Abstract
Prostate inflammation is a common condition in men characterized by swelling of the prostate gland, often associated with other prostate diseases. Understanding the role of chronic inflammation in prostatic diseases is important due to the changes in prostatic cells and the persistence when undiagnosed. The evaluation and management of chronic prostatitis (CP) and chronic pelvic pain (CPP) involve specific diagnostic tests. This study aimed to evaluate the efficiency of the polymerase chain reaction (PCR) detection kits that use multiplex real-time PCR in comparison to standard microbiology sperm culture for detecting pathogens in individuals with CP and CPP. This retrospective observational study analyzed data from a database of 68 patients, aged 50.1 ± 17.8 years and treated at a secondary care urology center. PCR testing detected at least one microorganism in 63/68 samples (92.6%), while conventional culture yielded positive results in 12/68 cases (17.6%). The most detected microorganisms by PCR were the Bacteroides/Porphyromonas/Prevotella group (61.8%). Most of the samples were found to be polymicrobial, with the most common high-order combination consisting of Anaerococcus spp., Atopobium cluster, Bacteroides/Porphyromonas/Prevotella, Megasphaera/Veillonella/Dialister, and Peptostreptococcus/Parvimonas. This study concluded that PCR is more effective than traditional sperm culture in detecting organisms (p < .05), especially in identifying polymicrobial infections and fastidious microorganisms in patients with CP and CPP. PCR has higher sensitivity in detecting pathogens, including those often missed by standard culture techniques, leading to improved clinical outcomes, particularly in cases of polymicrobial infection.
Introduction
Prostatitis, or prostate inflammation, is a common condition in men characterized by swelling of the prostate gland, often associated with other prostate diseases. Prostatitis can be classified into four types: Acute Bacterial Prostatitis (Category 1), Chronic Bacterial Prostatitis (Category 2), Chronic Prostatitis or Chronic Pelvic Pain Syndrome (Category 3), and Asymptomatic Inflammatory Prostatitis (Category 4) (Nickel, 2011).
Evaluation and management of bacterial prostatitis involve specific diagnostic tests tailored to the type of prostatitis. Acute bacterial prostatitis requires urine analysis, culture, and imaging to rule out an abscess. Chronic bacterial prostatitis necessitates the 4- and/or 2-glass test for white blood cell counts and culture, with semen culture (Magri et al., 2025). Chronic prostatitis (CP) and chronic pelvic pain (CPP) involve symptom scoring, with optional tests like leukocyte analysis in ejaculate, prostate-specific antigen (PSA), ultrasound imaging, and Chlamydia trachomatis/Ureaplasma testing. Asymptomatic prostatitis may not require evaluation unless considering treatment for elevated PSA or infertility (Weidner & Anderson, 2008).
Chronic Pelvic Pain is characterized by symptoms such as pain and lower urinary tract symptoms (LUTS), making diagnosis and treatment challenging (Magri et al., 2025). The exact cause is still being investigated, but ruling out infections as the cause of prostate inflammation and identifying specific CPP symptoms can help differentiate it from microbial prostatitis (Pirola et al., 2019). Understanding the role of chronic inflammation in prostatic diseases is important due to the changes in prostatic cells (Oseni et al., 2023) and the persistence when undiagnosed (Anothaisintawee, 2011; Stamatiou et al., 2020).
This study aimed to evaluate the efficiency of the PCR Detection Kits, which use multiplex Real-Time Polymerase Chain Reaction in comparison to standard microbiology sperm culture for detecting pathogens in men with CP and CPP.
Materials and Methods
This prospective observational study analyzes data from a database of patients treated at a secondary care urology center between January 2024 and September 2025. All patients gave written consent for their clinical data to be processed and published anonymously. The study was approved by the institution’s ethics committee. Patients’ symptoms were evaluated using the International Prostate Symptom Score (IPSS), serum prostate-specific antigen (PSA) levels, urine peak flow rate (Qmax), post-micturition residual urine volume, and digital rectal examination of the prostate. Diagnostic imaging of the prostate, bladder, and seminal vesicles was conducted using trans-abdominal ultrasonography for each patient.
In all patients, microbiological analyses of the semen were performed in a single microbiology laboratory, including sperm culture and sperm PCR Detection analyses. A positive specimen was defined as a bacterial load of 103 colony-forming units/mL.
Results
A total of 68 ejaculates were analyzed using PCR Detection Kits and conventional sperm culture. The mean age of patients was 50.1 ± 17.8 years, with a median age of 51 years (IQR = 32–66). PCR detected at least one microorganism in 63 out of 68 samples (92.6%), while conventional culture yielded positive results in 12 out of 68 cases (17.6%) (Table 1). The most detected microorganisms by PCR were the Bacteroides/Porphyromonas/Prevotella group (42/68, 61.8%), Eubacterium spp. (41/68, 60.3%), Corynebacterium spp. (33/68, 48.5%), and Megasphaera/Veillonella/Dialister group (33/68, 48.5%). Streptococcus spp. (28/68, 41.2%) and Anaerococcus spp. (28/68, 41.2%) were also frequently observed. Classical sexually transmitted pathogens (Chlamydia trachomatis, Neisseria gonorrhoeae, Trichomonas vaginalis, Mycoplasma genitalium) were either absent or rare. The detailed prevalences can be found in Table 2 and Figure 1.
Baseline Characteristics of the Study Population.

Prevalence of Microorganisms Detected by Polymerase Chain Reaction Sperm Testing.


Prevalence of Microorganisms Detected by PCR.
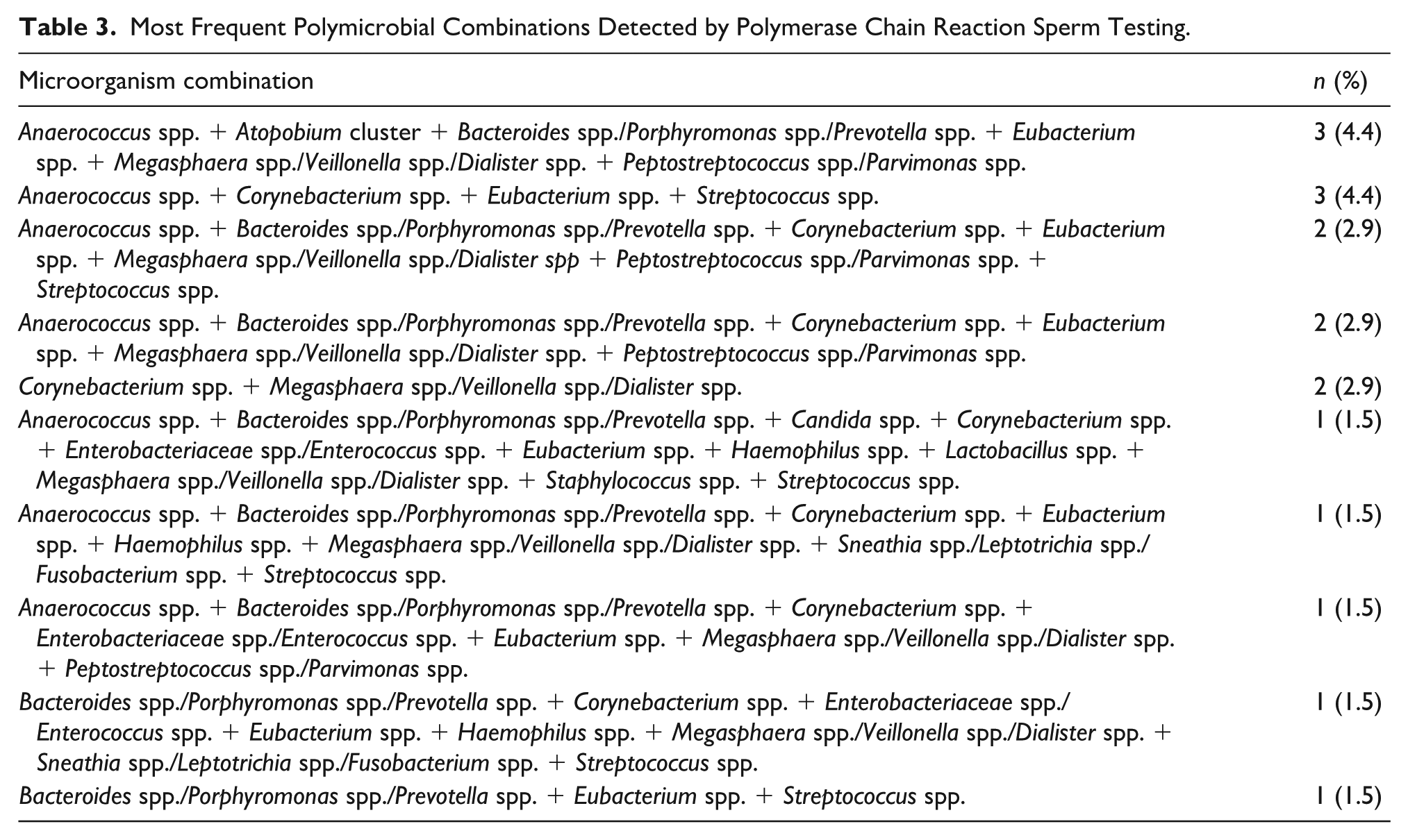
Most of the samples assessed with PCR were found to be polymicrobial, with the most common high-order combination (observed in three out of 68 samples, 4.4%) consisting of Anaerococcus spp., Atopobium cluster, Bacteroides/Porphyromonas/Prevotella, Megasphaera/Veillonella/Dialister, and Peptostreptococcus/Parvimonas. Examples of these combinations included Anaerococcus spp., Corynebacterium spp., Eubacterium spp., and Streptococcus spp. In two out of 68 samples (2.9%), other combinations were also noted, primarily comprising anaerobic flora (Table 3). The age distribution of the most commonly present microorganisms in PCR samples revealed an increased prevalence in younger and middle-aged men for the most common taxa and Eubacterium spp., as well as members of the Bacteroides/Porphyromonas/Prevotella group (median age of 45.5 and 46, years, respectively). Corynebacterium spp. and Megasphaera/Veillonella/Dialister were slightly more frequent in older subjects (median age of 56 and 60 years old), but the interquartile ranges overlapped to a significant extent, indicating that these organisms were present across the entire age range.
Most Frequent Polymicrobial Combinations Detected by Polymerase Chain Reaction Sperm Testing.
Univariate analysis of PCR profiles and sperm cultures showed a strong association between culture positivity and Enterobacteriaceae/Enterococcus spp. (50.0% of culture-positive vs. 14.3% of culture-negative samples; OR = 6.0, p = .012) (Figure 2). Pseudomonas/Ralstonia/Burkholderia spp. and Candida spp. were also associated with positive culture, but due to small case numbers, the estimates had wide confidence intervals (Table 4).

Association of the PCR Sample with Sperm Culture Positivity of Enterobacteriaceae/Enterococcus spp.
Univariate Analysis Showing the Association Between Microorganisms Detected by PCR and Sperm Culture Findings.

Discussion
Urologists globally have varying approaches to diagnosing chronic prostatitis (CP), often using less optimal methods due to a lack of consensus in the literature and practical challenges, and further research is needed to develop standardized diagnostic guidelines for CP (Stamatiou et al., 2020).
Molecular techniques such as polymerase chain reaction (PCR)-based Nucleic Acid Amplification Tests (NAATs) are replacing culture-based methods for identifying uropathogens (Leech et al., 2025). The PCR Detection Kits use multiplex Real-Time Polymerase Chain Reaction (PCR) to detect total bacterial DNA and specific pathogens in men’s urogenital tract. The threshold for a positive specimen was a bacterial load of 103 colony-forming units/mL based on the diagnostic criteria established by Naber and the Lomefloxacin Prostatitis Study Group (Naber, 2002) and other research groups (Pagliuca et al., 2021). These kits provide comprehensive diagnostics for genitourinary diseases in men, including Neisseria gonorrhoeae (GC), Chlamydia trachomatis (CT), Trichomonas vaginalis, and Mycoplasma genitalium. They also assess microbiota composition in the genitourinary tract and can be performed on various urogenital samples, such as urine, semen, and expressed prostate secretions (Elia et al., 2024).
The results of this study showed mostly polymicrobial presence, each dominated by a specific microorganism group, thus forming 10 microbiota clusters: Anaerococcus spp. (7/10), Bacteroides spp./Porphyromonas spp./Prevotella spp. (2/10), and Corynebacterium spp. (1/10). In the clusters led by Anaerococcus spp. (strictly anaerobes), the next bacteria in line were obligate anaerobes, which brings to light that anaerobes and obligatory anaerobes are most frequently present in patients with CP. These results are similar to the findings of Voroshilina et al., that the cluster led by obligate anaerobes was most frequent and characterized by the larger species diversity (Voroshilina et al., 2020).
Our study was conducted due to the need for research directly comparing PCR to semen culture in diagnosing CP and CPP, particularly in patients at higher risk of clinical progression and complications. Moreover, since the precise reasons for CPP remain unclear, it is essential to continue researching to reveal the mechanisms behind its onset (Hua et al., 2005). So far, the main pathogens linked to bacterial prostatitis and diagnosed by traditional microbiologic urine and semen cultures were Escherichia coli and Enterococcus spp., Pseudomonas spp., Proteus mirabilis, Klebsiella spp., and Serratia spp. (Sfanos & De Marzo, 2012). Historically, E. coli has been recognized as the primary cause of bacterial prostatitis, with a noticeable rise in cases of enterococcal prostatitis, as reported by other researchers (Cai et al., 2011; Magri et al., 2011). Given this trend, it is crucial to accurately identify bacterial infections in the prostate caused by Enterococci for effective patient treatment and public health measures.
The results of our study showed that Anaerococci, as strict anaerobes and obligate anaerobes (Bacteroides/Porphyromonas/Prevotella), are the most prevalent microorganisms detected by PCR in over 90% of samples that were not detected in traditional sperm cultures. If our study were based only on microbial sperm culture, the enterococcal infection would have been most prevalent as well. However, our results based on PCR testing align with research showing that PCR is more sensitive than conventional culture and sensitivity testing, as it can detect polymicrobial infections and fastidious or slow-growing organisms that are often overlooked by traditional methods (Kardjadj et al., 2025). Eventually, the results supported previous studies (Kapoor et al., 2024; Wagenlehner et al., 2020), indicating that traditional microbiologic culture techniques have limitations in identifying fastidious organisms and all causative organisms in polymicrobial infections, which can reduce the clinical effectiveness of sperm culture for prostate infections caused by a wider range of pathogens with increased antimicrobial resistance.
The interpretation complexity arises from detecting multiple organisms through sensitive PCR, making clinical interpretation challenging (Upadhyay et al., 2025). It is crucial to closely correlate the detected bacteria with the patient’s symptoms and clinical context, as some may be harmless or non-pathogenic. The study is limited by the lack of PCR testing in standard diagnostic protocols for chronic prostatitis and chronic pelvic pain in men, as well as the limited availability of the method in accredited diagnostic laboratories.
Conclusion
Based on statistical analysis of this research, polymerase chain reaction (PCR) testing is significantly more effective than traditional sperm culture testing (p < .05), especially in identifying polymicrobial infections and fastidious organisms in patients with bacterial chronic prostatitis and CPP. This improved pathogen detection leads to better clinical outcomes, particularly in cases of polymicrobial infection where incomplete detection by traditional sperm culture is associated with higher rates of clinical failure.
Footnotes
Acknowledgements
None.
Author Contributions
MSG designed the work and article draft; OT obtained the patient’s data; ST revised it critically for important intellectual content; SA analyzed and interpreted the data. All authors approved the publication of the version.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
