Abstract
Abnormal aromatase (CYP19A1) expression may participate in prostate cancer (PCa) carcinogenesis. However, the results of studies on the CYP19A1 gene polymorphisms and PCa are conflicting. This meta-analysis aimed to systematically evaluate the associations between the CYP19A1 Arg264Cys polymorphism and the (TTTA)n repeat polymorphism and PCa. Electronic databases (PubMed, EmBase, ScienceDirect, and Cochrane Library) were comprehensively searched to identify eligible studies. The strength of the association between the Arg264Cys polymorphism and PCa was assessed by pooled odds ratios (ORs) and 95% confidence intervals (95% CIs) in allelic, dominant, recessive, homozygous, and heterozygous genetic models. To analyze the impact of the (TTTA)n repeat polymorphism, we sequentially took the N-repeat allele (where N equals 7,8,10,11,12, and 13) as the minor allele and the sum of all the other alleles as the major allele. The ORs and 95% CIs were calculated in the allelic model; this analysis was performed individually for each repeat number. Pooled estimates of nine studies addressing the Arg264Cys polymorphism indicated that this polymorphism was not associated with PCa risk in the overall population or in the Caucasian or Asian subgroups. The 8-repeat allele in the (TTTA)n repeat polymorphism increased PCa risk in the overall population (OR = 1.34, 95% CI = 1.14–1.58, p = .001) and in the subgroup with population-based (PB) controls (OR = 1.41, 95% CI = 1.13–1.74, p = .002) as well as in the subgroup using capillary electrophoresis to identify this polymorphism (OR = 1.34, 95% CI = 1.09–1.65, p = .006).The meta-analysis indicated that the CYP19A1 (TTTA)n repeat polymorphism, but not the Arg264Cys polymorphism, may affect PCa risk.
Prostate cancer (PCa), the most common noncutaneous cancer in males, is the second leading cause of cancer death in the United States and represents a major public health challenge (Siegel et al., 2018). The carcinogenesis of PCa is multifactorial, and its details remain obscure. In addition to age, race, obesity, geography, smoking, certain lifestyle factors, radiation, and urinary tract infections, genetic susceptibility is also considered a candidate risk factor for PCa (Cuzick et al., 2014). Men whose first-degree relatives suffer from PCa have a higher risk of this malignancy than those without affected first-degree relatives, and the risk is even higher when these men are younger than 65 years (Kicinski et al., 2011). The prevalence and prognosis of PCa vary among racial groups, and the incidence and mortality are highest in men of African descent (Shenoy et al., 2016). These findings highlight the possible function of genetic factors in PCa. Multiple genes, including the RANSEL, MSR1, HOXB13, CHD5, and EPHB2 genes, have been reported to be associated with susceptibility to PCa (Boyd et al., 2012). Insights into genetic risk factors underlying PCa may help to identify the high-risk population.
PCa is androgen-dependent, and estrogens provide protection against PCa because of their anti-androgenic effects. In contrast to females, in whom estrogens are secreted by a central organ (ovaries), the main source of estrogens in males is the peripheral conversion of androgen precursors (Rahman et al., 2016). The enzyme aromatase (CYP19A1), mainly expressed in the gonads and peripheral tissues, including the prostate, is essential for the conversion of androgens to estrogens and represents an important therapeutic target in breast cancer (Rahman et al., 2016; Regan et al., 2011). Aberrant aromatase expression was detected in PCa tissue but not in benign prostate hyperplasia (BPH) tissue and may contribute to the recurrence of PCa (Celhay et al., 2010; Ellem et al., 2004; Gianfrilli et al., 2014). Liang and coworkers documented that CYP19A1 expression was significantly higher in castration-resistant prostate cancer (CRPC) than in primary PCa, and patients with elevated CYP19A1 expression had shorter overall survival after first hormone therapy than patients with lower expression (Liang et al., 2019). The level of CYP19A1 expression was also correlated with the Gleason score in primary PCa patients (Liang et al., 2019). These findings suggest that CYP19A1 might contribute to the onset, development, and prognosis of PCa and its response to treatment. CYP19A1 is encoded by the CYP19A1 gene mapped to chromosome 15q21.2 (Sebastian & Bulun, 2001). Although the polymorphism of the CYP19A1 gene is related to the levels of sex hormones, including testosterone, in men with and without PCa, the impact of polymorphisms of this gene on PCa remains controversial (Kanda et al., 2015; Travis et al., 2009). Several studies have focused on the associations between the Arg264Cys polymorphism (rs700519) in exon 7 and the tetranucleotide simple tandem (TTTA)n repeat polymorphism in intron 4 of the CYP19A1 gene and PCa, but the results were not unequivocal (Modugno et al., 2001; Onsory et al., 2008; Soni et al., 2012; Suzuki et al., 2003). Therefore, we performed a meta-analysis to investigate the effects of these two polymorphisms in PCa more comprehensively.
Methods
Literature Search
The electronic databases of PubMed, EmBase, ScienceDirect, and Cochrane Library were searched comprehensively and systematically by two independent investigators from inception to December 31, 2020 to identify relevant studies. The key words applied were “CYP19A1,” “aromatase,” “rs700519,” “Arg264Cys,” “R264C,” “TTTA” and “polymorphism” in combination with “prostate cancer” or “prostatic neoplasms.” Reference lists in the retrieved publications were also manually screened for potentially eligible studies.
Inclusion and Exclusion Criteria
The inclusion criteria were as follows: (1) case-control, cross-sectional, or nested case-control studies on the CYP19A1 Arg264Cys polymorphism and/or (TTTA)n repeat polymorphism in PCa; (2) studies published in English; and (3) studies with adequate allele and genotype data for both the cases and the controls to calculate the odds ratios (ORs) and 95% confidence intervals (CIs). In the case of overlapped subjects in different studies, only the most complete study was involved.
The exclusion criteria were as follows: (1) reviews, meta-analyses, or case reports; (2) case-only or family-based studies; (3) studies with intervention strategies that might modify the risk of PCa; (4) studies not providing allele/genotype frequencies or adequate data allowing their calculation; (5) studies on the polymorphisms of other genes or other prostate diseases; (6) duplicate publications; and (7) publications written in languages other than English.
Data Extraction and Quality Assessment
Two investigators independently extracted the following data from selected studies: the first author’s name, publication year, study design, ethnicity and country of the study population, source of controls, sample size, age of cases and controls, sample acquisition and genotyping method, allele and genotype distributions in each polymorphism (stated in the publication or calculated from the study data using standard formulae). Disagreements were resolved by discussing and consulting the third investigator. The Newcastle–Ottawa Scale (NOS) was used to assess the quality of case-control and nested case-control studies (Stang, 2010), and studies scoring six or more stars were considered to have a high quality (Piovezan et al., 2019).
Statistical Analysis
The meta-analysis was conducted with STATA software 12.0 (STATA Corp., College Station, TX, USA). In each study, the Hardy-Weinberg equilibrium (HWE) in the control group was tested by the Pearson chi-square test. To evaluate the strength of the association between the Arg264Cys polymorphism and PCa, pooled ORs with corresponding 95% CIs were calculated in five genetic models: allelic (T vs. C), dominant (CT + TT vs. CC), recessive (TT vs. CT + CC), homozygous (TT vs. CC), and heterozygous (CT vs. CC) models. For the (TTTA)n repeat polymorphism, the N repeat allele (with N equal to 7, 8, 10, 11, 12, and 13, sequentially) was considered the minor allele, and all the other alleles were added together as the major allele. The ORs and 95% CIs were calculated in the allelic model. The significance of ORs was determined by the Z-test, and p < .05 was considered statistically significant. Heterogeneity among studies was assessed by the I2 statistic. . When I2 was less than 50%, the fixed-effect model (the Mantel–Haenszel method) was applied. Otherwise, the random-effects model (the DerSimonian and Laird method) was used. Additionally, subgroup analyses were performed. Sensitivity analysis was conducted to assess the stability of the pooled results; individual studies were omitted one at a time, and the pooled results were recalculated. Begg’s funnel plots and Egger’s linear regression test were conducted to detect potential publication bias (Langhorne, 1998); an asymmetric funnel plot suggests a possible publication bias and p < .05 in Egger’s test indicates significant publication bias (Egger et al., 1997).
Results
Selection and Characteristics of Eligible Studies
According to the inclusion and exclusion criteria, a total of 14 studies (Fukatsu et al., 2004; Holt et al., 2013; Huang et al., 2007; Kachakova et al., 2016; Modugno et al., 2001; Onsory et al., 2008; Price et al., 2016; Sarma et al., 2010; Soni et al., 2012; Sonoda et al., 2010; Suzuki, Nakazato, Matsui, Koike, Okugi, Kashiwagi, et al., 2003; Suzuki, Nakazato, Matsui, Koike, Okugi, Ohtake, et al., 2003; Tang et al., 2011; Travis et al., 2009) were deemed eligible for this meta-analysis (Figure 1 details the study selection process). Among them, eight studies investigated only the Arg264Cys polymorphism (Fukatsu et al., 2004; Holt et al., 2013; Modugno et al., 2001; Onsory et al., 2008; Price et al., 2016; Sarma et al., 2010; Suzuki, Nakazato, Matsui, Koike, Okugi, Kashiwagi, et al., 2003; Travis et al., 2009), five studies explored only the (TTTA)n repeat polymorphism (Huang et al., 2007; Soni et al., 2012; Sonoda et al., 2010; Suzuki, Nakazato, Matsui, Koike, Okugi, Ohtake, et al., 2003; Tang et al., 2011), and one study focused on both polymorphisms (Kachakova et al., 2016). The main characteristics of these studies are listed in Tables 1 and 2. With the exception of the work of Fukastu et al. (2004) with a score of 5, all studies were considered of high quality based on NOS ≥ 6.
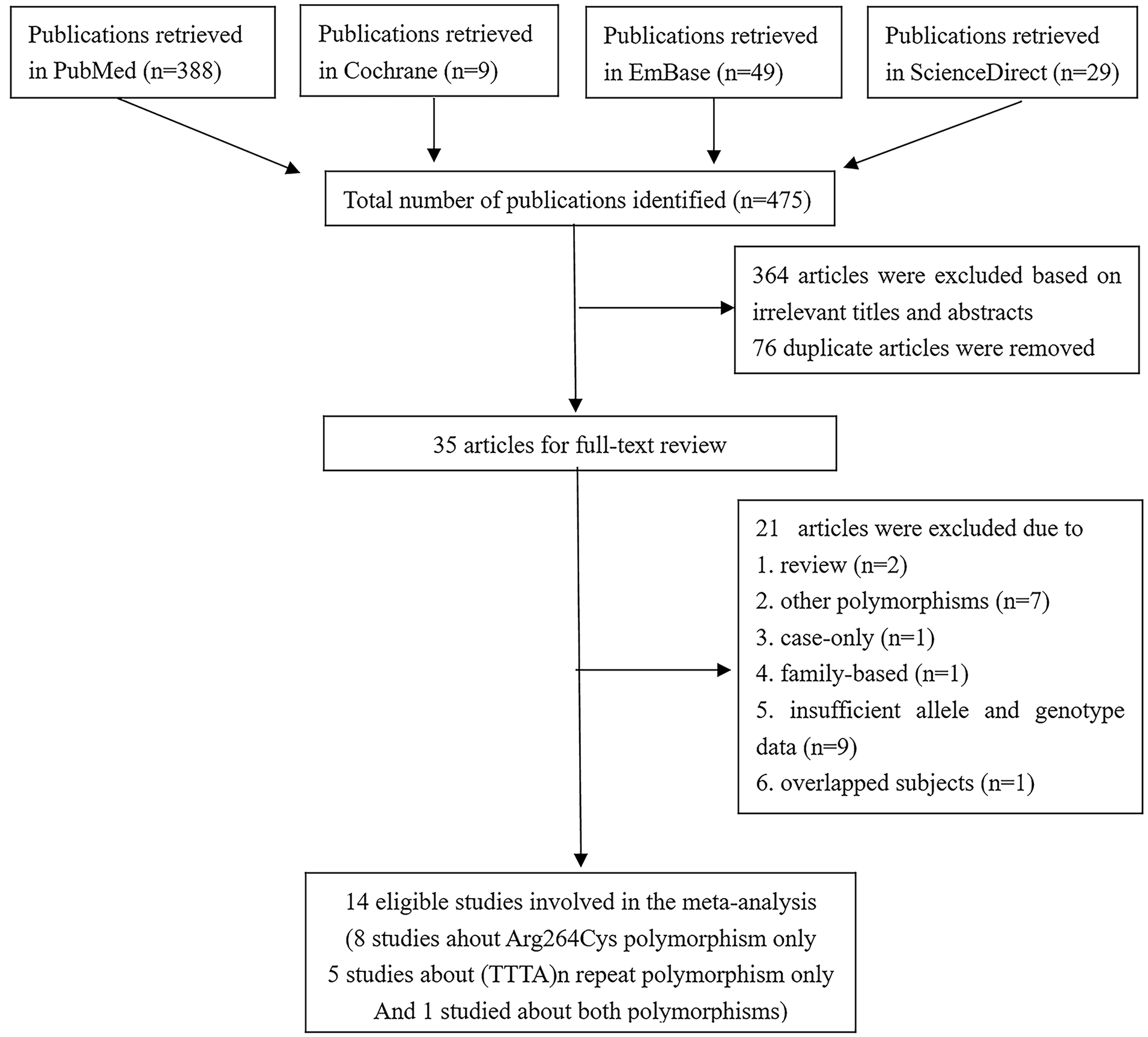
Selection of studies for this meta-analysis.
Main Characteristics of Studies About the CYP19A1 Arg264Cys Polymorphism and Prostate Cancer.

Note. PB = population based; HB = hospital based; HWE = Hardy-Weinberg equilibrium; PCR = polymerase chain reaction; RFLP = restriction-fragment length polymorphism; SSCP = single-strand conformation polymorphism; BPH = benign prostate hyperplasia; NA = not available; NOS = the Newcastle-Ottawa Scale.
A nested case-control study, in which the intervention measures might have influenced the risk of prostate cancer; only the placebo arm data were used.
Main Characteristics of Studies About the CYP19A1 (TTTA)n Repeat Polymorphism and Prostate Cancer.

Note. PB = population based; HB = hospital based; PCR = polymerase chain reaction; RFLP = restriction fragment length polymorphism; NA = not available; NOS = the Newcastle-Ottawa Scale.
A nested case-control study, in which the intervention measures might have influenced the risk of prostate cancer; only the placebo arm data were used. All other studies are case-control studies.
Association Between the CYP19A1 Arg264Cys Polymorphism and PCa Risk
Nine studies (seven case-control studies and two nested case-control studies), which recruited a total of 11,824 patients and 11,368 control subjects, addressed the relationship between the CYP19A1 Arg264Cys polymorphism and PCa (Table 1) (Fukatsu et al., 2004; Holt et al., 2013; Kachakova et al., 2016; Modugno et al., 2001; Onsory et al., 2008; Price et al., 2016; Sarma et al., 2010; Suzuki, Nakazato, Matsui, Koike, Okugi, Kashiwagi, et al., 2003; Travis et al., 2009). These studies enrolled Caucasians (four studies), Asians (two studies), Indians (one study), African-Americans (one study), and subjects of mixed ethnicities (one study). One study employed intervention measures (finasteride) that might confer PCa risk and might conceal the real situation of this polymorphism in PCa; therefore, only data in the placebo arm in this study were considered (Price et al., 2016). In two studies, genotype distributions in the control groups deviated from HWE (Onsory et al., 2008; Travis et al., 2009). Controls were hospital-based (HB) in three studies, population-based (PB) in five studies, and both HB and PB in one study. One study merely recruited cases with a family history of PCa in a first-degree relative (Suzuki, Nakazato, Matsui, Koike, Okugi, Kashiwagi, et al., 2003), one study described the ratio of familial cases in all participants (Sarma et al., 2010), and the remaining seven studies did not provide information about family history. Peripheral blood was used to detect this polymorphism in seven studies (Holt et al., 2013; Kachakova et al., 2016; Modugno et al., 2001; Price et al., 2016; Sarma et al., 2010; Suzuki, Nakazato, Matsui, Koike, Okugi, Kashiwagi, et al., 2003; Travis et al., 2009), while peripheral blood or frozen prostate tissues were used in the other two studies (Fukatsu et al., 2004; Onsory et al., 2008).
The allele and genotype frequencies in the overall population and all subgroups are summarized in Table 3. The overall pooled results and subgroup analyses are listed in Table 4. Neither the T allele nor the TT genotype was associated with PCa in the overall population, Caucasians, or Asians (Figure 2 displays a nonsignificant association between the T allele and overall PCa risk). Subgroup analyses could not be performed for African-Americans, Indians, or mixed ethnicities since each of these ethnicities was included only in one study. Subgroup analysis stratified by the source of controls was also conducted. When PB controls were considered, no evident association was found between the Arg264Cys polymorphism and PCa in any genetic model. Nevertheless, an association between the Arg264Cys polymorphism and PCa was observed under the dominant model (CT+TT vs. CC: OR = 1.35, 95% CI = 1.02–1.78, p = .04) and heterozygous model (CT vs. TT: OR = 1.40, 95% CI = 1.04–1.88, p = .03) in the HB subgroup. Subgroup analysis could not be accomplished for the PB+HB subgroup since only one study included PB+HB controls. We further conducted subgroup analyses based on the sample acquisition and genotyping method. No significant association was detected in these subgroups (Table 4).
Allele and Genotype Distribution in the CYP19A1 Arg264Cys Polymorphism of Subjects Included in the Meta-Analysis.

Note. PB = population-based; HB = hospital-based; PCR = polymerase chain reaction; RFLP = restriction fragment length polymorphism; SSCP = single-strand conformation polymorphism.
Meta-Analysis of the Association Between the CYP19A1 Arg264Cys Polymorphism and Prostate Cancer.

Note. PB = population-based; HB = hospital-based; PCR = polymerase chain reaction; RFLP = restriction fragment length polymorphism.
We couldn’d perform subgroup analyses for subgroups with only one study.

Forest plot for the association between the CYP19A1 Arg264Cys polymorphism and prostate cancer (T vs. C).
Association Between the CYP19A1 (TTTA)n Repeat Polymorphism and PCa Risk
Six studies (five case-control studies and one nested case-control study) that recruited a total of 1488 cases and 1621 controls were analyzed (Table 2) (Huang et al., 2007; Kachakova et al., 2016; Soni et al., 2012; Sonoda et al., 2010; Suzuki, Nakazato, Matsui, Koike, Okugi, Ohtake, et al., 2003; Tang et al., 2011). Peripheral blood was used in all these studies to detect the (TTTA)n repeat polymorphism. Three of these studies were conducted in Asians and one each in Caucasians, Indians and subjects of mixed ethnicities. The repeat numbers ranged from 7 to 14, but the 9-repeat allele was not detected in four studies (Huang et al., 2007; Sonoda et al., 2010; Suzuki, Nakazato, Matsui, Koike, Okugi, Ohtake, et al., 2003; Tang et al., 2011), and the 14-repeat allele was identified only in one study (Kachakova et al., 2016). Thus, the 7-, 8-, and 10–13-repeat alleles were considered in this meta-analysis. Soni and coworkers (Soni et al., 2012) did not find 10, 11, and 13 repeats either in patients or in controls, so their study could not be included in the analysis of these three alleles. For the same reason, the study of Tang (Tang et al., 2011) was not used in the analysis of the 13-repeat allele.
The pooled results are listed in Table 5. Only the 8-repeat allele was significantly associated with the risk of PCa in the overall population (OR = 1.34, 95% CI = 1.14–1.58, p = .001) (Figure 3), while all the other alleles appeared not related to the overall PCa risk. Among the six studies addressing the 8-repeat allele, one was performed in Caucasians, three in Asians, one in Indians, and one in subjects of mixed ethnicities. Therefore, the subgroup analysis only applied to Asians and yielded a nonsignificant result (OR = 1.27, 95% CI = 0.88–1.85, p = .17). In the subgroup analysis based on the source of controls, the 8-repeat allele was a risk factor for PCa in the PB subgroup (OR = 1.41, 95% CI = 1.13–1.74, p = .002). Although in the subgroup analysis stratified by the genotyping method, the 8-repeat allele increased PCa susceptibility in the subgroup of studies using capillary electrophoresis to investigate the (TTTA)n repeat polymorphism (OR = 1.34, 95% = 1.09–1.65, p = .006), the results should be treated cautiously because there were only two studies using this genotyping method.
Meta-Analysis of the Association Between the CYP19A1 (TTTA)n Repeat Polymorphism and Prostate Cancer.

Note. PB = population-based; HB = hospital-based; PCR = polymerase chain reaction; RFLP = restriction fragment length polymorphism.
We couldn’d perform subgroup analyses for subgroups with only one study.

Forest plot for the association between the 8-repeat allele in the CYP19A1 (TTTA)n repeat polymorphism and prostate cancer.
Heterogeneity and Sensitivity Analysis
Although there was considerable heterogeneity among the studies in certain subgroups in this meta-analysis (Tables 4 and 5), sensitivity analysis showed that the synthetic results were not materially altered by omitting any single study, indicating the stability of the results (Figures 4 and 5).

Sensitivity analysis of studies on the CYP19A1 Arg264Cys polymorphism and prostate cancer (T vs. C).

Sensitivity analysis of studies on the 8-repeat allele in the CYP19A1 (TTTA)n repeat polymorphism and prostate cancer.
Publication Bias
Begg’s funnel plots did not reveal any evidence of obvious asymmetry in studies on the Arg264Cys polymorphism (Figure 6), and Egger’s test with p > .05 further verified the absence of publication bias (Table 4). For the (TTTA)n repeat polymorphism, Begg’s funnel plots and Egger’s test indicated the absence of publication bias except for the 7-, 8-, and 13-repeat alleles (p < .05; Table 5).

Begg’s funnel plot for studies addressing the CYP19A1 Arg264Cys polymorphism in prostate cancer (T vs. C).
Discussion
The human CYP19A1 gene, a member of the cytochrome P450 superfamily of genes, spans approximately 123 base pairs (bp) and comprises nine translated exons (exons II–X) and one untranslated exon I spliced at the 5’ end and nine introns (Carreau & Hess, 2010; Corbin et al., 1988). Several distinct polymorphisms in the CYP19A1 gene have been studied in PCa, among which the Arg264Cys polymorphism and (TTTA)n repeat polymorphism are the most extensively investigated. Studies on these two polymorphisms in PCa did not provide consistent results, possibly due to the low statistical power of every single study or limited impact of these polymorphisms on PCa. A meta-analysis, a powerful statistical tool, may clarify the conflicting results of previous research and provide a more precise estimation of the impact of the CYP19A1 Arg264Cys and (TTTA)n repeat polymorphisms on PCa (Munafo & Flint, 2004). To the best of our knowledge, this is the first meta-analysis investigating the associations between these two polymorphisms of the CYP19A1 gene and PCa.
The C→T transition (rs700519) in exon 7 of the CYP19A1 gene leads to a single amino acid substitution of Arg by Cys at codon 264, and the T allele has been reported to enhance aromatase enzymatic activity, affecting the conversion of androgens. This modification may alter the risk of several disorders, such as breast cancer, polycystic ovary syndrome (PCOS), and endometrial cancer (Pan et al., 2016; Sun et al., 2015; Tao et al., 2007; Wang et al., 2011). The current meta-analysis failed to detect any association between the Arg264Cys polymorphism and the overall PCa risk. Given the well-established ethnic disparities in PCa incidence, a subgroup analysis based on ethnicity was conducted. No relationship was identified between the Arg264Cys polymorphism and PCa in Caucasians or Asians. Studies were also divided according to the differences between the enrolled control subjects, the sample acquisition and the genotyping method, and the CT+TT and CT genotypes were found to be associated with PCa susceptibility in the subgroup of HB controls. These results should be interpreted with caution, since the relatively low number of studies and small sample sizes in each subgroup prevent statistical interpretation with confidence and do not exclude the possibility that these correlations might be incidental. In addition, evident heterogeneity among the analyzed studies have also weakened the reliability of the results.
The (TTTA)n repeat polymorphism is located at the intron 4 of the CYP19A1 gene, and its effect on the enzymatic activity of aromatase has not been unequivocally established (Celhay et al., 2010; Hammoud et al., 2010). However, this polymorphism may affect mRNA stability or be in linkage disequilibrium with other polymorphisms, including a nearby 3-bp TCT deletion in conjunction with the 7-repeat only and a T/C change in the 3’-untranslated region in which the TT genotype is linked to 8 and longer repeats (Celhay et al., 2010; Olson et al., 2007). The repeat numbers reported in the literature range from 7 to 15, with two major peaks at 7 and 11 or 12 repeats (Fernandez et al., 2014; Hammoud et al., 2010; Xu et al., 2013). Luigi et al. discovered that a high (TTTA)n repeat genotype (10 or more repeats) might be associated with a higher serum level of estradiol (E2) in a group of Italian men older than 55 years (Luigi et al., 2004). Previous studies have demonstrated that the (TTTA)n repeat polymorphism might convert susceptibility to several diseases, including PCOS, gynecomastia, and breast cancer (Ahsan et al., 2004; Izabella et al., 2008; Miyoshi et al., 2000; Xu et al., 2013). In studies related to PCa, Sonoda reported that 11 or more repeats were associated with an increased risk of PCa (Sonoda et al., 2010), while Huang and coworkers identified a similar relationship for homozygous 7-repeats (Huang et al., 2007). Conversely, Soni reported no association between (TTTA)n polymorphism and PCa (Soni et al., 2012). These inconsistent findings might be attributed to discrepancies in ethnicity, country of origin, sample size, phenotypes, type of controls, and genotyping methods. The current meta-analysis uncovered a significant association between the 8-repeat allele and the overall risk of PCa, which was largely driven by studies utilizing PB controls and capillary electrophoresis to detect this polymorphism. On the other hand, the 7-, 10–13-repeat alleles seemed to have nothing to do with PCa risk. English language criterion in literature search, exclusion of studies without adequate amount of data, and the preferential publication of studies that achieved significance might lead to the publication bias detected by Egger’s test in studies on the 8-repeat allele, which is not unusual in meta-analyses (Elias & Joseph, 2008). However, the low heterogeneity and statistically robust findings in sensitivity analysis suggest that the obtained results are relatively reliable.
Several limitations should be taken into consideration. First, the number of eligible studies was limited and the sample sizes were restricted. The number of controls did not match the number of cases in two studies (Modugno et al., 2001; Sarma et al., 2010). These factors may affect the statistical power of the determination of the significance of associations. Second, because of the lack of access to the original data of analyzed studies, it was impossible to adjust for other PCa risk factors such as age, family history, obesity, lifestyle factors, and gene-gene or gene-environment interactions. Third, the inclusion of research published in English with enough data for calculation might have introduced publication bias. Fourth, the nonuniform selection of cases and controls across different studies may also bias the results. A similar effect might have been produced by the deviation from HWE noted in certain studies, possibly due to bias in the selection of control subjects or genotyping errors. Fifth, studies on the Arg264Cys polymorphism were mostly conducted in the Caucasian population. Studies on the (TTTA)n repeat polymorphism have mostly been conducted in Asian populations. These studies were not adequate to identify the genetic etiology of PCa, the prevalence of which varies among racial groups. Well-designed studies with larger sample sizes are warranted to verify the conclusion of the present study.
Conclusions
In summary, this meta-analysis provides evidence of the association between the 8-repeat allele in the CYP19A1 (TTTA)n repeat polymorphism and susceptibility to PCa. Conversely, the Arg264Cys polymorphism in the CYP19A1 gene appeared to have no impact on PCa risk. Further studies are necessary to fully understand the role of these two polymorphisms in PCa, and to facilitate early identification of subjects at high PCa risk.
Footnotes
Acknowledgements
Not applicable.
Authors’ Contributions
LG and YL carried out the data search, selection, and study quality assessment. LL and SS performed the statistical analysis. LG, YL, YC and JG drafted the manuscript. HN conceived the study and participated in critical revision of the manuscript. All authors have read and approved the final version of the manuscript and agreed with the order of presentation of the authors.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
