Abstract
High-throughput screening (HTS) approaches incorporating multiplexed, cell-based assays are increasingly common for identifying novel modulators of complex biological processes. To enable the discovery of small molecule modulators of mRNA processing, we developed a multiplexed, bead-based, high-throughput QuantiGene assay, leveraging the Luminex platform that is capable of simultaneously quantifying transcript levels of multiple independent target genes within a single HTS campaign. To address plate variability and potential screening artifacts, a pragmatic hit-calling pipeline was implemented utilizing both plate- and well-based normalization strategies. This dual-normalization approach reduced false negatives and produced consistent hit confirmation rates. Application of this methodology led to the identification of unique compounds selectively modulating individual target genes. Strikingly, among the three exemplary genes, only 5% of primary actives demonstrated activity across all three target genes, underscoring the assay’s capacity for detecting selective mRNA modulators. Chemical motif analysis of confirmed actives recovered known RNA privileged scaffolds as well as novel scaffolds that are uniquely enriched for individual targets screened. Validation screening using an orthogonal, four-point concentration-response real-time PCR (qPCR) assay in a disease-relevant cell line demonstrated high validation rates, supporting the robustness and translational relevance of this multiplexed HTS platform. These findings establish a scalable and reliable strategy for identifying selective small molecule modulators of mRNA processing, with broad applicability in early drug discovery.
INTRODUCTION
High-throughput screening (HTS) is a pillar of modern drug discovery, used to identify novel chemical matter that can initiate drug development campaigns either as a standalone approach or in conjunction with complementary strategies such as virtual screening. 1 In HTS, a wide array of in vitro technologies is available, which can broadly be classified as either biochemical or cell-based assays. 2 While biochemical assays provide direct insight into on-target activity, cell-based assays offer information on compound-induced functional or phenotypic responses. Notably, cell-based screens often require downstream target validation and deconvolution after the primary screen. An advantage of implementing cell-based methods is the option to multiplex and extract data on multiple targets simultaneously in an endogenous setting.
In a typical cell-based HTS campaign, compounds are initially tested at a single concentration. Hit compounds are identified by comparing their performance to controls and the rest of the compound library. These primary actives then undergo confirmation screens and are further validated using orthogonal assays that incorporate concentration-response evaluations. 3 Various statistical techniques have been developed to characterize hits from HTS campaigns, with methods such as z-score and robust z-score transformations being widely used for data normalization and hit selection.4–6 Robust z-scores, in particular, are effective in identifying actives from “hot” plates containing numerous active compounds. Another useful metric is the strictly standardized mean difference (SSMD), which quantifies the difference between treatment groups and negative controls.7,8
Advances in automation, miniaturization, and the integration of multiplexed readouts have considerably improved HTS methodologies. 9 These improvements increase throughput, reduce costs, and mitigate experimental errors. For example, fluorescence-based screens now deliberately include compounds known to cause interference—due to color, inherent fluorescence, or aggregation propensity—to flag potential false positives.10,11 Alongside these practical measures, established statistical methods, such as the Z’-factor and titration experiments, are often used to quantify assay robustness and sensitivity.12–14
The reliable identification of active compounds remains the central challenge in HTS. Assay noise and systematic variability stemming from technical glitches, procedural differences, or reagent inconsistencies can compromise data normalization and hit calling. Several methods have been introduced to address these issues, including the B-score method, which employs Tukey’s median polish algorithm to correct for positional effects on screening plates.12–15 This B-score approach standardizes the observed values by the median absolute deviation along rows and columns, trying to reduce spatial artifacts and improve the hit calling confidence. 13 In an HTS, failing to account for assay noise and variability can lead to a high false-positive rate, wasting resources on compounds that do not reproduce upon repeat testing, or worse, lead to a high false-negative rate, potentially overlooking promising leads.
This study utilized a Luminex QuantiGene Plex Gene Expression Assay, a bead-based platform capable of simultaneously monitoring the expression levels of multiple mRNA targets in a high-throughput format.16,17 This Luminex platform has been widely utilized primarily within the 96-well format, which remains the industry standard. The application of Luminex in a 384-well format, particularly at the scale employed in this study, is a significant advancement. To our knowledge, such large-scale applications of this technology have not been previously reported. This adaptation pushes the boundaries of Luminex technology and offers distinct advantages in multiplexed gene expression assays for HTS, conserving time and resources by enabling the parallel measurement of the expression levels of several genes, i.e., 12 in this article. We also present a pragmatic hit calling workflow that accounts for both systematic and random variability observed in compound activity and gene expression measurements. Our method involves a cluster-first approach, in which plates with similar positional effects are grouped together prior to a well-level normalization and z-score transformation for subsequent hit calling. This strategy ensures more accurate data normalization, preventing the inadvertent elimination of promising hits. The readouts across the exemplified target genes are not collinear as presented in this report. This led to the identification of novel and selective small molecule modulators of mRNA processing, evident in the uniquely active compounds discovered, and the unique scaffolds retrieved in the screen.
MATERIALS AND METHODS
Luminex QuantiGene™ Plex HTS
A large, cryopreserved stock of assay-ready HEK293T cells (CRL-3216, ATCC) was generated for this high-throughput screen. Cells were revived for 48 h and plated at 8,000 cells per well in 384-well plates. Compound treatments (10 µM final concentration) and vehicle controls (dimethyl sulfoxide [DMSO]) were dispensed using an ECHO acoustic liquid handler (Beckman Coulter). After 24 h of treatment, gene expression was measured using the QuantiGene™ Plex 384-well assay (Thermo Fisher Scientific), adapted for HTS with a “plex-on-plex” pooling method. The plex-on-plex approach involved pooling of spectrally unique bead sets targeting the same gene panel from two separate 384-well assay plates to increase readout throughput. Target probes were custom-designed to capture the target-specific transcripts selectively (Thermo Fisher Scientific). Fluorescence was quantified using the Luminex xMAP Intelliflex system.
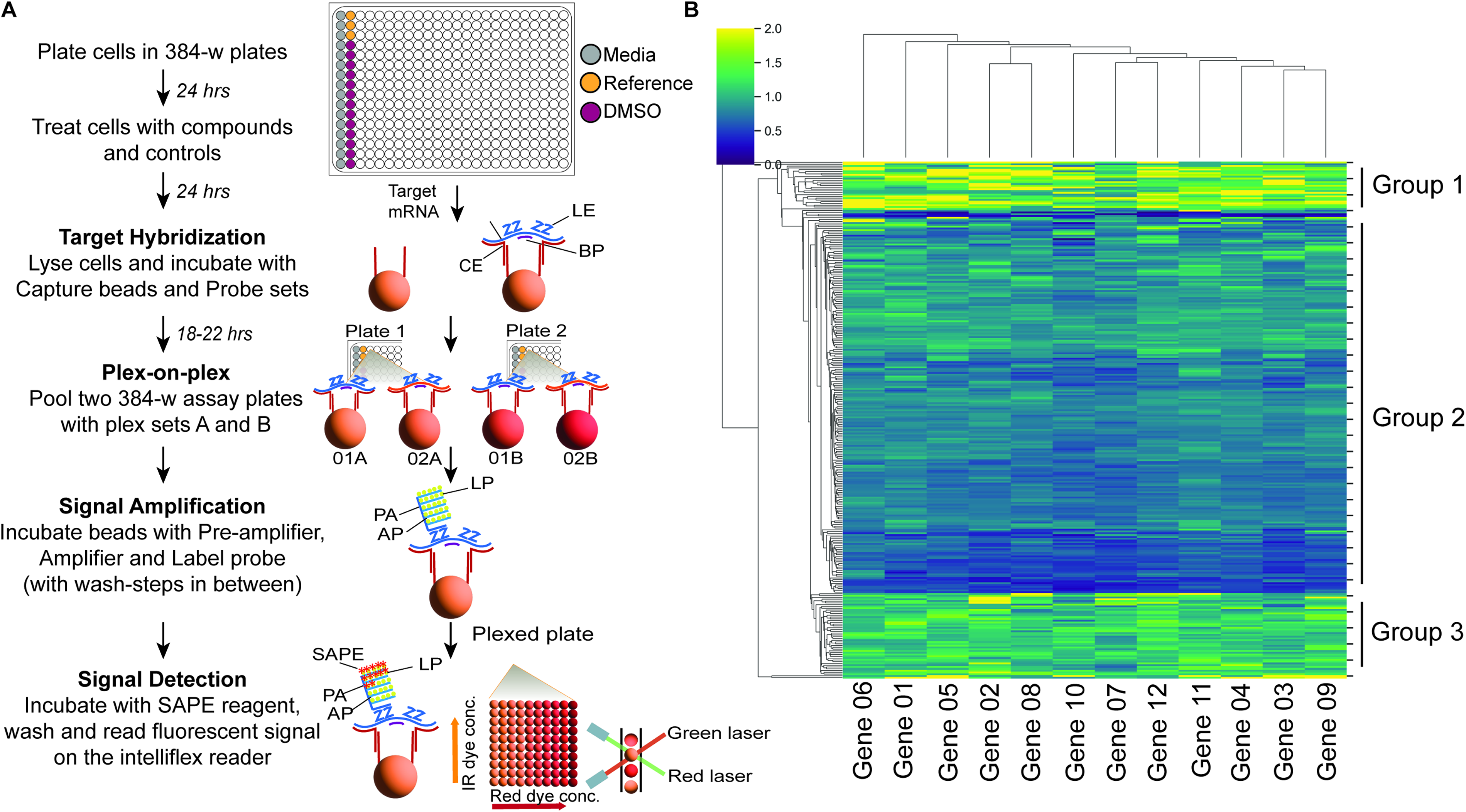
Media-only wells served as positive controls, and DMSO-treated cells as negative controls. A reference compound was included on each plate to monitor inter-plate variability. Gene expression data were normalized using the mean fluorescence intensity (MFI) of the housekeeping genes GUSB and HPRT1, which were selected for their consistent coefficient of variation (CV) and comparable MFI levels to target genes. MFI normalization was performed by dividing each target gene’s MFI by the geometric mean MFI of the housekeeping genes in the same well. Both the primary (single replicate) and confirmation (duplicate) screens were performed using this workflow, as illustrated in Figure 1A and detailed in Supplementary Table S1.
qPCR Validation Screen
Target-specific high-expressing cell lines were treated with confirmed active compounds in a four-point concentration response (30, 15, 7.5, and 3.75 µM) alongside vehicle controls (DMSO), using an ECHO acoustic liquid handler (Beckman Coulter). After 24 h, cells were lysed, and crude lysates were used directly for qPCR. TaqMan qPCR assays were performed using primers and probes targeting conserved regions of the specific mRNA transcript, designed (using PrimerQuest™) and synthesized by IDT. Probes were dual labeled with reporter and quencher dyes and multiplexed with a housekeeping gene. Reactions were run on the QuantStudio™ 7 Pro Real-Time PCR system and analyzed using QuantStudio Design and Analysis Software v2.6.0 (Thermo Fisher Scientific). The data analysis was done by first determining the ΔCt vs the housekeeper gene. This ΔCt was then normalized against the DMSO control (ΔΔCt) and converted to relative quantification using the 2−ΔΔCt equation. The response was then fitted to the four parametric logistic equations to evaluate the concentration dependence of compound treatment.
Chemical Library
In this screening effort, a subset of the proprietary Remix small molecule chemical library containing 115,249 compounds representing the overall diversity of the collection was screened. The Remix library is a custom-built screening collection of small molecules assembled to survey features of chemical space targeting RNA, RNA-binding proteins, RNA-protein complexes, and upstream proteins important in RNA processing. Notably, approximately 15% of the library is comprised of compounds synthesized exclusively in-house, thereby extending beyond the chemical space represented in commercial libraries. Additionally, the Remix library adheres to widely accepted lead-like property space descriptors, including molecular weight, polar surface area, and a fraction of sp3 hybridized carbon atoms.18–20 Finally, the library was curated to exclude compounds containing potentially problematic substructures and undesirable functional groups. 21
Quality Control Methods
Quality control (QC) plays a crucial role in ensuring the accuracy and reliability of HTS data analysis.4,7,15,22–25 In our screening process, each 384-well plate was outfitted with 16 positive and 13 negative control wells. The positive controls (media) were used to mimic conditions in which the target gene mRNA is completely depleted, while the negative controls (DMSO) reflected baseline levels of the target gene mRNA in the absence of compound treatment. QC metrics were implemented using the Python programming language (Python Software Foundation, https://www.python.org/). High-level descriptions of the QC metrics implemented here can also be found in the Supplementary Data.
Hit Calling
Following the primary screen, data were pre-processed and normalized to correct for systematic variability. Fold change in target gene mRNA levels relative to DMSO controls was calculated for each compound. Two normalization strategies were applied: plate-based and well-based. In the plate-based method, z-score normalization was applied across compounds within each 384-well plate. For well-based normalization, plates with similar well position patterns were clustered using k-means clustering, and z-score normalization was applied to the same well positions across the plates in each cluster.
Compounds with a z-score ≤−3 in either normalization scheme were classified as primary actives, indicating a significant reduction in mRNA expression relative to the total compound collection screened. These primary actives were re-tested in duplicate in biologically distinct samples. Confirmed actives were defined as compounds that reproducibly caused ≥20% mRNA reduction. Confirmed actives were further validated in orthogonal four-point qPCR assays. Compounds showing a concentration-dependent decrease in mRNA levels upon visual curve inspection were designated as validated hits.
RESULTS AND DISCUSSION
Luminex QuantiGene™ Plex HTS
We adopted the Luminex QuantiGene gene expression assay system to measure the impact of compound treatment on the mRNA levels of endogenously expressed genes to identify RNA processing modulators. This cell-based system encompasses the full complement of RNA processing machinery and enables simultaneous multiplexed readouts. The assay workflow, illustrated in Figure 1A, was implemented to profile mRNA levels of 12 independent target genes. Two spectrally distinct fluorophore-labeled beads per target were used (plex sets A and B), allowing samples from two 384-well plates to be pooled and assayed concurrently for increased throughput.
We observed comparable MFIs for the two plex sets, A and B, across the 12 tested genes (Supplementary Fig. S1). mRNA levels were measured after a 24-hour treatment period at a 10 μM concentration using HEK293T cells. Fluorescent bead counts were classified based on the internal dye of each bead, allowing differentiation among target-specific beads. Target-specific probe sets with signal amplification through branched chain DNA (bDNA) provided specificity and a good fluorescence signal with low background. MFIs for each compound treatment were calculated and then normalized to the housekeeper genes from the corresponding sample well. Finally, the fold change was determined relative to the negative DMSO control wells on the same plate. The fold change distributions before and after normalization behaved as expected: One notable observation is that compound treatment causes minimal changes in the mRNA fold changes of housekeeping genes compared to the tested target genes, as evidenced by narrower fold change distributions for the housekeeping genes relative to those of the target genes (Supplementary Fig. S2).
Visual inspection of the Z/Z’ factor values across the 12 tested mRNAs allowed categorization of the primary HTS plates into three groups (Fig. 1B). In Group 1, control samples were more variable (Z/Z’ > 1). In Group 2, compound treatment samples showed increased variability (Z/Z’ < 1). In Group 3, control and compound treatment samples exhibited similar variability (Z/Z’ = 1). As anticipated, many of the plates in our study fell into Group 2—the middle, large cluster in the heatmap—indicating higher variability in compound treatment compared to control wells.
Assay Quality Control
Several QC metrics—outlined in the Supplementary Data—were monitored for every plate and gene combination (Fig. 2). The box plots indicate that these parameters are within the expected ranges for each target. For instance, the signal-to-background (S/B) window ranged from 0.9 to 1 for the tested genes, which is adequate for discriminating true positives from false positives arising from non-specific or background effects. However, the S/B values for some genes (01, 06, 08, and 11) were more variable and lower.
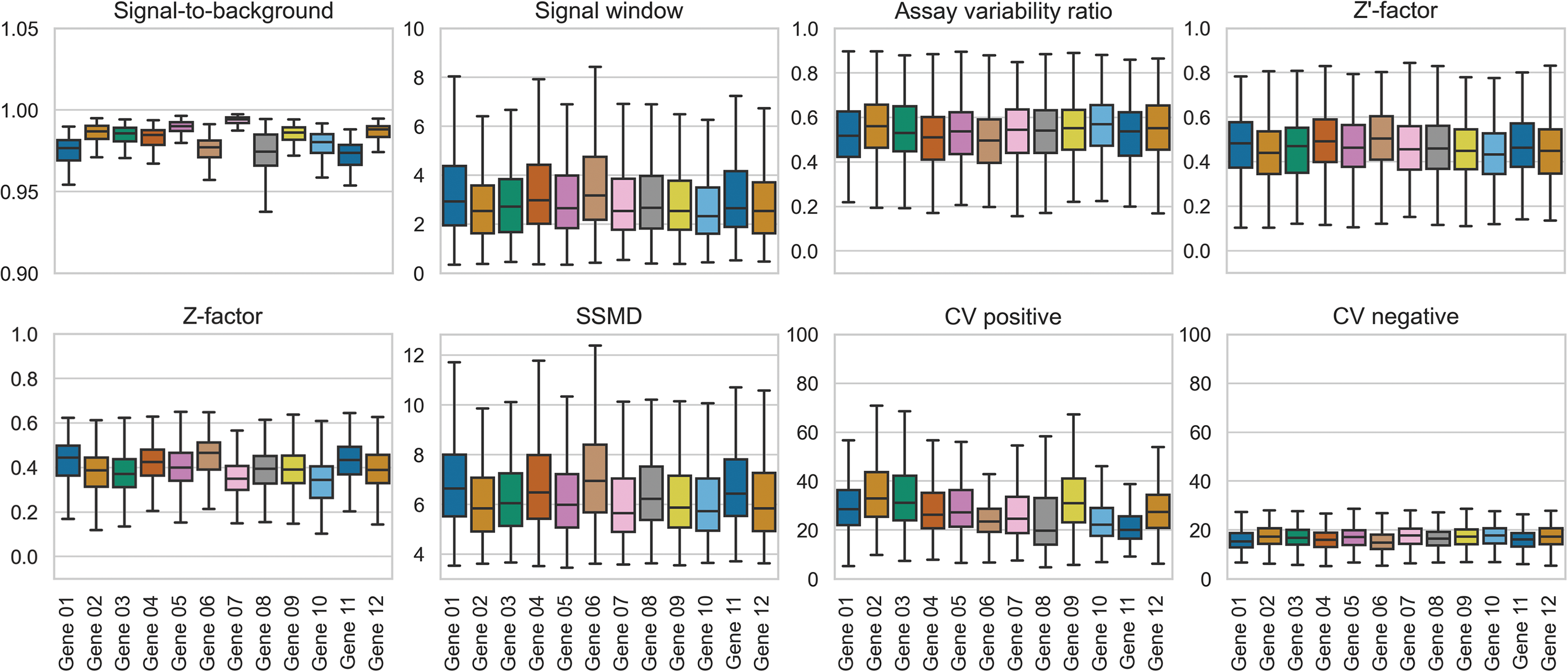
Visual distribution of HTS quality control parameters derived from target gene mean fluorescence intensity (MFIs): The signal-to-background (S/B) shows the ratio of the mean signal of the positive vs. negative control wells. Higher values indicate a better window to identify active compounds. The signal window (SW) reflects the assay’s dynamic range; a larger SW indicates better discrimination between active and inactive samples. The assay variability ratio (AVR) quantifies variability in positive and negative controls relative to their means, with lower AVR values indicating higher reproducibility. Z’-factor incorporates both means and standard deviations of positive and negative controls to evaluate assay robustness, where values closer to 1 indicate higher reliability. Modified Z-factor was also calculated using positive control and all compound-treated wells to assess overall screen quality. The strictly standardized mean difference (SSMD) represents the effect size between the positive and negative controls in units of standard deviation, where higher values reflect stronger signal separation. Coefficients of Variation (CV) for both positive and negative controls provide a measure of signal consistency, with lower values indicating less variability.
Over half of the HTS plates showed a robust signal window (SW) of three or greater, with over 60% displaying an SW of at least two. The average assay variability ratio (AVR) across the tested genes was below 0.6. Although this value is slightly higher than desired, it remains acceptable given the highly multiplexed nature of the HTS. Z’-factors, commonly used in assay development to quantify quality, generally indicate a robust assay when above 0.7. In our screen, the median Z’-factor ranged between 0.43 and 0.51, while some plates achieved values as high as 0.78–0.83. Plates displaying consistently low Z’-factors across multiple genes were re-run to enhance data quality and the reliability of hit calling.
In general, we observed SSMD scores of at least 5, which effectively separated the negative and positive control samples. The CV—a measure of the relative variability in the control readouts—was also closely monitored, where lower CV values indicate that the data are tightly clustered around the mean, suggesting greater precision and accuracy in the control measurements used to normalize compound treatment data. Our target gene panel and the housekeeping genes—which were selected during the assay optimization stage based on their comparable expression level to the 12 genes of interest—exhibited a robust gap between the negative and positive control samples (Supplementary Fig. S3). However, a subset of genes—specifically genes 02, 03, and 09—demonstrated increased variability in their positive control CVs. The MFI distributions for both control groups corroborate the general trends observed from the CV parameters (Supplementary Fig. S3). Although the primary HTS encompassed target Genes 01–12, for clarity and succinctness, the remainder of the work presented here is focused on Genes 01–03. Genes 04–12 remain high-value targets for future drug discovery efforts.
Data Normalization and Hit Calling
HTS data analysis and hit calling can be confounded by both plate- and well-position effects. These artifactual readouts can exhibit as alternating high—low signal rows or columns. Another common plate effect is elevated or reduced signals at the plate edges. These artifacts are often readily visible by plotting the data in a plate format. In traditional HTS campaigns, plates exhibiting obvious artifacts are candidates for re-screening. However, in multiplexed screens, the degree of plate effects can vary across measured readouts. For instance, a noticeable plate effect seen for one target gene may not be present for others, such as the example plates for target Genes 01–03 (Supplementary Fig. S4). In this study, fold change readouts were first clustered separately for each measured gene (Supplementary Fig. S5), revealing distinct, position-specific patterns. Based on these patterns, plates were grouped according to well-position effects, and normalization was performed to account for plate-to-plate variability.
For target Genes 01–03, primary active compounds were determined by applying the z-score threshold and categorized into one of three normalization categories: Primary actives meeting the z-score threshold using (1) both well- and plate-based normalization (labeled as both plate/well-navy), (2) those identified only by plate-based normalization (labeled as plate only-green), and (3) lastly those identified only by well-based normalization (labeled as well only-yellow). For target Genes 01–03, the number of primary active compounds was compared across these 3 categories (Fig. 3). Approximately 47%–55% of the primary active compounds met the hit-calling threshold after applying both the well- and plate-based methods. The plate-based and well-based categories made up an additional 17%–22% and 27%–33% of primary active compounds, respectively.
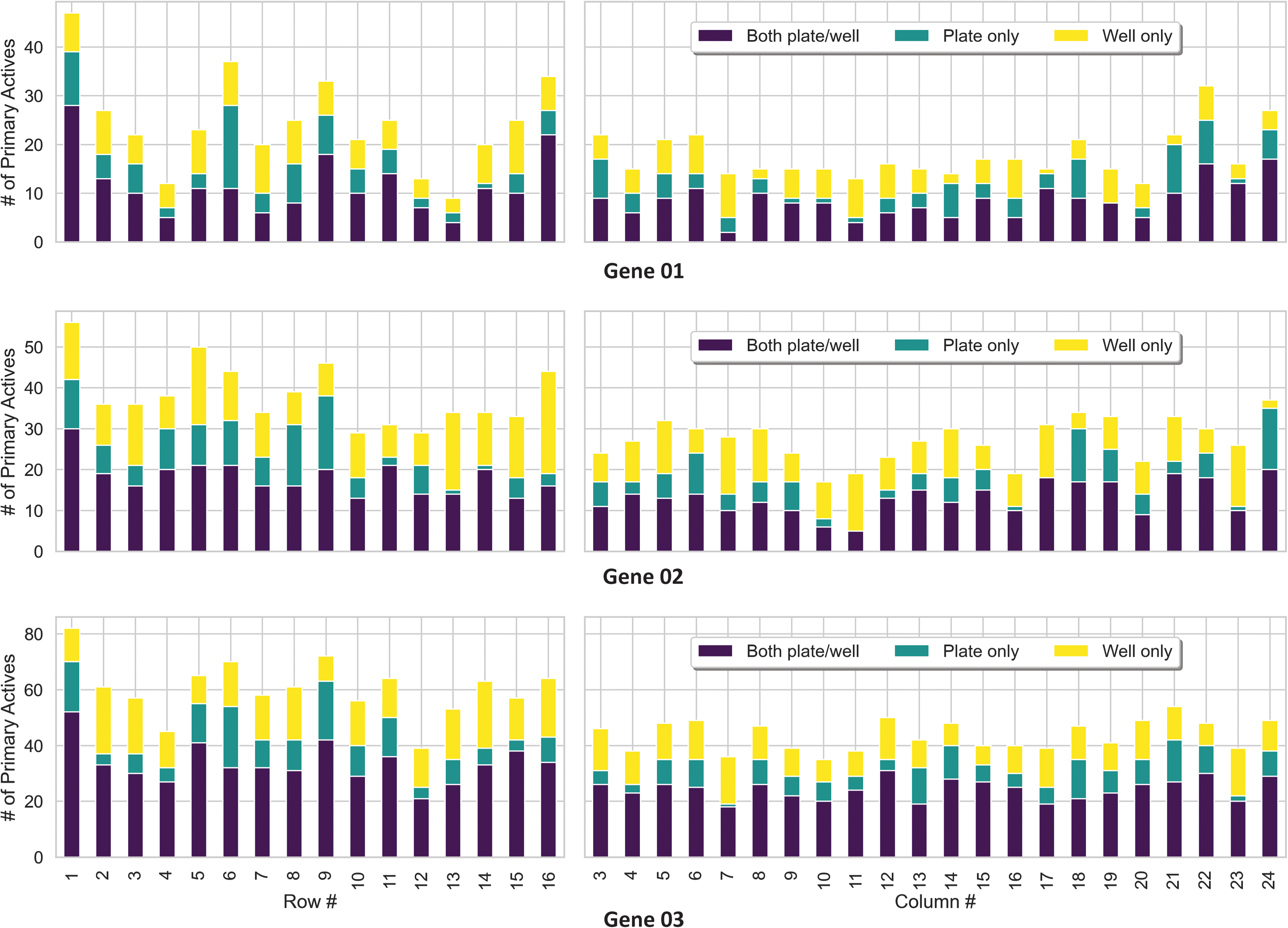
For Genes 01–03, a visual representation of the primary active distribution across the HTS plate row and column positions. Colors encode whether the hit-criteria were met using both plate- and well-based normalization (navy), plate-based normalization only (green), and well-based normalization only (yellow). Many active compounds are identified across the rows and columns using both plate and well-based normalization. Well-only normalization complements the plate-only normalization analysis by rescuing actives in rows/columns that could be missed by plate-only level analysis. This is particularly impactful when plate-level normalization yields few additional hits, as observed for Gene 02 in columns 11, 12, 16, 17, and 23.
While the primary actives identified by both plate and well-based normalization methods are more potent. The well-based category captures primary actives with fold change profiles comparable to those obtained with the plate-based approach (Fig. 4A). This highlights the importance of employing multiple normalization schemes during HTS data analysis to minimize false negatives. The mean fold change values for the primary actives across Genes 01–03 were 0.62 ± 0.10, 0.62 ± 0.13, and 0.49 ± 0.10, respectively. The primary hit rates for these genes were 0.81%, 1.08%, and 1.71%, respectively (Supplementary Table S2). Notably, there was a low overlap between the primary actives across these genes—only 5% of the hits showed activity against all three targets (Fig. 4B). Lastly, the primary readouts across tested genes exhibited low correlation, with Pearson correlation coefficients ranging from 0.4 to 0.46 (Supplementary Table S3 and Supplementary Fig. S6).
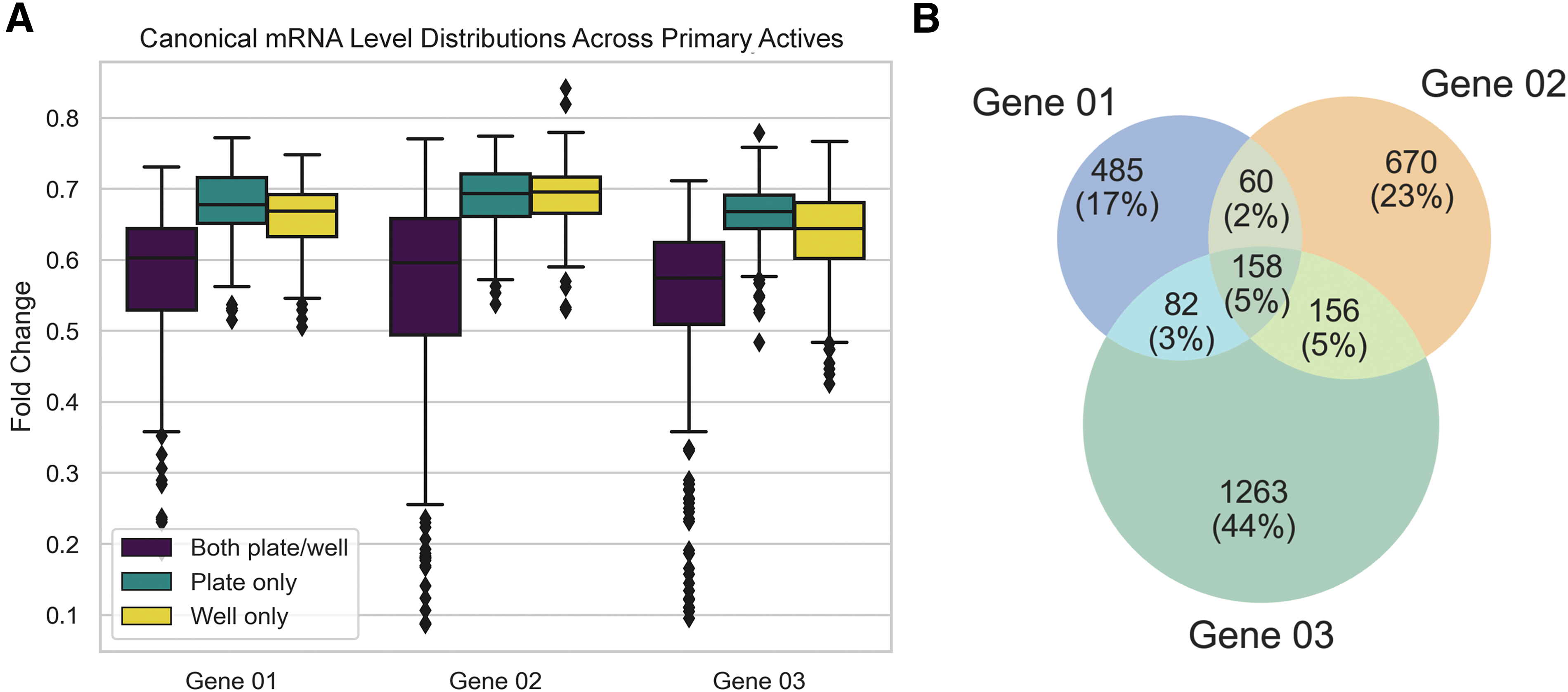
Confirmation screening was also performed using the Luminex QuantiGene Plex technology using freshly prepared samples. Compounds displaying larger fold changes during the primary screen were more likely to confirm upon retesting (Fig. 5A), with confirmation rates ranging from 20% to 46% across the three normalization categories (Supplementary Table S2). Further breakdown of the confirmation rates based on the normalization scheme used during the primary screen showed only qualitative differences (Fig. 5B). The only statistically significant difference was observed for Gene 02, where hits identified by both plate/well-based normalization schemes were more likely to confirm than those identified solely via plate-based normalization (p < 0.01, Fisher Exact Test).26,27
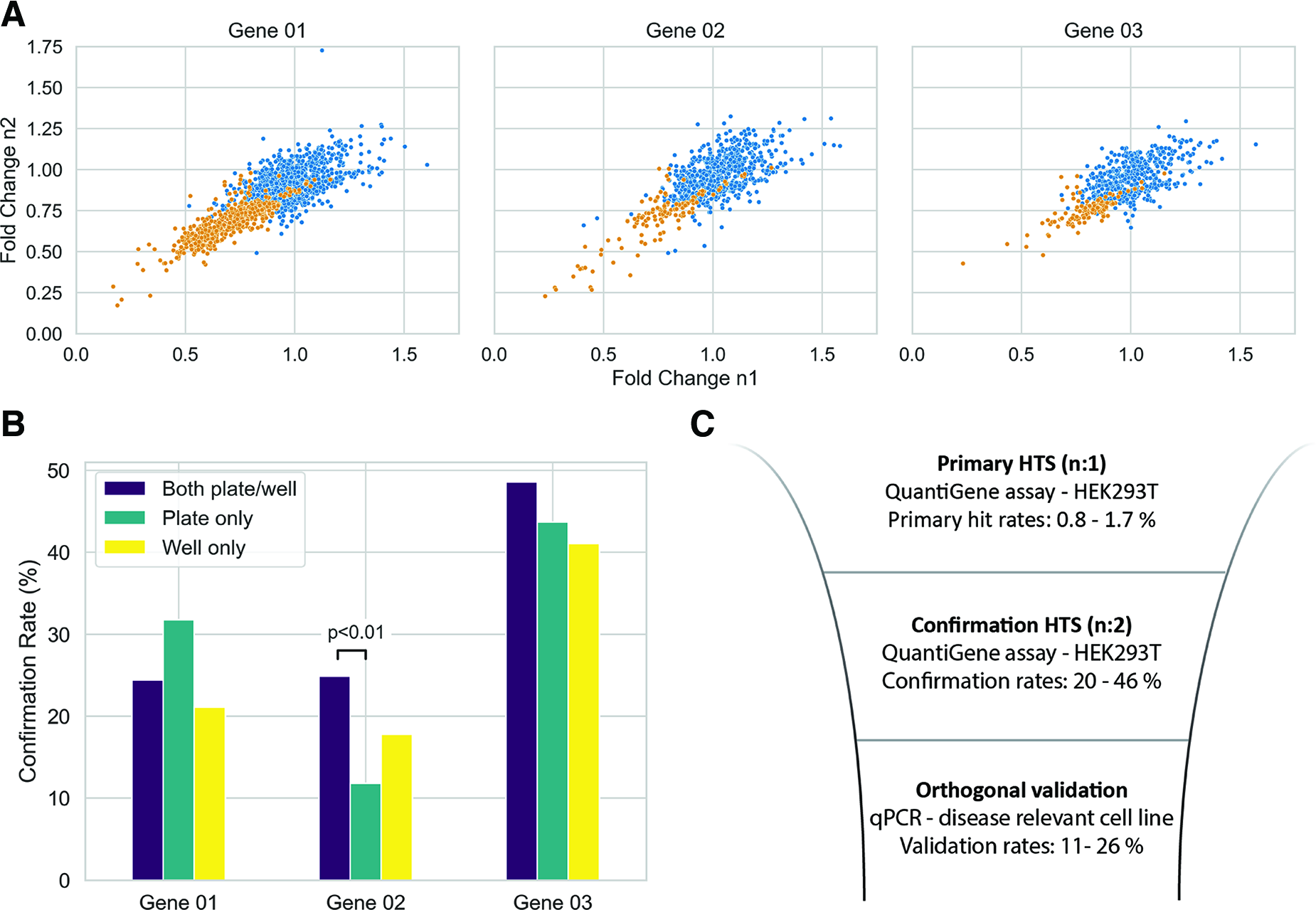
In summary, the primary hit rates for Genes 01–03 ranged from 0.8% to 1.7%, while the confirmation rates upon retesting varied between 20% and 46% (Fig. 5C and Supplementary Table S2). The confirmed hits were then subjected to further validation using a four-point concentration response qPCR assay in a high-expressing cell line, yielding an overall validation rate of 11%–26% (Fig. 5C and Supplementary Table S2). Representative qPCR profiles of compounds demonstrating robust concentration-dependent response in canonical mRNA decrease for Genes 01–03 are shown in Supplementary Figure S7.
Lastly, we ran a substructure motif enrichment analysis to determine if any chemotypes were enriched among the confirmed actives. We saw significant enrichment of known RNA privileged chemotypes28–30 as well as some novel motifs (Fig. 6). Our screen recovered substructure motifs enriched across all tested targets with known RNA binding/modulating activity, such as pyridazine and piperazine substructures. We also identified purine analogs—an RNA-privileged scaffold—as motifs that were strongly enriched across the tested target gene, as well as motifs uniquely enriched for Genes 01–03. Inspection of confirmed active scaffolds identified multiple heteroaromatic and non-aromatic structural motifs. For example, pyrimidine and pyrazolo-pyrimidine substructures show strong enrichment in the Gene 03 confirmed active compounds. Through the implementation of this workflow, unique chemical matter that modulates mRNA processing was identified, providing promising starting points for further hit validation and expansion efforts.
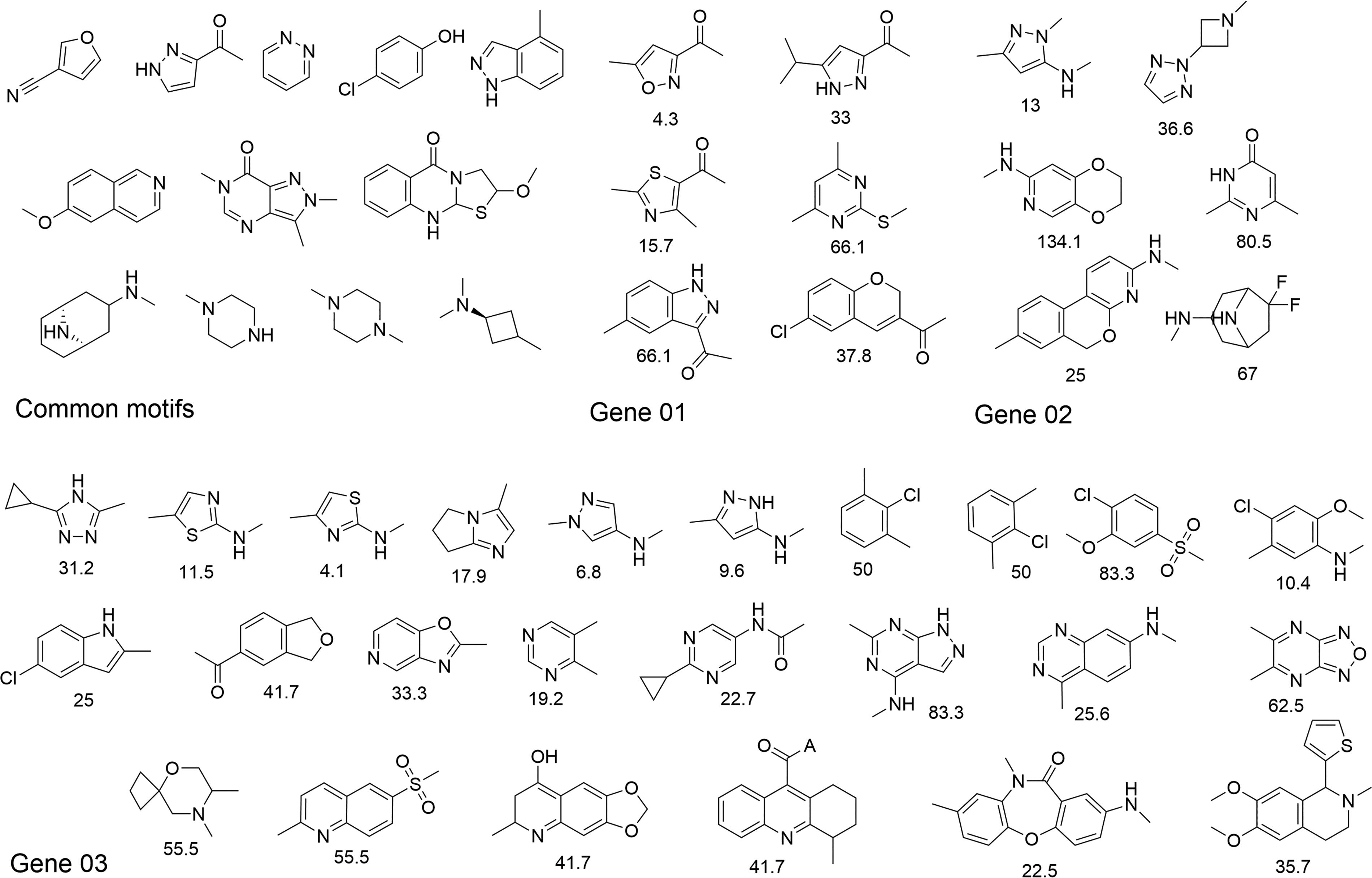
Substructure motifs significantly enriched among confirmed active modulators for Genes 01–03. Multiple scaffolds, including heteroaromatic structures, are enriched in the confirmed hit lists compared to their frequency in the primary screening library. Values represent enrichment ratios for each uniquely enriched substructure per target, indicating structural features associated with target-specific activity.
CONCLUSIONS
Here, we employed the bead-based QuantiGene Plex Gene Expression technology to concurrently assess the effects of compound treatment on canonical mRNA levels for 12 distinct gene targets. This technology was selected for its ability to monitor transcript levels in a highly multiplexed and parallel format, enabling simultaneous quantification of multiple analytes within a single well. By applying this assay in a HTS context using 384-well plates, we identified lead-like compounds capable of modulating target gene mRNA expression. The outcomes across the target genes displayed limited collinearity and minimal overlap, suggesting selective modulation of these target genes at the mRNA transcript level.
To address potential false negatives arising from assay artifacts and plate effects, traditional plate-based normalization was supplemented with a well-based normalization scheme. This strategy was designed to rescue false-negative compounds that might otherwise have been overlooked. Recognizing the potential for increased false positives with well-based normalization, a clustering step prior to normalization was also incorporated. This allowed plates exhibiting similar effects to be grouped together, thereby improving the accuracy of well-based z-score normalization and hit calling.
Our data demonstrates comparable hit confirmation rates between plate- and well-based normalization strategies. Notably, approximately one-third of the confirmed actives were uniquely identified through the well-based normalization, with no appreciable drop in the confirmation rates. For example, compounds B and D that were further validated by qPCR (Supplementary Fig. S7) were only considered active in the primary HTS using the well-based strategy. This further demonstrates the value of this strategy to rescue compounds that would otherwise have been considered inactive. The workflow we present is adaptable to other HTS formats, including multiplexed in vitro biophysical and cell-based screens.31,32 Our approach identified structurally distinct bioactive compounds for multiple target genes with robust hit confirmation rates. Furthermore, a follow-up orthogonal qPCR validation study—featuring a four-point concentration-response analysis of confirmed actively—yielded high validation rates, underscoring the reliability and robustness of our screening and analysis strategy.
AUTHORS’ CONTRIBUTIONS
S.S.: Software, data curation, formal analysis, writing—original draft, writing—review and editing, and visualization. N.S.S.: Methodology, validation, formal analysis, investigation, data curation, writing—original draft, and supervision. M.W.: Methodology, validation, formal analysis, investigation, data curation, and writing—review and editing. J.M.: Formal analysis and writing—review and editing. D.J.R.: Conceptualization, resources, writing—review and editing, supervision, and funding acquisition. F.H.V.: Conceptualization, methodology, resources, writing—review and editing, supervision, and funding acquisition.
Footnotes
ACKNOWLEDGMENTS
All Remixers involved in cell culture, compound treatment, HTS assay, and follow-up qPCR assay performance are thanked for their contributions. Thermo Fisher Scientific is thanked for probe design, bioinformatic support, and technical assistance during the HTS.
FUNDING INFORMATION
This work was funded by
DISCLOSURE STATEMENT
S.S., N.S.S., M.W., D.J.R., and F.H.V. are employees of
