Abstract
Background. Epigenetic biomarkers have the potential to explain outcome heterogeneity following traumatic brain injury (TBI) but are largely unexplored. Objective. This exploratory pilot study characterized brain-derived neurotrophic factor (BDNF) DNA methylation trajectories following severe TBI. Methods. Brain-derived neurotrophic factor DNA methylation trajectories in cerebrospinal fluid (CSF) over the first 5 days following severe TBI in 112 adults were examined in association with 3- and 12-month outcomes. Results. Group-based trajectory analysis revealed low and high DNA methylation groups at two BDNF cytosine-phosphate-guanine (CpG) targets that showed suggestive associations (P < .05) with outcomes. Membership in the high DNA methylation groups was associated with better outcomes after controlling for age, sex, and injury severity. Associations of age × trajectory group interactions with outcomes at a third CpG site revealed a pattern of the same or better outcomes with higher ages in the high DNA methylation group and worse outcomes with higher ages in the low DNA methylation group. Conclusions. Although no observed associations met the empirical significance threshold after correcting for multiple comparisons, suggestive associations of the main effect models were consistent in their direction of effect and were observed across two CpG sites and two outcome time points. Results suggest that higher acute CSF BDNF DNA methylation may promote recovery following severe TBI in adults, and this effect may be more robust with higher age. While the results require replication in larger and racially diverse independent samples, BDNF DNA methylation may serve as an early postinjury biomarker helping to explain outcome heterogeneity following TBI.
Keywords
Introduction
Globally, approximately 5.5 million people sustain a severe traumatic brain injury (TBI) each year, with ∼15% mortality and a high number of survivors living with chronic functional disability.1,2 Current prediction models explain only a third of the variance in patient outcomes. 3 The remaining unexplained heterogeneity in outcomes is a critical barrier to accurate prognostication and hinders the development of evidence-based interventions.4,5 The investigation of personal biological factors or “biomarkers” is a promising avenue that may explain additional variance and identify mechanistic targets for treatment development. The field of epigenetics has become central to studying the modulation of genetic phenotypes by environmental factors. The most well-studied epigenetic mechanism is DNA methylation (DNAm), which involves the addition of a methyl group to a cytosine-phosphate-guanine (CpG) site, with higher methylation often rendering the gene less transcriptionally active. 6 Characterizing DNAm representing the central nervous system (CNS) environment following TBI has great potential to elucidate mechanistic differences that may explain outcome heterogeneity.
Experimental models indicate robust DNAm changes in various brain regions and cell types post-TBI. These dynamic methylomic changes have been observed in both pediatric and adult TBI rat models during the acute/sub-acute7–11 and chronic12,13 periods following brain injury. Both differentially hypomethylated and hypermethylated loci, and associated changes in gene expression, have been observed relative to sham controls. Systems biology approaches have identified enrichment of differentially methylated genes following experimental TBI in cell death and survival, nervous system development and function, inflammation, metabolism, and cell communication pathways.10,12
The present exploratory pilot study aimed to characterize methylation trajectories in DNA extracted from serial cerebrospinal fluid (CSF) samples collected over the first 5 days following severe TBI in adults and examine associations of inferred trajectory groups with short- and long-term patient outcomes. We chose a candidate gene approach, targeting the brain-derived neurotrophic factor (BDNF) gene. Brain-derived neurotrophic factor is a well-studied neurotrophin growth factor that is released in an activity-dependent fashion from pre- and post-synaptic neurons. Synthesized as pro-BDNF and subsequently cleaved to mature BDNF, pro-BDNF and mature BDNF function antagonistically to regulate neuroplasticity. 14 BDNF gene expression is regulated in part by DNAm, with higher methylation associated with lower gene expression 15 and neuronal synthesis of BDNF. 16
The examination of BDNF DNAm in CSF in the acute period following severe TBI may provide new information regarding the role of BDNF in outcome heterogeneity and the epigenetic influence of the CNS environment on BDNF gene expression. We characterized BDNF DNAm trajectories in CSF over the first 5 days following severe TBI in adults and examined associations of inferred trajectory group membership with patient outcomes 3- and 12 months post-injury. Given prior TBI studies suggesting differential effects of BDNF with age,17,18 we also explored the moderating effect of age on BDNF DNAm trajectories and their association with outcomes.
Methods
Study Design and Participants
We derived data from a prospective, longitudinal, observational study of adults with severe TBI. The study was approved by the University of Pittsburgh Institutional Review Board and consent obtained from legal authorized proxy individuals and re-consent obtained when appropriate. Inclusion criteria included severe head injury, defined as hospital-admission Glasgow Coma Scale (GCS) score ≤8 prior to the administration of paralytics or sedatives; age 16–80 years; and CSF sampling via external ventricular drain (EVD) during standard care. Individuals with cardiovascular disease or conditions predisposing them to cardiovascular disease were excluded to control for differential DNAm relating to these comorbidities 19 rather than TBI and to control for the effect of such comorbidities on TBI recovery. Additional exclusion criteria were penetrating head injury or brain death.
DNA Methylation Data Collection
Genome-wide DNAm data were collected from bisulfite-converted DNA extracted from CSF collected via EVD into a sterile ventriculostomy bag over the first 6 days post-TBI. Cerebrospinal fluid samples were centrifuged, and the cellular pellet and aliquoted supernatant were stored in a −80° freezer. DNA was extracted from the cellular pellet using the QIAamp DNA extraction kit from Qiagen Corp (Qiagen, Valencia, CA, USA).
The Infinium HumanMethylation450 BeadChip and Illumina iSCAN (Illumina, Incorporated, San Diego, CA, USA) were used to generate and scan genome-wide DNAm data at the Center for Inherited Disease Research. As part of laboratory quality control (QC), technical replicates and DNAm controls were included, participants were randomly distributed across the batches, and all samples from a participant were run on the same chip and within the same batch.
Genome Studio Software (Illumina, Incorporated, San Diego, CA, USA) was used to analyze the raw genome-wide DNAm data. Data cleaning and QC were performed as previously described. 20 Briefly, we used the Minfi, 21 ENmix, 22 and funtooNorm 23 packages in R to remove poorly performing samples and probes and perform functional normalization. Our final dataset included DNAm data for 419 895 probes in 120 participants at up to 3 cross-sectional time points. Following data cleaning, BDNF DNAm data were extracted from the gene transcript region ±5000 kb upstream and downstream (GRCh37/hg19, chr11:27671442-27727600).
Patient Outcomes
Outcomes of interest examined at 3 and 12 months post-injury included mortality, global outcome as measured by the Glasgow Outcome Scale (GOS), 24 neurobehavioral dysfunction as measured by the Neurobehavioral Rating Scale-Revised (NRS-R), 25 and functional outcome as measured by the Disability Rating Scale (DRS). 25
Statistical Analyses
Preliminary Analyses
Statistical analyses were conducted using R (version 4.0.1, Vienna, Austria) and SAS (version 9.4, SAS Institute Incorporated, Cary, NC, USA). The NRS-R was treated as continuous. The DRS was collapsed into categories and treated as ordinal (0, no disability; 1–11, mild disability; 12–29, severe disability; 30, death), consistent with prior studies 26 and to maintain clinical meaningfulness. Both GOS and death were treated as binary after dichotomizing GOS (good, 4 to 5; poor, 1 to 3).27,28 GCS was dichotomized (severe, 6–8; very severe, 3–5) due to small cell sizes.28,29
For DNAm data, M values were used, and participants with only one DNAm data observation were removed. Standard descriptive statistics and data screening were performed. DNAm observations identified as an outlier >3 times the interquartile range (IQR) within each day were score adjusted. Regression assumptions were evaluated, and data were transformed to normality as necessary.
Group-Based Trajectory Analyses
DNAm trajectory groups were inferred for each CpG site using group-based trajectory analysis (GBTA) and censored normal models implemented with the Proc TRAJ macro in SAS.30,31 We largely automated our analyses using R, SAS, and an objective protocol as previously described. 32 Briefly, our criterion for best model selection included evaluation of Bayesian Information Criterion, a model simplicity score, and posterior model selection QC including requirement of an average posterior probability of >.7, odds of correct classification >5, and estimated group membership of >5%. 32
Cell-Type Heterogeneity
Differences in cell types within biospecimens can vary between individuals, across time and tissues, and can confound relationships in DNAm studies. 33 The overall DNAm level represents a composite value of cell-type–specific methylation levels weighted by proportion. In cases where genome-wide DNAm data are generated, existing algorithms can be used to infer cell-type proportions for individual samples so this heterogeneity can be controlled for. Because there are no reference DNAm data to infer cell-type heterogeneity (CTH) found in CSF, we used Houseman’s reference-free method and the RefFreeEWAS package in R 34 to generate CTH data as percentages of major cell types. While this method has been shown to have high accuracy resulting in putative “cell types” defined by cell-type–specific methylomes, 34 and cell types found in peripheral blood are observed within CSF following TBI, the true cell-type identities cannot be confirmed. However, it is currently unlikely that genome-wide and CTH data would be generated for a clinical biomarker due to cost, time, and laboratory burden. To explore the potential clinical utility of BDNF DNAm unadjusted for CTH, as well as potential confounding by cell type, our GBTA protocol was implemented twice for all CpG sites to compute trajectories both unadjusted and adjusted for CTH.
Association Analyses
Following GBTA, for CpG sites with trajectory models passing posterior model QC described above, the relationships between inferred trajectory group assignment and patient outcomes were tested in R while controlling for age, sex, and dichotomized GCS using linear (NRS-R), ordinal logistic (DRS), and binary logistic (GOS and death) regression. P-values <.05 were considered suggestive. Given the correlation between patient outcomes selected for this study, we did not adjust for multiple testing. However, we did compute an empirical significance threshold, P emp , using 10 000 permutations as previously described. 32 For CpG sites associated with outcomes, participant characteristics by trajectory groups were compared using t-tests for continuous variables and chi-square or Fisher’s exact test for categorical variables. Given prior TBI studies suggesting differential effects of BDNF with age,17,18 we also explored the moderating effect of age on BDNF DNAm trajectories and their association with outcomes. Analyses mirrored the above but also included an age–trajectory group interaction term for each model.
Results
Preliminary Analyses
Sample Characteristics.

Abbreviations: DAI, diffuse axonal injury; Death-3, death at 3 months; death-12, death at 12 months; DRS-3; Disability Rating Scale at 3 months; DRS-12, Disability Rating Scale at 12 months; GOS-3, Glasgow Outcome Scale at 3 months; GOS-12, Glasgow Outcome Scale at 12 months; ICH, intracranial hemorrhage; IVH, intraventricular hemorrhage; NRS-R-3, Neurobehavioral Rating Scale at 3 months (untransformed, raw data); NRS-R-12, Neurobehavioral Rating Scale at 12 months (untransformed, raw data); SAH, subarachnoid hemorrhage; SD, standard deviation.
a Secondary hemorrhage patterns were present for many participants, but information was not available for this sample.
From the genome-wide DNAm data, data for 45 CpG sites were included in our analyses. During DNAm outlier screening, 43 observations across the 5 days were identified as extreme outliers >3 times the IQR and score adjusted. NRS-R data were transformed to normality using a square root transformation. In bivariate analyses, age was associated with square root–transformed NRS-R at 3 and 12 months (P = .005 and .006, respectively), DRS at 3 and 12 months (P = .003 and P < .00005, respectively), GOS at 12 months (P = .002), and death at 3 and 12 months, (both P < .00005), and worse GCS was associated with DRS at 3 months (P = .02). No associations between sex and outcomes were observed.
Group-Based Trajectory Analyses
Of the 45 CpG sites available, DNAm trajectory groups were inferred at 32 CpG sites unadjusted for CTH and 39 CpG sites adjusted for CTH as shown in Supplementary Table 1. Trajectory plots for all CpG sites with models passing QC have been included in the Supplementary Material. Of the models that passed QC, 14 unadjusted models and 36 CTH-adjusted models consisted of only one trajectory group and were excluded from further analyses. Therefore, of the models passing QC, 18 unadjusted and 3 CTH-adjusted models with more than one trajectory group were carried forward for patient outcome association tests.
Association analyses
Primary Analyses
Associations between inferred trajectory group assignment and patient outcomes were tested using regression while controlling for age, sex, and dichotomized GCS. Given the scale of this study, we have only presented results for CpG sites with suggestive associations with patient outcomes (cg26057780 and cg25328597). Results for all CpG sites can be found in Supplementary Table 1. At cg26057780 and cg25328597, two distinct trajectory groups were inferred at each CpG site including a “low DNAm” group and a “high DNAm” group. Inferred participant group assignment for each site included cg26057780 (n = 105 [93.8%] low DNAm, n = 7 [6.2%] high DNAm) and cg25328597 (n = 98 [87.5%] low DNAm, n = 14 [12.5%] high DNAm). Observed and estimated DNAm trajectories are shown in Figure 1. Trajectory plots for CpG sites cg26057780 and cg25328597 unadjusted for cell-type heterogeneity. Abbreviations: CSF, cerebrospinal fluid; TBI, traumatic brain injury; (A) cg26055780; (B) cg25328597.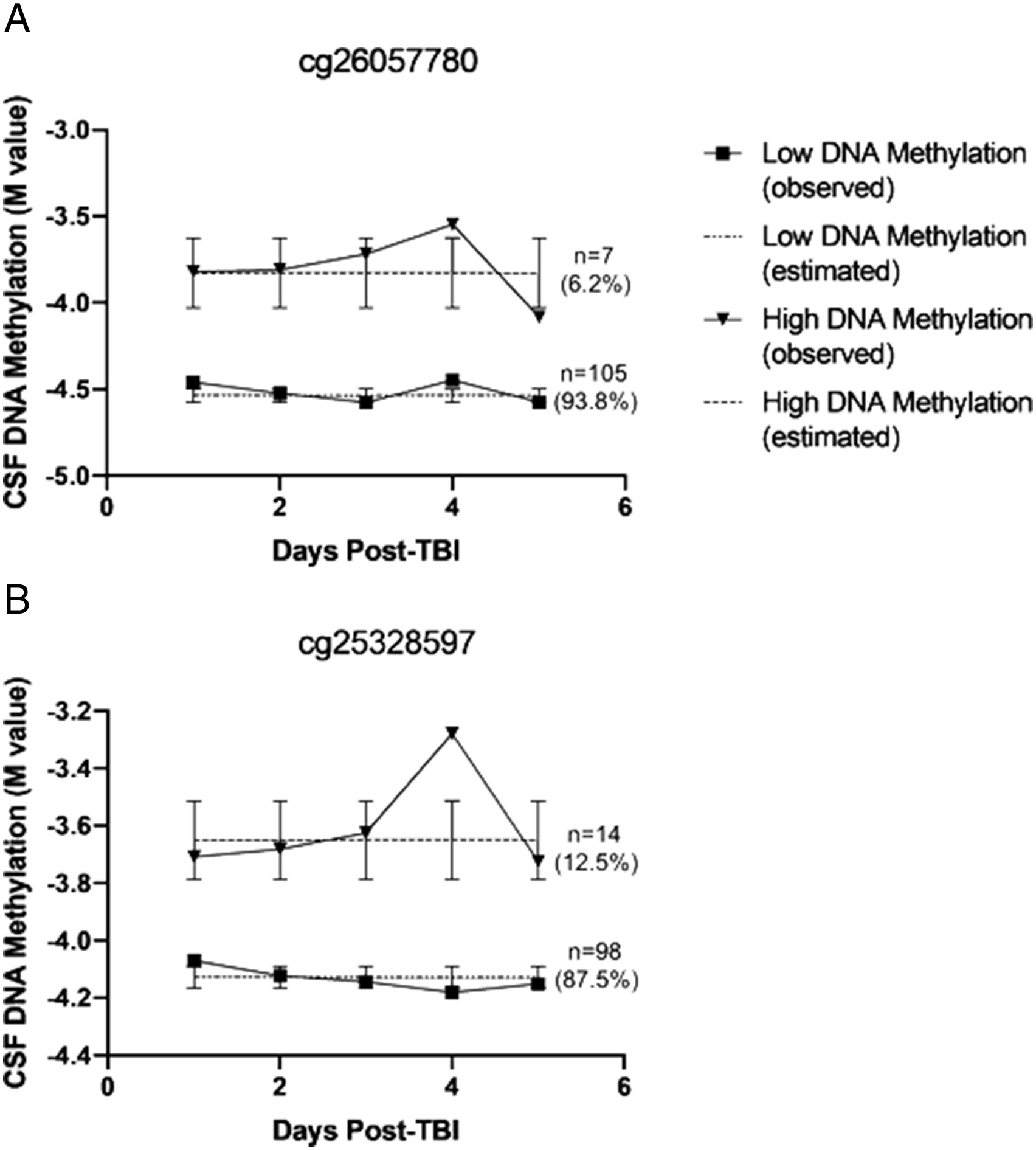
Trajectory Group Characteristics for cg26057780 and cg25328597.
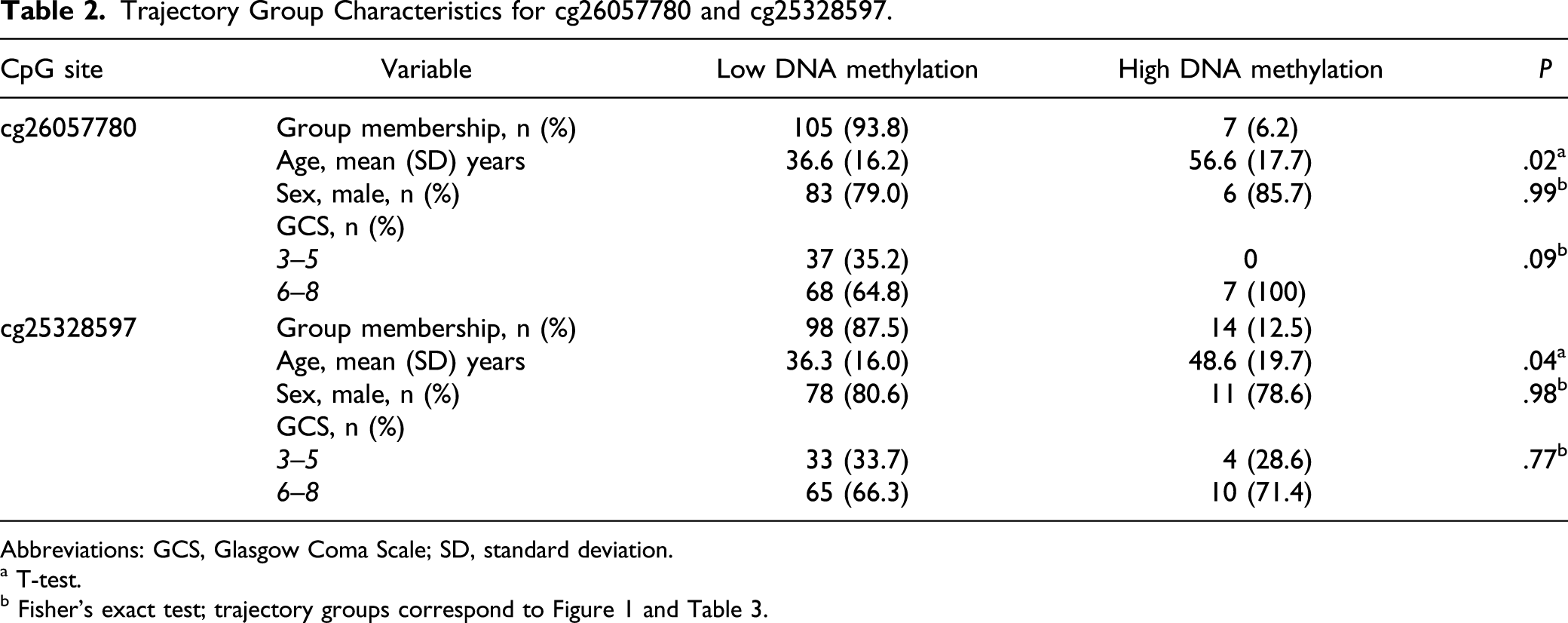
Abbreviations: GCS, Glasgow Coma Scale; SD, standard deviation.
a T-test.
Results of Trajectory Group–Patient Outcome Association Tests While Controlling for Age, Sex, and Dichotomized Glasgow Coma Score.
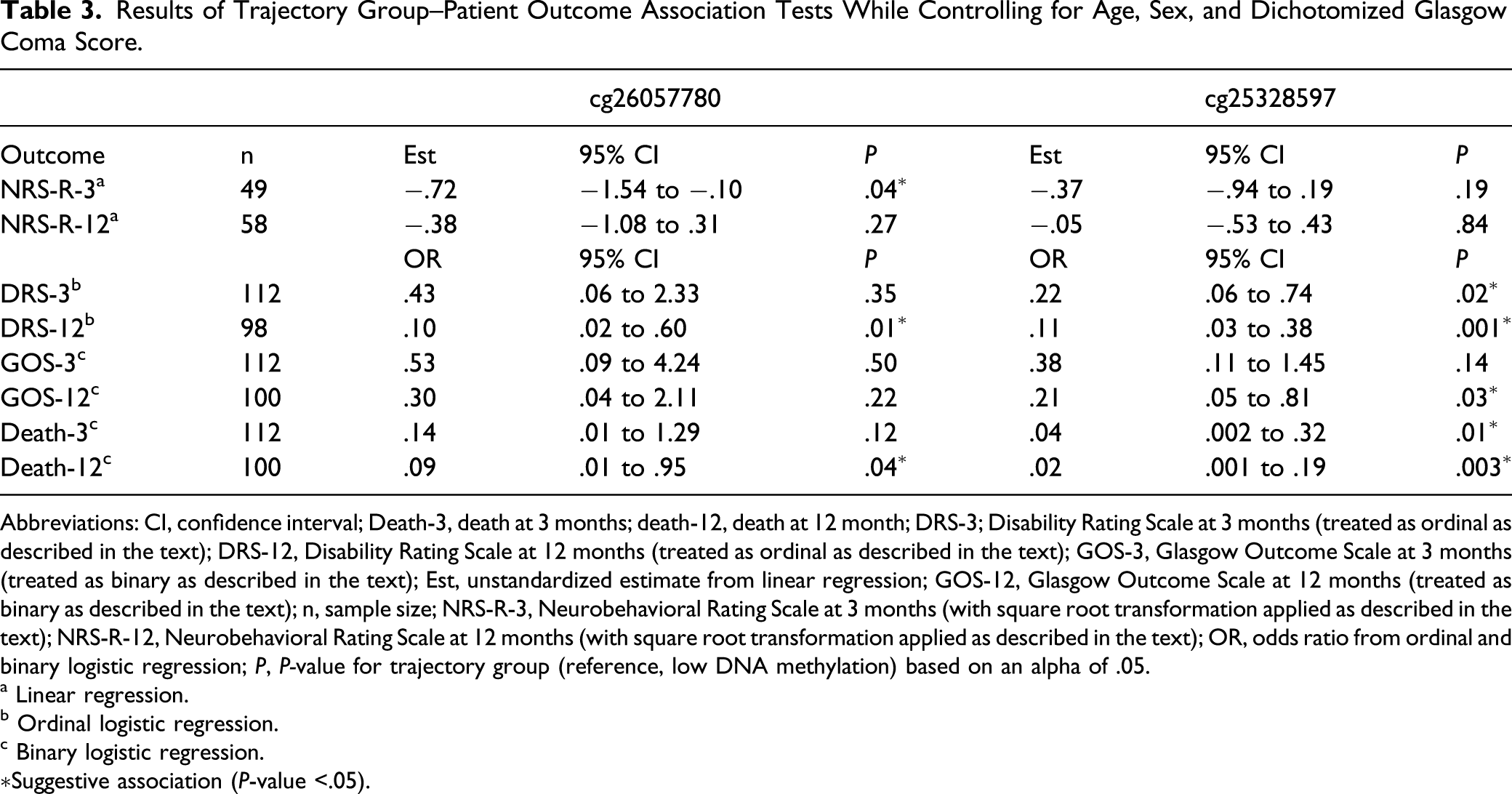
Abbreviations: CI, confidence interval; Death-3, death at 3 months; death-12, death at 12 month; DRS-3; Disability Rating Scale at 3 months (treated as ordinal as described in the text); DRS-12, Disability Rating Scale at 12 months (treated as ordinal as described in the text); GOS-3, Glasgow Outcome Scale at 3 months (treated as binary as described in the text); Est, unstandardized estimate from linear regression; GOS-12, Glasgow Outcome Scale at 12 months (treated as binary as described in the text); n, sample size; NRS-R-3, Neurobehavioral Rating Scale at 3 months (with square root transformation applied as described in the text); NRS-R-12, Neurobehavioral Rating Scale at 12 months (with square root transformation applied as described in the text); OR, odds ratio from ordinal and binary logistic regression; P, P-value for trajectory group (reference, low DNA methylation) based on an alpha of .05.
a Linear regression.
b Ordinal logistic regression.
c Binary logistic regression.
Suggestive association (P-value <.05).
Interaction Analyses
To explore potential interactions between age and DNAm, we used regression while controlling for age, sex, dichotomized GCS, trajectory group, and age × trajectory group interaction as described above. We observed suggestive associations for age × trajectory group interactions with patient outcomes at one CpG site (cg25962210). At cg25962210, two distinct trajectory groups were inferred (n = 75 [67.0%] low DNAm, n = 37 [33.0%] high DNAm) with no change over time as shown in Figure 2. Participant characteristics by inferred trajectory groups are presented in Table 4. No differences in participant characteristics were observed by trajectory groups. Detailed results for all CpG sites can be found in Supplementary Table 2. Trajectory plots for CpG sites cg26057780 and cg25328597 unadjusted for cell-type heterogeneity. Abbreviations: CSF, cerebrospinal fluid; TBI, traumatic brain injury. Participant Characteristics by DNA Methylation Trajectory Group for cg25962210. Abbreviations: GCS, Glasgow Coma Scale; SD, standard deviation. a T-test.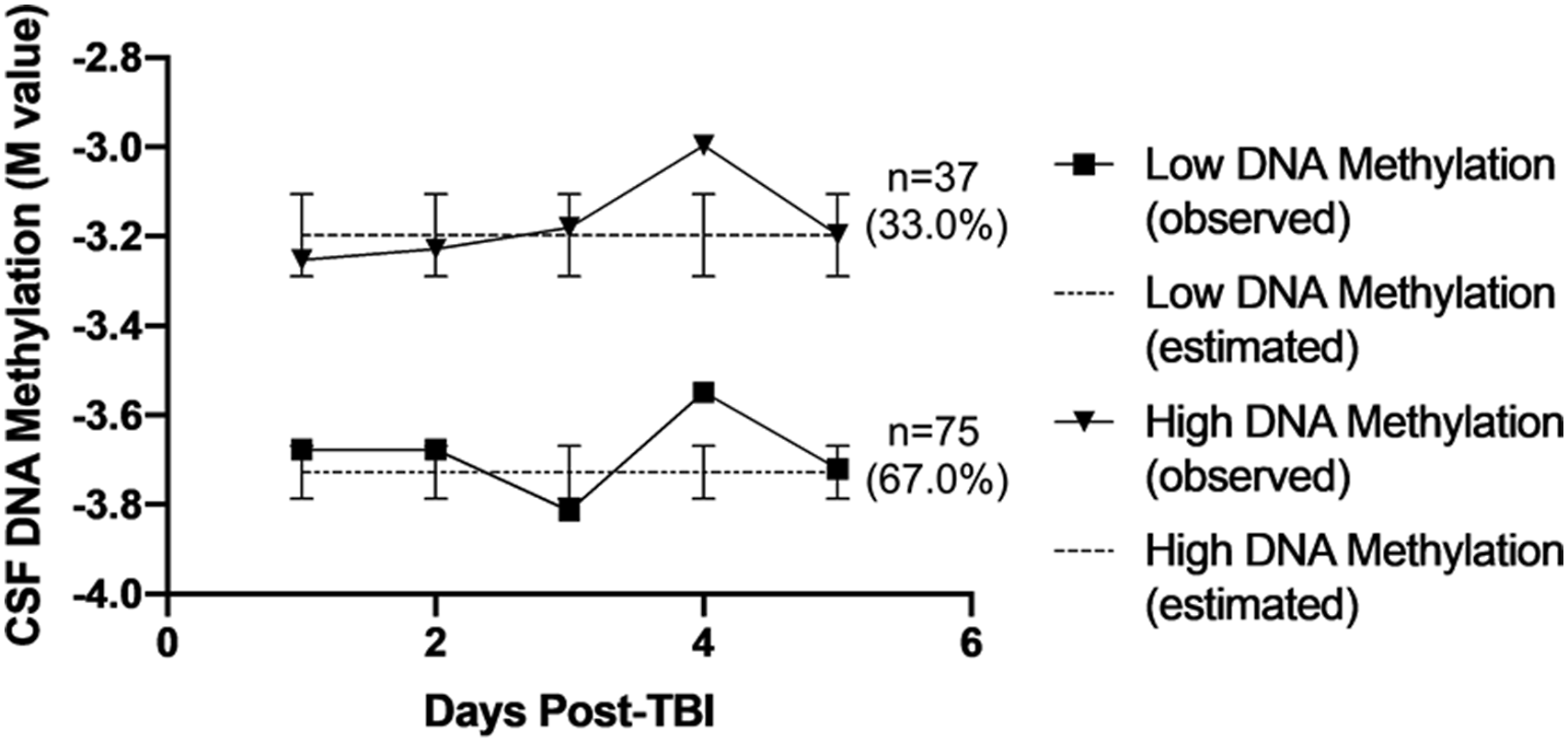
Results of Age–Trajectory Group Interaction Analyses While Controlling for Age, Sex, DNA Methylation Trajectory Group, and Dichotomized Glasgow Coma Score.
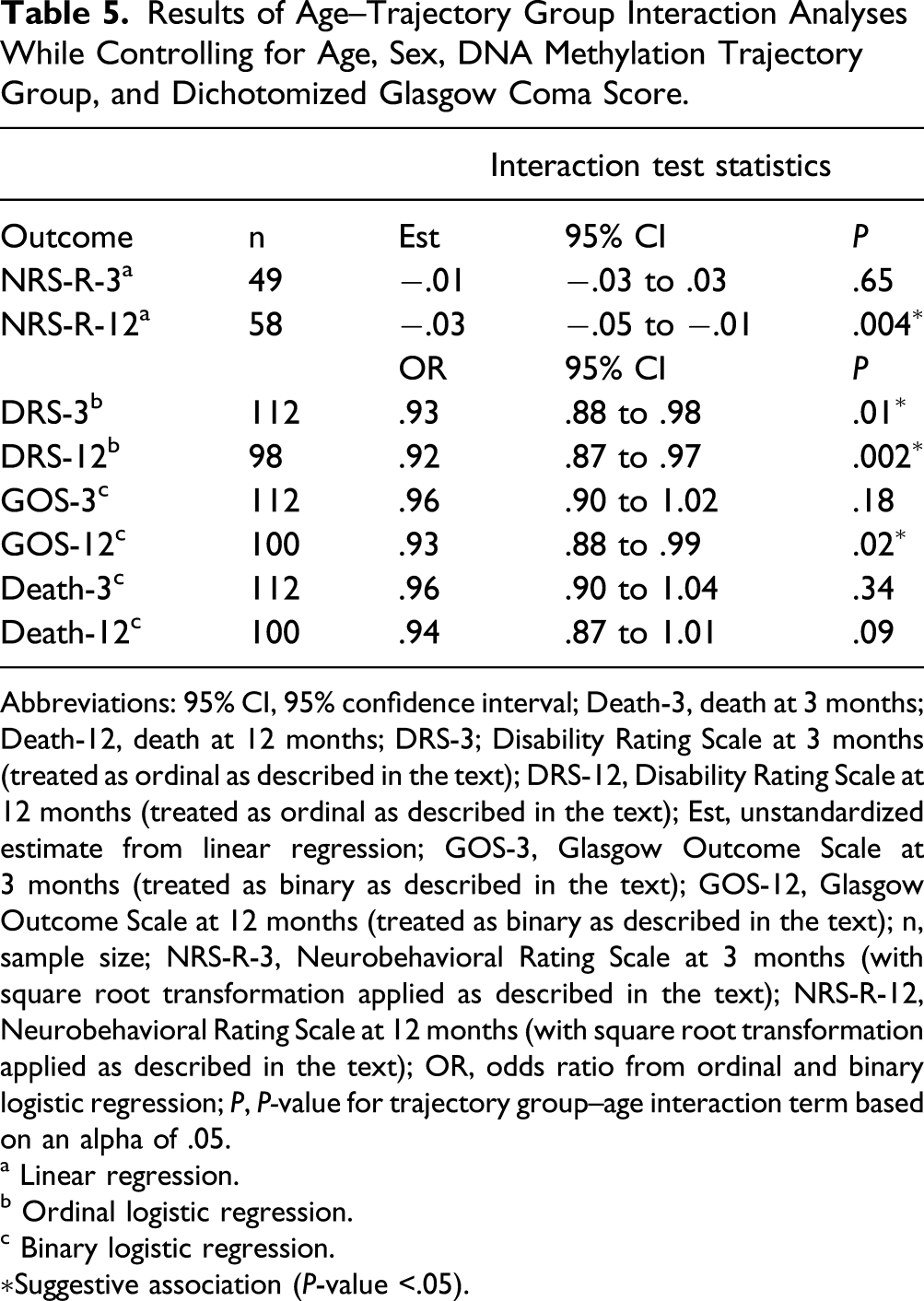
Abbreviations: 95% CI, 95% confidence interval; Death-3, death at 3 months; Death-12, death at 12 months; DRS-3; Disability Rating Scale at 3 months (treated as ordinal as described in the text); DRS-12, Disability Rating Scale at 12 months (treated as ordinal as described in the text); Est, unstandardized estimate from linear regression; GOS-3, Glasgow Outcome Scale at 3 months (treated as binary as described in the text); GOS-12, Glasgow Outcome Scale at 12 months (treated as binary as described in the text); n, sample size; NRS-R-3, Neurobehavioral Rating Scale at 3 months (with square root transformation applied as described in the text); NRS-R-12, Neurobehavioral Rating Scale at 12 months (with square root transformation applied as described in the text); OR, odds ratio from ordinal and binary logistic regression; P, P-value for trajectory group–age interaction term based on an alpha of .05.
a Linear regression.
b Ordinal logistic regression.
c Binary logistic regression.
Suggestive association (P-value <.05).
To visualize these interactions, an age × trajectory group interaction plot for NRS-R at 12 months is shown in Figure 3, and interaction plots for all outcomes are shown in the Supplementary Material. In general, participants in the high DNAm group had the same or better outcomes at higher ages while participants in the low DNAm group had worse outcomes at higher ages. Age × DNA methylation interaction plot for NRS at 12 months at CpG site cg25962210. Abbreviation: NSR-R, Neurobehavioral Rating Scale-Revised (square root transformed).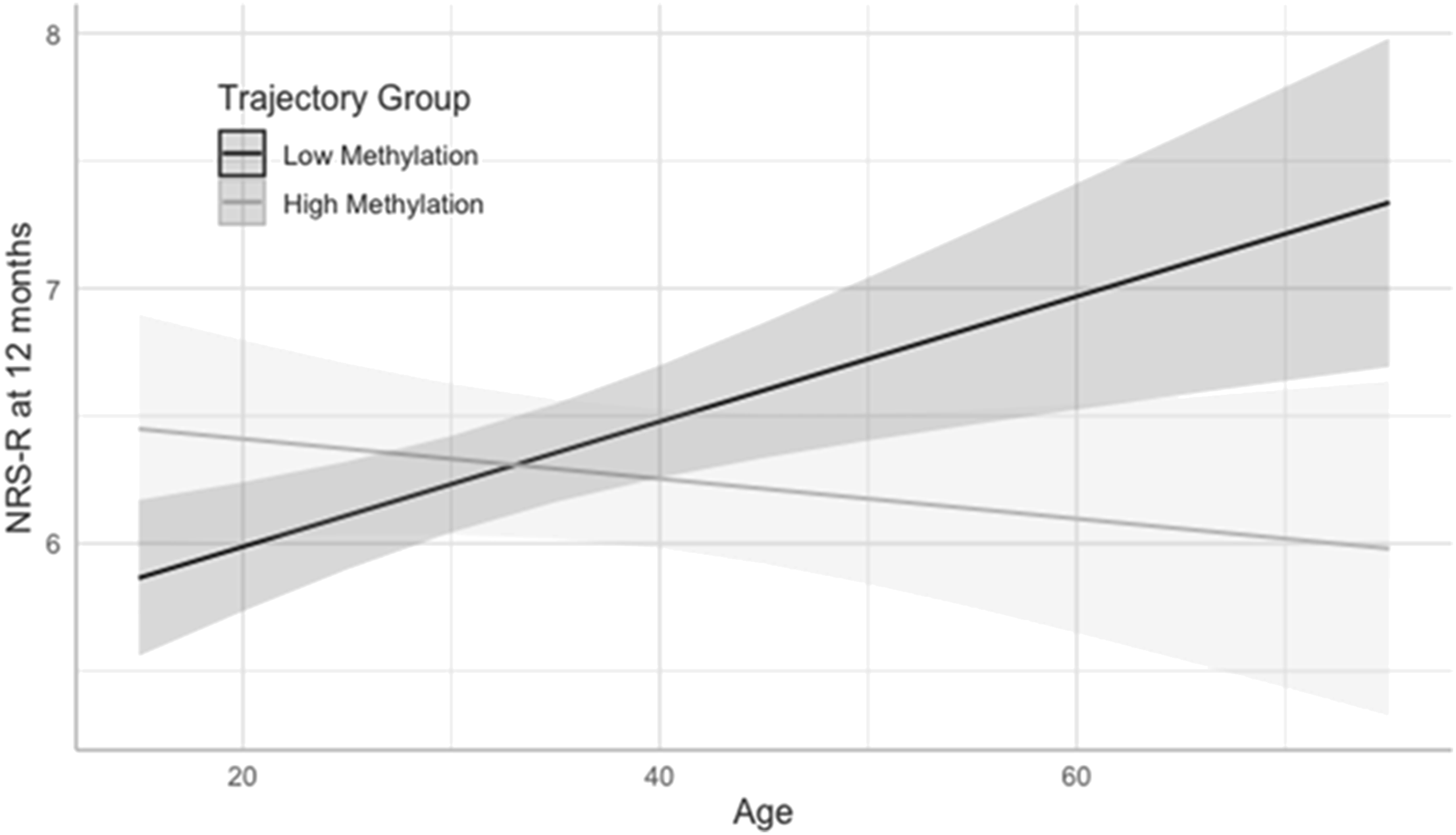
Discussion
The present exploratory pilot study characterized BDNF DNAm trajectories in CSF over the first 5 days following severe TBI in 112 adults and examined associations of inferred trajectory group membership with patient outcomes 3 and 12 months post-injury. Trajectory analysis revealed low and high DNAm groups at two BDNF CpG targets (cg26057780 and cg25328597) that showed suggestive associations (P < .05) with patient outcomes in models unadjusted for CTH. DNAm trajectories at these two sites were stable across the first 5 days post-injury. At both CpG sites, a small minority of participants had inferred group assignment in the high DNAm trajectory, and participants in this trajectory group were significantly older. Despite their older age, results showed suggestive associations between the high DNAm trajectory groups and better outcomes at 3 and 12 months relative to participants assigned to the low DNAm trajectory groups after controlling for age, sex, and injury severity (GCS). When we explored the moderating effect of age, a third CpG site showed suggestive associations with patient outcomes in models unadjusted for CTH. Specifically, at cg25962210, we observed a similar trajectory pattern to that described above of low and high DNAm trajectories with no change over time. Associations of age × trajectory group interactions with outcomes revealed a relatively consistent pattern of the same or better outcomes with higher age in the high DNAm group and worse outcomes with higher age in the low DNAm group. Although no observed associations met the empirical significance threshold after correcting for multiple comparisons, suggestive associations of the main effect models were consistent in their direction of effect and were observed across two CpG sites and two outcome time points. No associations were identified with patient outcomes in trajectory models adjusted for CTH.
The results suggest that during the acute phase following severe TBI in adults, higher BDNF DNAm in CSF, which is assumed to be associated with lower BDNF expression 15 and neuronal synthesis, 16 may promote recovery; and conversely, lower BDNF DNAm, with presumably higher BDNF expression and synthesis, may hinder recovery. Further, our results suggest that this effect may be more robust with higher age at a third suggestive CpG site. The findings are generally consistent with prior BDNF protein level and genetic studies in cohorts of adults with moderate to severe TBI. Relative to non–brain-injured controls, some studies have shown that BDNF levels during the first week following TBI in adults are higher in CSF18,35 and significantly lower in serum.18,36 Along the same lines, CSF and serum BDNF levels are marginally negatively correlated. 18 Higher acute BDNF levels in CSF, and lower acute and chronic levels in serum/plasma, have been associated with poorer outcomes, including mortality, 18 global functioning,36,37 and long-term functional cognition and depression.35,38 Similarly, initial genetic studies have shown a protective effect of the BDNF Val66Met Met allele (associated with reduced activity-dependent secretion of BDNF) on outcomes following moderate to severe TBI.39,40
Together, these emerging BDNF biomarker studies demonstrating poorer outcomes with greater BDNF action in the CNS suggest that the apoptotic action of pro-BDNF predominates over the survival signaling of mature BDNF in adults with TBI. The risk conferred by greater BDNF action, however, may be moderated by age, possibly related to reductions in BDNF levels 41 and regional balances of BDNF receptor ratios across the lifespan,42,43 as has been previously suggested.17,18 It is interesting that the detrimental effect of higher BDNF action found in adults with TBI contrasts with most evidence in non–brain-injured populations in which lower BDNF action appears to confer risk for psychiatric and neurological conditions.44–56 This suggests brain-injury–specific involvement of BDNF, possibly related to shifts in BDNF receptor balances in response to injury.17,18,57
Regarding location and functionality of our suggestive CpG sites, all three are annotated to a BDNF promoter region (GRCh37/hg19, depending on the transcript of interest: cg26057780, chr11:27721277 north shore in gene body, 5’ untranslated region [UTR], or transcription start site [TSS]; cg25328597, chr11:27722638 CpG island in gene body, 5’UTR, or TSS; cg25962210, chr11:27721222 north shore in gene body, 5’UTR, or TSS). Examining the functionality of these CpG sites is outside the scope of the present study, and reviewing prior literature for functionality of specific CpG sites is challenging as specific CpG sites are inconsistently included in searchable content (e.g., title, abstract, and keywords) in literature databases. Nevertheless, we found several studies reporting significant associations of our three suggestive CpG sites with brain-related outcomes or correlates. DNAm in blood at both cg25328597 and cg25962210 was positively associated with age in a cohort of low-income, primarily African American adults. 58 In another study, DNAm in blood at cg25328597 was differentially methylated in pre-symptomatic dementia cases versus controls. 59 In a third study, DNAm in placenta at cg25962210 was positively associated with maternal exposure to war trauma and maternal chronic stress. 60 Finally, using the match algorithm and TRANSFAC matrices in geneXplain (genexplain.com), we identified putative transcription binding sites at each of our three suggestive CpG sites (see Supplementary 3), suggesting that these sites may be functionally relevant for gene transcription. More specifically, our third CpG site at which we observed a suggestive interaction of age × trajectory group is situated at a binding site for transcription factor MeCP2, which is a known transcriptional regulator of BDNF, affecting BDNF-induced myelin repair mechanisms. 61
Our study selected CSF as the tissue of interest. Because DNAm patterns are tissue-specific and reflective of the local environment of each cell type, and examination of brain tissue is not feasible in patients who survive their injuries, determining which types of tissues serve as effective proxies for the CNS environment is an important methodological consideration. Several studies have shown that DNAm in peripheral tissues, including blood and saliva, is correlated with DNAm in brain tissue or CSF,62–64 for several target genes, including BDNF.65,66 To our knowledge, however, no studies have investigated concordance of DNAm in CSF and the brain. Although it is difficult to make inferences about the brain itself without such understanding, to be useful as a clinical biomarker, it is also essential that markers be found in readily accessible tissues or cells. Cerebrospinal fluid is often accessible in individuals with severe TBI via collection from an EVD placed for intracranial pressure monitoring and CSF diversion as standard of care. However, because the blood brain barrier is disrupted during injury, CSF from individuals with TBI is likely to contain blood, further exacerbating CTH. The influence of CTH in CSF DNAm remains understudied and is likely of critical importance in CSF, in particular, because of injury- and recovery-specific temporal changes. An interesting observation in this study was that, once we controlled for CTH during GBTA, trajectory groups were inferred with high accuracy at a larger number of sites compared with the unadjusted analyses (i.e., 39 vs 32; see Supplementary Table 1). Further, a majority of these CTH-adjusted models included only one group and, of the three carried forward for patient outcome association testing, no associations were identified. These observations suggest that CTH is likely an important factor to variability of CSF BDNF DNAm and should be considered in future studies. Of note, however, the examination of CTH is only possible in genome-wide investigations and not in targeted DNAm studies.
Limitations
Several important limitations of the present study should be considered. First, because of the relatively small sample size of the present pilot study, we were unable to examine potential effects of injury heterogeneity such as brain regions affected and type of injury (e.g., diffuse axonal injury and subdural hematoma) on the results. Next, by selecting DNAm targets within the relatively narrow window of ±5000 kb upstream and downstream of the BDNF gene region, we could have missed potentially important regulatory regions of the gene. In addition, BDNF genotype, which has been inconsistently shown to modulate BDNF DNAm,47,67–69 was not examined. We focused our exploratory pilot study on epigenetic variability in BDNF to the exclusion of other potentially relevant candidate genes for TBI recovery. Interpretability of the results is limited without considering interactions with a larger genetic network. Future investigations examining the effects of injury heterogeneity, expanding the region of interest to include additional CpG sites, examining whether BDNF genotype interacts with DNAm in affecting outcomes, and exploring the epigenetic effects of other candidate genes are warranted. Although, to our knowledge, this is the largest clinical study of DNAm following TBI to date, the sample size was relatively small. Relatedly, the interpretability of the results is limited by the large number of tests conducted. It is imperative that this study be replicated in a larger sample to confirm these findings. Finally, race was self-reported by study participants, and use of this variable limited the ability to stringently control for population substructure. Moreover, our sample was homogeneous with all participants reporting white race. Thus, the results may not generalize to individuals with non-white ancestries. Future efforts are needed to replicate these findings in diverse samples.
Conclusion
The investigation of epigenetic biomarkers has the potential to explain outcome heterogeneity following TBI. Results of the present exploratory pilot study suggest that patients’ trajectory of BDNF DNAm in CSF over the first 5 days following severe TBI may be associated with short- and long-term outcomes over and above traditional models accounting for age, sex, and injury severity. While the present results require replication in larger and more racially diverse independent samples, BDNF DNAm from accessible biosamples may serve as an early postinjury biomarker that more accurately predicts TBI outcomes, thereby allowing for earlier identification of patients at greatest risk for poor recovery and the earlier provision of targeted intervention.
Supplemental Material
sj-pdf-1-nnr-10.1177_15459683211028245 – Supplemental Material for Acute Brain-Derived Neurotrophic Factor DNA Methylation Trajectories in Cerebrospinal Fluid and Associations With Outcomes Following Severe Traumatic Brain Injury in Adults
Supplemental Material, sj-pdf-1-nnr-10.1177_15459683211028245 for Acute Brain-Derived Neurotrophic Factor DNA Methylation Trajectories in Cerebrospinal Fluid and Associations With Outcomes Following Severe Traumatic Brain Injury in Adults by Amery Treble-Barna, Lacey W. Heinsberg, Ava M. Puccio, John R. Shaffer, David O. Okonkwo, Sue R. Beers, Daniel E. Weeks and Yvette P. Conley in Neurorehabilitation and Neural Repair
Supplemental Material
sj-pdf-2-nnr-10.1177_15459683211028245 – Supplemental Material for Acute Brain-Derived Neurotrophic Factor DNA Methylation Trajectories in Cerebrospinal Fluid and Associations With Outcomes Following Severe Traumatic Brain Injury in Adults
Supplemental Material, sj-pdf-2-nnr-10.1177_15459683211028245 for Acute Brain-Derived Neurotrophic Factor DNA Methylation Trajectories in Cerebrospinal Fluid and Associations With Outcomes Following Severe Traumatic Brain Injury in Adults by Amery Treble-Barna, Lacey W. Heinsberg, Ava M. Puccio, John R. Shaffer, David O. Okonkwo, Sue R. Beers, Daniel E. Weeks and Yvette P. Conley in Neurorehabilitation and Neural Repair
Supplemental Material
sj-tif-3-nnr-10.1177_15459683211028245 – Supplemental Material for Acute Brain-Derived Neurotrophic Factor DNA Methylation Trajectories in Cerebrospinal Fluid and Associations With Outcomes Following Severe Traumatic Brain Injury in Adults
Supplemental Material, sj-tif-3-nnr-10.1177_15459683211028245 for Acute Brain-Derived Neurotrophic Factor DNA Methylation Trajectories in Cerebrospinal Fluid and Associations With Outcomes Following Severe Traumatic Brain Injury in Adults by Amery Treble-Barna, Lacey W. Heinsberg, Ava M. Puccio, John R. Shaffer, David O. Okonkwo, Sue R. Beers, Daniel E. Weeks and Yvette P. Conley in Neurorehabilitation and Neural Repair
Footnotes
Acknowledgments
We thank Dr Ansuman Chattopadhyay and the University of Pittsburgh Health Sciences Library System Molecular Biology/Bioinformatics Information Service for assistance obtaining information contained in ![]() . GeneXplain software licensed through the Molecular Biology Information Service of the Health Sciences Library System, University of Pittsburgh, was used for data analysis for Supplementary 3.
. GeneXplain software licensed through the Molecular Biology Information Service of the Health Sciences Library System, University of Pittsburgh, was used for data analysis for Supplementary 3.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Research reported in this publication was supported by the National Institutes of Health under Award Numbers R21NR015142 (YPC), P50NS030318 (DOO), K01HD097030 (ATB), F31NR017311 (LWH), and TL1TR001858 (LWH). The content is solely the responsibility of the authors and does not represent the official views of the National Institutes of Health. No competing financial interests exist.
Supplemental Material
Supplementary material for this article is available on the Neurorehabilitation & Neural Repair website along with the online version of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
