Abstract
The ubiquitin-proteasome system (UPS) is intricately linked to the growth and metastasis of numerous types of cancer. However, current analysis on the function and impact of UPS in bladder cancer (BLCA) is still sparse. First, univariate Cox regression analysis was applied to screen survival-associated UPS-related genes (UPSGs). Then, based on the expression of survival-related UPSGs, BLCA patients were divided into three clusters through unsupervised cluster analysis, and the overall survival rate of Cluster 1 and Cluster 3 was higher than that of Cluster 2, but there was no survival difference between Cluster 1 and Cluster 3. Therefore, Cluster 1 + Cluster 3 (Cluster A) and Cluster 2 (Cluster B) were selected for subsequent analysis. The differentially expressed genes of the two clusters were predominantly enriched in signaling pathways such as the Calcium signaling pathway and Cytokine-cytokine receptor interaction. An immune microenvironment analysis revealed that Cluster B patients were characterized by highly infiltrated immunosuppressive cells (M2 macrophages and Tregs) as well as high expression of immune checkpoint genes (such as BTLA and CTLA4). Furthermore, Cluster B exhibited a higher TIDE score. Somatic mutations demonstrated that the mutation rate of Cluster B was higher than that of Cluster A. In addition, candidate drugs for two clusters of patients were predicted, with Lapatinib, Doramapimod, and SCH772984 may be potential drugs for Cluster A patients. Luminespib, Staurosporine, and Dasatinib may be more suitable for Cluster B patients. The study provides a reference for guiding the clinical treatment of BLCA patients.
Introduction
Bladder cancer (BLCA) ranks among the 10 most prevalent cancers in the world. According to statistics, in 2022, there will be about 613,000 new BLCA cases and 220,000 deaths worldwide (Bray et al., 2024). BLCA is a heterogeneous disease with two subtypes: non-muscle-invasive bladder cancer (NMIBC) and muscle-invasive bladder cancer (MIBC), where NMIBC accounts for approximately 75% of BLCA cases (Richters et al., 2020). Despite advancements in radical cystectomy and platinum-based neoadjuvant therapy, the prognosis for BLCA patients remains suboptimal, with higher rates of recurrence and metastasis (Wang et al., 2022). The recurrence rate of NMIBC patients within 5 years after surgery exceeds 50%, and 10% to 30% of patients may progress to MIBC (Kamat et al., 2016). MIBC, due to strong invasiveness, is likely to invade blood vessels and lymph nodes, leading to distant metastasis and metastatic BLCA. 5-year survival rate is relatively low, ranging from 5-36% (Lenis et al., 2020). At present, the standard diagnostic method used clinically is invasive surgery, such as cystoscopy and biopsy (Varchulova Novakova et al., 2023). However, some histopathological patterns may exhibit a deceptive appearance (S. Chen et al., 2021). Recently, more and more scholars have found that molecular markers are conducive to the early diagnosis of BLCA patients, and can stratify patients for risk, thereby improving patient prognosis (Varchulova Novakova et al., 2023). Clearly, there is an urgent need to better understand the molecular basis of BLCA in order to develop effective treatments for BLCA.
The ubiquitin-proteasome system (UPS) plays a central part in controlling protein homeostasis, responsible for degrading short-lived, misfolded, as well as non-essential proteins (Colberg et al., 2020). UPS is an intricate system that includes E1, E2, and E3 ubiquitinating enzymes, deubiquitinating enzymes, and 26S proteasome (Park et al., 2020). UPS is implicated in a multitude of biological processes, including cell apoptosis, cell cycle regulation, DNA damage repair, and chromatin remodeling, and it participates in cancer-related cellular signaling (Cetin et al., 2021; Dagar et al., 2023). For example, Wnt signaling is instrumental for cell survival, cell cycle, proliferation, and metastasis in endometrial cancer. UCH37 can activate Wnt signaling and affect the expression of target genes (c-Myc, cyclin D1, and β—catenin), thereby facilitating tumor growth (D. Liu et al., 2020). an intimate connection between UPS dysfunction and the development and progression of malignant tumors has been revealed (Yerlikaya et al., 2021). Z. Wang et al. (2021) asserted that USP7 (Ubiquitin specific processing protein 7) is overexpressed in gastric cancer tumors, and expression level is positively linked with PD-L1 expression, suggesting that USP7 may help stabilize PD-L1 protein levels and enhance tumor immune escape. Furthermore, ubiquitin-specific protease 2 (USP2) is greatly upregulated in BLCA tissues and cells. By stabilizing EZH2, USP2 silences the expression of SOX1, thereby boosting the growth, migration, and invasion of BLCA cells (F. Xu et al., 2024). Although the function of UPS in various cancers has been discovered, further research and exploration are necessary to probe into the function and clinical significance of UPS-related genes (UPSGs) in BLCA.
Therefore, this study attempted to obtain BLCA gene expression data through TCGA database, obtained UPSGs related to BLCA survival based on univariate Cox regression analysis, and then divided the TCGA-BLCA cohort clustering into three clusters. Then, functional enrichment, tumor microenvironment characteristics and mutation differences among subtypes were analyzed. In addition, we predicted small molecule drugs. Therefore, our study aimed to provide better insights into disease mechanisms by identifying different subtypes of BLCA and analyzing their underlying molecular mechanisms. In addition, the prediction of small molecule drugs provides new ideas for the development of effective treatments.
Materials and Methods
Data Acquisition
The gene expression profile and clinical data of BLCA were retrieved and downloaded from the TCGA (“Genomic Data Commons Data Portal, n.d.”), with 431 samples (412 tumor samples; 19 normal samples).
UPSGs were obtained from the literature, including 837 genes (Supplemental Table S1) (J. Wang et al., 2021).
Cluster Analysis of TCGA-BLCA Dataset Based on UPSGs
Based on the TCGA-BLCA cohort, we screened UPSGs associated with BLCA survival through univariate Cox regression analysis. Subsequently, based on survival-related UPSGs, the TCGA-BLCA queue was subjected to cluster analysis using the “ConsensusClusterPlus” R package (D. Xu et al., 2023), with 1,000 repetitions needed to ensure classification stability. The TCGA-BLCA queue was ultimately clustered into three clusters. The survival disparities between the three clusters were analyzed by employing the “survminer” R package (Shi et al., 2023). Cluster 1 and Cluster 3 were merged as Cluster A because of insignificant survival differences. Cluster 2 was renamed as Cluster B.
Enrichment of Functional Pathways
The “edgeR” package (P. Zhao et al., 2023) was employed to analyze the differential expression in Cluster A and Cluster B (FDR <0.05 and |log FC| >1), with differentially expressed genes (DEGs) yielded. Then, the “clusterProfiler” and “enrichplot” R package (Y. Zhao et al., 2022) were utilized in the Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) analyses of DEGs.
Immune Landscape
Based on TCGA-BLCA, an immune-related algorithm was employed to analyze the immune landscapes between Cluster A and Cluster B. The immune cell activity of each sample was evaluated by using the quantiseq algorithm in the “IOBR” R package (R. Liu et al., 2023). Moreover, expression levels of human leukocyte antigen (HLA)-related genes and immune checkpoint-related genes were assessed within various clusters.
As is well known to all, higher tumor immune dysfunction and exclusion (TIDE) scores have a bearing on poorer immune checkpoint blockade therapy and shorter survival (Qiu et al., 2021). TIDE scores from different clusters of BLCA patients were used to analyze differences in immunotherapy response (“Tumor Immune Dysfunction and Exclusion, n.d.”).
Gene Mutation
Based on TCGA somatic mutation data, gene mutation analysis was undertaken by using the “maftools” R package (Mayakonda et al., 2018). An analysis was conducted on the mutations in Cluster A and Cluster B, followed by the creation of waterfall plots showcasing the top 20 mutated genes.
Small Molecular Drugs
According to the Genomics of Drug Sensitivity in Cancer (GDSC) database, which included IC50 values and expression profiles of various drugs for each sample, the “oncoCredit” R package (Maeser et al., 2021) was utilized to input TCGA-BLCA data and predict the drug sensitivity values of each sample. Further evaluation of the potential small molecular drugs in patients with Cluster A and Cluster B was processed. Wilcox test assessed the difference in drug sensitivity between the two clusters.
Results
Cluster Analysis Based on UPSGs
Through univariate Cox regression analysis on 837 UPSGs, 123 UPSGs associated with BLCA survival were generated (Supplemental Table S2). Subsequently, the TCGA-BLCA samples were subjected to clustering analysis based on the UPSGs, resulting in three clusters: Cluster 1 (n = 138), Cluster 2 (n = 151), and Cluster 3 (n = 71) (Figure 1A–C). Kaplan–Meier (K-M) survival curve analysis manifested significant differences in survival between Cluster 2 and the other two clusters (Clusters 1 and 3), with Cluster 2 having an overall survival rate significantly lower than those of Clusters 1 and 3 (p < .05). There was no survival difference between Cluster 1 and Cluster 3 (Figure 1D and E). Therefore, we conducted further research using Cluster 1 + Cluster 3 (Cluster A) and Cluster 2 (Cluster B).

Consensus Clustering Analysis Based on the Expression of UPSGs in TCGA-BLCA Samples.
Biological Functional Analysis
In the TCGA-BLCA cohort, DEGs in Cluster A and Cluster B were subjected to KEGG and GO enrichment analyses. GO enrichment manifested that DEGs were mainly linked with GO entries such as Epidermis development, Humoral immune response, Peptidase inhibition activity, and Collagen-containing extracellular matrix (p < .05) (Figure 2A). KEGG analysis uncovered that these DEGs were significantly enriched in pathways such as Cytokine-cytokine receptor interaction, neuroactive ligand-receptor interaction, and calcium signaling pathway (p < .05) (Figure 2B).
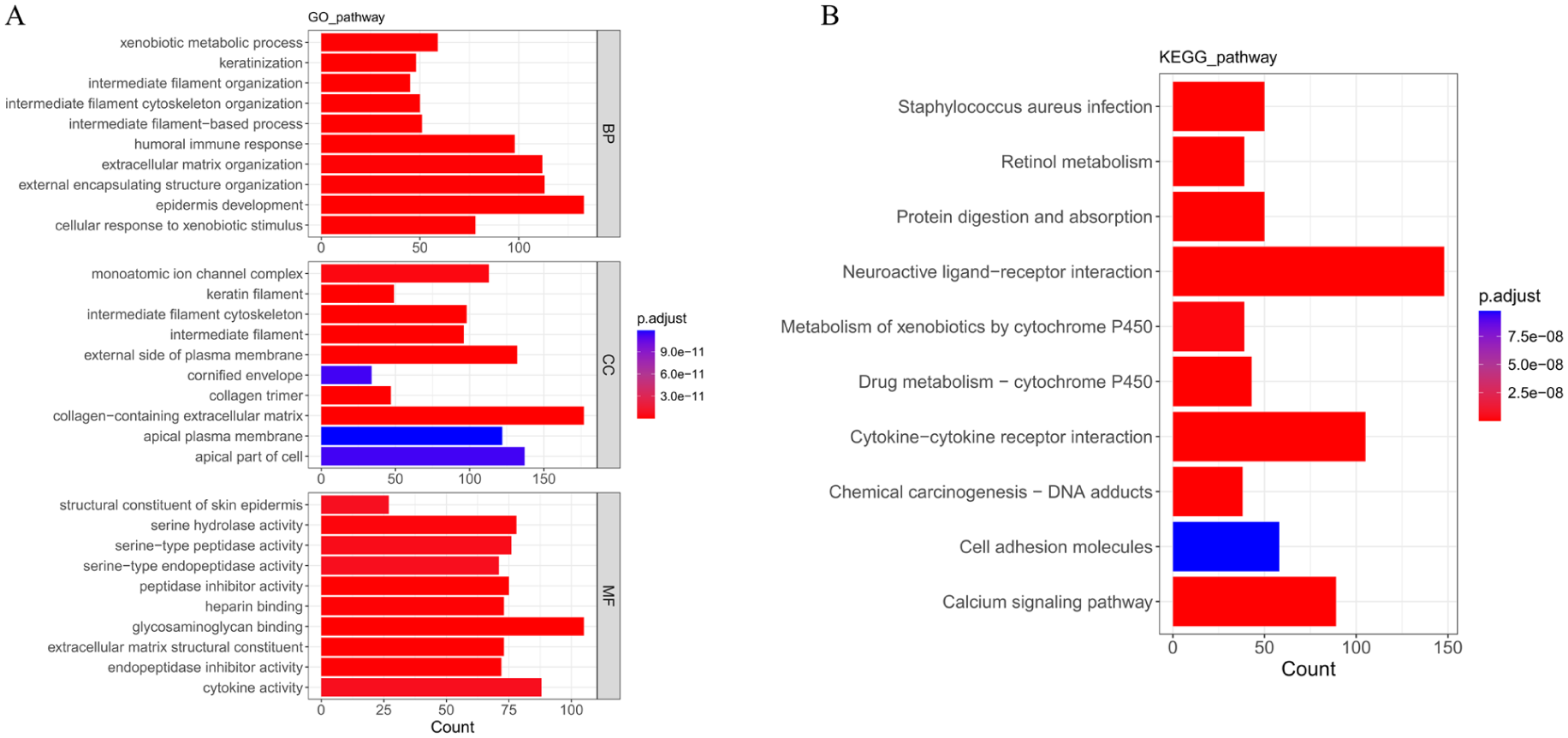
GO and KEGG Enrichment Analyses.
Tumor Microenvironment Analysis
The differences in tumor microenvironment between the two clusters were evaluated through multiple immune-related feature analyses. Significant differences were detected in immune infiltrating cells between the two clusters. Compared with Cluster A, Cluster B was mainly linked with a higher abundance of immunosuppressive cells, including M2 Macrophages and Tregs, as well as high infiltration of B cells, M1 Macrophages, CD8+T cells, and Monocytes (p < .05) (Figure 3A). In terms of HLA gene expression, the results manifested that the expression of HLA-DMB, HLA-DMA, HLA-A, HLA-DRB1, and HLA-DQA1 genes in Cluster B was significantly higher than in Cluster A (p < .05) (Figure 3B). Subsequently, the evaluation of the expression levels of immune checkpoint genes revealed that BTLA, LAG3, CTLA4, and CD27 were significantly higher in Cluster B compared to Cluster A (p < .05) (Figure 3C). In addition, according to the TIDE algorithm applied in predicting the response of BLCA patients to immunotherapy, the TIDE score of Cluster B patients was higher than that of Cluster A patients (Figure 3D). Furthermore, the number of immune-responsive samples in Cluster B patients was significantly lower than that of Cluster A (p < .001) (Figure 3E).

Tumor Microenvironment Landscape of Cluster A and Cluster B Patients.
Somatic Mutations
To dissect the differences in gene mutations between Cluster A and Cluster B, we downloaded SNV data from the TCGA database. Missense mutations were the most common type of mutation in both clusters, and the single nucleotide variation (SNV) type was mainly C>T (Figure 4A and B). Further discovery revealed the top 20 genes with the highest mutation rates in Cluster A and Cluster B. The top 5 genes with the highest mutation rates in Cluster A were TTN (43%), TP53 (40%), KDM6A (31%), MUC16 (27%), and ARID1A (25%). The top 5 genes with the highest mutation rates in Cluster B were TP53 (59%), TTN (44%), KMT2D (27%), RB1 (27%), and ARID1A (25%). The gene mutation rate of Cluster B (96%) was higher than that of Cluster A (92.68%) (Figure 4C and D).

Somatic Mutations in Cluster A and Cluster B.
Prediction and Screening of Small Molecular Drugs
A further prediction of potential small molecular drugs in Cluster A and Cluster B patients revealed that Cluster B patients were more sensitive to AZ960, Luminespib, Staurosporine, Dasatinib, IGF1R3801, and RVX.208. Cluster A patients exhibited heightened sensitivity to TAF1_5496, SB505124, Acetalax, Lapatinib, Doramapimod, and SCH772984 (Figure 5).

Sensitivity Analysis of Cluster A and Cluster B Patients to Different Drugs.
Discussion
As one of the most prevalent urinary system malignancies, BLCA causes a high incidence rate and mortality and brings a heavy burden to the medical care system (Jubber et al., 2023; van Hoogstraten et al., 2023). Despite advances in surgery and systemic treatments (chemotherapy and immunotherapy), overall survival rates for BLCA patients have not seen remarkable improvement (Chung et al., 2023; Robertson et al., 2023). Consequently, effective therapeutic measures remain elusive for BLCA patients. The abnormal expression of UPS is linked with the onset and progression of BLCA. For instance, the E3 ubiquitin ligase RNF144A can repress the occurrence and development of BLCA tumors. It can boost the apoptosis of tumor-initiating cells in an early phase, curb tumor cell proliferation at a later phase, and reduce tumor immune evasion (S. R. Ho et al., 2021). There is currently a lack of research utilizing UPSG expression to identify BLCA patients and optimize treatment methods. Consequently, this work undertook clustering analysis on the TCGA-BLCA samples based on survival-related UPSGs. The evaluation of the tumor microenvironment and small molecular drugs between different clusters was undertaken. Overall, this study aimed to develop novel treatment strategies to guide the treatment of BLCA.
We herein identified three molecular subtypes using the consensus clustering method and merged Cluster 1 and Cluster 3 which had insignificant survival differences into a Cluster A for analysis. We discovered great discrepancies between Cluster A and Cluster B in terms of TME. The infiltration of immune cells in TME reflects the immune landscape in tumors (W. J. Ho et al., 2020). Different immune cells serve diverse functional roles. Typically, B cells, M1 macrophages, CD8+ T cells, and dendritic cells (DCs) have an anti-tumor effect, while regulatory T cells (Tregs) and M2-type macrophages tend to motivate tumor progression. The roles played by monocytes and neutrophils in tumor development are more complex (Batalha et al., 2021; Y. Zhang & Zhang, 2020). Compared to Cluster A, patients in Cluster B had a lower survival rate, which was accompanied by a high infiltration of immunosuppressive cells, such as M2 macrophages and T regulatory cells, along with an overall increase in immune checkpoint expression. These observations suggested the existence of an immunosuppressive microenvironment in Cluster B. Specifically, macrophages, as the dominant immune infiltrating cells in tumors, can polarize into either inflammatory (M1) or immunosuppressive (M2) phenotypes depending on external stimuli (Cassetta et al., 2019; Chevrier et al., 2017). In TME, M2 macrophages curb T-cell proliferation by producing anti-inflammatory cytokines (IL-10) and interact with T cells’ PD-1 by expressing PD-L1, suppressing T-cell growth and inducing apoptosis, thereby leading to immunological resistance (Basak et al., 2023; W. Li et al., 2022). M2 macrophages further reinforce tumor progression through their facilitation of fatty acid oxidation, mitochondrial respiration, and angiogenesis (M. Li et al., 2023). Tregs suppress the anti-tumor immune effects of effector T cells and natural killer cells by releasing inhibitory cytokines such as IL-10, TGF-β, and IL-35 (Cuende et al., 2015; Sawant et al., 2019). Furthermore, Tregs upregulate costimulatory molecules (GITR, ICOS, and CD27), thereby enhancing their vitality and growth. Tregs also make various immune checkpoint molecules (PD-1, LAG-3, TIGIT, and VISTA) overexpressed to hinder anti-tumor effector cells (Qin et al., 2024). Patients in Cluster B exhibited weaker responses to immunotherapy, which may be attributed to the presence of an immunosuppressive microenvironment to suppress effective immune reactions. These conclusions indicated the crucial function of immune cell infiltration in tumor progression and treatment response. Further exploration of the specific functions of immune cells is essential for innovating new immunotherapy strategies.
Gene mutations are crucial for the occurrence, progression, and treatment of BLCA (X. Chen et al., 2021; Zhu et al., 2020). In this study, the mutation rates of TP53 and TTN were the highest in both Cluster A and Cluster B populations. The TP53 gene is widely recognized as an inhibitory gene for various tumors, playing a central part in modulating the DNA repair, cell differentiation, cell cycle, and apoptosis (Tornesello, 2025). Mutations in TP53 and related signaling pathways can serve as driving factors for BLCA, boosting progression and affecting cancer prognosis and treatment strategies. Cells harboring TP53 gene mutations may exhibit enhanced sensitivity to certain chemotherapeutic drugs such as mitomycin-C, doxorubicin, and gemcitabine (Wu et al., 2019). The protein encoded by the TTN gene is Titin, and mutations have also been found in certain types of cancer (Han et al., 2022; Qi & Li, 2020). In BLCA patients, upregulation of TTN-AS1 (Titin-antisense RNA1) is linked with shorter disease progression–free survival. It can also boost tumor cells to grow and invade by modulating the level of downstream factor ATF2 (Xiao et al., 2021). Genetic mutations play a pivotal part in the progression of BLCA. However, our understanding of their exact mechanisms within the disease remains limited. Future research must delve deeper into how genetic mutations influence the biological properties of BLCA, as well as how these mutations guide patient treatment selections.
In this work, we also predicted small molecular drugs that may favor BLCA patients, including Dasatinib, Lapatinib, and Luminespib. Dasatinib is a multi-target tyrosine kinase inhibitor primarily targeting SRC family kinases, BCR-ABL, and c-KIT (Heo et al., 2017; Sarno et al., 2023). Dasatinib exerts targeted anti-proliferative and pro-apoptotic effects on BD-138T PDCs, suggesting that it is a promising candidate for an antineoplastic drug in the treatment of MIBC with epidermal growth factor receptor (EGFR) gene amplification and PTEN deficiency (Chang et al., 2016). Lapatinib is a dual inhibitor that can dampen the activity of EGFR and HER-2, curbing tumor cell proliferation and survival by repressing the binding and autophosphorylation of adenosine triphosphate (Dolatabadi et al., 2024; Maeda et al., 2022). Buss et al. (2019) demonstrated that nanocapsules loaded with Lapatinib (NC-LAP) can exert anti-tumor effects on HER-positive BLCA cells by triggering cell cycle arrest and apoptosis. In mice with BLCA tumor cell lines, lapatinib enhances radiation sensitivity by reducing DNA damage-induced apoptosis in radiation-induced EGFR and HER-2 activation (Mu & Sun, 2018). Luminespib (also known as AUY922 or NVP-AUY922) is a heat shock protein 90 (Hsp90) inhibitor that has demonstrated anti-tumor activity in multiple cancers (Augello et al., 2019; Epp-Ducharme et al., 2021). In BLCA, Luminespib may potentially exhibit therapeutic efficacy in patients (L. Zhang & Peng, 2022). Therefore, future studies must delve into verifying the efficacy and safety of these medications in BLCA patients, as well as exploring their potential for combination therapy with other treatment modalities.
In conclusion, this study used the expression data of UPSGs related to the survival of BLCA to cluster BLCA samples into three clusters, which were finally analyzed as Cluster A and Cluster B. The results showed that Cluster B had a relatively high degree of immune infiltration compared with Cluster A. In addition, immunotherapy in Cluster A was more effective. At the same time, we also found that Cluster B had a higher mutation rate than Cluster A. In addition, the results of enrichment analysis and drug prediction provide a new reference direction for the subsequent investigation of BLCA mechanism and the development of therapeutic drugs. Therefore, we believe that based on UPSGs, we can effectively classify BLCA patients into subtypes and identify potential markers of response to treatment. In addition, this may also help clinicians in the precision treatment of BLCA patients. Some limitations persist in our work. First, as a retrospective analysis of the TCGA database, this work lacks a large-scale prospective study to validate our findings. Second, we need to gather more sample data to further compare the differences in immune therapy response among different patients. Finally, although we have screened potential small molecule drugs, we still need to do further in vitro and in vivo experiments to verify the efficacy of these drugs.
Supplemental Material
sj-xlsx-1-jmh-10.1177_15579883251410519 – Supplemental material for Harnessing Ubiquitin-Proteasome System-Related Genes to Identify Subtypes of Bladder Cancer and Reveal Immune Landscape
Supplemental material, sj-xlsx-1-jmh-10.1177_15579883251410519 for Harnessing Ubiquitin-Proteasome System-Related Genes to Identify Subtypes of Bladder Cancer and Reveal Immune Landscape by Guanyun Shi, Xianfei Zhou, Fan Yang and Tianxing Wang in American Journal of Men's Health
Supplemental Material
sj-xlsx-2-jmh-10.1177_15579883251410519 – Supplemental material for Harnessing Ubiquitin-Proteasome System-Related Genes to Identify Subtypes of Bladder Cancer and Reveal Immune Landscape
Supplemental material, sj-xlsx-2-jmh-10.1177_15579883251410519 for Harnessing Ubiquitin-Proteasome System-Related Genes to Identify Subtypes of Bladder Cancer and Reveal Immune Landscape by Guanyun Shi, Xianfei Zhou, Fan Yang and Tianxing Wang in American Journal of Men's Health
Footnotes
Ethical Considerations
Not applicable.
Informed Consent
Not applicable.
Author Contributions
Guanyun Shi and Xianfei Zhou conceived, designed the study, and collected the data. Fan Yang analyzed and interpreted the data. Guanyun Shi and Tianxing Wang contributed to the manuscript writing and editing. All authors read and approved the final manuscript.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The data used to support the findings of this study are available from the corresponding author upon request.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
