Abstract
The development of information and telecommunication technologies has given rise to new platforms for e-Health. However, some difficulties have been detected since each manufacturer implements its communication protocols and defines their data formats. A semantic incongruence is observed between platforms since no common healthcare domain vocabulary is shared between manufacturers and stakeholders. Despite the existence of standards for semantic and platform interoperability (e.g. openEHR for healthcare, Semantic Sensor Network for Internet of Medical Things platforms, and machine-to-machine standards), no approach has combined them for granting interoperability or considered the whole integration of legacy Electronic Health Record Systems currently used worldwide. Moreover, the heterogeneity in the large volume of health data generated by Internet of Medical Things platforms must be attenuated for the proper application of big data processing techniques. This article proposes the joint use of openEHR and Semantic Sensor Network semantics for the achievement of interoperability at the semantic level and use of a machine-to-machine architecture for the definition of an interoperable Internet of Medical Things platform.
Keywords
Introduction
Over the past years, the technological development has led to a continuous redefinition of daily life. Better quality services have taken advantage of information and telecommunications technologies, as the Internet, which simplifies the remote access to them. Users can pay bills, buy a product, call a taxi, and check traffic from a smartphone or a personal computer using the Web and mobile applications.
Apart from conventional services, daily life objects/devices have been connected to the Internet for remote interaction with their users and even with other objects/devices with no human intervention. Such devices, called “things,” are part of a more complex definition known as the Internet of Things (IoT), described by Minerva et al. 1 as systems composed of computing devices and applications embedded in objects that monitor/actuate over the environment interconnected by the Internet. IoT has been widely accepted and has influenced several domains, such as agriculture, the environment, security, education, and healthcare.
Particularly in the healthcare domain, IoT has been given a special definition. Manogaran et al. 2 defined the Internet of Medical Things (IoMT) (also known as healthcare IoT) as the global infrastructure consisting of medical devices and applications interconnected by the Internet, where temperature, carbon dioxide, ECG/EEG/EMG, pressure, gyroscope, blood oxygen saturation, humidity, respiration, and blood pressure sensors enable the observation and monitoring of patients’ health in a continuous manner. As a consequence, healthcare providers can offer more accurate services and improve the quality of life of patients. However, their integration into an interoperable environment has been challenging.
Recent IoT-related studies3–5 have highlighted interoperability issues, for example, each manufacturer defines its architecture, protocols, and data formats; therefore, applications must be redeveloped or adapted to interoperate with each IoT platform (IoMT platform in the healthcare environment). Desai et al. 6 named such issues “Vertical Silos” (VSs) problem in IoT. A VS is defined as an IoT platform that uses a custom data representation format, devices, and protocols and demands the development of new features for granting interoperability between different systems.
Besides the VSs problem, other issues on compatibility with current healthcare systems remain open. Healthcare systems are based on Electronic Health Records (EHR), which store patients’ clinical history and hospital administrative data, but ignore the existence of IoMT platforms and integration with them. Standards, such as OpenEHR,7,8 have been proposed for the definition of EHR models according to a unified medical vocabulary, while others, as HL7, 9 have been developed for the exchange of records between different institutions. 10
The lack of semantic integration between EHRs and IoMT platforms exerts a negative impact on the development of modern healthcare services since measurements collected by IoMT platforms may comprehend a large volume of heterogeneous data. Moreover, as observed by Dineshkumar et al., 11 physicians cannot handle such data directly and highlighted the importance of applying big data techniques for knowledge extraction. If the data are not based on a platform that promotes semantic interoperability, the application of big data algorithms may be hampered since data can be unreadable, incorrectly parsed, or misinterpreted. Therefore, semantic interoperability in IoMT is a necessary condition for big data techniques to support decision-making processes.
A platform for IoMT that provides EHR functionalities should propose solutions to the following challenges: (a) IoMT platforms must consider the semantics and the same vocabulary defined in EHRs; (b) the format of data handled in the IoMT environment may differ and must be translated to an EHR standard one; and (c) the IoMT platform should include a definition of the individual IoMT devices for identifying the data sent by them, their origin, and the way they are collected.
Our literature review identified a lack of proposals for the solution of interoperability between EHRs and IoT/IoMT. Our approach is based on the interoperability between an openEHR standard and a Semantic Web-based IoMT platform; it considers both healthcare and IoT domains in an interoperable environment and the Semantic Sensor Network (SSN) ontology to extend the IoT semantics to the openEHR context.
The joint use of such standards provides an interoperable environment with a standardized query mechanism; it follows the Semantic Web principles and uses standardized languages, as SPARQL. Moreover, a machine-to-machine (M2M) communications standard (provided by the European Telecommunications Standards Institute (ETSI) 12 ) can be useful for the definition of the platform architecture and enable communication between components of IoMT platforms.
The main contributions of this research involve (a) an interoperable IoMT platform for e-Health Applications; (b) an alignment of an openEHR-based ontology with the SSN ontology for IoT platforms; and (c) an extension of M2M ETSI standard that follows semantic technologies to grant IoMT and EHR interoperability.
The remainder of the article is organized as follows: section “Relevant standards and ontologies” addresses some standards and ontologies that play a crucial role in our proposal; section “Related work” discusses some related work; section “Alignment of SSN and OpenEHR-based ontologies” focuses on aspects related to the ontology alignment process; section “Semantic IoMT platform for e-Health” introduces our approach for a semantic IoT platform for e-Health; section “Example of a use case” describes a use case focused on respiratory screening; finally, section “Conclusion” reports the conclusions and outlines some future work.
Relevant standards and ontologies
Several organizations, such as openEHR, the Internet Engineering Task Force (IETF), the World Wide Web Consortium (W3C), and the ETSI have proposed useful standards and ontologies to achieve the objectives outlined in this article. This section presents some of these standards and ontologies on which the proposed IoMT platform is based.
OpenEHR7,8 is a standard that formalizes the modeling of EHR. It is defined by a three-level modeling approach which considers (a) reference model (RM), (b) reusable content element definitions, and (c) context-specific data set definitions.7,8 RM contains definitions of data structures, types, security aspects, and an identifier attribute that supports the indexation of each RM. The reusable content definitions formalize clinical content data points and groups in the form of archetypes written in Archetype Definition Language (ADL). Archetypes define specific concepts related to the healthcare domain and are maintained by physicians (e.g. blood pressure observation). The context-specific data set definitions comprehend forms, documents, messages, and so on, created from the combination of required elements of relevant archetypes into openEHR templates. Templates are defined as the joint use of archetypes to conform to definitions of healthcare concepts, as screening of hypertension diseases which considers blood pressure observation.
OpenEHR assumes the IT environment as a fixed model of information designed to store models of content (the knowledge models) using archetypes, templates, and user interface definitions. A change in the content models affects the information system marginally. This process results in a very short time-to-operation cycle and simplifies the handling of constant changes of clinical information representation induced by progress in medical research, legal requirements, and so on.
As shown in Figure 1, developers define a data schema based on an archetype model which considers the data types defined in the RM. The schema is used in the implementation of the EHR system for the definition of the structure of the data persistence model (EHR Repository) and storage and retrieval of archetyped data regardless of their content. For example, domain specialists or institutions can create or modify archetypes and templates, and the information system remains compatible. Moreover, openEHR enables the use of standardized terminologies, such as SNOMED, for the definition of archetypes and templates, which is an advantage in the presentation of concepts to users via input forms or when an EHR is displayed.

OpenEHR multi-level modeling and software engineering.
Apart from the modeling of clinical data, openEHR models demographic data of patients, people, organizations, among other entities. Regarding security, it promotes the sharing of open data since the demographic model is separated from the clinical model in such a way the clinical data enable no identification of the patient that generated them (section “Data published in openEHR-extended as individuals”).7,8 In informational terms, an EHR system based on openEHR consists of an EHR repository, an archetype repository, a terminology, and a demographic repository, as shown in Figure 2.

EHR and demographic separation.
A demographic repository can act as a front end to an existing repository (i.e. a hospital database with patients already registered), or in its own right (i.e. deployed by the EHR system). In both ways, its functions are standardization of demographic information structures and versioning. An EHR contains references to entities in whichever demographic repository configured for use in the environment. In this sense, an EHR taken in isolation contains few or no clue on the identity of the patient it belongs to. This approach offers several security benefits and, in particular, those related to anonymity.
FHIR (Fast Healthcare Interoperability Resources) is another standard which enables the exchange of information for the delivery of healthcare in a wide variety of settings. The specification is based on Web resources and adapts modern and widely used RESTful practices for the delivery of integrated medical care across a broad range of teams and organizations. 13 However, different reasons led us to not consider FHIR in our platform. FHIR cannot be used directly A mapping between ADL and FHIR resources would be proposed toward the use of the application programming interface (API); however, this is outside the scope of this research. We aim at sharing data openly and not specifically among healthcare institutions, in a context of smart city applications (for example). In this sense, the use of ontologies, rather than an API-based architecture, seems to be advantageous since semantic web-based solutions (e.g. based on Linked data) have been widely accepted and a large number of tools focuses on it.
On the other hand, Sensor Measurement Lists (SenML) defines media types to represent simple sensors measurements and device parameters. 14 The representations are provided in JavaScript Object Notation (JSON) and eXtensible Markup Language (XML), which shares the common SenML data model. Amrutur et al. 15 proposed its extension using semantic annotations over the sensor data to describe the data collection process, the meaning of a collected value, and the platform used.
W3C 16 defined the SSN ontology for the description of sensors, networks, observations, features, samples, properties, and procedures. It is based on a core ontology called Sensor, Observation, Sample and Actuator (SOSA), which describes the main classes, namely sensor, observation, sample and actuator, and the axioms that constrain or relate them.
Finally, Lucic et al. 17 discussed the standardization of M2M as the direct communication between devices through a mobile or fixed network. In the M2M architecture, the components below play a fundamental role, that is, the Gateway and Network Service Capabilities Providers (GSCP and NSCP, respectively) enable communication between the IoMT devices and the M2M Applications and also provide functions for IoMT device discovery, device identification, and binding between device and applications. The M2M application located in the Gateway is mostly focused on patients, data preprocessing, and their forwarding to the Network Domain. The Network Domain M2M Applications use the prepossessed data for a specific purpose, for example, knowledge extraction, statistical applications, or simply their storage on a database.
For an e-Health application, the M2M devices (similar to IoMT devices) sense the different parameters of the patients or their environments and forward the data to the final applications at the top layer of the M2M architecture through M2M Gateways that enable the integration of medical sensors to the platform.
Related work
This section addresses a literature review of architectures for IoMT platforms and studies related to semantic integration of healthcare data and clinical knowledge in IoMT.
IoMT architecture design
Several manuscripts that discuss and propose possible IoMT architectures, commonly based on multi-layering, have been published.
Irfan and Ahmad 18 observed that two to five layers had been proposed toward the abstraction of the complexities of IoMT architectures and concluded that a three-layered structure was presumably ideal for a logical fragmentation of IoMT architectures composed of Things, Intermediate, and Integrated Application layers. The Things layer is composed of heterogeneous IoMT objects that communicate through various communication protocols and networks. The Intermediate layer is represented by a Middleware or Gateway and is supposed to handle the IoMT devices and process data at a local stage; it is implemented by Multi-agent, Service Oriented, or Publish-Subscribe architectures and enables fog computing technologies. The Integrated Application layer stores huge amounts of data and processes them through several applications for the control of medical processes. Some of the proposals that implement such a three-layer pattern are discussed below.
Rahmani et al. 19 proposed an IoT-based health monitoring system composed of the following layers: Smart Devices, which include the Sensor Networks and IoMT devices; Edge/Fog, which contains the Gateways, whose primary function is to forward the health data to the Cloud and provide services for the discovery and control of IoMT devices; and Cloud, where the data are processed, stored, and consumed by applications. In this study, the Gateway plays the key role of enabling Fog Services for local data preprocessing and storage improving mobility and accessibility issues. However, the communications protocols, data format used, and integration with external applications are not clear.
Rashed et al. 20 proposed another IoT-based healthcare platform for remote monitoring in ambient-assisted living. It also considers three layers, namely Perception, where data are perceived and aggregated to be further transmitted to the next layer, Network/Gateway, which acts as a bridge between the sensor network (Perception) and the backend servers, and Integrated Application, which provides the end user with deep insights on the sensed data. IoMT devices communicate with the Gateway using the Message Queue Telemetry Transport (MQTT) protocol. 21 Each IoMT device acts as a publisher and the applications hosted in the Gateway, which implements an MQTT broker, act as subscribers.
Antonić et al. 22 proposed a Cloud-based Publish-Subscribe middleware (CUPUS) using a smartphone as a Gateway to connect the IoMT devices with a Cloud-based Publish-Subscribe engine. The data inside the messages are not semantically constrained and proposals that use such protocols might be incompatible due to incongruences in data format and meaning of the fields in the messages.
Datta et al. 23 discussed the sensors compatibility problem through the use of the M2M Gateway defined in ETSI 12 and the data representation issue through the implementation of the SenML standard proposed by Jennings et al. 14 According to SenML, the Gateway receives the sensed data in a sensor custom format and transcodes them following the SenML specification. However, the SenML-encoded data are sent to the upper layers, and other aspects that affect interoperability are disreagrded. For example, different units of measure can be associated with the same observation type and, therefore, sensors can handle the data in different units (i.e. the distance can be measured in meters, kilometers, miles, etc.). Since SenML defines only one unit of measure per observation type (i.e. meters are used for distance representation), a conversion mechanism is required. Moreover, sampling rates, observation method, or numerical precision, in scenarios that involve more than one platform, may lead to a wrong data interpretation. The authors did not address semantic interoperability.
Kim et al. 24 studied patents and the future directions of IoT and observed that ETSI standards were broadly adopted by many companies. Particularly, the M2M architecture and communication protocols defined in the ETSI TS 102 690 standard 12 have been mostly used and the Network Domain can be covered by the Internet in the IoT context.
Vidanagama 25 considered the M2M standard in which IoT devices communicate with the M2M Gateway through Representational State Transfer (REST) APIs and forward the data to the M2M Service Capabilities layer for storage and future queries of data by M2M applications. Although the study demonstrates why the M2M standard does not grant the interoperability required, the authors do not address semantic interoperability and follow no common data format for data representation.
The previous studies do not focus on the importance of semantic definitions of EHRs. EHRs play a key role for healthcare service providers, and integrated IoMT platforms should follow the same EHR semantic definition and vocabulary for the interchange of data with such types of systems. The next sub-section addresses relevant work on those issues.
Semantic integration of healthcare data
The semantic integration of healthcare data represents an essential component for an adequate interoperability of the different parts of a healthcare information system. Below are some possible solutions to this integration.
Pereira et al. 26 proposed an M2M-based platform that grants semantic interoperability and a common data interchange format. The authors considered M2M devices for data capture and a smartphone as an M2M Gateway that employs the HL7 message format in the communication with an openEHR-based EHR storage. Although the platform implements HL7 services for data interchange between healthcare systems, it neither addresses interoperability problems at sensor level nor considers IoT/IoMT scenarios with a large number of heterogeneous sensors. No tools have been developed for data interchange with external applications, which is a key aspect for the application of knowledge extraction techniques or big data processing.
Boutros-Saikali et al. 27 designed an IoMT platform focused on the development of healthcare monitoring applications. Following the concepts of data adapters, it enables the retrieval of data provided by different EHRs’ data sources through a unified set of REST services. Although the approach enables applications to query data from different EHR definitions, it avoids direct access to sensor-related meta-data and ignores the integration of isolated IoMT devices. Besides, the authors do not follow any standardized EHR definition and the healthcare semantics cannot be retrieved by applications.
Villanueva-Miranda et al. 28 highlighted that the use of ontologies can significantly enhance semantic interoperability in an environment with multiple IoMT platforms. The authors analyzed two main approaches, namely “One-to-One Ontology Alignment” (dynamic alignment of ontologies over the same domain) and “Central Ontology” (predefined ontology resulting from the merge of domain related ontologies), and concluded the former was better for interpretation and scalability.
Abinaya et al. 29 developed a “Central Ontology”-based system through which physicians can update the semantic definitions of diseases. The system stores the records following such definitions as ontology individuals. The semantic relationships help decision-making processes by either big data techniques or human analyses. Therefore, the integration with IoMT semantic devices is not clear, and the ontology follows no standardized EHR definition.
Gomez et al. 30 designed an ontology-based platform for health monitoring of patients with chronic diseases. Differently from the previous scheme, the platform uses a smartphone as a Gateway to integrate the IoMT devices and defines ontology classes to represent IoMT platforms. However, the definition disregards the existence of ontologies that have already covered this topic, therein promoting the heterogeneity of definitions and increasing the complexity of integration with existing IoT/IoMT platforms.
Table 1 shows a summary of the main characteristics of the proposals.
Summary of related work.
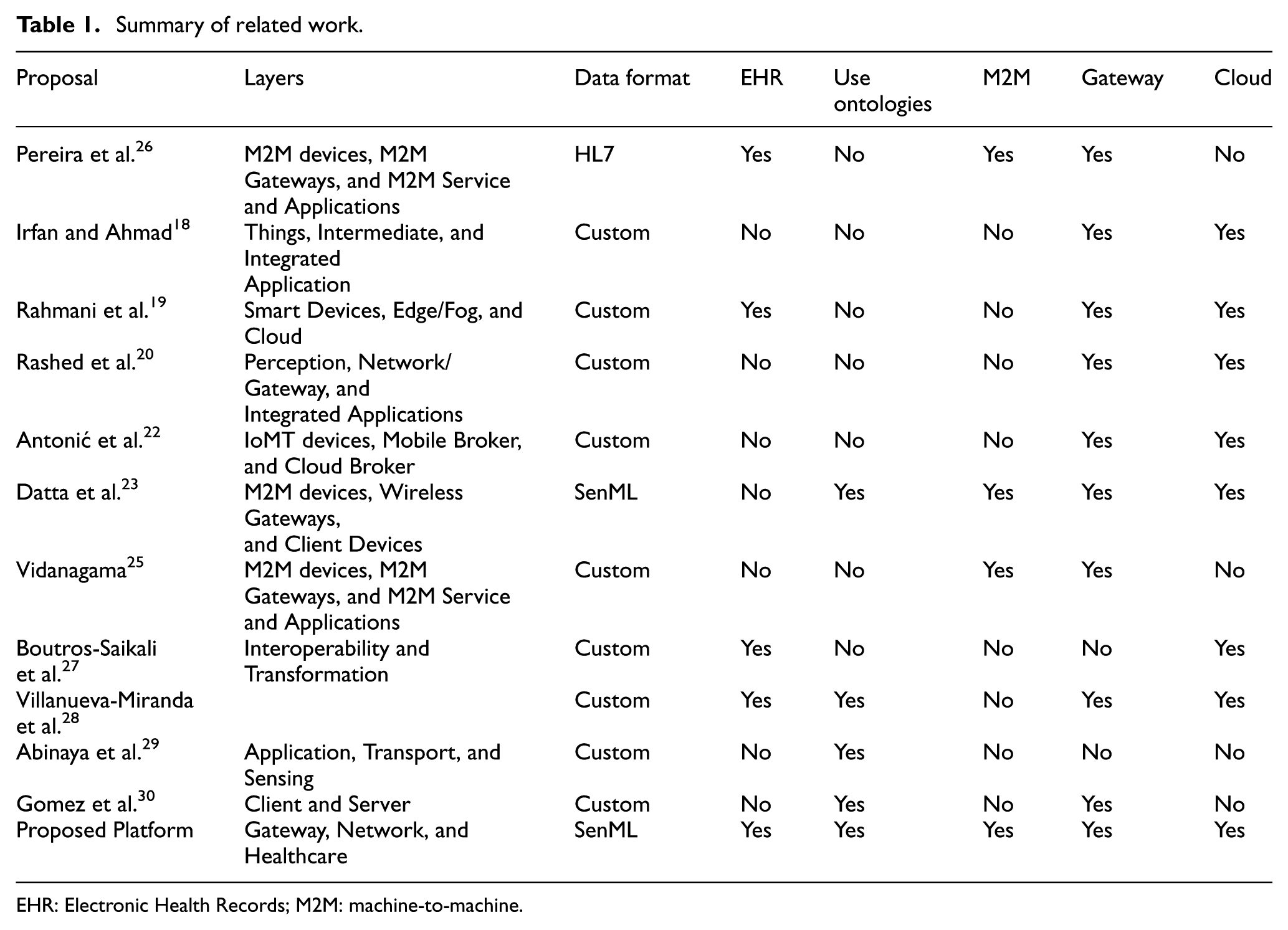
EHR: Electronic Health Records; M2M: machine-to-machine.
In addition, some work on languages and tools for semantic interoperability in the healthcare domain is available. Román 31 provided a set of Ontology Web Language (OWL) definitions that represents the openEHR RM as an ontology. However, the resulting ontology does not consider the other two layers (Archetypes and Templates) in openEHR.
Legaz-García et al. 32 described an OWL-based framework for the handling of EHR archetyped data as OWL ontologies. Although it enables the application of reasoning techniques and the automatic generation of inferred relations, it promotes no extraction of the OWL definition.
Lezcano 33 developed an open source translator for the conversion of ADL archetypes to an OWL representation. It includes a translation library composed of two packages (adl2owl.parser and adl2owl.translator) and a standalone application with a graphical user interface for the selection of an ADL file and its translation to an OWL one.
Discussion on related work
According to Table 1, many studies have adopted customized data formats for the data interchange between sensors and platforms, which has led to problems of interoperability, since the lack of standardization hampers the development of sensors specifically focused on the target platform. We considered SenML a mature standard, and IoT-related studies conducted by Su et al., 34 Fan et al., 35 Datta and Bonnet, 36 and Datta et al. 37 validate its relevance for the standardization of heterogeneous data sources.
Regarding the use of EHRs for the representation and storage of clinical records, some proposals19,27,28 implemented their storage systems based on EHRs. However, they do not address the simplification of exchange of clinical data between health institutions through the use of standardized EHRs.
In terms of use of ontologies, Datta et al., 23 Villanueva-Miranda et al., 28 Abinaya et al., 29 and Gomez et al. 30 showed an enrichment of the application domain for IoMT. According to the authors, ontologies enable both the application of computational reasoning techniques, which yield predictive results used as support to decision making, and the standardized representation of data in formats understandable by humans and computer programs. This characteristic is vital when interoperability in IoMT is considered.
Regarding functional architecture, the adoption of a standardized approach, based, for example, on the ETSI M2M standard,23,25 is more suitable than a customized one.18,19,22 The number of new sensors that have been developed demands mechanisms that simplify integration with health systems. Such an integration is facilitated by the use of a single standard, rather than the integration of a device with heterogeneous medical systems based on different standards.
Almost all the studies considered the use of Gateways and Cloud services.
In this sense, our IoMT platform was defined and implemented considering
A SenML data interchange format inside an M2M architecture;
Use of openEHR as an EHR representation;
Use of openEHR and SSN to provide ontologies for IoMT platform and healthcare data representation;
Use of a Gateway as IoMT devices data relay.
However, some issues have arisen from the integration of such standards. For example, openEHR covers concepts related to measures, observations, and healthcare domain knowledge, and SSN is one of the most used ontologies for the coverage of features specific to the IoT context. According to Ganzha and Paprzycki, 3 openEHR and SSN are some of the most promising semantic definitions in their areas. However, they cannot be joined directly since the archetypes and templates of openEHR are written in ADL and SSN is written in OWL. If openEHR is specified as an OWL ontology, it will have overlapped concepts with SSN, which requires an ontology alignment process.
Román et al., 31 Legaz-García et al., 32 and Lezcano 33 proposed some useful tools for solving the first, as previously described.
Regarding the second issue, this article proposes an alignment of SSN and openEHR ontologies, as described in the next section.
Alignment of SSN and OpenEHR-based ontologies
This section addresses an alignment between openEHR and SSN ontologies toward covering the healthcare and IoT concepts in the same IoMT vocabulary. The alignment process involves the translation of openEHR archetypes to OWL and identification of the overlapped concepts presented in openEHR and SSN. In addition, aspects of user demography, which are represented outside the openEHR-based resulting ontology and must be considered for the establishment of a user- device-observation relationship, are clarified.
We adopted the tool provided by Lezcano 33 to transform ADL archetypes files to OWL ones. The resulting OWL files were updated to the openEHR RM provided by Román. 31 The overall resulting ontology was called openEHR-OWL.
Since openEHR-OWL and SSN are available in the same language (OWL), an ontology alignment process must identify their overlapping concepts. Our resulting ontology is called openEHR-Extended and considers the mapping between the openEHR RM and archetypes and SSN Ontology. In OpenEHR, concepts defined above the RM and the archetype model (i.e. templates, versioning, services, among others) do not specify or define concepts related to devices. In this sense, the proposed mapping does not consider such a type of definitions since SSN does not have overlapped concepts with them. For example, the vocabulary and concepts defined by physicians in the upper layer (i.e. templates) are automatically covered by openEHR-OWL and require no alignment with SSN.
In openEHR-Extended, openEHR basic data types (Figure 3), as “text,” are mapped to “string” in SSN. The same occurs for numerical values that have direct matching. Dates and datetimes are mapped to “owl:datetime.” Other types, such as “openehr:paragraph,” which have no direct representation in SSN, are mapped to “ssn:string.” More complex types resulting from the combination of basic types are mapped to SSN data types, according to the definition provided by W3C Unicode Consortium, 38 for example, an “openehr:count” is mapped to an “rdf:count,” an “openehr: quantity” is mapped to an “rdf:measurement.”

openEHR basic data types.
The observation concept is defined in openEHR RM as the base class that represents the phenomenon of an observation or a sampling process. Any class derived from observations has common attributes, that is, date, duration, events, event time, information provider, subject, and workflow. Particularly, the events attribute of an observation could define a unique “in point of time” measure or a set of samples observed in a time interval represented as (“start datetime,”“start datetime”+“duration”).
In SSN, the observation concept is defined for a punctual measure of a sensor value and the sampling concept refers to a set of samples that followed a “sample procedure.” An “ssn:observation” was mapped to an “openehr:observation,” where the events attribute is forced to be null and the duration is always 0. “ssn:sampler” was mapped to an “openehr:observation,” where the events are gathered from “ssn:samples.”
SSN defines the sensor and platform classes regarding devices. A sensor refers to the entity that can observe an observable property. Sensors generate an observation and a sample. In openEHR, this concept is treated as an agent that might be a device that senses a parameter for making an observation. Therefore, an “ssn:sensor” will act as an agent in the openEHR domain. The platform class groups sensors into a platform (i.e. a smartphone has an accelerometer, gyroscopic and other sensors). The concept of platform is not considered in openEHR, and we propose extending the openEHR definition so that a platform can make an observation.
The other challenge refers to the identification of the user “patient” to whom an observation or a sampling is related. Since the ontology will be used for the sharing of clinical data following a semantic web approach, the patients’ privacy must be maintained. However, concomitantly, EHRs that belong to the same patient must be identifiable. openEHR separates EHR and demographic information in such a way no information is available in EHR for the identification of the patient’s identity.7,8 The EHR only defines the subject class whose attribute subject Id is related to the patient in a separated demography repository. In this sense, each EHR implementation can define its custom structure (the openEHR demography model does not need to be followed) for the patient representation. The binding of subjects with patients is performed by the Id at the software layer. On the other hand, SSN is not focused on observations over individuals. Indeed, the concept of patient or user is not defined. We extended the “ssn:observation” and “ssn:sampling” classes through the definition of an Id that identifies the patient under monitoring. Such Ids must be handled by IoMT applications when the individuals are published for the ontologies.
The next section addresses how to obtain patients’ Ids and publication of the ontology with the use of the data gathered by IoMT devices.
Semantic IoMT platform for e-Health
This section describes the IoMT platform and considerations taken in the development stage. Particularly, an extension of the functional architecture of ETSI M2M is proposed for providing semantic interoperability at the Gateway and Network layers. The section also describes the way SenML was used and enabled data model interoperability between the Gateway and Network layers and addresses the platform behavior.
The proposal of a Semantic IoMT Platform for e-Health followed communication protocols, network standards and services, according to the ETSI 12 M2M Functional Architecture specification, and was applied to the e-Health use cases defined by ETSI. 39 The architecture considers three domains, namely Gateway, Network, and Healthcare. Similar to the platform proposed by Pereira et al., 26 the Gateway domain is composed of IoMT devices that sense different parameters of the patients connected to Gateways through the M2M Area Network (i.e. Bluetooth, WiFi, or ZigBee). A Gateway provides access to the Internet and enables capabilities for device controller (i.e. device discovery and management) and sensed data management. The network domain provides capabilities for communication between the Gateways and the M2M Applications through the access network (i.e. Cellular Long Term Evolution (LTE)) and core networks (i.e. Internet). The Healthcare Domain covers the storage of the data and their semantics and availability to be consumed by the final applications. It also considers legacy EHRs and the way they will be maintained.
In contrast to Pereira et al., 26 our approach enables data sharing according to a semantic Web paradigm, considers user privacy issues, and uses more adequate data formats for the IoMT context.
Figure 4 shows the definition of the proposed architecture; blue and brown highlight the unmodified entities of ETSI M2M and the semantically enabled ones, respectively. M2M e-Health devices represent IoMT devices/platforms that collect patient-related data and connect to a smartphone Gateway through the M2M Area Network. The Semantic e-Health Application (SeHA) deployed in the M2M smartphone Gateway grants semantic interoperability in the Gateway domain and functions for both device discovery and data trans-coding. In the network domain, observation messages sent from the gateway to the NSCP are forwarded to the semantic controller, which populates the individuals in the openEHR-Extended OWL Ontology. openEHR-Extended OWL provides functions for the insertion, update, and deletion of individuals regarding the semantic controller and retrieval of data through SPARQL queries by New Semantic Applications. The applications can use the data for numerous purposes, for example, training of machine-learning models for predictions, implementation of fuzzy logic techniques for decision support, among others.

Platform architecture.
The openEHR storage controller grants compatibility with legacy hospital applications by maintaining openEHR-based EHRs updated. Once it receives an observation message, it updates the corresponding EHR record. Therefore, legacy web platforms and applications that use EHRs as data providers are not affected.
Figure 5 shows an example of the platform behavior. The M2M e-Health devices are appropriately defined in openEHR-Extended and an Internationalized Resource Identifier (IRI) pointing to this resource is available. The process starts when the patient logs into the SeHA hosted in the smartphone Gateway. The application communicates with the service capability provider to authenticate the user and retrieve the unique Identifier (pId) stored in the EHR. A process for the discovery of the M2M e-Health devices is then started. The devices are discovered and configured according to the ETSI definition. The GSCP handles both network interfaces and the discovery process. Parameters such as observation rate, time alignment for start and end times are set in this phase.
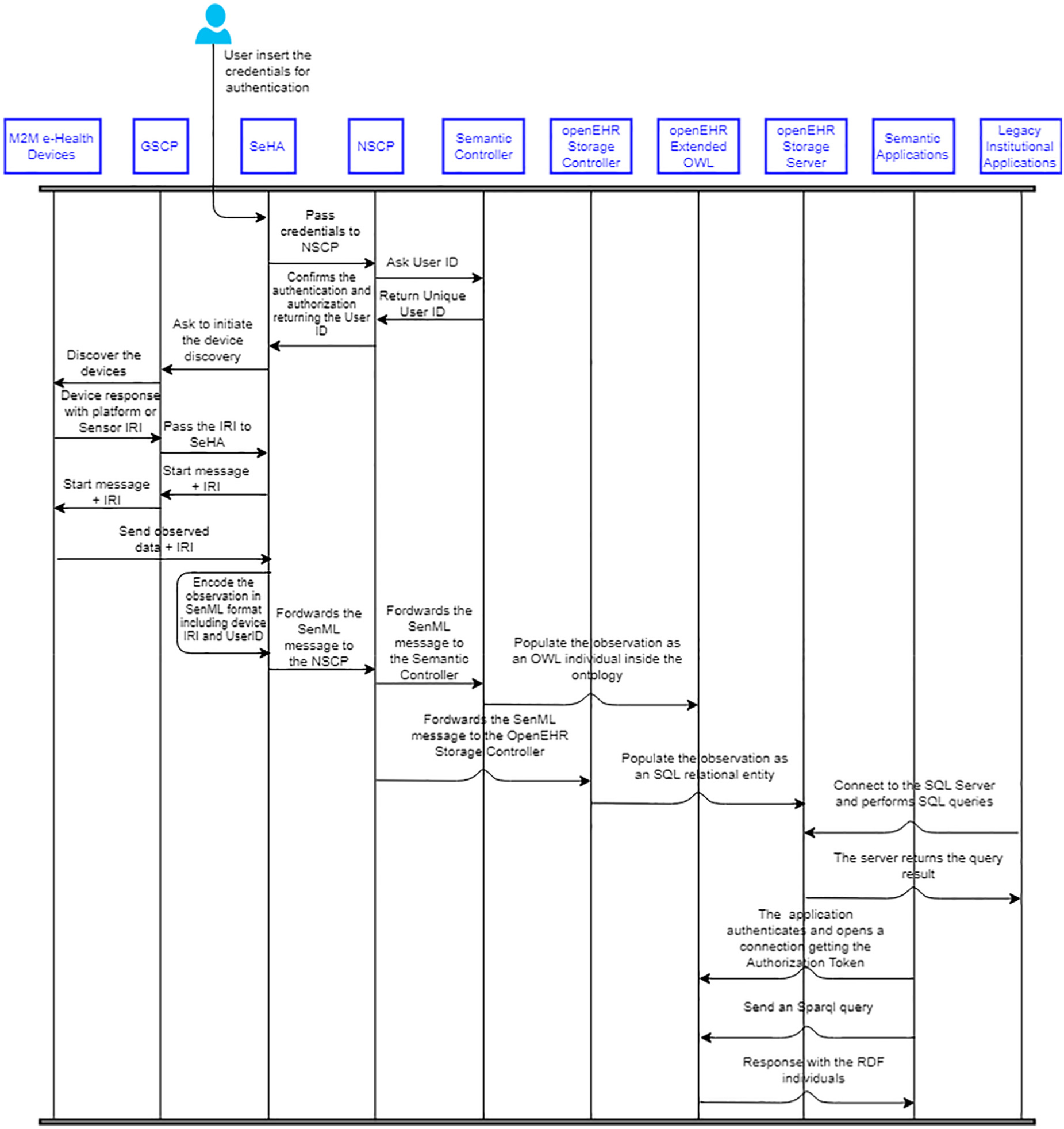
Message sequence diagram.
Each e-Health device discovered is identified by the corresponding IRI of the class in openEHR-Extended, which describes the sensor or the platform. The M2M e-Health device starts to transmit the observations to the SeHA located in the Gateway. When an observation message has arrived, the SeHA translates it to a SenML encoded one following the definition below:
{“n”: “IRI of the sensor in openEHR-Extended”+“/Uid”,
“t”: “time at which the observation was received”,
“u”: “IRI of the unit_of_measure in openEHR Extended”,
“v”: “Numeric value obtained in the observation”,
“vs”: “String encoded value, if it exists” }
Once the Message has been encoded, the SeHA forwards it to the NSCP according to a REST-based web services approach. NSCP transmits the received JSON SenML message to both semantic controller and openEHR storage controller.
The semantic engine application obtains the sensor IRI and User Id (Uid) from field “n” and asks openEHR-extended OWL ontology for the creation of the individual. The openEHR storage controller updates the EHR database. Finally, semantic applications can query data stored in the ontology, and applications that use the EHR database as an information provider continue using the same approach with no modifications.
Our approach does not grant interoperability between non-M2M compliant sensors. Each sensor uses its custom network interface over the M2M Gateway network according to its proprietary communication protocols. Furthermore, the SeHAs hosted in the Gateways must be customized to map the data provided by sensors in SenML messages. The platform provides the skeleton of the e-Health semantic application where interfaces must be implemented for the creation of a data converter. Libraries for integration with upper layers are provided inside the skeleton as a framework for the development of Android applications.
If a device does not know their IRI inside the platform, the devices can request an IRI from the SeHA. Such a request must contain an archetypeId parameter, which refers to one of the observation archetypes already defined in the openEHR definition. The archetypeId is transmitted to the semantic controller, which creates the sensor on the ontology and associates it with the type of observation defined by archetypeId. Then, it returns the IRI to the SeHA, which forwards it to the sensing device.
Implementation details
Several technologies were involved in the platform implementation, as shown in Figure 6.

Technologies used for implementation.
CookingHack e-Health Sensor Platform Complete Kit V2.0 40 was considered for the Proof of Concept (PoC) and played the role of IoMT/M2M e-health devices. The kit comprehends sensors, such as airflow, ECG, SpO2, electromyography, blood pressure, glycaemia (blood sugar), body temperature, and an accelerometer, that detect the patient’s position (sitting, lying, or standing). They are integrated to an Arduino development platform by a shield board. The connection of the devices with the Gateway is provided by a Bluetooth module.
As in ETSI, 12 Arduino hosts the Device Applications (DA) for each sensor, which communicates through GSCP services with the SeHA hosted by the M2M Gateway. DAs were developed in C++ and in compliance with the ETSI definition. They connect to the M2M Gateway and receive, parse, and store the sensor IRI, used for the identification of the device in future messages. Each DA developed follows the same sampling definition, that is, 60 Hz sampling frequency, and the samples are transmitted separately to the SeHA over the Bluetooth interface. DAs encode the samples following the SenML format before forwarding them to SeHAs. A C++ skeleton of a DA is provided as a framework for the integration of other devices different from those used in our PoC.
An Android smartphone was used for the implementation of the M2M Gateway. Both GSCP and SeHAs were developed in Java, and the SeHAs were developed as normal Android mobile applications. A Java framework for the integration of SeHAs with NSCP enables the development of new local features, such as filtering, data visualization, and offline buffering.
The development of NSCP, controllers, and storage followed a cloud paradigm focused on the Microsoft Azure cloud service provider. NSCP functions, developed as NodeJS REST APIs and deployed in an Azure App Service, forward the SenML messages to both controllers developed and deployed as Azure WebJobs. An instance of both webjobs is created for each message received, thus enabling parallel processing.
Finally, the openEHR-Extended OWL ontology is managed by an Apache Jena server deployed on an Azure Virtual Machine Service. The implementation of the openEHR storage server followed an SQL-based approach and used the Microsoft Azure SQL Service.
The openEHR storage controller is configured with the connection string and uses the openEHR specification to infer both the database schema and mapping rules between the message and the database structure. On the other hand, the semantic controller uses the openEHR-Extended ontology to map the SenML message to OWL resources maintained on the Apache Jena server.
For the sake of security, all entities communicate over TLS using certificates assigned by a public authority, automatically managed by the platform and issued periodically. They enable encryption, authentication, and authorization of applications. The openEHR-extended OWL entity provides a set of services for token authentication.
Figure 7 shows the usage workflow to be followed by developers for adding a new sensor and application and for the correct use of the platform. The process involves the instantiation of the platform, definition of openEHR archetypes that represent the observations, registration of the sensor/platform and archetype on the openEHR-Extended OWL ontology, modification of DAs and the gateway application (GA), and development/integration of the healthcare application that will consume the data. The next section reports on a use case and provides an example of the process.

Development workflow.
Example of a use case
The use case analyzed considers the remote monitoring of the respiratory waveform, electrocardiography signal, and percentage of oxygen in the blood of a patient in his house. The application focuses on the development of sleep studies conducted by doctors remotely. In addition, the system enables the detection of parameters outside the intervals considered healthy and implements an alert system to notify the healthcare professionals. The workflow (Figure 7) comprises the following steps:
The instantiation of the platform followed the technologies shown in Figure 6;
The openEHR observation archetypes considered were Pulse Oximetry for SpO2 GA and Respiration for Airflow GA and ECG result, available in the openEHR Clinical Knowledge Manager.
Three DAs were configured to handle each sensor connected to an Arduino UNO development platform, and PostObservation, Start, Stop, and Connect functions were adapted to fulfill each sensor’s requirement. Particularly, the Connect function was rewritten toward a WiFi network connection through the setting of network parameters, such as M2M Gateway IP address and GSCP port;
A GA was instantiated on the M2M Gateway and a visual interface for patient login and sensor controller was developed;
A semantic application was developed for querying Apache Jena Server resources relative to the patient’s Id, in 3 s intervals. It monitors if parameters are outside boundaries considered healthy and alerts a medical team;
A Web application was developed for the registration of patients and review of clinical reports.
Figure 8 shows the run-time workflow for the use case. The process considered the registration of the patient and his or her demographic data, installation of the platform components on the user’s smartphone (as the M2M Gateway), manual monitoring of health parameters by the medical team, and an automatic monitoring approach that uses the semantic application that alerts the medical team on at least one parameter outside the recommended boundaries.

Run-time workflow.
All IRIs considered for sensors follow prefix https://openehrextended.blob.core.windows.net/sensor# and those considered for observations follow prefix https://openehrextended.blob.core.windows.net/observations#. Below are some examples of the way data involved in this use case are stored and queried by the applications.
Data published in openEHR-extended as individuals
The example below shows the sensors represented as individuals inside the ontology:
<ECG7838> a sosa:Sensor;
rdfs:label “3 Level ECG Sensor”;
openEHRExtended:Observes
“openEHR-EHR-OBSERVATION.ecg_result.v0”;
<AF2107> a sosa:Sensor;
rdfs:label “Airflow Sensor”;
openEHRExtended:Observes
“openEHR–EHR–OBSERVATION.respiration.v1”;
<SPO29877> a sosa:Sensor;
rdfs:label “Pulse Oximeter Sensor”;
openEHRExtended:Observes
“openEHR-EHR-OBSERVATION.pulse_oximetry.v1”;
Each sensor is defined by the SSN sensor class defined in the SOSA ontology, but extended to reference the Id of the openEHR observation the sensor can measure.
An observation inside the ontology can be represented through the use of the specialized observation class related to one of the three observable properties considered in this use case. For example, the OWL fragment below shows an ECG-related observation made by sensor “ECG7838”:
<Observation/1> a sosa:Observation;
sosa:hasFeatureOfInterest “ECG”
openEHR:openEHR–EHR–OBSERVATION.ecg_ result.v0>;
sosa:madeBySensor <ECG7838>;
openEHR:agent <ECG7838>;
openEHR:has_subject <UID2384>
sosa:hasResult <ResultECG1>;
openEHR:data <ResultECG1>;
openEHRExtended: at_datetime “10/23/2018 10:13:24” owl:datetime ;
openEHRExtended: has_data <Data/1> openEHR: data;
<ResultECG1> a openEHR:ecgdata
om:numericValue “0.9”;
openEHRExtended:for_property “Lead_I”
openEHR:propertyValue;
The sensor was defined twice in the previous observation, that is, once as in SSN, and once as in openEHR, which enabled the design of queries according to the SSN or the openEHR definitions. The value of the observation is represented by the “ecgdata” class as an SSN “result” pointing to the millivolt “unit_of_ measure.” Property “openEHRExtended:for_property” identified what generated the observation result.
Example of SPARQL queries performed by the semantic application
We consider a semantic Web application interested in querying the data sends a GET request to a REST API, passing the SPARQL query string as a parameter. For example, the query below obtains observations of a sensor in a date range:
SELECT ? observation ? result
WHERE
{
?observation rdf:type
sosa:Observation.
?sensor rdf:type sosa:Sensor.
?result rdf:type openEHR:data.
?result ssn:belongs_to_observation
?observation.
?observation . sosa:madeBySensor
?sensor.
?sensor refs:label
“3 Level ECG Sensor”
?observation openEHR:data ?Result
?observation openEHRExtended:at_datetime
?date.
FILTER (?date >= “2018–10–23
00:00:00”^^owl:datetime) .
FILTER (?date < “2018–10–24
00:00:00”^^owl:datetime)
}
The query of the measurement relative to a patient is also possible. The fragment below shows the way it can be designed for the user ID “UID2384”:
SELECT ?observation ?result
WHERE
{
?observation rdf:type
sosa:Observation.
?subject rdf:type
openEHR:subject.
?observation openEHR:has_subject
?subject.
?subject refs:label
“UID2384”.
?result rdf:type openEHR:data.
?result ssn:belongs_to_observation
?observation.
?observation
}
Similar to the examples above, all measurements of a patient airflow sensor can be obtained in a time interval for the computation of the average value, or filters can be applied for the retrieval of the most recent SpO2 observation. In summary, the use of the ontology to represent the data and SPARQL as the query language enables the retrieval of data in multiple ways, hence the development of more granulated e-Health applications.
Considerations on the web application and openEHR storage server
Once the connection string for the openEHR storage server has been configured on the Web application, stored procedures can be invoked for the accomplishment of the following tasks:
Registration of a patient;
Registration of a doctor;
Query of EHRs by patient and date;
Navigation on EHR pages;
Retrieval of hierarchical documents.
All such functions were defined in SQL, rather than in AQL, toward a simpler development of new applications for e-Health since SQL is a common-use query language. However, many tools use AQL files to generate healthcare forms automatically. In this sense, an inverse mapping from SQL results to AQL files can be performed, thus leveraging the automation of a visual interface generation. Frade et al. 41 reviewed studies on different openEHR storage implementations, mappings, and automation techniques for data visualization.
Conclusion
This article has presented an IoMT platform that relies on Semantic Web Concepts and M2M communications to simplify and standardize the development and integration of healthcare systems. Differently from other proposals, it considers data structures for the storage and transmission of data related to healthcare observations, which enables interoperability regarding data representation formats, IoMT platforms, and domain language (healthcare vocabularies, in our study).
The main contribution of the platform is the openEHR-Extended ontology, which aligns the Healthcare domain (openEHR) with the IoMT technical domain (SSN). It serves as a data storage model following a semantic WEB approach, but also identifies sensors and automatically translates SenML sensed data to OWL individuals, thus granting semantic coherence between both domains.
Another key aspect of this research regards the semantic extension of the openEHR model to the M2M Gateway Domain, which enabled definitions of heterogeneous IoMT devices inside a unique data model through a relation between the openEHR Observation archetype ID observed by the device and the device definition inside SSN. Differently from other semantic-based approaches, our platform enables the management and query of both clinical data and IoMT platforms that collect them.
Moreover, M2M capable IoMT devices can use their custom message format to forward sensed data over the M2M environment. Our platform addresses this problem and provides tools for the implementation of data converters in the SeHA deployed on the M2M Gateway, which converts heterogeneous data into a semantically coherent SenML format.
Future research includes performance tests and extensions to new scenarios, as Smart Cities (SC), where healthcare data play a vital role in the creation of statistical indicators on the quality of life and cities performance. Moreover, for an SC implementation, the integration and interoperability among different verticals (transport, energy, environment, healthcare, security, among others) represent an area to be addressed by the extension of our proposal.
Supplemental Material
DSN-19-0235.R3_Latex_files – Supplemental material for Interoperable Internet of Medical Things platform for e-Health applications
Supplemental material, DSN-19-0235.R3_Latex_files for Interoperable Internet of Medical Things platform for e-Health applications by Jesús Noel Sárez Rubí and Paulo Roberto de Lira Gondim in International Journal of Distributed Sensor Networks
Footnotes
Handling Editor: Ghufran Ahmed
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
