powder. Fine flour was divided into five replicates and each replicate contained 30 g flour and one pair of T. castaneum. After 10 days, adults were removed and data were recorded weekly for life cycle attributes, that is, larvae, pupae, and F1 adults. Data were compared with the control (0.1 M phosphate buffer; pH 7.0) to observe the effectiveness against T. castaneum.
Statistical Analysis
Data of insecticidal activities were examined by using analysis of variance and means were measured by Turkey’s test (HSD) using the software Statisticx 8.1. 38
Results
Purification and Identification of PsV
Purified PsV produced a single 50 kDa band on SDS-PAGE (Figure 1) under both reduced and non-reduced conditions. LC-MS/MS spectrometry generated three random sequences which showed high homology with vicilins of Pisum sativum and Cicer arietinum (Table 1). SDS-PAGE is showing banding pattern of freshly purified PsV loaded on to gel. From right to left; Lane 1 is the standard protein ladder (Thermo Scientific™, 26 616) while Lane 2 is containing crude, Lane 3 is showing supernatant of 60% ammonium sulfate precipitation, Lane 4 is the pooled fractions of anion exchange chromatography and Lane 5 is highly purified PsV after gel filtration. Protein Identification was Performed via Blast of LC-MS/MS Generated Sequences in UniProtKB Database.

Predicted Model of PsV
Predicted PsV model showed three monomeric polypeptide chains (Chains A, B and C) associated in a head-to-tail fashion and form a homotrimeric structure (Figure 2). Predicted model validity was checked via VERIFY3D analysis which showed 83.3% of the residues had an average 3D-1D value greater than 0.2 (S1 Figure) and ERRAT score of 88.97% (S2 Figure). Ramachandran plot showed 87.3% residues in most favored region, 10.5% in allowed region, 1.2% in generously allowed region and 0.9% in the disallowed region (S3 Figure). Predicted 3D ribbon model of PsV which is comprised of three chains (A, B, and C) in head-tail association via hydrogen bonding.
Site Map Analysis and Molecular Docking
Site Map Analysis of Pisum sativum Vicilin (PsV) Model.

aDruggability Score.

PsV Site map analysis is showing five probable sites for chitin oligomer binding. Stronger site 4 is located at front surface of two monomer of PsV.
Molecular Dynamic Simulation
Protein-ligand complex (PsV-Chitin oligomer) stability was evaluated based on RMSD fluctuations during MD simulation. The RMSD values of protein and ligand have remained in optimal range (3.0 Å) which confirmed that the complex was stable during simulation. RMSD initially showed higher fluctuation values but after attaining the equilibrium, the fluctuation of protein and ligand was kept constant up to 0.3 and 0.8 Å, respectively (Figure 4) and such lower values of RMSD are indicating the structural stability of the complex during simulation experiment. It was noticed that most of the interacting residues of PsV were residing in β-strands and have shown low RMSF values (S5 Figure). Chitin oligomer rotation was evaluated by radius of gyration (rGyr) which ranged between 6.50 and 6.60 Å. Other ligand properties like MolSA, SASA, and PSA values ranged from 650 to 665 Å, 450 to 600 Å2, and 500 to 550 Å2, respectively. Moreover, all of these values ensured the stability of ligand during simulation studies (S6 Figure). PsV trajectory frame was produced during simulations which indicated the residues which developed more than one contact with chitin oligomer during time period of 50 ns (S7 Figure). Additionally, MD simulations clearly indicated the types of atomic interactions between two molecules (Figure 5A) parallel to all those specific PsV residues which have shown significant strong interactions (>30% of the time trajectory) with corresponding atoms of chitin oligomer (Figure 5B). Time-dependent protein–ligand root mean square deviation plot (Å) is exhibiting the fluctuations of the chitin oligomer in PsV binding pocket. Molecular dynamics simulation is generating very important information about interacting atoms of 2 molecules. (A) Histogram is showing the strong proportion of water bridges followed by H-bonds of corresponding PsV residues. (B) Protein-ligand schematic diagram is highlighting all those specific residues which have consumed more than 30% of the simulation time.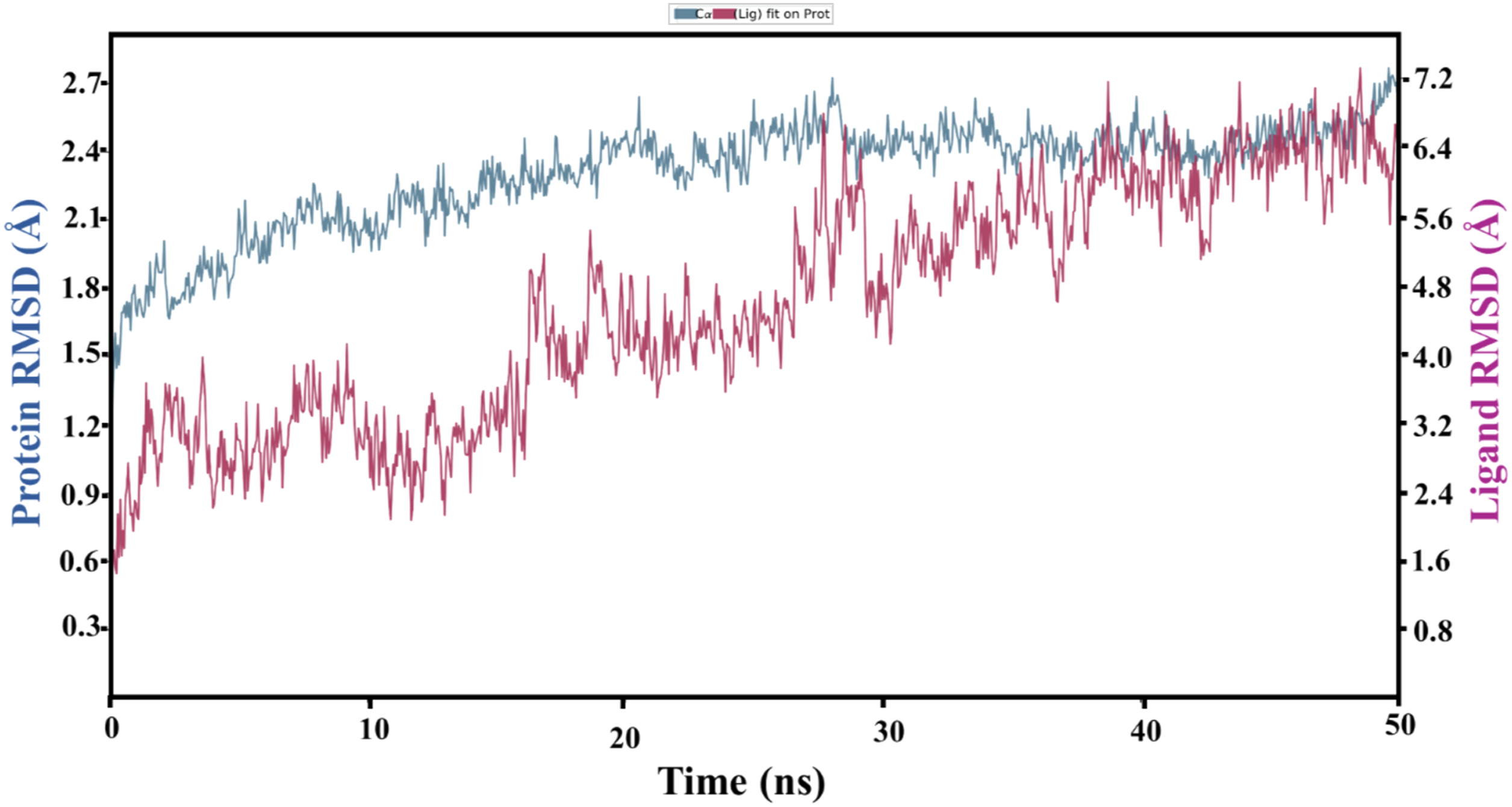

Antifungal Activity of PsV
PsV showed strong antifungal activity against A. flavus, A. niger, and F. oxysporum as shown in Figure 6. PsV concentration of 60 μg produced 50% reduction in mycelial growth after 24 hours of incubation but with no significant improvement of inhibition at 70 μg. The disc diffusion assay also validates the data showing the same growth inhibition pattern at 60 μg after 48 hours of incubation. These results are confirming that PsV has a strong potency against phytopathogenic fungi. Antifungal activity of PsV against three phytopathogenic fungal strains (A. flavus, A. niger, and F. oxysporum). Time course study has indicated 60 μg concentration inhibited 50% conidial germination after 24 hours. 100 mM phosphate buffer produced maximum conidial germination of all tested fungi. Similarly, 60 μg soaked disc 3 inhibited fungal growth after 48 hours of inoculation on petri dishes.
Insecticidal Activity
Efficacy of PsV Against Brevicoryne brassicae
Efficacy of PsV was determined against Brevicoryne brassicae (Cabbage aphid). All three concentrations (1, 2, and 3 mg/mL) produced significantly high aphid toxicity as compared to control (Figure 7). Between treatments, it was observed that 3 mg/mL produced maximum insect mortality but it was statistically non-significant as compared to 2 mg/mL. Insect mortality was observed high up to 48 hours at 2 mg/mL and after which protein effect was found non-significant between treatments. Mortality graph is showing time based PsV strong insecticidal activity against Brevicoryne brassicae (Cabbage aphid).
Efficacy of PsV Against Tribolium castaneum
Similarly, 3 mg PsV showed a significant population reduction of T. castaneum larvae (30.2 ± 1.5). Moreover, pupae population was reduced to 18 ± 2 at 3 mg, 53.6 ± 1 at 2 mg, and 58.2 ± 1.6 at 1 mg treatments. Maximum pupae population was observed in the control set of the experiment (219 ± 3). 3 mg concentration of PsV showed significant population decrease of adults (12.2 ± 2.4). Highest mean population of T. castaneum was observed in control at all stages of the life cycle (Figure 8). Mean population data is showing the effect of PsV on different life stages of T. castaneum. Mean population ±SE was significantly (P < .001) reduced at 3.0 mg dose against larvae, pupae and adults in comparison to control set.
Discussion
Chitin oligomers have already been reported as difficult candidates to characterize for their in-vivo binding affinities with different proteins. Hence, a potent alternative is the use of computational analysis for the development of in-silico studies mimicking the real time interactions. Previous reports have indicated the efficacy of vicilins against different pathogens which generated a lot of interest to seek the real time interactions between vicilin and chitin molecules in order to have sequential glimpses of in-vivo scenarios. Predicted model of PsV showed three monomers that are assembled by non-covalent interactions in a triangle-like arrangement with head-to-tail association. 26 VERIFY3D analysis and ERRAT score showed satisfactory and high quality of the predicted model. 39 Our site map analysis finds a new chitin-binding pocket in comparison to previously reported ones. Schrodinger site map analysis tool located site 4 at the junction of two monomers (chain A and chain C) and is residing at the front surface of the triangle. Interestingly, the three GlcNAc monomers were lying in relatively deep pocket between chains A & C stabilized mainly via hydrogen bonding while the fourth one is hanging inside the cavity of the triangle and is stabilized via hydrophobic interactions contributed by the chain A residues as mentioned in Figure S3. Some previous reports have indicated the chitin-binding pockets at different positions, for example, Miranda et al., reported the location of chitin-binding pocket at α-helix 8 comprising of residues between Arg-208 to Lys-216 (REQIRELMK). However, they have mentioned the more authenticity of binding domain for monomeric form as compared to homotrimeric configuration of cowpea vicilin. 40 Similarly, Rocha et al., reported the presence of three potential sites located at vertices of three chains on the interface of head-to-tail configuration inside trimeric form of cowpea vicilin in their in-silico studies. 41 Interestingly, as comparative studies, we have observed that their indicated binding pockets are also overlapping with our reported low probable sites (5 in number, Figure 3). However, our studies did not support them much in comparison to site 4 due to their low D-score (Table 2). Moreover, they indicated that pockets are residing at vertices of three chains which is again in different location than our highlighted pocket 4.
Apart from site location contradiction, it is interesting to note that common residues were found for the architectural makeup of sites. It was observed that charged and polar residues made major contribution in stabilizing the interactions between two molecules as previously reported. 41 PsV binding site is comprised of eight residues (DKEDRNEN) where negatively charged Glu (2) and Asp (2) contributed mainly in the development of stable molecular interactions similar to homotrimeric cowpea vicilin that share the same residual distribution. 41 Another report described cowpea vicilin monomer where binding site is comprised of nine residues (REQIRELMK) and interestingly they have also indicated the main molecular interactions contributed by the negatively charged residues. 40 Some other reports have also indicated that protein-glucose interactions of binding sites are comprised of a high propensity of negatively charged glutamate and aspartate along with polar and aromatic residues. 42 It is important to note that all reports are highlighting the importance of negatively charged residues for stabilizing molecular interactions between vicilins/proteins and sugars; especially like chitins which are cationic polysaccharides and even nonacidic sugars. 43 Moreover, we are also reporting the zero contribution of tryptophan as there is no such residue present inside the primary sequence of PsV and tyrosine showed little interaction with chitin oligomer up to 30 ns and after that, it does not play any further role (Figure S7) and these findings are in line with previous reports.40,41
In-silico generated molecular interaction data between PsV and chitin oligomer were further experimentally verified where vicilin showed strong efficacy against different pathogenic fungal and insect species (Figures 6-8). It has been reported that vicilin binds with the fungal cell wall and disturbs the conidia germination. 44 In another report, vicilin binds to the fungal cell wall and such binding interferes the plasma membrane which leads to inhibition of H+ pumping and causes cell death due to the acidity inside the cell. 44 Vicilin binding to glycol-conjugates of membrane surface containing GlcNAc residues, led to the death of yeast cells. 44 Many plant vicilins from Fabaceae have been reported as antifungal agents, for example, Vigna unguiculata, Vigna aconitifolia, Vigna radiata, and Glycine max.10,45,46 Vicilin isolated from Vigna unguiculata showed growth inhibition against Saccharomyces cerevisiae, Fusarium oxysporum, Fusarium solani, Ustilago madis, and Neurospora crassa. Similarly, another vicilin from cheese weed inhibited the growth of Fusarium graminearum and Phytophthora infestans. 47 Immunocytochemical assay showed the interaction of vicilin with chitin-like components that leads to fungal growth restriction. Similarly, in many reports, insecticidal activity of plant vicilins was also indicated where the immunodetection and fluorescence localization techniques were used to confirm the binding affinity between vicilin isolated from Erythrina velutina seeds and chitin structures present in Ceratitis capitata digestive system. The vicilin binding to chitin moieties of peritrophic membrane of gut epithelial cells caused deleterious effects in C. capitata digestive tract. 48 On the other hand, some authors have shown that cowpea vicilins cause detrimental effects on Callosobruchus maculatus larvae and these vicilin variants are not digested by the bruchid midgut proteinases. 39 Vicilins of various plants, such as Albizia lebbeck showed chitin-binding activity. 49 Another report has shown the vicilin binding to peritrophic membrane of Diatraea saccharalis and Tenebrio molitors, leading to inhibitory larval development.8,9 It has been reported that the toxic effect of vicilin ingestion is chronic rather than acute and midgut physiology disturbs due to vicilin vesicle trafficking which minimized the larval development and leading to death. 50 The investigation and characterization of such kind of molecules may help toward the control of pathogens and stored grains pests.
Conclusions
A combined study including both in-silico characterization and in-vivo functional analysis was performed for P. sativum seed vicilins. In-silico studies reported a new chitin-binding domain of PsV via site map analysis which was further supported by Docking and MD simulations. Aspartate and Glutamate were the main interacting vicilin residues that played significant role for the development of stable interactions between protein and chitin mimicking as the replica of real fungal cell wall and insect gut epithelial peritrophic membrane parts. Interacting residues and types of interactions involved in complex stabilization have been characterized which lead to the triggering of downstream steps for the suggested killing of pathogens. Similar significant results were found against pathogenic bacterial (gram-positive and negative) strains (data not shown, but provided in supplementary file Figure S8). Further nano-formulations of the vicilin are under process in order to enhance its potency or efficacy against different insect pests and pathogens.
Supplemental Material
sj-pdf-1-dos-10.1177_15593258221108280 – Supplemental Material for In-silico Studies Calculated a New Chitin Oligomer Binding Site Inside Vicilin: A Potent Antifungal and Insecticidal Agent
Supplemental Material, sj-pdf-1-dos-10.1177_15593258221108280 for In-silico Studies Calculated a New Chitin Oligomer Binding Site Inside Vicilin: A Potent Antifungal and Insecticidal Agent by Ahsan Saeed, Zahra Rafiq, Muhammad Imran, Qamar Saeed, Muhammad Q. Saeed, Zahid Ali, Rana K. Iqbal, Saber Hussain, Binish Khaliq, Sohaib Mehmood and Ahmed Akrem in Dose-Response
Footnotes
Author Contributions
Ahsan Saeed: Drafting and Bioinformatics. Zahra Rafiq: Wet lab. Muhammad Imran: Protein Purification. Qamar Saeed: Insecticidal assay and statistical analysis. Muhammad Q. Saeed: Microbiological Data. Zahid Ali: Review and Editing. Rana K. Iqbal: Formal analysis. Saber Hussain: Collection of Insecticidal data. Binish Khaliq: Mass Spectrometry. Sohaib Mehmood: Proof Editing. Ahmed Akrem: Group Leadter, conceptualization and Write up Improvement. Authors have read and agreed to the published version of the manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
